Abstract
Key message
A whole-genome resequencing-derived SNP dataset used for genome-wide association analysis revealed 196 loci significantly associated with drought stress based on root traits. Candidate genes identified in the regions of these loci include homologs of known drought resistance genes in A. thaliana.
Abstract
Drought is the main abiotic constraint of the production of common bean. Improved adaptation to drought environments has become a main goal of crop breeding due to the increasing scarcity of water that will occur in the future. The overall objective of our study was to identify genomic regions associated with drought resistance based on root traits using genome-wide association analysis. A natural population of 438 common bean accessions was evaluated for root traits: root surface area, root average diameter, root volume, total root length, taproot length, lateral root number, root dry weight, lateral root length, special root weight/length, using seed germination pouches under drought conditions and in well-watered environments. The coefficient of variation ranged from 11.24% (root average diameter) to 38.19% (root dry weight) in the well-watered environment and from 9.61% (root average diameter) to 39.05% (lateral root length) under drought stress. A whole-genome resequencing-derived SNP dataset revealed 196 loci containing 230 candidate SNPs associated with drought resistance. Seventeen candidate SNPs were simultaneously associated with more than two traits. Forty-one loci were simultaneously associated with more than two traits, and eleven loci were colocated with loci previously reported to be related to drought resistance. Candidate genes of the associated loci included the ABA-responsive element-binding protein family, MYB, NAC, the protein kinase superfamily, etc. These results revealed promising alleles linked to drought resistance or root traits, providing insights into the genetic basis of drought resistance and roots, which will be useful for common bean improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common bean (Phaseolus vulgaris L.), with high protein and rich nutrition, is the most important legume crop species for direct human consumption and is particularly important in many developing countries (Broughton et al. 2003; Beebe et al. 2013; Dwivedi et al. 2012; Wu et al. 2020). However, drought usually causes severe losses in common bean production. Approximately 60% of production regions for common bean undergo intermittent or terminal drought (Thung et al. 1999), making drought the second largest contributor to reduced production after disease (Rao et al. 2001). In Africa, approximately 40% of the production area of common bean is affected by frequent droughts, resulting in approximately 300,000 Mt of yield losses every year (Asfaw et al. 2012). In Latin America, the production loss of common bean can reach 73% under drought stress. Moreover, only 7% of planting areas are not affected by drought (Broughton et al. 2003). Common bean is generally cultivated on marginal lands that are susceptible to abiotic and biotic stresses (Beebe et al. 2012). Due to insufficient rainfall and limited irrigation equipment, drought has become the main cause of yield losses of common bean in China. Moreover, climate models predict that the main production area of common bean in China will become successively drier during the next several decades (Piao et al. 2010). Molecular breeding, which requires deep insight into the genetic mechanism of drought resistance, is considered to be an effective and powerful strategy for drought resistance improvement (Li et al. 2019a).
Many studies have been performed on the drought resistance of common bean, mainly focusing on the selection of drought-tolerant germplasm, but limited numbers of drought-related QTLs have been identified using different molecular markers. In the absence of an effective linkage map, Schneider et al. (1997) identified drought-related random amplification polymorphism DNA (RAPD) markers using two recombinant inbred line (RIL) populations under stress and non-stress conditions. A genetic map integrating 186 amplified fragment length polymorphism (AFLP), RAPD and simple sequence repeat (SSR) markers was generated using a RIL population from BAT477 × DOR364 (Blair et al. 2012) and was used to detect drought-related QTLs. Five, 5, 16, 3 and 6 QTLs were associated with yield, yield per day, seed weight, days to flowering and days to maturity, respectively, under drought conditions in this study, showing that drought influences production and development. The same population was used to study the genetics of drought resistance in different regions (Asfaw et al. 2012), and drought-related QTLs based on yield have been mapped to Pv04 and Pv08 under drought stress. Drought resistance analyses of common bean have mainly been carried out by genetic linkage analysis (Mukeshimana et al. 2014; Trapp et al. 2015; Boris et al. 2017). Recently, genome-wide association studies (GWASs) have also become powerful tools for dissecting the genetic basis of complex quantitative traits and have been successfully applied for discovering drought-related genes in many crop species, such as maize and wheat (Mao et al. 2015; Wang et al. 2016; Li et al. 2019a). In common bean, GWASs have also been performed to identify genetic loci associated with drought resistance and drought-related traits (Emiliano et al. 2015; Hoyos-Villegas et al. 2017; Berny et al. 2019). Hoyos-Villegas et al. (2017) detected eight drought-related SNPs, one of which was associated with the degree of wilting. All these studies were carried out in the field based on aboveground traits at the maturity stage; however, the underlying genetic basis of root traits related to drought resistance remains to be elucidated.
Plant drought resistance involves four major mechanisms: drought avoidance, drought tolerance, drought escape and drought recovery (Fang et al. 2015). Drought avoidance is the capability of plants to maintain fundamental normal physiological processes under mild or moderate drought stress conditions by adjusting certain morphological structures or growth rates to avoid the negative effects caused by drought stress (Fang et al. 2015). A deep root system has been proposed to be an important drought avoidance response of common bean because it can increase water-uptake efficiency (Sponchiado et al. 1989; Frahm et al. 2004). This water uptake depends on the size (length or mass), activity and spatial distribution of roots (Huang et al. 2000). However, root growth and development are slow or stagnant under drought stress and can even die when soil experience severe drought in the field (Smucker et al. 1991). A healthy and thriving root system can maintain plant water status, improve defense against environmental stresses such as insect pests, and ultimately stabilize production under drought conditions (Mun˜oz-Perea et al. 2006; Kamoshita et al. 2008). The importance of root traits in adapting to drought stress is well recognized in common bean (Rao et al. 2001; Beebe et al. 2008). However, the filed measurement of root traits is a time-consuming and laborious process. In addition, root damage is easy to occur in the sampling process, leading to root losses. There are some alternative ways to better measure and evaluate the root system, such as PVE sleeve tube, germination pouch. Root traits, such as root depth, root length, root volume, average root diameter and specific root length, were measured using PVC sleeve tubes in the greenhouse under drought and non-stress conditions, and eight root-related QTLs mapped to Pv01, Pv10 and Pv11 (Asfaw and Blair 2012). This strategy has been utilized in gaining insights of the genetic basis of root traits in improving drought tolerance in other crop, such as rice (Li et al. 2017), wheat (Li et al. 2019b). However, root QTL with effect on drought resistance has not been reported in common bean at bud stage as far as we know. It is generally believed that crops are most sensitive to environmental changes during germination, and drought resistance at bud stage is one of the important contents to study drought resistance of crops (Yordanov et al. 2000; Ashraf 2010;). Thus, the genetic mechanism of drought resistance related to root traits at bud stage needs investigation.
In this study, 438 common bean accessions were phenotyped for nine different root traits at the bud stage under drought stress and unstressed conditions. GWASs were conducted to identify loci associated with drought resistance based on the relative value of nine root traits and a dataset consisting of 4.8 M SNPs. Our study aimed to (a) identify potential trait-marker associations in response to drought stress and (b) identify drought-related candidate genes within the mapped regions.
Materials and methods
Plant material and genotypic data
A total of 438 common bean accessions representing broad genetic diversity, including accessions in China and elsewhere, were screened for a GWAS (Table S1). This common bean panel was derived from the whole-genome resequencing panel of 683 accessions as previously described (Wu et al. 2020). Genotypic data generated from the whole-genome resequencing project contained 4,811,097 SNPs. For the GWAS, nucleotide variations were further filtered based on the missing rates (≥ 10%) and a minor allele frequency of < 0.05, yielding 3,783,033 qualified SNPs.
Experimental design and drought stress treatment
For the GWAS of root traits at the seedling stage, 438 common bean accessions were grown in seed germination pouches (height:width = 30 cm:25.5 cm) (www.phytotc.com, Fig. S1) (German et al. 2000; Liao et al. 2001). The seed germination pouches served as inexpensive, space-saving replacements for pots and similar containers used to grow plants, and the growth of the root system can be observed visually. Uniform seeds of common bean with the same degree of fullness were subjected to surface sterilization and germinated in distilled water, with a germination box (12 × 12 × 6 cm) used as a bed for the seeds. After 2–3 d, seedlings with radicle lengths of 2 ± 0.1 cm and undifferentiated primary roots were transferred to the seed germination pouches and placed in the trough of the paper wick. The pouches were then fixed to a support plate using clips, and then placed on an acrylic shelf, which helps maintain the pouches upright. The shelf was ultimately placed in an incubator, and the conditions were set as follows: 12 h of light at 25°C; 12 h of darkness at 22°C. In this study, we simulated drought stress using −0.7 MPa PEG 6000 solution (19.6%) (Li et al. 2013), with distilled water treatment as a control. Before transferring seedlings to the pouch, 100 ml of distilled water or PEG 6000 solution was added to the pouch, and the paper wick was completely moistened. To keep the paper wick moist, an additional 10 ml of distilled water or PEG 6000 solution was added to the trough of the pouch every 24 h. In addition, to reduce moisture evaporation and keep the paper wick moist, we attached 2–3 clamps to the opening of the pouch. The common bean seedlings were grown in the incubator for 6 d, and the related phenotypes were evaluated. For each sample, three seedlings were placed in the same seed germination pouch, and six seedlings were subjected to well-watered conditions or a drought stress environment in the same experiment. The location of each pouch in the incubator was random. The experiment was repeated three times.
Phenotypic data measurement and analysis
The root traits were measured after incubation for 6 days. To measure these traits, the plants were removed from the pouches, and the roots were washed clean with water. A total of nine root-related traits were phenotyped in this study, including root surface area (RSA), root average diameter (RAD), root volume (RV), total root length (TRL), taproot length (TL), lateral root number (LRN), root dry weight (RDW), lateral root length (LRL) and special root weight/length (SRL). The roots were spread out without overlap or crossing and scanned with a CI-600 root scanning system (CID Inc, America), which generated images in TIF format. From the scanned images, RSA (cm2), RAD (mm), RV (cm3) and TRL (cm) were analyzed using WinRHIZO Tron MF image analysis software (Regent Instruments Inc., Canada) (Armengaud et al. 2009). The TL (cm), which represents the length of the taproot from the root tip to the root node, was measured with a ruler. The primary lateral roots longer than 2 cm were considered LRNs. The roots were cut from the basal node of the plant, and the dry weight of the roots (RDW, mg) was measured after the roots were heated at 105°C for 30 min followed by 75°C for 48 h. LRL (cm) represents length of all lateral roots. The SRL (mg/cm) represents the root dry weight per unit root length. The traits investigated for each treatment were represented by the suffix “W” or “D”: “W” represents the well-watered treatment, and “D” represents the drought stress treatment. For instance, RSAD denotes the root surface area under drought stress, and RSAW denotes the root surface area under well-watered condition. The tolerance of the genotype to drought for each trait was evaluated using a relative value (Li et al. 2019a), represented by the prefix “R” in this study and calculated as the ratio of the mean value of the investigated trait under drought stress conditions to that under water-well conditions; for instance, RRSA denotes the drought stress tolerance based on the root surface area and the formula is as follows: RRSA = RSAD/RSAW.
The mean values were used for descriptive statistical analysis. Statistical analysis and significance testing were performed using IBM SPSS 19.0 software. Correlations among different traits were evaluated using the R package based on the Pearson's correlation coefficient.
Genome-wide association study
In the present study, we focused on the drought resistance of common bean accessions. Thus, nine root traits and 3,783,033 SNPs were included in a GWAS. Principal component analysis (PCA) was performed using PLINK 1.9 software based on all SNPs after filtration. The first three principal components were used as the principal component (PC) matrix for the association method. The kinship matrix (K matrix) was calculated using Tassel 5.0. Two GWAS methods, the general linear model (GLM) and the compressed mixed linear model (cMLM), were used to detect the associations between markers and traits using Tassel 5.0. The P value threshold of the entire population was 1 × 10–6. Given the trade-off between dismissing false positives and accepting false negatives (Li et al. 2019a), the SNPs that exceeded the threshold (1 × 10–6) in both models were considered associated SNPs in this study. The chromosome region with an associated SNP ± LD decay distance was considered a candidate segment or associated locus (represented by prefix “Locus_”), and the LD decay distance was different for different chromosomes as previously described (Table S2, Wu et al. 2020). When two associated loci overlapped, the overlapping part was considered a new locus. The peak SNPs (SNPs with the lowest P value) for each locus were defined as candidate SNPs.
Locus identification and candidate gene prediction
We searched for previously published QTLs that are related to root or drought resistance and whose physical region was determined by the left border marker and the right border marker or the physical position of markers ± the LD decay distance on the PHASEOLUS GENES website (http://phaseolusgenes.bioinformatics.usdavis.edu) and in the NCBI database (https://www.ncbi.nlm.nih.gov/). An overlapping locus was identified if the physical region of the association locus overlapped with the physical region of any previously reported QTL or SNP for root traits or drought resistance traits.
We annotated the genes of all candidate segments using the annotation file of common bean accession G19833 v1.0 (Schmutz et al. 2014) and searched for homologous genes in A. thaliana according to the NCBI database (https://www.ncbi.nlm.nih.gov/). A candidate gene was claimed if (1) it was a stress-responsive transcription factor, such as a NAC, MYB, or bHLH transcription factor, and (2) its homologous genes in A. thaliana were identified in a previous study as being involved in the drought response, osmotic response, ABA response or root development.
Results
Phenotypic analysis of root traits under stress and non-stressed conditions
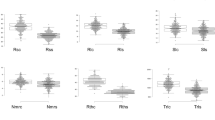
The seedlings of 438 common bean accessions were subjected to drought stress (D) or well-watered (W) conditions using germination pouches. A total of nine root traits were measured after treatment. All the root traits showed significant differences between drought stress and the well-watered treatments (Fig. 1). Drought stress simulated by PEG 6000 solution had a negative influence on all root traits except RAD and SRL. The range of phenotypic variation differed among the traits; the highest coefficient of variation (CV) was observed for LRLD (39.05%), followed by RDWW (38.19%), and the lowest coefficient of variation was observed for RADD (9.61%) (Table S3). In addition, the descriptive statistics of the relative value of the nine root traits were summarized (Table S4). The RLRL ranged from 3.51% to 44.28% with an average of 19.15% and the highest coefficient of variation of 35.79%, followed by the RTRL, which ranged from 8.34% to 59.63% with an average of 23.00% and coefficient of variation of 31.30%; the RRAD ranged from 82.62% to 146.96% with an average of 106.36% and the lowest coefficient of variation of 8.26% (Table S4). The histograms demonstrated that the relative values of each all the root traits conformed to normal distributions, indicating that the phenotypic data are reliable for further analysis (Fig. S2).
Boxplots of the means of the root traits of 438 common bean accessions. The middle line indicates the median, the box indicates the range of the 25th and 75th percentiles of the total data, the whiskers indicate the interquartile range, and the outer dots are outliers. The different letters indicate statistically significant differences at the level of p < 0.01, Student’s t test, two-tailed. D drought stress (PEG 6000 solution), W well-watered (ddH2O), RSA root surface area, RAD root average diameter, RV root volume, TRL total root length, TL taproot length, LRN lateral root number, RDW root dry weight, LRL lateral root length SRL special root weight/length
Phenotypic trait correlations
To study the relationships between different root traits, the correlations of the root traits under different treatments were analyzed. The root traits were significantly correlated between the drought stress and well-watered treatments (Fig. 2). Pairwise positive correlations (p < 0.01) were detected for TL, TRL and LRL between the different treatments, perhaps, because they are related to the same type of trait: root length. The same result was detected between the root weight traits—RDW and SRL. The RDW was positively correlated (p < 0.01) with the three root length traits, but the SRL was negatively correlated (p < 0.01) with them, suggesting that the rate of increase in root length was greater than that of root weight regardless of the treatment. We also noticed that RAD was negatively correlated with root length but positively correlated (p < 0.01) with RDW, RV and SRL. For the relative value (Fig. S3), pairwise positive correlations (p < 0.01) were noticed among RTL, RTRL and RLRL, each of which was positively correlated (p < 0.01) with RRDW, RRSA and RRV but negatively correlated (p < 0.01) with RSRL. The two root weight traits, RRDW and RSRL, were positively correlated (p < 0.01), the RRDW was positively correlated (p < 0.01) with all the relative traits, but the RSRL was negatively correlated (p < 0.01) with all the relative traits except RRAD. RRV was positively correlated (p < 0.01) with all traits except RSRL, and RRSA was positively correlated (p < 0.01) with all traits except RRAD.
Heat map of the correlations between the means of the root traits. The blue color indicates a positive correlation, the red color indicates a negative correlation, and the darkness of the color corresponds to the value of the correlation coefficient according to the legend. The size of the circle indicates the absolute value of the correlation coefficient; the larger the circle is, the greater the value. D drought stress (PEG 6000 solution), W well-watered (ddH2O), RSA root surface area, RAD root average diameter, RV root volume, TRL total root length, TL taproot length, LRN lateral root number, RDW root dry weight, LRL lateral root length, SRL special root weight/length
A GWAS revealed marker-trait associations for drought resistance
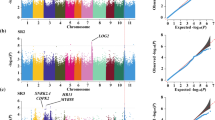
A GWAS was conducted using the phenotypic data of root traits from drought stress and well-watered trials using the general linear model (GLM) and compressed mixed linear model (cMLM). A total of 2542 and 408 significant SNPs were detected by GLM and cMLM, respectively (Table S5, Table S6). A total of 389 SNPs were simultaneously detected by both models and were considered associated SNPs. As described, the chromosomal region with associated SNPs ± LD decay distance was considered a candidate segment or an associated locus. When two associated loci overlapped, the overlapping part was considered a new locus instead of the two loci, and the peak SNPs (SNPs with the lowest P value) for each locus were defined as candidate SNPs. A total of 196 loci were found for 9 quantitative traits, containing 230 candidate SNPs (Table S7). Among these 196 loci, in total, 104, 47, 45, 23, 19, 4, 3 and 3 loci were associated with RTRL, RSRL, RRSA, RRV, RLRL, RTL, RRAD and RRDW, respectively (Table S7). There were 41 loci associated with more than one trait (Table S10). In particular, Locus_138 on Chr08 was associated with five traits (RTRL, RLRL, RSRL, RRSA and RRV), and Locus_134 on Chr08 was associated with four traits (RTRL, RSRL, RRSA and RRV). Moreover, six loci were simultaneously associated with three traits.
The information and physical location of QTLs previously reported to be associated with drought resistance were obtained from the NCBI database and compared with the GWAS results. Eleven loci colocated with loci previously reported to be related to drought resistance (Blair et al. 2012; Emiliano et al. 2015; Trapp et al. 2015; Hoyos-Villegas et al. 2017) (Table S7).
Among these candidate SNPs, 16 were associated with two traits (Fig. 3a), one SNP (Chr02__3484349) was simultaneously associated with three traits (RLRL, RRSA and RRV) (Fig. 3b), and this SNP was present in the intron of the Phvul.002G034800 gene, which encodes a protein kinase superfamily member. Further analysis demonstrated that 130 of 230 candidate SNPs were present in intergenic regions; 57, up/downstream (within 2 kb of the genes) regions or 3′/5′ UTRs of genes; and 43, genic regions (exonic or intronic sequences) (Table S7). Combined gene expression profiles constructed from contrasting RNA-seq-based drought-resistant (Long 22–0579) and drought-sensitive (Naihua) genotypes from different gene pools under drought stress and non-stressed conditions (Wu et al. 2014) demonstrated that four SNPs were located within up-regulated genes, and 22 SNPs were located within down-regulated genes in both varieties under a drought condition compared with under a well-watered condition (Table S7). Among these 57 candidate SNPs located in up/downstream regions or in 3′/5′ UTRs of genes, 36 candidate SNPs were present upstream of genes, of which one was upregulated and seven were downregulated in both varieties under drought conditions compared with well-watered conditions (Table S7, Wu et al. 2014), indicating a causal relationship between candidate SNPs and gene expression. Additionally, 7 of the 57 SNPs were located in reported genes whose homologs in A. thaliana are involved in the response to ABA, drought and osmotic stresses and in root development (Strizhov et al. 1997; Li et al. 2002; Cantero et al. 2006; Lin et al. 2011; Xi et al. 2012; Sura et al. 2017; Podia et al. 2018) (Table S8). Among the 43 candidate SNPs located in genic regions (exonic or intronic sequences), 12 were located in the exonic sequences of genes, one of which is located in a gene whose homolog in A. thaliana is reportedly involved in the response to ABA (Li et al. 2007) (Table S8, S9). These results verified that the candidate SNPs were associated with drought resistance.
Consistent significant SNPs for the relative value of nine root traits. a Network plot of traits and consistent significant SNPs. The line indicates the significant association between a marker and a trait. b Local Manhattan plot surrounding the significant SNP (Chr02__3484349) for the RRSA, RRV, and RLRL
Identification of drought-related candidate genes
Predicting and annotating candidate genes at candidate SNP-harbored loci could be an effective method for identifying causal genes (Li et al. 2019a; Wu et al. 2020). Therefore, the putative candidate genes of all loci were annotated. A total of 2962 genes in 196 loci were annotated (Table S11); we subsequently searched the NCBI database to determine the function of these genes in A. thaliana. A total of 164 genes at 88 loci were considered candidate genes, including genes of transcription factor families that respond to drought stress, such as members of the NAC, MYB, bHLH and AREB families (Mao et al. 2015; Wang et al. 2017a, b; Li et al. 2019c), as well as membrane proteins, protein kinases and other types of proteins (Table S12). Using previous transcriptome information (Wu et al. 2014), we determined that the expression of 22 genes was upregulated and that of 32 was downregulated in both varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014). WRKY 51, which was found at Locus_35, has been shown to promote lateral root formation and is induced by drought and ABA in wheat (Wang et al. 2013; Hu et al. 2018). In the present study, Locus_35 was associated with RTRL and RSRL and contained three associated SNPs (RTRL-associated Chr02__43041529, RSRL-associated Chr02__43108109 and Chr02__43108142) (Fig. 4a) with close physical distance (66.61 kb). Chr02__43041529 was linked genetically to Chr02__43108109 (r2 = 0.95) but was not linked genetically to Chr02__43108142. Based on the variation in three SNPs, two haplotypes were identified within the germplasm population. We noticed that the RTRL of Hap1 was significantly higher than that of Hap2, but the RSRL of Hap1 was significantly lower than that of Hap2 (Fig. 4b). These results further suggested that these loci were related to drought resistance.
Colocalization of RTRL-associated SNPs (Chr02__43041529) and RSRL-associated SNPs (Chr02__43108109 and Chr02__43108142) at the same locus. a Local Manhattan plot of the region of Locus_35 (top) and a linkage disequilibrium heatmap (bottom) surrounding the SNP for the RTRL and RSRL on chromosome 2. The dotted lines indicate the position of the SNPs, the darkness of color of each box corresponds to the R2 value according to the legend, and the red lines indicate Chr02__43041529, Chr02__43108109 and Chr02__43108142. b Bar diagrams for RTRL and RSRL based on the haplotypes (Hap), which were formed by the variation in the marker alleles in Chr02__43041529, Chr02__43108109 and Chr02__43108142. The different letters indicate statistically significant differences at the level of p < 0.05, Student’s t test, two-tailed. RTRL relative total root length, RSRL relative special root weight/length
Longer and deeper roots are essential for plants under drought stress, and they can efficiently absorb water from soil that is dry at the surface but retains moisture in deep layers (Gupta et al. 2020). Therefore, RTRL, RTL and RLRL can reflect the influence of drought stress on common bean. For RTRL, 98 candidate genes were found (Table S12). The strongest association signal (Chr07__41151877, p = 5.04E-07) at Locus_118 was located in the upstream region of Phvul.007G176000, whose homolog in A. thaliana encodes histone H2A 11 and is expressed in response to osmotic stress (Sura et al. 2017) (Table S8). The candidate SNP (Chr03__46736106, p = 9.68 × 10–7) at Locus_53 was localized in the CDS region of gene Phvul.003G243300 (Fig. 5a, b), which was annotated as mitogen-activated protein kinase kinase kinase kinase 21 (MAPKKK21); this SNP was a missense mutation (Fig. 5b). We observed that the RTRL of accessions with the T allele was significantly higher than that of accessions with the C allele (Fig. 5c). The gene expression profiles showed that the genes were downregulated in both varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014). Furthermore, Locus_53 harbored another two candidate genes, Phvul.003G242900 and Phvul.003G243800. Phvul.003G242900 encodes a bHLH DNA-binding superfamily protein and was upregulated in both varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014). Phvul.003G243800 encodes zeaxanthin epoxidase (ZEP) (ABA1), whose homolog in A. thaliana is involved in osmotic stress tolerance (Xiong et al. 2002; Park et al. 2008), which was downregulated in both varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014). For RTL, two candidate genes were found (Table S12). Phvul.010G114800, which was annotated as basic helix-loop-helix (bHLH) DNA-binding superfamily protein, was found at Locus_175. The lowest association (Chr10__38036952, p = 2.75E-07) of this region was found upstream of the gene. For RLRL, 41 candidate genes were found (Table S12). The strongest association signal (Chr08__59307769, p = 6.15E-07) at Locus_145 was found in the upstream region of Phvul.008G289400, whose homolog in A. thaliana encodes a portal protein and is involved in the response to drought stress (Podia et al. 2018) (Table S8). Another candidate gene at this locus, Phvul.008G290200, encodes UDP-glucosyl transferase 71B6, which is involved in the response to ABA (Priest et al. 2010). The gene expression profile showed that these two genes were downregulated in both varieties under drought conditions compared with under well-watered conditions (Table S12, Wu et al. 2014). Locus_120 was associated with RTRL and RLRL and contained two candidate SNPs (RTRL-associated Chr07__42273823, RTRL- and RLRL-associated Chr07__42296614) (Fig. 6a). Interestingly, the SNP (Chr07__42296614, p = 7.76E-07) was found in the upstream region of Phvul.007G186700. Its homologous gene (AtGT2L) in A. thaliana encodes a duplicated homeodomain-like superfamily protein, and its expression increased after ABA treatment (Xi et al. 2012). Further analysis showed these two SNPs were physically close to each other (22.79 kb) and were linked genetically (r2 = 0.93). Based on the variation in these two SNPs, two haplotypes were identified within the germplasm population. We noticed that the RTRL and RLRL of Hap1 were significantly lower than those of Hap2 (Fig. 6b).
Analysis of significant RTRL-associated SNPs (Chr03__46736106) associated with RTRL. a Local Manhattan plot of the Phvul.003G243300 region on chromosome 3. The dotted arrow indicates the position of a significant SNP (Chr03__46736106). b Gene structure of Phvul.003G243300 and the mutated base and amino acid. The filled black box represents the coding sequence. The three letters represent the codon containing the mutated base. The dotted arrow and red letters indicate the mutated base (T-G). The brackets indicate the change in the amino acid c boxplot for RTRL based on the haplotypes (Hap), which were formed by the variation in the marker allele at Chr03__46735442. The middle line indicates the median, the box indicates the range of the 25th and 75th percentiles of total data, the whiskers indicate the interquartile range, and the outer dots are outliers. *Indicates p < 0.05, Student’s t test, two-tailed. RTRL relative total root length
Colocalization of consistent significant SNPs for RTRL and RLRL. a Local Manhattan plot of the region of Locus_120 (top) and a linkage disequilibrium heatmap (bottom) surrounding the SNPs for RTRL and RLRL on chromosome 7. The dotted lines indicate the position of SNPs, the darkness of color of each box corresponds to the R2 value according to the legend, and the red lines indicate Chr07__42273823 and Chr07__42296614. b Boxplots for RTRL and RLRL based on the haplotypes (Hap), which were formed by the variation in the marker allele in Chr07__42273823 and Chr07__42296614. The middle line indicates the median, the box indicates the range of the 25th and 75th percentiles of total data, the whiskers indicate the interquartile range, and the outer dots are outliers. *Indicates p < 0.05. **Indicates p < 0.01, Student’s t test, two-tailed. RTRL relative total root length, RLRL relative lateral root length
The surface area, volume and diameter of the root system are also indicators that can reflect root development. For RRV, 18 candidate genes were found (Table S12). We found that the strongest association signal (Chr02__42267364, p = 1.90E-07) at Locus_34 was found in the upstream of Phvul.002G255700, whose homolog in A. thaliana encodes annexin 6, which is involved in the drought stress response (Cantero et al. 2006) (Table S8). The gene expression profile showed that the gene was downregulated in both varieties under drought conditions compared with well-watered conditions (Table S12, Wu et al. 2014). For RRSA, 17 candidate genes were found (Table S12). The strongest signal associated with RRSA (Chr10_2488342, p = 1.84E-09) was present at Locus_160, which was found in the upstream region of Phvul.010G015400, whose homolog in A. thaliana encodes delta 1-pyrroline-5-carboxylate synthase 2 (P5CS2), which is involved in the response to ABA and drought stress (Strizhov et al. 1997). In common bean, P5CS2 is induced by drought stress and salt stress (Chen et al. 2013). The gene expression profile showed that P5CS2 was upregulated in both varieties under a drought condition compared with a well-watered condition (Table S8, Wu et al. 2014). For RRAD, 2 candidate genes were found (Table S12). Phvul.002G224900, which was annotated as NAC domain-containing protein 25, was found at Locus_32, and the gene expression profile showed that this gene was downregulated in both varieties under drought compared with a well-watered condition (Table S12, Wu et al. 2014).
An increase in dry matter accumulation under drought stress is a manifestation of the drought resistance of plants. For RRDW, seven candidate genes were detected (Table S11). The strongest association signal (Chr02__44861559, p = 2.87E-08) was present at Locus_37. Five genes were found at Locus_37: Phvul.002G283600, Phvul.002G285000, Phvul.002G285100, Phvul.002G285800 and Phvul.002G286100. Candidate SNPs were found in the Phvul.002G285100 exonic gene, whose homolog in A. thaliana encodes hydroxysteroid dehydrogenase 1 and is involved in the response to ABA (Li et al. 2007). Phvul.002G285000 was the same as Phvul.002G285100 and encodes the same protein. Phvul.002G283600 and Phvul.002G285800 encode NAC domain-containing protein 83 and WRKY DNA-binding protein 75, respectively, whose homologous genes in A. thaliana are, respectively, involved in the ABA response and root development (Yang et al. 2011; Devaiah et al. 2007). Phvul.002G286100 encodes gamma carbonic anhydrase-like 2, whose homologous gene is involved in the ABA response in A. thaliana (Zhang et al. 2012). The gene expression profile showed that Phvul.002G283600 was downregulated and that Phvul.002G285000, Phvul.002G285100 and Phvul.002G285800 were upregulated in both varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014).
For RSRL, 14 candidate genes were found (Table S12). We found that the strongest association signal (Chr01__23283094, p = 3.22E-08) at Locus_6 was present in the upstream region of the Phvul.001G100900 gene, whose homolog in A. thaliana encodes UBP1-associated protein 2A and is involved in the ABA-activated signaling pathway (Li et al. 2002) (Table S8). At the same locus, another three candidate genes, Phvul.001G100500, Phvul.001G100600 and Phvul.001G101000, were found, which encode A. thaliana NAC domain-containing protein 87, Myb/SANT-like DNA-binding domain protein and ABA-responsive element-binding protein 3, respectively. Phvul.001G101000 (AREB 3) is expressed in response to salt and drought stresses (Uno et al. 2000). At another locus (Locus_27, which is associated with RSRL and RRV), the candidate SNP (Chr02__29371010, p = 6.24E-07) was found in the downstream region of Phvul.002G152900, which encodes expansion A7, and its homolog in A. thaliana promotes root hair elongation (Lin et al. 2011) (Table S8).
Among the loci associated with more than one trait, Locus_138 was associated with five traits (RTRL, RLRL, RSRL, RRSA and RRV). Phvul.008G179800, whose homolog in A. thaliana was annotated as histone deacetylase3 and has been reported to promote root development, was found at this locus (Li et al. 2017). Locus_136 was simultaneously associated with three traits (RTRL, RLRL and RSRL). Phvul.008G178400, whose homolog in A. thaliana encodes a DNAJ heat shock family protein and has been reported to be involved in the ABA response, was located at this locus (Park et al. 2014; Wang et al. 2014). Locus_26 was associated with three traits (RRV, RRSA and RTRL), Phvul.002G146400, whose homologous genes in A. thaliana encode AINTEGUMENTA-like 6 and promote root development, was found at this locus (Galinha et al. 2007). The gene expression profile showed that Phvul.002G146400 was upregulated in both varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014). Locus_150 was associated with RTRL and RSRL and contained seven candidate genes, including four transcription factors: Phvul.009G060900 (bZIP44), Phvul.009G062700 (MYB77), Phvul.009G065500 (AREB2) and Phvul.009G065700 (bHLH), Phvul.009G061000 (which encodes a mitogen-activated protein kinase 1), Phvul.009G063300 (which encodes a transmembrane protein) and Phvul.009G064500 (which encodes a calmodulin like 37). Phvul.009G062700, Phvul.009G065500, Phvul.009G061000, Phvul.009G063300 and Phvul.009G064500have been reported to be involved in ABA or drought stress response in A. thaliana (Table S12, Ortiz-Masia et al. 2007; Shin et al. 2007; Chapman et al. 2019). In addition, only one gene (Phvul.001G079700) was predicted to be present at Locus_4, which harbored one RRSA-associated SNP (Chr01__12623037). Phvul.001G079700 was annotated as a plant protein of unknown function (DUF247). Taken together, all the above results further reflected the relatively high resolution of the GWAS, providing a reference for gene function exploration.
Discussion
Common bean plays a crucial role in sustaining food security in Africa and Latin America but is greatly threatened by drought stress worldwide. To understand the genetic basis of drought resistance in common bean and identify candidate genes underlying drought resistance, we investigated nine root traits under drought stress and well-watered conditions at the bud stage using seed germination pouches and conducted a GWAS to identify trait-marker associations. In the present study, the largest haplotype map available, consisting of 4.8 M SNPs, was used for the GWAS; this map was created by resequencing 683 common bean landraces and breeding lines representing both Andean and Mesoamerican gene pools from China and elsewhere in the national GenBank collection (Wu et al. 2020). The panel of 438 common beans used in the study was derived from the resequencing population by random sampling. Both the number of individuals and the density of markers are relatively large in genome-wide association analyses of common bean. We focused on drought resistance represented by relative values of nine root traits at the bud stage for the GWAS was conducted based on relative values. In the present study, 196 association loci containing 230 candidate SNPs were identified as being linked to drought resistance, with individual candidate SNPs explaining between 6.68% and 11.86% of the phenotypic variation (Table S7). Individual QTLs explained 8–22% of the phenotypic variation in a similar study (Blair et al. 2012). A set of candidate genes were identified as being related to drought resistance. These genes encoded NAC, WRKY, etc., stress-responsive transcription factors; protein kinase superfamily members; and ABA-responsive element-binding proteins. Together, these genes provide reference information for research on the drought resistance of common bean.
In common bean, previous studies have reported several root QTLs associated with different traits, such as rhizosphere acidification (Yan et al. 2004), root rot resistance (Hagerty et al. 2015), low phosphorus tolerance (Liao et al. 2004; Beebe et al. 2006; Ochoa et al. 2006; Cichy et al. 2009) and response to auxin (Remans et al. 2008). However, few studies have reported root QTLs associated with the response to drought in common bean. Asfaw and Blair et al. (2012) detected QTLs associated with rooting patterns of plants grown in PVC tubes in a greenhouse under drought stress and well-watered conditions. Four QTLs were identified as being associated with root traits, such as rooting depth, total root length and root volume, of which one QTL was associated with total root length, fine root length, length of thick roots, root volume, and root dry weight and did not overlap with any yield or yield component QTLs detected in the field. The root-associated locus identified in the present study overlapped with previously reported yield and yield components. Locus_99 overlapped with seed weight-associated BM187 (Blair et al. 2012), Locus_178 overlapped with seed yield-associated SNP46337 (Trapp et al. 2015), and Locus_154 overlapped with hundred-seed weight-associated ss71564685 and ss715646847 (Hoyos-Villegas et al. 2017) as well as SNP55 and SNP20 (Emiliano et al. 2015), indicating the underlying relationship between drought resistance and yield-related traits. In addition, SNP90/SNP91, SNP17, SNP120, SNP80, and SNP54 were associated with days to flowering and the reproduction period in a previous report (Emiliano et al. 2015) and colocated with Locus_19, Locus_54, Locus_57, Locus_100, and Locus_103, respectively, identified in the present study (Table S7). These results suggested that strong drought resistance can enhance seedling vitality early in the reproductive cycle and further influence production and the reproduction period.
Transcription factors, including NAC, bHLH, MYB, DREB, AP2/ERF, WRKY and bZIP members, etc., compose an important class of genes (Lata et al. 2014; Liu et al. 2014; Mao et al. 2015; He et al. 2016; Wang et al. 2017a, b). Transcription factors have an essential role in the response to many types of abiotic and biotic stresses, including drought. The features of transcription factors that respond to drought stress, such as ZmNAC111 (Mao et al. 2015), TaNAC67 (Mao et al. 2014), ZmMYB84 (Wang et al. 2017a, b), OsMYB48-1 (Xiong et al. 2014), OsDREB2A (Cui et al. 2011), OsAP21 (Jin et al. 2012), AtWRKY64 (Ding et al. 2014) and AtWRKY93 (Qin et al. 2015), have been studied in various crop species. In our study, 48 transcription factor genes, including 15 bHLH genes, 10 WRKY genes, 10 MYB genes, 7 NAC genes and 6 bZIP genes, were identified as candidate genes within the locus region. A gene encoding MYB94 was located at Locus_51. The TaMYB94 transcription factor activates cuticular wax biosynthesis in A. thaliana and is important in the plant response to environmental stress, including drought (Lee et al. 2015). Other genes that encode MYB73, MYB30 and MYB77 were located at Locus_3, Locus_129 and Locus_150, respectively. AtMYB77 is involved in lateral root development (Shin et al. 2007), and AtMYB73 is also involved in lateral root development (Zhao et al. 2014), as well as the response to salt stress (Kim et al. 2013). AtMYB30 is involved not only in root cell elongation (Mabuchi et al. 2018) but also in the response to ABA (Zheng et al. 2012). Locus_37 contained two transcription factors, NAC83 and WRKY75. In A. thaliana, NAC83 is an abscisic acid (ABA)-responsive NAC transcription factor that regulates a subset of genes responsive to dehydration (Yang et al. 2011). WRKY75 in A. thaliana is a member of the WRKY transcription factor family reported to be involved in regulating root development (Devaiah et al. 2007). In poplar, WRKY75 is downregulated during the early stages of salt and osmotic stresses and can reduce both reactive oxygen scavenging and the accumulation of proline under stresses and can positively regulate the water loss rate of leaves (Zhao et al. 2019). Locus_102 contained three transcription factors: WRKY14, WRKY48 and MYB78. The expression of WRKY48 is induced by osmotic stress in A. thaliana (Xing et al. 2008). Taken together, these results suggested that these transcription factors might regulate drought resistance in common bean; therefore, further studies on these genes using molecular biology methods could help explain the molecular mechanism underlying drought resistance and the genetic relationship between drought and roots in common bean.
Many other kinds of genes may be involved in the response to drought stress or osmotic stress, including kinase family, ABA-responsive element-binding factor, ARM repeat superfamily protein and G-box binding factor members identified in the present study. Their homologous genes in A. thaliana have been identified as being involved in the response to osmotic stress, water deprivation, drought stress and ABA. Genes encoding protein kinase superfamily members belonging to protein kinases in the SNF1-related protein kinase 2 family were detected at Locus_33, Locus_36 and Locus_42. These genes have critical functions in the response to osmotic stress and drought stress (Fujii et al. 2011). Locus_6 and Locus_150 contained genes encoding the ABA-responsive element-binding factors AREB2 and AREB3, respectively. These genes are members of the basic leucine zipper (bZIP)-type ABRE-binding protein family, of which AREBs regulate novel ABRE-dependent ABA signaling, enhancing the drought resistance of vegetative tissues (Uno et al. 2000; Fujita et al. 2005). A gene encoding an ARM repeat superfamily protein located at Locus_20 was upregulated in response to drought, salt, cold and abscisic acid (ABA) in A. thaliana. Overexpression of this gene led to insensitivity to ABA, while its down-regulation enhanced drought resistance in A. thaliana (Liu et al. 2011). A G-box binding factor was located at Locus_101 and Locus_174. GBF is conserved and functions in the same way in response to drought and other abiotic stresses to confer tolerance in several different plant species (Ramegowda et al. 2017). However, its function is still unknown in common bean.
Analysis of differential gene expression has been proposed as a promising approach to verify candidate genes in the GWAS results (Sun et al. 2017; Wen et al. 2018). In this study, a total of 164 genes were considered candidate genes, among these genes, the expression of 54 genes was changed (22 up-regulated genes and 32 down-regulated gene) in both common bean varieties under drought condition compared with under well-watered condition (Table S12, Wu et al. 2014). RTRL-associated candidate gene, Phvul.010G117200, was found at Locus_177, which encodes homeobox 7, and is the most up-regulated gene with log2(FC) = 9.8. The homolog of in Phvul.010G117200 A. thaliana is in response to water deprivation and ABA (Söderman et al. 1996; Ré et al. 2014; Pruthvi et al. 2014). In summary, 72 percent of up or down regulated genes (39 of 54) were identified to involved in ABA, drought stress response or root development (Table S12). These results further suggested that these genes may be associated with drought resistance, and the function of these genes need to verify.
This study is the first report to identify root or drought resistance-related QTLs on a large scale in common bean. The results of our study revealed a large number of drought-associated loci, greatly enriching the genetic information of drought resistance and root traits and laying a foundation for crop improvement using molecular marker-assisted selection. In addition, a set of genes were identified as prospective candidate genes involved in drought response and root development, providing insights into the genetic character of plant mechanisms in response to drought and providing data for future functional verification of drought resistance genes in common bean.
Availability of data and material
Data supporting the current study can be obtained by contacting the corresponding author (wangshumin@caas.cn).
Code availability
Not applicable.
Abbreviations
- GWAS:
-
Genome-wide association study
- SNP:
-
Single nucleotide polymorphisms
- CHR:
-
Chromosome
- LD:
-
Linkage disequilibrium
- QTL:
-
Quantitative trait loci
- ABA:
-
Abscisic acid
- RSA:
-
Root surface area
- RAD:
-
Root average diameter
- RV:
-
Root volume
- TRL:
-
Total root length
- TL:
-
Taproot length
- LRN:
-
Lateral root number
- RDW:
-
Root dry weight
- LRL:
-
Lateral root length
- SRL:
-
Special root weight/length
References
Armengaud P, Zambaux K, Hills A, Sulpice R, Pattison RJ, Blatt MR, Amtmann A (2009) EZ-Rhizo: integrated software for the fast and accurate measurement of root system architecture. Plant J 57:945–956
Asfaw A, Blair MW (2012) Quantitative trait loci for rooting pattern traits of common beans grown under drought stress versus non-stress conditions. Mol Breed 30:681–695
Asfaw A, Blair MW, Struik PC (2012) Multienvironment quantitative trait loci analysis for photosynthate acquisition accumulation and remobilization traits in common bean under drought stress. G3 2:579–595
Ashraf M (2010) Inducing drought tolerance in plant: recent advances. Biotechnol Advs 28:169–183
Beebe S (2012) Common bean breeding in the tropics. Plant Breed Rev 36:357–426
Beebe SE, Rojas-Pierce M, Yan X, Blair MW, Pedraza F, Munoz F, Tohme J, Lynch JP (2006) Quantitative trait loci for root architecture traits correlated with phosphorus acquisition in common bean. Crop Sci 46:413–423
Beebe S, Rao IM, Cajiao C, Grajales M (2008) Selection for drought resistance in common bean also improves yield in phosphorus limited and favorable environments. Crop Sci 48:582–592
Beebe SE, Rao IM, Blair MW, Acosta-Gallegos JA (2013) Phenotyping common beans for adaptation to drought. Front Physiol 4:1–20
Berny Mier Y, Teran JC, Konzen ER, Medina V, Palkovic A, Ariani A, Tsai SM, Gilbert ME, Gepts P (2019) Root and shoot variation in relation to potential intermittent drought adaptation of Mesoamerican wild common bean (Phaseolus vulgaris L.). Ann Bot 124:917–932
Blair MW, Galeano CH, Tovar E, Torres MCM, Castrillón AV, Beebe SE (2012) Development of a Mesoamerican intra-gene pool genetic map for quantitative trait loci detection in a drought tolerant. Mol Breed 29:71–88
Boris B, Cardoso PJMK, Santa RJ, Denis B, Ribeiro GJG, Caléo A (2017) Mapping QTLs for drought tolerance in a SEA5×AND277 common bean cross with SSRs and SNP markers. Genet Mol Bio 40:813–823
Broughton WJ, Hernandez G, Blair MW, Beebe SE, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp.)—model food legumes. Plant Soil 252:55–128
Cantero A, Barthakur S, Bushart TJ, Chou S, Morgan RO, Fernandez MP, Clark GB, Roux SJ (2006) Expression profiling of the Arabidopsis annexin gene family during germination, de-etiolation and abiotic stress. Plant Physiol Biochem 44:13–24
Chapman K, Taleski M, Ogilvie HA, Imin N, Djordjevic MA (2019) CEP-CEPR1 signalling inhibits the sucrose-dependent enhancement of lateral root growth. J Exp Bot 70:3955–3967
Chen JB, Yang JW, Zhang ZY, Feng XF, Wang SM (2013) Two P5CS genes from common bean exhibiting different tolerance to salt stress in transgenic Arabidopsis. J Genet 92:461–469
Cichy KA, Snapp SS, Blair MW (2009) Plant growth habit, root architecture traits and tolerance to low soil phosphorus in an Andean bean population. Euphytica 165:257–268
Cui M, Zhang W, Zhang Q, Xu Z, Zhu Z, Duan F, Wu R (2011) Induced over-expression of the transcription factor OsDREB2A improves drought tolerance in rice. Plant Physiol Biochem 49:1384–1391
Devaiah BN, Karthikeyan AS, Raghothama KG (2007) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143:1789–1801
Ding Z, Yan J, Xu X, Yu D, Li G, Zhang S, Zheng S (2014) Transcription factor WRKY46 regulates osmotic stress responses and stomatal movement independently in Arabidopsis. Plant J 79:13–27
Dwivedi SL, Sahrawat KL, Rai KN, Blair MW, Pfeiffer W (2012) Nutritionally enhanced staple food crops. Plant Breed Rev 36:169–291
Emiliano VP, González-Chavira Mario M, Patricia GC, Acosta-Gallegos JA, Juan C-P (2015) Identification of novel drought-tolerant-associated SNPs in common bean (Phaseolus vulgaris). Front Plant Sci 6:546
Fang Y, Xiong L (2015) General mechanisms of drought response and their application in drought resistance improvement in plants. Cell Mol Life Sci 72:673–689
Frahm MA, Rosas JC, Mayek-Pérez N, López-Salinas E, Acosta-Gallegos JA, Kelly JD (2004) Breeding beans for resistance to terminal drought in the lowland tropics. Euphytica 136:223–232
Fujii H, Verslues PE, Zhu JK (2011) Arabidopsis decuple mutant reveals the importance of SnRK2 kinases in osmotic stress responses in vivo. Proc Natl Acad Sci U S A 108:1717–1722
Fujita Y, Fujita M, Satoh R, Maruyama K, Parvez MM, Seki M, Hiratsu K, Ohme-Takagi M, Shinozaki K, Yamaguchi-Shinozaki K (2005) AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 17:3470–3488
Galinha C, Hofhuis H, Luijten M, Willemsen V, Blilou I, Heidstra R, Scheres B (2007) PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 449:1053–1057
German MA, Burdman S, Okon Y, Kigel J (2000) Effects of Azospirillum brasilense on root morphology of common bean (Phaseolus vulgaris L.) under different water regimes. Biol Fert Soils 32:259–264
Gupta A, Rico-Medina A, Caño-Delgado AI (2020) The physiology of plant responses to drought. Science 368:266–269
Hagerty CH, Cuesta-Marcos A, Cregan PB, Song Q, McClean P, Noffsinger S, Myers JR (2015) Mapping and root rot resistance and root architecture quantitative trait loci in common bean. Crop Sci 55:1969–1977
He G, Xu J, Wang Y, Liu J, Li P, Chen M, Ma Y, Xu Z (2016) Drought-responsive WRKY transcription factor genes TaWRKY1 and TaWRKY33 from wheat confer drought and/or heat resistance in Arabidopsis. BMC Plant Biol 16:116
Hoyos-Villegas V, Song Q, Kelly JD (2017) Genome-wide association analysis for drought tolerance and associated traits in common bean. Plant Genome. https://doi.org/10.3835/plantgenome2015.12.0122
Hu Z, Wang R, Zheng M, Liu X, Sun Q (2018) TaWRKY51 promotes lateral root formation through negatively regulating ethylene biosynthesis in wheat (Triticum aestivum L.). Plant J 96:372–388
Huang B, Gao H (2000) Root physiological characteristics associated with drought resistance in tall fescue cultivars. Crop Sci 40:196–203
Jin X, Xue Y, Wang R, Xu R, Bian L, Zhu B, Han H, Peng R, Yao Q (2012) Transcription factor OsAP21 gene increases salt/drought tolerance in transgenic Arabidopsis thaliana. Mol Biol Rep 40:1743–1752
Kamoshita A, Babu RC, Manikanda Boopathi N, Fukai S (2008) Phenotypic and genotypic analysis of drought-resistance traits for development of rice cultivars adapted to rainfed environments. Field Crops Res 109:1–23
Kim JH, Nguyen NH, Jeong CY, Nguyen NT, Hong SW, Lee H (2013) Loss of the R2R3 MYB, AtMyb73, causes hyper-induction of the SOS1 and SOS3 genes in response to high salinity in Arabidopsis. J Plant Physiol 170:1461–1465
Lata C, Mishra AK, Muthamilarasan M, Bonthala VS, Khan Y, Prasad M (2014) Genome-wide investigation and expression profiling of AP2/ERF transcription factor superfamily in foxtail millet (Setaria italica L.). PLoS ONE 9:e113092
Lee SB, Suh MC (2015) Cuticular wax biosynthesis is up-regulated by the MYB94 transcription factor in Arabidopsis. Plant Cell Physiol 56:48–60
Li J, Kinoshita T, Pandey S, Ng KY, Gygi SP, Shimazaki KI (2002) Modulation of an RNA-binding protein by abscisic-acid-activated protein kinase. Nature 418:793–797
Li F, Asami T, Wu X, Tsang EW, Cutler AJ (2007) A putative hydroxysteroid dehydrogenase involved in regulating plant growth and development. Plant Physiol 145:87–97
Li L, Wang L, Wu J, Jing R, Wang S (2013) Drought tolerance in common bean germplasm at bud stage. J Plant Genet Res 14:600–605
Li X, Guo Z, Lv Y, Cen X, Ding X, Wu H, Li X, Huang J, Xiong L (2017) Genetic control of the root system in rice under normal and drought stress conditions by genome-wide association study. PLoS Genet 13:e1006889
Li L, Mao X, Wang J, Chang X, Matthew R, Jing R (2019a) Genetic dissection of drought and heat-responsive agronomic traits in wheat. Plant Cell Environ 42:1–14
Li L, Peng Z, Mao X, Wang J, Chang X, Matthew R, Jing R (2019b) Genome-wide association study reveals genomic regions controlling root and shoot traits at late growth stages in wheat. Ann Bot 124:993–1006
Li Z, Liu C, Zhang Y, Wang B, Ran Q, Zhang J (2019c) The bHLH family member ZmPTF1 regulates drought tolerance in maize by promoting root development and abscisic acid synthesis. J Exp Bot 70:5471–5486
Liao H, Rubio G, Yan X, Cao A, Brown KM, Lync JP (2001) Effect of phosphorus availability on basal root shallowness in common bean. Plant Soil 232:69–79
Liao H, Yan X, Rubio G, Beebe S, Blair M, Lynch J (2004) Genetic mapping of basal root gravitropism and phosphorus acquisition efficiency in common bean. Funct Plant Biol 33:959–970
Lin C, Choi HS, Cho HT (2011) Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis. Mol Cells 3:393–397
Liu Y, Wu Y, Huang X, Sun J, Xie Q (2011) AtPUB19, a U-box E3 ubiquitin ligase, negatively regulates abscisic acid and drought responses in Arabidopsis thaliana. Mol Plant 4:938–946
Liu W, Tai H, Li S, Gao W, Zhao M, Xie C, Li W (2014) bHLH122 is important for drought and osmotic stress resistance in Arabidopsis and in the repression of ABA catabolism. New Phytol 201:1192–1204
Mabuchi K, Maki H, Itaya T, Suzuki T, Nomoto M, Sakaoka S, Morikami A, Higashiyama T, Tada Y, Busch W, Tsukagoshi H (2018) MYB30 links ROS signaling, root cell elongation, and plant immune responses. Proc Natl Acad Sci U S A 115:E4710–E4719
Mao X, Chen S, Li A, Zhai C, Jing R (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in Arabidopsis. PLoS ONE 9:e84359
Mao H, Wang H, Liu S, Li Z, Yang X, Yan J, Li J, Tran LS, Qin F (2015) A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat Commun 6:8326
Mukeshimana G, Butare G, Cregan P, Blair MW, Kelly JD (2014) Identification of quantitative trait loci associated with drought tolerance in common bean using SNP markers. Crop Sci 54:923–938
Mun˜oz-Perea CG, Teran H, Allen RG, Wright JL, Westermann DT, Singh SP (2006) Selection for drought resistance in dry bean landraces and cultivars. Crop Sci 46:2111–2120
Ochoa I, Blair MW, Lynch J (2006) QTL analysis of adventitious root formation in common bean under contrasting phosphorus availability. Crop Sci 46:1609–1621
Ortiz-Masia D, Perez-Amador MA, Carbonell J, Marcote MJ (2007) Diverse stress signals activate the C1 subgroup MAP kinases of Arabidopsis. FEBS Lett 2581:1834–1840
Park MY, Kim SY (2014) The Arabidopsis J protein AtJ1 is essential for seedling growth, flowering time control and ABA response. Plant Cell Physiol 55:2152–2163
Park HY, Seok HY, Park BK, Kim SH, Goh CH, Lee BH, Lee CH, Moon YH (2008) Overexpression of Arabidopsis ZEP enhances tolerance to osmotic stress. Biochem Biophys Res Commun 375:80–85
Piao S, Ciais P, Huang Y, Shen Z, Peng S, Li J (2010) The impacts of climate change on water resources and agriculture in china. Nature 467:43–51
Podia V, Milioni D, Martzikou M, Haralampidis K (2018) The role of Arabidopsis thaliana RASD1 gene in ABA-dependent abiotic stress response. Plant Biol (Stuttg) 20:307–317
Priest DM, Ambrose SJ, Vaistij Fabián E, Elias L, Bowles DJ (2010) Use of the glucosyltransferase UGT71B6 to disturb abscisic acid homeostasis in Arabidopsis thaliana. Plant J 46:492–502
Pruthvi V, Narasimhan R, Nataraja KN (2014) Simultaneous expression of abiotic stress responsive transcription factors, AtDREB2A, AtHB7 and AtABF3 improves salinity and drought tolerance in peanut (Arachis hypogaea L.). PLoS One 9:e111152
Qin Y, Tian Y, Liu X (2015) A wheat salinity-induced WRKY transcription factor TaWRKY93 confers multiple abiotic stress tolerance in Arabidopsis thaliana. Biochem Biophys Res Commun 464:428–433
Ramegowda V, Gill US, Sivalingam PN, Gupta A, Gupta C, Govind G, Nataraja KN, Pereira A, Udayakumar M, Mysore KS, Senthil-Kumar M (2017) GBF3 transcription factor imparts drought tolerance in Arabidopsis thaliana. Sci Rep 7:9148
Rao IM (2001) Role of physiology in improving crop adaptation to abiotic stresses in the tropics: the case of common bean and tropical forages. In: Pessarakli M (ed) Handbook of plant and crop physiology. Marcel Dekker Inc, New York, pp 583–613
Ré DA, Capella M, Bonaventure G (2014) Arabidopsis AtHB7 and AtHB12 evolved divergently to fine tune processes associated with growth and responses to water stress. BMC Plant Biol 14:1–14
Remans R, Beebe S, Blair MW, Manrique G, Tovar E, Rao IM, Croonenborghs A, Torres-Gutierrez R, El-Howeity M, Michiels J, Vanderleyden J (2008) Physiological and genetic analyses of root responsiveness to auxin-producing plant growth-promoting bacteria in common bean (Phaseolus vulgaris L.). Plant Soil 302:149–161
Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J, Jenkins J, Shu S, Song Q, Chavarro C, Torres-Torres M, Geffroy V, Moghaddam SM, Gao D, Abernathy B, Barry K, Blair M, Brick MA, Chovatia M, Gepts P, Goodstein DM, Gonzales M, Hellsten U, Hyten DL, Jia G, Kelly JD, Kudrna D, Lee R, Richard MM, Miklas PN, Osorno JM, Rodrigues J, Thareau V, Urrea CA, Wang M, Yu Y, Zhang M, Wing RA, Cregan PB, Rokhsar DS, Jackson SA (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713
Schneider KA, Brothers ME, Kelly JD (1997) Marker-assisted selection to improve drought resistance in common bean. Crop Sci 37:51
Söderman E, Mattsson J, Engstrm P (1996) The Arabidopsis homeobox gene ATHB-7 is induced by water deficit and by abscisic acid. Plant J 10:375–381
Shin R, Burch AY, Huppert KA, Tiwari SB, Murphy AS, Guilfoyle TJ, Schachtman DP (2007) The Arabidopsis transcription factor MYB77 modulates auxin signal transduction. Plant Cell 19:2440–2453
Smucker AJM, Nunez-Barrios A, Ritchie JT (1991) Root dynamics in drying soil environments. Below-ground Ecol 1:1–5
Sponchiado BN, White JW, Castillo JA, Jones PG (1989) Root growth of four common bean cultivars in relation to drought tolerance in environments with contrasting soil types. Exp Agr 25:249–257
Strizhov N, Abrahám E, Okrész L, Blickling S, Zilberstein A, Schell J, Koncz C, Szabados L (1997) Differential expression of two P5CS genes controlling proline accumulation during salt-stress requires ABA and is regulated by ABA1, ABI1 and AXR2 in Arabidopsis. Plant J 12:557–569
Sun Z, Wang X, Liu Z, Gu Q, Zhang Y, Li Z, Ke H, Yang J, Wu J, Wu L (2017) Genome-wide association study discovered genetic variation and candidate genes of fibre quality traits in Gossypium hirsutum L. Plant Biotechnol J 15:982–996
Sura W, Kabza M, Karlowski WM, Bieluszewski T, Kus-Slowinska M, Pawełoszek Ł, Sadowski ŁJ, Ziolkowski PA (2017) Dual role of the histone variant H2A.Z in transcriptional regulation of stress-response genes. Plant Cell 29:791–807
Thung M, Rao IM (1999) Integrated management of abiotic stresses. In: Singh SP (ed) Common bean improvement in the twenty-first century. Kluwer, Dordrecht, pp 331–370
Trapp JJ, Urrea CA, Cregan PB, Miklas PN (2015) Quantitative trait loci for yield under multiple stress and drought conditions in a dry bean population. Crop Sci 55:1596
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci U S A 97:11632–11637
Wang R, Wu H, Zhang M, Ni Z, Sun Q (2013) Cloning, characterization and transgenic function analysis of wheat (Triticum aestivum L.) TaWRKY51 gene. JAB 21:1019–1027
Wang X, Jia N, Zhao C, Fang Y, Lv T, Zhou W, Sun Y, Li B (2014) Knockout of AtDjB1, a J-domain protein from Arabidopsis thaliana, alters plant responses to osmotic stress and abscisic acid. Physiol Plant 152:286–300
Wang X, Wang H, Liu S, Ferjani A, Li J, Yan J, Yang X, Qin F (2016) Genetic variation in ZmVPP1 contributes to drought tolerance in maize seedlings. Nat Genet 48:1233–1241
Wang Y, Wang Q, Liu M, Bo C, Cai R (2017a) Overexpression of a maize MYB48 gene confers drought tolerance in transgenic Arabidopsis plants. J Plant Biol 60:612–621
Wang N, Zhang W, Qin M, Li S, Qiao M, Liu Z, Xiang F (2017b) Drought tolerance conferred in soybean (Glycine max. L) by GmMYB84, a novel R2R3-MYB transcription factor. Plant Cell Physiol 58:1764–1776
Wen Z, Tan R, Zhang S, Collins P, Yuan J, Du W, Gu C, Ou S, Song Q, An Y (2018) Integrating GWAS and gene expression data for functional characterization of resistance to white mould in soya bean. Plant Biotechnol J 16:1825–1835
Wu J, Wang L, Li L, Wang S (2014) De novo assembly of the common bean transcriptome using short reads for the discovery of drought-responsive genes. PLoS ONE 9:e109262
Wu J, Wang L, Fu J, Chen J, Wei S, Zhang S, Zhang J, Tang Y, Chen M, Zhu J, Lei L, Geng Q, Liu C, Wu L, Li X, Wang Q, Wang Z, Xing S, Zhang H, Blair M, Wang S (2020) Resequencing of 683 common bean genotypes identifies yield component trait associations across a north-south cline. Nat Genet 52:118–125
Xi J, Qiu Y, Du L, Poovaiah BW (2012) Plant-specific trihelix transcription factor AtGT2l interacts with calcium/calmodulin and responds to cold and salt stresses. Plant Sci 185–186:274–280
Xing DH, Lai ZB, Zheng ZY, Vinod KM, Fan BF, Chen ZX (2008) Stress- and pathogen-induced Arabidopsis WRKY48 is a transcriptional activator that represses plant basal defense. Mol Plant 1:459–470
Xiong L, Lee H, Ishitani M, Zhu JK (2002) Regulation of osmotic stress-responsive gene expression by the LOS6/ABA1 locus in Arabidopsis. J Biol Chem 277:8588–8596
Xiong H, Li J, Liu P, Duan J, Zhao Y, Guo X, Li Y, Zhang H, Ali J, Li Z (2014) Overexpression of OsMYB48-1, a novel MYB-related transcription factor, enhances drought and salinity tolerance in rice. PLoS ONE 9:e92913
Yan X, Liao H, Beebe SE, Blair MW, Lynch JP (2004) QTL mapping of root hair and acid exudation traits and their relationship to phosphorus uptake in common bean. Plant Soil 265:17–29
Yang SD, Seo PJ, Yoon HK, Park CM (2011) The Arabidopsis NAC transcription factor VNI2 integrates abscisic acid signals into leaf senescence via the COR/RD genes. Plant Cell 23:2155–2168
Yordanov I, Velikova V, Tsonev T (2000) Plant responses to drought, acclimation, and stress tolerance. Photosyntheetica 38:171–186
Zhang J, Guo X, Li X, Xiang F, Zhou B, Yu D, Tang D, Liu X (2012) The genetic and physiological analysis of late-flowering phenotype of T-DNA insertion mutants of AtCAL1 and AtCAL2 in Arabidopsis. Mol Biol Rep 39:1527–1535
Zhao Y, Xing L, Wang X, Hou YJ, Gao J, Wang P, Duan CG, Zhu X, Zhu JK (2014) The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci Signal 7:ra53
Zhao K, Zhang D, Lv K, Zhang X, Cheng Z, Li R, Zhou B, Jiang T (2019) Functional characterization of poplar WRKY75 in salt and osmotic tolerance. Plant Sci 289:110259
Zheng Y, Schumaker KS, Guo Y (2012) Sumoylation of transcription factor MYB30 by the small ubiquitin-like modifier E3 ligase SIZ1 mediates abscisic acid response in Arabidopsis thaliana. Proc Natl Acad Sci U S A 109:12822–12827
Funding
This work was supported by grants from the National Key R & D Program of China (2018YFD1000700/2018YFD1000704), the Ministry of Agriculture of China [the earmarked fund for the China Agriculture Research System (CARS-08)], the Agricultural Science and Technology Innovation Program of CAAS.
Author information
Authors and Affiliations
Contributions
L W, YJ C performed QTL analysis and characterized candidate genes. L W, LF W, J W and SM W designed and conducted the field experiments and generated phenotype data. L W, YJ C and J W wrote and finalized the manuscript. SM W planned and organized this study, and co-wrote the manuscript. All authors contributed to and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Janila Pasupuleti.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, L., Chang, Y., Wang, L. et al. Genetic dissection of drought resistance based on root traits at the bud stage in common bean. Theor Appl Genet 134, 1047–1061 (2021). https://doi.org/10.1007/s00122-020-03750-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03750-6