Abstract
Key message
We demonstrated the effect of heading time genes on reproductive transition and yield components under an identical genetic background using CRISPR/Cas9 gene-editing technology, and we propose that the elite allele will provide a new breeding strategy for rice breeding in high-latitude regions.
Abstract
Heading date is a factor closely associated with grain yield in rice (Oryza sativa L). In recent decades, a number of genes responsible for heading time have been identified, the variation of which contributes to the expansion of the rice cultivation area. However, it is difficult to compare the phenotypic effects of these genes due to the different genetic backgrounds. In this study, we generated 14 heading time mutants using CRISPR/Cas9 gene-editing technology and marker-assisted selection with a japonica Sasanishiki wild-type (WT) genetic background. Photoperiod sensitivity, the relationship between days to heading (DTH), and yield components of mutants were investigated. We found that the yield increases with increases in DTH, but eventually plateaus at maximum and then began to decrease, whereas the biomass continued to increase. The mutants exerted distinctly different effects on DTH and yield components. The convergent double mutants had severe yield reduction compared with single mutants, even with a DTH that was similar to that of single mutants. We also found that an elite mutant of se14 achieved a yield equal to that of the WT, but with heading occurring 10 days earlier. A sequence analysis of 72 cultivars collected from the japonica cultivated zone shows that elite se14 mutants have not been applied to rice breeding. Our study demonstrates the effect of heading time genes on reproductive transition and yield components under an identical genetic background. These results may provide new insights into rice breeding using heading time mutants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Major advancements in crop domestication often come from the naturally occurring genetic variation in the universal florigen pathway (Park et al. 2014). In rice (Oryza sativa L.), heading (or flowering) time is closely associated with grain yield. The appropriate heading time guarantees successful reproduction and balances vegetative and reproductive growth. The genetic regulation of heading time is a gene network of an antagonistic regulation of accelerated pathways under short-day (SD) conditions and repressed pathways under long-day (LD) conditions (Komiya et al. 2009). The Early heading date 1 (Ehd1) gene functions as a signal integrator under both SD and LD conditions (Doi et al. 2004). Ehd1 enhances the expression of Heading date 3a (Hd3a) and Rice FT-like 1 (RFT1), two florigen-like genes in rice (Takahashi et al. 2009), and while edh1 mutant delays heading under both SD and LD conditions (Doi et al. 2004). The hd3a and rft1 double mutant did not undergo heading 200 days after sowing, which indicates the absence of other florigen-like genes in rice genome (Komiya et al. 2003). Delayed heading in rice under LD conditions can be caused by several factors that inhibit Ehd1. The Grain number, plant height and heading date 7 (Ghd7) gene, which encodes a CCT (CO, CO-like, and TIMING OF CAB1)-domain protein (Xue et al. 2008), and DTH8, which encodes the OsHAP3 subunit of a CCAAT-box binding protein (HAP complex) (Dai et al. 2012; Wei et al. 2010; Yan et al. 2011) suppresses the expression of Ehd1 in LD conditions. Photoperiod sensitivity-13 (Se13) encodes a phytochromobilin synthase involved in phytochrome activity, and the se13-deficient mutants were insensitive to day length changes owing to the lack of functional phytochromes (Saito et al. 2011; Yoshitake et al. 2015). Another phytochrome associated gene, Phytochrome B (PHYB), achieves a similar insensitivity when inactivated by a frame shift (Ishikawa et al. 2011). Heading date 1 (Hd1) is a gene regulator that delays and promotes heading under LD and SD conditions, respectively. Moreover, this regulation is independent of Ehd1 (Yano et al. 2000). The Ef7 gene is rice ortholog of Arabidopsis EARLY FLOWERING 3 (ELF3), which delayed heading under both SD and LD conditions. Ef7 negatively regulates the expression of Ghd7 (Saito et al. 2012; Yuan et al. 2009). Additionally, Se14, which encodes a JumonjiC (JmjC) domain-containing protein, regulated the expression of RFT1 through Ehd1 pathway (Yokoo et al. 2014).
Heading time genes have pleiotropic effects on yield components. Different combinations of Hd1 and Ehd1 displayed branch number diversity in panicles (Endo-Higashi and Izawa 2011). The deficient mutant of ghd7 and dth8 reduces the grain number per panicle (Wei et al. 2010; Xue et al. 2008). There are multiple changes in vegetative organs in transgenic plants bearing defects in mutated hd3a (Tamaki et al. 2007). We previously found a reduction in panicles in the se13 mutant, a decrease in setting rate in the hd1 mutant, and a decrease in grain number per panicle in ghd7 mutant (Xu et al. 2014). However, the genetic effects of these genes have not been investigated under an identical genetic background to determine how they affect yield.
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-associated (Cas) systems have been successfully used as an efficient genome editing tool in a numbers of species (Li et al. 2016; Ma et al. 2015; Shan et al. 2013). In the present study, we generated 14 single, double, and triple mutants using CRISPR/Cas9 gene-editing technology and marker-assisted selection (MAS). We accessed the heading time, photoperiod sensitivity, and yield components, revealing distinctly different mutant effects on yield components and heading time. We aimed to gain insight into heading time, with a goal to provide better information and germplasm for rice breeding.
Methods
CRISPR/Cas9 vector construction and plant transformation
The experiment was conducted under the genetic background of Japanese commercial japonica cultivar obtained from Kyoto University (Kyoto, Japan). We linked the codon optimized hSpCas9 (Cong et al. 2013) to the maize ubiquitin promoter (UBI) in an intermediate plasmid, and then inserted this expression cassette into a binary pCAMBIA1300 (Cambia, Australia) that contains the hygromycin B phosphotransferase (HPT) gene. Subsequently, we removed the original BsaI site in the pCAMBIA1300 backbone using a point mutation kit (Transgen, China). To produce the CRISPR/Cas9 binary vector pBGK032, we inserted a fragment containing an OsU6 promoter (Feng et al. 2013), a negative selection marker gene ccdB flanked by two BsaI sites, and a sgRNA derived from pX260 (Cong et al. 2013) into the vector using an in-fusion cloning kit (Takara, Japan). We used Escherichia coli strain DB3.1 to maintain this binary vector (Supplementary Fig. 1). The 23-bp targeting sequences (including PAM) were selected within the target genes, and their targeting specificity was confirmed using a BLAST search against the rice genome (http://blast.ncbi.nlm.nih.gov/Blast.cgi) (Hsu et al. 2013). The designed targeting sequences (Fig. 1) were synthesized and annealed to form the oligo adaptors. Vector pBGK032 was digested by BsaI and purified using a DNA purification kit (Tiangen, China). A ligation reaction (10 µL) containing 10 ng of the digested pBGK032 vector and 0.05 mM oligo adaptor was conducted and directly transformed into E. coli competent cells to produce CRISPR/Cas9 plasmids. The CRISPR/Cas9 plasmids were introduced into A. tumefaciens strain EHA105. Rice transformation was performed as previously described (Nishimura et al. 2006). Genomic DNA was extracted from the transformed plants, and primer pairs flanking the designed target site were used for PCR amplification. We sequenced the 200–500 bp PCR products directly using the Degenerate Sequence Decoding method (Ma et al. 2015).
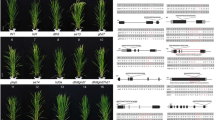
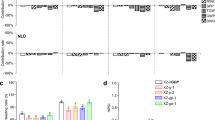
CRISPR/Cas9-induced single heading time-related mutants and their phenotype. The schematic map of the genomic region of the heading time genes, the sgRNA target sites and the sequence alignment between wild type (WT) and mutants is shown to the right of the plant image. Arrows indicate the emerged panicle. The images were taken when mutants or WT reached heading for each line. The scale bars are equal to 20 cm
Analysis of photoperiodic response
The wild type (WT) and mutants were used to evaluate the photoperiod response in the greenhouse. The LD condition was set as 14.5 h light and 9.5 h dark, and the SD condition was set as 9.5 h light and 14.5 h dark. Supplemental artificial light from incandescent lamps (3.24 Wm22 at soil surface) was used for the LD conditions. We transplanted at least five plants for each line, 14 days after sowing. The experiment was conducted on May 21, 2017, with two replicates. We recorded the heading time when the first panicle emerged from the sheath of the flag leaf for each line.
Field experiments
Field experiments were conducted in the experimental farm of Shenyang Agricultural University, Shenyang, China (N41°, E123°) in 2017. The seeds of 72 cultivars were obtained from the Academy of Agricultural Sciences for each province. Seeds were sown in the seedling nursery on 24 April, and one seedling was transplanted per hill on 23 May. The experiments were arranged in a randomized block design with three replicates. Each plot was 5.4 m2 and included 120 plants with planting densities (30 cm × 15 cm intervals). The cultivation method and field management were as described previously (Li et al. 2018).
DNA extraction, sequencing
We extracted the genomic DNA of 72 cultivars from fresh-frozen leaves using the CTAB method (Doyle 1991). The samples were sequenced on the Illumina HiSeq 2500 according to the manufacturer’s instructions, and the sequencing libraries were constructed. We aligned the sequencing reads to the japonica reference genome (http://rapdb.dna.affrc.go.jp/) using BWA software (Li and Durbin 2009). In total, 1638.72 Gb of clean data were generated across all 72 cultivars, with approximately 53-fold depth for each cultivar.
Data analysis
The genomic sequences and protein sequences were aligned using ClustalX 2.0.0, and these alignments were used as an input format into TASSEL V2.1. Nucleotide diversity and Tajima’s D statistics were calculated using the DnaSP 5.0 program. Statistical analysis was performed using STATISTICA software (StatSoft 1995). The evolutionary relationship among the haplotypes was inferred using the UPGMA method, and phylogenetic analyses were conducted using MEGA5 software.
Results
Heading time gene investigation of receipt for transformation
The japonica rice variety Sasanishiki (WT) was used as receipt for transformation. Sasanishiki is a typical Japanese variety which was derived from an ancestor variety ‘Gimbozu,’ and widely cultivated in Northeast of China before 1980s. Sasanishiki is a major backbone parent for rice breeding in China, and many currently utilized varieties in Northeast China share the genomic pedigree of Sasanishiki (Supplementary Fig. 2). Thus, Sasanishiki is an ideal receipt for transformation not only for function demonstration of heading time genes, but also for the genetic improvement in rice varieties in Northeast China. Ten core heading time genes of Sasanishiki were investigated (Table 1). The results revealed a similar genotype between Sasanishiki and Nipponbare. Only Hd1 was slightly different, harboring a 43 bp deletion at the first exon of Hd1 compared to Nipponbare (Supplementary Fig. 3).
Generation of heading time gene mutants using CRISPR/Cas9
To compare the effects of heading time genes on rice reproductive transition and yield components, we mutated ten core heading time genes (Table 1) using CRISPR/Cas9 to specifically induce mutagenesis. We generated the mutant of then heading time genes using CRISPR/Cas9 as described (Li et al. 2017). We examined the mutation efficiency in the T0 generation, and 20 plants of each mutant were sequenced. Homozygous mutations were determined using Sanger sequence results. Single peaks in Sanger sequence results were considered as homozygous mutation, while the double peaks represented the heterozygous mutations (Supplementary Fig. 4). On average, mutations occurred in 50% of plants, and 13% of sequenced plants had a putative homozygous mutation (Fig. 1 and Table 2). We obtained more than one type of mutation for each heading time gene, except Ehd1. For most genes, the different types of mutations exhibited similar phenotypes, except ghd7-2 and dth8-2. Combinations of the Ghd7, Ghd8/DTH8, and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice (Zhang et al. 2015), and hence, we crossed the hd1, ghd7, and dth8 mutants to generate double and triple mutant lines. We successfully selected hd1ghd7, ghd7dth8, hd1dth8 double mutants and the hd1ghd7dth8 triple mutant in the F2 population (Fig. 2). In total, 14 lines (10 single mutants, three double mutants, and one triple mutant) with homozygous mutations were used in this study. Forty plants per mutant were sown in the experimental field of Shenyang Agricultural University (N41°, E123°) on April 23, 2017. The heading time was recorded when the first panicle emerged from the sheath of the flag leaf. Ten early mutants underwent significantly earlier heading than the WT: phyb, se13, ghd7, dth8, se14, hd3a, hd1ghd7, ghd7dth8, hd1dth8, and hd1ghd7dth8. Three late ef7, ehd1, and rft1 mutants were significantly delayed in heading time compared with WT.
Characteristics of photoperiod response in mutants
Rice development before heading is divided into two successive phases: The basic vegetative growth phase, which is photoperiod insensitive, and the photoperiod sensitive phase (Yoshitake et al. 2015). The photoperiod sensitive phase is defined as the days to heading (DTH) under LD conditions minus the days to heading under SD conditions. To analyze the photoperiod response of mutants, we planted 14 mutants and the WT under SD and LD conditions. We found that the early mutants had similar basic vegetative growth phase compared to the WT, whereas the photoperiod sensitive phase was significantly shorter than that of the WT. In particular, the se13 and phyb mutants completely lost the photoperiod sensitive phase. The double mutant ghd7dth8 and triple hd1ghd7dth8 mutant had an additive effect in the regulation of the photoperiod sensitive phase. On the other hand, late mutants had similar photoperiod sensitive phase compared with the WT, but the basic vegetative growth phase was extended in all but the rft1 mutant. The rft1 mutant underwent heading at the same time as the WT under the SD condition, but had a significantly longer photoperiod sensitive phase than that of the WT. Under natural day length, SD and LD conditions, the hd1 mutant reached heading slightly earlier than the WT (Fig. 3). Interestingly, the hd3a mutant showed an earlier heading phenotype under LD and natural day length conditions.
The heading date of the WT and mutants under SD and LD conditions. The blue color indicates the heading date under SD conditions, the orange color indicates the photoperiod sensitive stage, and the blue color plus the orange color indicates the heading date under LD conditions. The black and gray colors indicate the WT variety
Yield-related traits of mutants
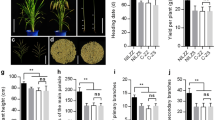
Numerous studies have demonstrated that heading time genes directly regulate yield components. Consequently, we conducted yield-related trait investigations of both mutants and the WT (Table 3). The results showed that the yield increased with an increase in the number of days to heading, but reaching a limit at 115 days to heading (Fig. 4). Early mutants had lower biomass and grain yield per plant; however, the sources of yield loss were different. Among the single mutants, phyb exhibited a decrease in the number of grains per panicle and the setting rate, se13 had fewer panicles, se14 had a smaller grain size, and ghd7 and dth8 showed a decrease in both panicle number and grain number per panicle. The double mutant dth8/ghd7 and triple mutant hd1/dth8/ghd7 had severe yield reduction compared with that of dth8 and ghd7 single mutants, and even had lower yield than phyb mutant and se13 single mutants. The hd3a mutant had reduced yield from decreases in panicle number and setting rate. Among the late mutants, the ef7 mutant increased grain yield, whereas rft1 significantly reduced yield by decreasing grain number per panicle. The ehd1 mutants had a higher grain number per panicle compared with the WT, however, a lower number of panicles placed similar to that of the WT.
Elite allele in mutants
Most of the improved varieties are insensitive to photoperiod by the introduction of the photoperiod-insensitivity gene and can be grown during any season and in most tropical and subtropical countries (Khush 2001). We investigated photoperiod-insensitivity mutants for potential uses in rice breeding. Among the photoperiod insensitive mutants, only se14 had a yield comparable to the WT, but underwent heading 10 days earlier than the WT under natural field conditions (Fig. 4). An expression analysis showed that Se14 was ubiquitous but was more highly expressed in the flag leaf, stem, and panicle 5 days after heading and spikelet production, but no significant difference between the WT and se14 mutant was observed (Supplementary Fig. 5). Our previous study identified Se14 from artificial γ radiation mutants (Yokoo et al. 2014); however, the natural variation of the Se14 locus remains unclear. We collected 72 cultivars that were widely released from 1960 to 2000 in the japonica cultivated area in China (Supplementary Table 1): 17 cultivars from Heilongjiang Province (HLJ), 19 cultivars from Liaoning Province (LN), 16 cultivars from Shandong Province (SD), and 20 cultivars from Jiangsu Province (JS) (Fig. 5). Then, we sequenced the Se14 genes for the 72 cultivars, revealing 11 SNPs and 5 INDELs (Fig. 5). Using this information, we constructed 3 haplotypes from the 72 cultivars; all of the cultivars in HLJ belonged to Hap1. There were 17 Hap1, 1 Hap2 and 1 Hap3 among LN cultivars; 9 Hap1 and 7 Hap2 occurred in SD; and 16 Hap1, 3 Hap2, and 1 Hap3 occurred in JS. We compared the heading time and yield of these cultivars in their cultivated area; however, no significant difference was found among haplotypes. As no frame shift occurs among haplotypes compared to the reference genome (Nipponbare and 9311), all three haplotypes likely carry a functional Se14 allele. In addition to Se14, we found that the plants carrying the dth8 mutant showed significant increases in the 1000-grain weight compared with that of the WT. However, the decrease in panicle number and grain number per panicle caused the yield reduction.
Discussion
CRISPR/Cas9 is an efficient target mutation method for many species (Cong et al. 2013; Feng et al. 2013; Ma et al. 2015; Shan et al. 2013; Zhou et al. 2018). In this study, we successfully mutated ten heading time genes in a single cultivar. This finding will facilitate the pyramiding of useful genes into a single cultivar for either breeding purposes or the dissection of gene regulatory networks. Novel phenotypes were characterized in this study, which differed from previous works. Hd1 functions in the promotion of heading under SD conditions and in inhibition under LD conditions (Yano et al. 2000). Sasanishiki carries a mutant hd1 allele which has a 43 bp deletion at the 3′ terminus of the first exon; our study generated a mutation at the 5′ end of Hd1. The hd1 gene-edited plant reached heading 5 days earlier than Sasanishiki. Thus, the Sasanishiki-type allele of Hd1 may have a weak function, while the CRISPR/Cas9 type Hd1 alleles is nonfunctional. Hd3a is a florigen of rice and promotes rice heading (Kojima et al. 2002). The present study showed a loss-of-function allele of hd3a as an earlier mutant under LD and natural day length conditions, and a later heading under SD conditions. These results indicate that Hd3a may mainly promote heading under SD conditions, and the other florigen of rice, RFT1, may primarily accelerate heading under LD conditions. However, further research is needed to dissect the molecular mechanisms behind earlier heading in the hd3a mutant under LD and natural day length conditions. The phytochrome-related mutants, se13 and phyb, completely lost their photoperiod sensitive phase and exhibited the same heading date under SD and LD conditions. The double mutant dth8/ghd7 has similar DTH as se13 and phyb mutants, indicating that ghd7 and dth8 contribute to the photoperiod sensitivity in Sasanishiki. Our previous study identified earlier heading phenotype se14 mutant as Os03g0151300 from mutant pool which was generated by γ radiation treatment. The se14 mutant had a 23-bp deletion at the first exon of Os03g0151300. The transcript experiments showed that Se14 contains cDNA of two gene locus Os03g0151300 and Os03g0151400 (Yokoo et al. 2014). Thus, the Se14 encodes a protein containing Jumonji N (JmjN), JmjC, and four copies of C2H2-type zinc finger (ZnF) domains, which share high homology with ELF6 in Arabidopsis. The present study confirmed the function of Se14 in heading time regulation. Moreover, we found that unlike other earlier mutant genes such as ghd7 and dth8, the earlier heading of the se14 mutant did not cause a decrease in yield.
Late heading is associated with a long growth period, which, in turn, is associated with larger biomass, which may guarantee higher yields. Our study confirms that an insufficient basic vegetative growth phase significantly decreases the yield, and all the early heading mutants had reduced yield (Fig. 4). However, the yield reached a maximum limit with an increase in the number of days to heading. After the yield peaked, the biomass continued to increase with increase in the number of days to heading, whereas the yield began to decrease (Table 2). The temperature of high-latitude areas drops dramatically after August; thus, late heading may cause a plant to suffer from low temperature stress at the pollination or filling stage, leading to a decrease in the setting rate and the 1000 grain weight.
The rice genome contains 20 Jumonji C (jmjC) domain encoding genes, and jmjC domain-containing proteins may function as histone demethylases (Tsukada et al. 2006). The defective jmj706 mutant caused to increased di- and trimethylations of H3K9, and the floral morphology was changed in mutant (Sun and Zhou 2008). The expression of JMJ705 is induced by stress signals and during pathogen infection (Li et al. 2013). A previous study demonstrated that Se14 encodes a jmjC domain-containing protein, and RFT1 and Ehd1 were up-regulated in the se14 mutant, which led to an early heading phenotype (Yokoo et al. 2014). In this study, we found the se14 mutant underwent heading 10 days earlier than the WT but had the same yield. This elite character will not only save resources in terms of labor, water and arable land but will also improve the adaptive ability of the variety. The sequence analysis of Se14 among the japonica cultivars shows that a nonfunctional Se14 allele may not yet have been used in rice breeding. However, the function of se14 needs to be tested under diverse genetic backgrounds to further elucidate its effects. Taken together, these results may provide new insights into rice breeding for the development of an early heading, high-yielding variety.
Conclusions
We analyzed the relationship between heading time and yield under an identical genetic background by using various heading time mutants developed by a CRISPR/Cas9 system. Heading time is often associated with yield-related traits and biomass, yet we found that yield increased with delayed time to heading, but reaches a maximal level before decreasing. An unexpected elite performance of the se14 mutant was a striking finding that could be applicable in rice breeding to generate early heading varieties with yield equal to standard varieties.
Author contribution statement
QX and ZX conceived the original screening and research plans; YC and MZ performed most of the experiments; QX conceived the project and wrote the article with contributions of all the authors; ZX supervised and complemented the writing. QX agrees to serve as the author responsible for contact and ensures communication. All authors have read and approved the manuscript.
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article and its additional files.
References
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 15:819–823
Dai X, Ding Y, Tan L, Fu Y, Liu F, Zhu Z, Sun X, Gu P, Cai H, Sun C (2012) LHD1, an allele of DTH8/Ghd8, controls late heading date in common wild rice (Oryza rufipogon). J Integr Plant Biol 54:790–799
Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A (2004) Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev 18:926–936
Doyle J (1991) DNA protocols for plants. In: Hewitt GM, Johnston AWB, Young JPW (eds) Molecular techniques in taxonomy. NATO ASI series (Series H: Cell Biology), vol 57. Springer, Berlin, Heidelberg
Endo-Higashi N, Izawa T (2011) Flowering time genes Heading date 1 and Early heading date 1 together control panicle development in rice. Plant Cell Physiol 52:1083–1094
Feng Z, Zhang B, Ding W, Liu X, Yang DL, Wei P, Cao F, Zhu S, Zhang F, Mao Y (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23:1229–1232
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31:827–832
Ishikawa R, Aoki M, Kurotani K, Yokoi S, Shinomura T, Takano M, Shimamoto K (2011) Phytochrome B regulates Heading date 1 (Hd1)-mediated expression of rice florigen Hd3a and critical day length in rice. Mol Genet Genom 285:461–470
Khush GS (2001) Green revolution: the way forward. Nat Rev Genet 2:815–822
Kojima S, Takahashi Y, Kobayashi Y, Monna L, Sasaki T, Araki T, Yano M (2002) Hd3a, a rice ortholog of the arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol 43:1096–1105
Komiya R, Ikegami A, Tamaki S et al (2003) Hd3a and RFT1 are essential for flowering in rice. Development 135:767–774
Komiya R, Yokoi S, Shimamoto K (2009) A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Development 136:3443–3450
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li T, Chen X, Zhong X, Zhao Y, Liu X, Zhou S, Cheng S, Zhou D-X (2013) Jumonji C domain protein JMJ705-mediated removal of histone H3 lysine 27 trimethylation is involved in defense-related gene activation in rice. Plant Cell 25:4725–4736
Li M, Li X, Zhou Z, Wu P, Fang M, Pan X, Lin Q, Luo W, Wu G, Li H (2016) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front Plant Sci 7:377
Li W, Zhu Z, Chern M, Yin J, Yang C, Ran L, Cheng M, He M, Wang K, Wang J (2017) A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170:114–126
Li X, Wu L, Wang J, Sun J, Xia X, Geng X, Wang X, Xu Z, Xu Q (2018) Genome sequencing of rice subspecies and genetic analysis of recombinant lines reveals regional yield- and quality-associated loci. BMC Biol 16:102
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284
Nishimura A, Aichi I, Matsuoka M (2006) A protocol for Agrobacterium-mediated transformation in rice. Nat Protoc 1:2796
Park SJ, Jiang K, Tal L, Yichie Y, Gar O, Zamir D, Eshed Y, Lippman ZB (2014) Optimization of crop productivity in tomato using induced mutations in the florigen pathway. Nat Genet 46:1337–1342
Saito H, Okumoto Y, Yoshitake Y, Inoue H, Yuan Q, Teraishi M, Tsukiyama T, Nishida H, Tanisaka T (2011) Complete loss of photoperiodic response in the rice mutant line X61 is caused by deficiency of phytochrome chromophore biosynthesis gene. Theor Appl Genet 122:109–118
Saito H, Ogiso-Tanaka E, Okumoto Y, Yoshitake Y, Izumi H, Yokoo T, Matsubara K, Hori K, Yano M, Inoue H, Tanisaka T (2012) Ef7 encodes an ELF3-like protein and promotes rice flowering by negatively regulating the floral repressor gene Ghd7 under both short- and long-day conditions. Plant Cell Physiol 53:717–728
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31:686–688
Sun Q, Zhou D-X (2008) Rice jmjC domain-containing gene JMJ706 encodes H3K9 demethylase required for floral organ development. Proc Natl Acad Sci USA 105:13679–13684
Takahashi Y, Teshima KM, Yokoi S, Innan H, Shimamoto K (2009) Variations in Hd1 proteins, Hd3a promoters, and Ehd1 expression levels contribute to diversity of flowering time in cultivated rice. Proc Natl Acad Sci 106:4555–4560
Tamaki S, Matsuo S, Wong HL, Yokoi S, Shimamoto K (2007) Hd3a protein is a mobile flowering signal in rice. Science 316:1033
Tsukada Y, Fang J, Erdjumentbromage H, Warren ME, Borchers CH, Tempst P, Zhang Y (2006) Histone demethylation by a family of JmjC domain-containing proteins. Nature 439:811–816
Wei X, Xu J, Guo H, Jiang L, Chen S, Yu C, Zhou Z, Hu P, Zhai H, Wan J (2010) DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol 153:1747–1758
Xu Q, Saito H, Hirose I, Katsura K, Yoshitake Y, Yokoo T, Tsukiyama T, Teraishi M, Tanisaka T, Okumoto Y (2014) The effects of the photoperiod-insensitive alleles, se13, hd1 and ghd7, on yield components in rice. Mol Breed 33:813–819
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X, Zhang Q (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yan W, Wang P, Chen H, Zhou H, Li Q, Wang C, Ding Z, Zhang YS, Yu SB, Xing YZ, Zhang QF (2011) A major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height, and heading date in rice. Mol Plant 4:319–330
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2484
Yokoo T, Saito H, Yoshitake Y, Xu Q, Asami T, Tsukiyama T, Teraishi M, Okumoto Y, Tanisaka T (2014) Se14, encoding a JmjC domain-containing protein, plays key roles in long-day suppression of rice flowering through the demethylation of H3K4me3 of RFT1. PLoS ONE 9:e96064
Yoshitake Y, Yokoo T, Saito H, Tsukiyama T, Quan X, Zikihara K, Katsura H, Tokutomi S, Aboshi T, Mori N (2015) The effects of phytochrome-mediated light signals on the developmental acquisition of photoperiod sensitivity in rice. Sci Rep 5:7709
Yuan Q, Saito H, Okumoto Y, Inoue H, Nishida H, Tsukiyama T, Teraishi M, Tanisaka T (2009) Identification of a novel gene ef7 conferring an extremely long basic vegetative growth phase in rice. Theor Appl Genet 119:675–684
Zhang J, Zhou X, Yan W, Zhang Z, Lu L, Han Z, Zhao H, Liu H, Song P, Hu Y (2015) Combinations of the Ghd7, Ghd8 and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice. New Phytol 208:1056–1066
Zhou X, Jiang G, Yang L, Qiu L, He P, Nong C, Wang Y, He Y, Xing Y (2018) Gene diagnosis and targeted breeding for blast-resistant Kongyu 131 without changing regional adaptability. J Genet Genom 45:539–547
Funding
The National Natural Science Foundation of China (31501284) supported this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Albrecht E. Melchinger.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cui, Y., Zhu, M., Xu, Z. et al. Assessment of the effect of ten heading time genes on reproductive transition and yield components in rice using a CRISPR/Cas9 system. Theor Appl Genet 132, 1887–1896 (2019). https://doi.org/10.1007/s00122-019-03324-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03324-1