Abstract
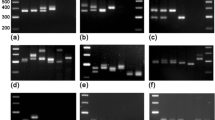
Fusarium oxysporum f. sp. ciceri, the causal agent of chickpea wilt, is an important fungal pathogen in India. Thirteen oligonucleotide probes complementary to microsatellite loci, in combination with 11 restriction enzymes, were used to assess the potential of such markers to study genetic variability in four Indian races of the pathogen. Hybridisation patterns, which were dependent upon both the restriction enzyme and oligonucleotide probe used, revealed the presence of different repeat motifs in the F. oxysporum f. sp. ciceri genome. Among the restriction enzymes used, hexa-cutting enzymes were more informative than tetra- and penta-cutting enzymes, whereas tetranucleotide and trinucleotide repeats yielded better hybridisation patterns than dinucleotide repeats. Dependent upon the levels of polymorphism detected, we have identified (AGT)5, (ATC)5 and (GATA)4 as the best fingerprinting probes for the F. oxysporum f. sp. ciceri races. The distribution of microsatellite repeats in the genome revealed races 1 and 4 to be closely related at a similarity index value of 76.6%, as compared to race 2 at a similarity value of 67.3%; race 3 was very distinct at a similarity value of 26.7%. Our study demonstrates the potential of oligonucleotide probes for fingerprinting and studying variability in the F. oxysporum f. sp. ciceri races and represents a step towards the identification of potential race diagnostic markers.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 12 March 2000 / Accepted: 14 April 2000
Rights and permissions
About this article
Cite this article
Barve, M., Haware, M., Sainani, M. et al. Potential of microsatellites to distinguish four races of Fusarium oxysporum f. sp. ciceri prevalent in India. Theor Appl Genet 102, 138–147 (2001). https://doi.org/10.1007/s001220051629
Issue Date:
DOI: https://doi.org/10.1007/s001220051629