Abstract
Key message
New strategy of breeding by modulating key heading date gene Ehd1 to enhance the variations of heading date regardless of genetic background for better adaptation to local environment in rice.
Abstract
Flowering time (or heading date) is an important quantitative trait in rice (Oryza sativa) that determines its adaptation to specific cultivation areas and growing seasons. However, breeding of flowering time is currently relying on laborious selections and combinations of different alleles of various genes. Here, we cloned a cis-variant allele of Ehd1 that regulated not only heading date but also yield potential. Genetic analysis revealed that Ehd1 acted downstream of Ghd7 as a negative regulator of yield potential, and expression divergence of Ehd1 negatively correlates with phenotype variations including heading date and grain yield. Moreover, regardless of genetic background, manipulations of the expression of a single gene, Ehd1, are sufficient for recreating beneficial heading date variations which could be subjected to the selection of best suitable lines for local environment conditions. Beyond a deeper understanding of transcriptional control of quantitative traits, this study provided an effective and flexible strategy for breeding rice cultivars to maximize grain production for any region of cultivation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Heading date determines the regional and seasonal adaptation of rice, and is closely related to the grain yield. In the last few decades, a complicated heading date regulatory network has been built in the rice genome (Hori et al. 2016). Early heading date 1 (Ehd1), a B-type response regulator, acts as a flowering time activator by inducing the florigen genes Heading date 3a (Hd3a) and Rice Flowering Locus T 1 (RFT1) in both long-day (LD) and short-day (SD) conditions (Doi et al. 2004; Zhao et al. 2015). And Ehd1 protein forms homomer to induce flowering time, which is inhibited by rice Response Regulator 1 (OsRR1) through binding to Ehd1 and forming a heterodimer (Cho et al. 2016). Grain number, plant height and heading date 7 (Ghd7) encoding a CCT (CONSTANS (CO), CO-like (COL), TIMING OF CAB EXPRESSION 1) domain-containing protein strongly suppresses flowering time by inhibiting the expression of Ehd1 under LD conditions (Itoh et al. 2010; Xue et al. 2008). Notably, many other genes, such as Ghd7.1/Days to heading 7 (DTH7)/Oryza sativa Pseudo-Response Regulator37 (OsPRR37) (Gao et al. 2014; Koo et al. 2013; Yan et al. 2014), Ghd8/DTH8 (Wei et al. 2010; Yan et al. 2011), Ghd2 (Liu et al. 2016b), rice CONSTANS-like 4 (OsCOL4) (Lee et al. 2010), OsCOL9 (Liu et al. 2016a), OsCOL10 (Tan et al. 2016), OsCOL13 (Sheng et al. 2016) and Ehd4 (Gao et al. 2013), also act upstream of Ehd1, either as inducers or suppressors. Ehd1 was first reported to work in parallel with Heading date 1 (Hd1) (Doi et al. 2004), but the latest results have revealed that Hd1 can also act as a repressor of Ehd1 by biologically interacting with Ghd7 (Nemoto et al. 2016; Zhang et al. 2017). Therefore, Ehd1 likely functions as a central signal integrator of floral transition in rice (Shrestha et al. 2014). Notably, most Ehd1 regulators have pleiotropic effects on not only heading date but also grain yield and plant height. For example, functional alleles of Ghd7 (Xue et al. 2008), Ghd8/DTH8 (Wei et al. 2010; Yan et al. 2011) and Ghd7.1/DTH7 (Gao et al. 2014; Yan et al. 2014) as well as overexpression of Ghd2 (Liu et al. 2016b), OsCOL9 (Liu et al. 2016a) and OsCOL10 (Tan et al. 2016) contribute to increased grain yield. Moreover, Ehd4 mutation delays heading date and increases plant height and the number of grains per panicle (Gao et al. 2013). However, whether these genes also regulate yield potential through Ehd1 remains unknown.
Rice originates from southern China (Huang et al. 2012) and has been domesticated and disseminated to tropical, subtropical and temperate regions (Gómez-Ariza et al. 2015; Zhang et al. 2015). During this process, domestication of the heading date has been crucial because day length gradually lengthens with increasing latitude, so local varieties must have an optimal heading date to complete the life cycle before the unfavorable seasons.
Recently, molecular evidence for rice heading date domestication was revealed via the re-sequencing of hundreds of varieties cultivated worldwide (Gómez-Ariza et al. 2015; Zhang et al. 2015). Abundant natural variation in key heading date regulators enables different varieties to be bred with distinct heading dates. Lost or attenuated function of LD suppressors, such as Ghd7, Ghd8, Ghd7.1 and Hd1, results in an early heading date (Gómez-Ariza et al. 2015; Zhang et al. 2015), and combinations of different alleles of these suppressors along with heading date inducers as Ehd1 and RFT1 in different varieties greatly contributed to the expansion of rice cultivars to higher-latitude regions (Gao et al. 2014; Takahashi et al. 2009; Yan et al. 2014; Zhang et al. 2015; Zhao et al. 2015). However, how to efficiently select and combine these alleles remain a challenge in breeding. Additionally, traditional approaches as well as the marker-assisted selection (MAS) method are time-consuming. Thus, there is an urgent need for a more efficient and predictable method of developing cultivars with an expected heading date by manipulation of known important genes.
In this study, we cloned a cis-variant allele of Ehd1, and experimentally revealed that Ehd1 acted downstream of Ghd7 to regulate several agronomic traits including heading date, yield potential and plant height. We generated series of Ehd1-knockdown transgenic lines in two japonica varieties by RNA interference strategy, and demonstrated that beneficial heading date and yield variations could be artificially induced and selected for local environmental conditions. Moreover, we proposed an effective and flexible strategy for breeding rice cultivars to maximize grain production for any region of cultivation regardless of genetic background.
Materials and methods
Plant materials
A quantitative trait locus, qEhd10 (Earlyheadingdate 10) was previously mapped on chromosome 10 using a recombinant inbred line population derived from the cross between Zhenshan 97 (ZZ) and Zhongzao 18 (ZZ) (Kovi et al. 2015). Following a trait-performance-derived nearly isogenic line (NIL) strategy (Zhang et al. 2006), an F7 inbred line carrying a heterozygous fragment of qEhd10 was self-crossed to develop NILs (NIL-ZZ and NIL-ZS) (Fig. S1a). An F2 population of 4800 plants deriving from the cross between NIL-ZZ and NIL-ZS was used to fine map qEhd10 (Fig. S1a).
To define the relationship between Ghd7 and Ehd1, we screened a ghd7 mutant with a G to A mutation resulting in a premature stop codon (Figs. S1b and S2) in the M2 generation of an ethyl methanesulfonate-treated japonica rice cultivar, Zhonghua 11 (ZH11, Oryza sativa L. ssp. japonica). For further analysis of the effects of Ehd1 on both heading date and yield, a series of materials were generated with transgenic method in the backgrounds of ZH11, ghd7 mutant or Nipponbare (Figs. S1b, S3). In ZH11 and ghd7 mutant backgrounds, Early heading date 1 (Ehd1) was knocking out with clustered regularly interspaced short palindromic repeats (CRISPR) strategy, which generated ehd1 single-mutant Ehd1-CR and ghd7 and ehd1 double-mutant ghd7/Ehd1-CR (Fig. S1b). Transgenic line overexpressing of Ghd7 in ZH11 background (Ghd7-ox) was previously described (Weng et al. 2014). Overexpression of Ehd1 in ZH11 and Ghd7 overexpression line backgrounds resulted in Ehd1 overexpression line (Ehd1-ox) and Ghd7 Ehd1 double overexpression line (Ghd7-ox/Ehd1-ox) (Fig. S1b).
Phenotype investigation
Heading dates under each condition were recorded as the number of days from germination to the emergence of the first panicle. Plant height was measured from the ground to the top of the tallest tiller of the plant before harvesting. Grains were harvested individually within each line and dried under sunlight for 5 days. Then the yield and yield component traits, including spikelets per panicle, spikelets on the main panicle, panicle length, number of primary branches and number of secondary branches, were investigated individually. Among these traits, panicle length, number of primary branches and number of secondary branches were scored with the mean of the three longest panicles of each individual (Fig. S3).
Vector construction and transformation
To construct the complementary vectors, the 2220-bp upstream regulatory sequence of Ehd1 from NIL-ZS was amplified with primers E1301-pro-F and E1301-pro-R (Table S1) and then introduced in pCAMBIA 1301 at the SmaI site, which resulted in an intermediate vector. Thereafter, the coding sequences of NIL-ZZ and NIL-ZS amplified with primers E1301-CDS-F and E1301-CDS-R were inserted at downstream of 2220-bp upstream regulatory sequence of Ehd1 in the intermediate vector (Table S1), resulting in two constructs, C-ZZ and C-ZS, respectively.
To construct the Ehd1 overexpression vector, the Ehd1 coding sequence from NIL-ZS was amplified using the primers E2301-UF and E2301-UR (Table S1), and then cloned into a KpnI-linearized pU2301. This construct was transformed into ZH11 and Ghd7-ox plants resulting in Ehd1 overexpression (Ehd1-ox) line and Ghd7 and Ehd1 double overexpression (Ghd7-ox/Ehd1-ox) line, respectively.
To construct the Ehd1 RNA interference (Ri) vector, 202 bp from the Ehd1 coding sequence of NIL-ZS was amplified with primers ERI-F (containing SpeI and KpnI digestion sites) and ERI-R (containing SacI and BamHI digestion sites), then cloned into a pGEM-T vector (Promega). The fragments were then excised with KpnI and BamHI, as well as SacI and SpeI, respectively, and were cloned into a pDS1301 vector at the corresponding sites (Fig. S4). With this construct, we developed RNA interference lines in ghd7 mutant (ghd7/Ehd1-Ri) and Nipponbare (Nip-Ehd1-Ri) (Fig. S3).
To construct the CRISPR-Cas9 vector for Ehd1, the target sequence was designed online [http://crispr.hzau.edu.cn/CRISPR2/ (Lei et al. 2014)] and fused in the ECR-F and ECR-R primers (Table S1). With a segment-overlapping PCR followed by a Gibson assembly reaction (Gibson et al. 2009), the sgRNA scaffold (containing target sequence) driving by rice U3 promoter sequence was cloned into a pCXUN-Cas9 vector (Sun et al. 2016). All these vectors (except the pGEM-T vector) were constructed with Gibson assembly reaction (Gibson et al. 2009). These vectors were induced into corresponding acceptor or specific lines with Agrobacterium (EH105)-mediated transformation (Hiei et al. 1994).
Plant growth conditions
The rice plants examined under natural field conditions were grown in Wuhan (Huazhong Agricultural University, 114°21′E, 30°28′N) and Hainan Island (Lingshui County, 110°01′E, 18°30′N), China. In Wuhan, natural long-day (NLD) conditions were from mid-May to August (more than 13.5 h), and natural day length (ND) conditions were from late June to late September (declining day length from approximately 14 to 12 h). In Hainan, natural short-day (NSD) conditions were from early December to late April (day length was about approximately 12 h). The NIL plants used to analyze flowering time genes were grown in controlled environment chambers under long-day (LD, 14-h light/10-h dark) and short-day (SD, 10-h light/14-h dark) conditions. The Nip-Ehd1-Ri plants were grown in a greenhouse under artificial LD (10-h light/14-h dark) conditions.
RNA sampling and gene expression analysis
For the NILs in controlled LD and SD conditions, leaves from 35-day-old plants that undergo photoperiodic responses were sampled every 4 h within a 24-h period, and three different individuals were used as biological replicates. For the ghd7/Ehd1-Ri lines and Nip-Ehd1-Ri plants, leaves were sampled at the corresponding times in the field under ND conditions and in the greenhouse under artificial LD conditions, respectively (Fig. S3). Total RNA was isolated with TRIzol reagent (TransGen Biotech). For reverse transcription quantitative PCR (RT-qPCR), first-strand cDNA was synthesized using reverse transcriptase (Invitrogen), and qPCR was then performed using gene-specific primers, SYBR Master Mix reagent (Roche), and a Quant-Studio 6 Flex Real-Time PCR System (Life Science), according to the manufacturer’s instructions. The PCR conditions were as follows: 10 min at 95 °C followed by 40 cycles of 10 s at 95 °C and 30 s at 60 °C. PCR amplifications were conducted in triplicate for each sample from three independent biological replicates, and a rice ubiquitin gene (Os02g0161900) was used for normalization. To quantify the expression of Hd1, Ehd1, Hd3a and RFT1, we used the specific primers listed in Table S1.
Statistical analysis
Statistical analyses were performed using Microsoft Excel 2010 or GraphPad Prism 6. The statistical differences in phenotypic values between NIL-ZZ and NIL-ZS were examined by the two-tailed Student’s t test. Correlation between expression levels of flowering time gene and agronomic traits in core collection varieties, ghd7/Ehd1-Ri lines and Nip-Ehd1-Ri plants were examined by Pearson’s correlation coefficient test.
Results
Pleiotropic effects of qEhd10
A major quantitative locus Earlyheadingdate 10 (qEhd10) was mapped on chromosome 10 previously (Kovi et al. 2015). In this study, nearly isogenic lines (NILs) of qEhd10 were developed. Compared with NIL-ZS, NIL-ZZ delayed heading by 12.2 d (19.9%), increased plant height by 9.3 cm (11.8%) and produced 5.6 g (29%) more grains per plant under natural long-day (NLD) conditions in Wuhan (Fig. 1a–g). Additionally, NIL-ZZ primarily improved the yield by increasing the number of spikelets through the production of more primary and secondary branches (Fig. 1h–j).
Phenotypes of the nearly isogenic lines (NIL-ZZ and NIL-ZS) and complementary plants. a Phenotypes of the NIL-ZZ (left) and NIL-ZS (right) plants grown in Wuhan (114°21′E, 30°28′N) under natural long-day (NLD, from mid-May to August) conditions when NIL-ZS reached maturity. b The main culms of NIL-ZZ (left) and NIL-ZS (right); hollow arrows indicate the nodes of the culms. c The main panicles of NIL-ZZ (left) and NIL-ZS (right). d Grains from whole plants of NIL-ZZ (left) and NIL-ZS (right). e–j Agronomic traits of the NIL-ZZ, NIL-ZS and complementary plants (lines transformed with upstream regulation sequence form NIL-ZS fused with coding sequence from NIL-ZZ (C-ZZ) and that from NIL-ZS (C-ZS)), under NLD conditions. Heading date and plant height, NIL-ZZ, n = 39; NIL-ZS, n = 39; C-ZZ, n = 25; C-ZS, n = 17. Yield per plant, spikelets on the main panicle, number of primary branches and number of secondary branches, NIL-ZZ, n = 21; NIL-ZS, n = 19; C-ZZ, n = 13; C-ZS, n = 13. Data represent the mean ± standard deviation (s.d.); **p < 0.01, two-tailed Student’s t tests. Scale bar, 25 cm in a and b; 5 cm in c and d
Differential expression of Ehd1 affecting heading date and grain yield
To isolate qEhd10, heading date was investigated in an NIL-F2 population of 4800 individuals (Fig. S1a), from which 672 plants with extremely late heading date were chosen for qEhd10 fine mapping. Then, qEhd10 was narrowed down to a 101-kb genomic region between the markers Indel-2 and RM25527, co-segregating with marker E1-IN (Fig. S5a). Fourteen open reading frames (ORFs) were predicted in this region (Fig. S5b), among which ORF7 was the previously cloned gene Ehd1 (Doi et al. 2004). Comparative sequencing identified five single-nucleotide polymorphisms (SNPs) in the coding sequence of Ehd1 between NIL-ZZ and NIL-ZS (Fig. S5c), and 19 SNPs and three indels were found in the 2220-bp upstream regulatory sequence from the initiation codon ATG (Table S2). To demonstrate the contribution of Ehd1 to the phenotypic change, a fragment derived from NIL-ZS containing the 2220-bp upstream regulatory sequence was fused with the coding region sequence of NIL-ZZ (C-ZZ) and NIL-ZS (C-ZS) and transformed to NIL-ZZ. Both C-ZZ and C-ZS restored the heading date, plant height, yield per plant and yield component phenotypes in NIL-ZZ (Fig. 1e–j). These results indicated that the difference in the coding region was not the causal variation and that the difference in Ehd1 expression was more important. Indeed, the expression level of Ehd1 in NIL-ZS was significantly higher than that of NIL-ZZ under both LD and SD conditions (Fig. S6a, d). The florigen gene Hd3a showed similar expression patterns to that of Ehd1 (Fig. S6c, f), but the level of Hd1 expression showed no differences under both conditions (Fig. S3b, e). These results suggested that differential expression of Ehd1 affected heading date and yield in the background of NILs.
Ehd1 acts downstream of Ghd7
We previously reported that Ghd7 delayed flowering time by repressing Ehd1 expression under LD conditions (Xue et al. 2008), but it is not clear whether Ghd7 also contributes to yield potential through the Ehd1 pathway. To study the relationship between these genes, a series of transgenic lines were generated (Fig. S1b). Ghd7 and Ehd1 double overexpression line (Ghd7-ox/Ehd1-ox) showed similar agronomic phenotype performances to Ehd1 overexpression line (Ehd1-ox) including heading date, yield per plant, plant height and other yield component traits (Fig. 2a–j). Furthermore, a ghd7 mutant, ehd1 single mutant (Ehd1-CR) and ehd1 ghd7 double mutant (ghd7/Ehd1-CR) were generated and used for further investigation (Figs. S1b, S2 and S7). Ehd1-CR showed strongly delayed heading and increased plant height and yield per plant (Fig. 2a–j), and the double-mutant ghd7/Ehd1-CR exhibited a phenotype similar to that of the single-mutant Ehd1-CR (Fig. 2a–j). Collectively, these results clearly supported Ghd7 regulating heading date as well as yield through Ehd1 suppression.
Ehd1 acts downstream of Ghd7 and has pleiotropic effects on an array of traits. a, b Phenotypes of the whole plants (a) and main panicles (b) of Ghd7 and Ehd1 double overexpression line (Ghd7-ox/Ehd1-ox), Ehd1 overexpression line (Ehd1-ox), ghd7 mutant, ZH11 wild type, Ghd7 overexpression line (Ghd7-ox), Ehd1 knocking out lines in ZH11 (Ehd1-CR) and ghd7 mutant (ghd7/Ehd1-CR) backgrounds, plants grown in Wuhan under NLD conditions; host variety: cv. ZH11 (Oryza sativa L. ssp. japonica). c Expression levels of Ehd1 at dawn in Ghd7-ox/Ehd1-ox, Ehd1-ox, ghd7, ZH11, Ghd7-ox, Ehd1-CR and ghd7/Ehd1-CR plants grown under NLD conditions; data represent the mean ± S.D. of three replicates; n.d., not determined. Scale bar, 25 cm in a and 5 cm in b. d–j Agronomic traits of Ghd7-ox/Ehd1-ox, Ehd1-ox, ghd7, ZH11, Ghd7-ox, Ehd1-CR and ghd7/Ehd1-CR plants grown under NLD conditions. Heading date and plant height, Ghd7-ox/Ehd1-ox, n = 7; Ehd1-ox, n = 6; ghd7, n = 30; ZH11, n = 29; Ghd7-ox, n = 26; Ehd1-CR, n = 27; ghd7/Ehd1-CR, n = 20. Yield per plant, plant height, spikelets on the main panicle, panicle length, number of primary branches and number of secondary branches, Ghd7-ox/Ehd1-ox, n = 7; Ehd1-ox, n = 6; ghd7, n = 12; ZH11, n = 10; Ghd7-ox, n = 14; Ehd1-CR, n = 6; ghd7/Ehd1-CR, n = 7. Data represent the mean ± S.D. k–m Correlations between Ehd1 expression level and heading date (k), yield per plant (l) and plant height (m) in the 45 varieties core germplasm collection under NLD conditions. Ehd1 levels at dawn in leaves of 35-day-old (when rice plants undergo photoperiodic responses) plants under NLD conditions were determined by quantitative real-time PCR (qRT-PCR) and are shown as natural logarithms. R indicates the Pearson’s correlation coefficient; ***p < 0.001, **p < 0.01
Negative correlations between Ehd1 expression level and heading date as well as grain yield
To obtain further insights into the role of Ehd1 in the control of heading date and yield, materials generated in the ZH11 background were subjected to a deep phenotypic characterization. Negative correlations were detected between Ehd1 expression level and several agronomic traits including heading date, grain yield and panicle architecture (Fig. 2c–j). Compared with the wild-type ZH11, the Ehd1 expression in the Ehd1-ox, Ghd7-ox/Ehd1-ox and ghd7 lines was significantly promoted, with the highest expression level in Ehd1-ox followed by Ghd7-ox/Ehd1-ox. Accordingly, Ehd1-ox plants had the earliest heading date, shortest panicle length, least spikelets on the main panicle and primary and secondary branches, and ultimately the lowest grain yield per plant (Fig. 2c–j). In contrast, Ghd7-ox exhibited reduced Ehd1 expression and thus performed oppositely, namely delayed heading date and enhanced grain yield (Fig. 2c–j).
To further confirm the negative correlations, 45 varieties from a core germplasm collection were investigated under NLD conditions (Table S3). As expected, the expression level of Ehd1 showed significant negative correlations with heading date and yield but not with plant height (Fig. 2k–m). Similar results were detected for Hd3a but not RFT1 (Fig. S8a–r).
Enhancing heading date variation in ghd7 mutant via manipulating Ehd1 expression
According to the negative correlations between Ehd1 expression level and heading date and grain yield, we hypothesized that beneficial variations of heading date and yield could be artificially induced by simply modulating Ehd1 expression. To test this hypothesis, 23 Ehd1 suppression lines in the ghd7 background (ghd7/Ehd1-Ri) were generated with RNA interference strategy (Fig. 3a, b, Fig. S2). Under natural day length conditions (ND, day length decreasing from approximately 14 to 12 h) from late June to late September in Wuhan, the expression levels of Ehd1 and two florigen genes, Hd3a and RFT1, were investigated at 5 a.m., when the expression levels reached their peaks (Fig. S9). Differential suppression of the Ehd1 expression level in distinct transgenic lines was detected and accompanied by gradually delayed heading date with a wide distribution from 52.5 to 90.9 days (Fig. 3c, d), and the average yield accordingly gradually increased from 10 to 32.8 g per plant (Fig. 3e). In general, lines with lower Ehd1 expression had more delayed heading dates and increased grain yield. In detail, an approximately two-fold decrease in Ehd1 expression level in line #21 extended the heading date by approximately 10 days and doubled the yield per plant compared with those in mutant ghd7 (Fig. 3d, e). Lines, such as #3 and #18, with extreme Ehd1 suppression by approximately 3.7-fold resembled the phenotypes of the Ghd7-ox, Ehd1-CR and ghd7/Ehd1-CR lines, which showed a delay in heading date of more than one month and a trebling of yield per plant compared with ghd7 (Fig. 3d, e). Furthermore, the relationship between agronomic traits and heading date genes was analyzed within all the ghd7/Ehd1-Ri plants. As expected, the heading date as well as the yield per plant, plant height and yield components all showed significantly negative correlations with the level of Ehd1 expression (Fig. 4a–h, Fig. S10a–p). Notably, heading date was significantly and positively correlated with yield per plant, indicating the possibility of improving yield by extending the heading date (Table S4).
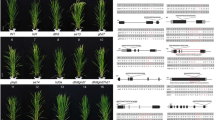
Phenotypes of Ehd1 RNA interference (Ri) T2 lines in the ghd7 mutant background (ghd7/Ehd1-Ri). a, b Phenotypes of the whole plants (a) and main panicles (b) of ghd7 and ghd7/Ehd1-Ri transgenic lines #9, #21, #23, #4, and #3 grown in Wuhan under natural day (ND, from late June to late September) length conditions. Scale bar, 25 cm in (a) and 10 cm in (b). c–e The distribution of Ehd1 expression level (c), heading date (d) and yield per plant (e) of ghd7, ZH11, ghd7/Ehd1-CR, Ehd1-CR, Ghd7-ox and all the ghd7/Ehd1-Ri transgenic lines, #3, #4, #12, #13, #15, #17, #18, #20, #21, n = 5; #6, #8, #9, #23, ghd7/Ehd1-CR#1, ghd7/Ehd1-CR#2, Ehd1-CR#2, n = 6; #2, #22, n = 7; #5, #7, #11, #16, #19, Ehd1-CR#1, n = 8; #10, #14, #20, Ghd7-ox, n = 10; ghd7, n = 15. Boxes represent interquartile ranges, and the middle line indicates the median. cEhd1 levels at 5 a.m. in the leaves of 35-day-old plants under ND conditions were determined by qRT-PCR and are shown as natural logarithms; n.d., not determined. d Heading dates are distributed in sequentially increasing order. e Yield per plant of each line is distributed according to the order of the lines in d
Ehd1 expression level is negatively correlated with multi-phenotype within ghd7/Ehd1-Ri lines. a–h Correlation of Ehd1 expression level of ghd7/Ehd1-Ri T2 plants with heading date (a), plant height (b), grain yield per plant (c), spikelets per panicle (d), panicle length (e), spikelets on the main panicle (f), number of primary branches (g), and number of secondary branches (h). Ehd1 levels in leaves of 35-day-old plants grown under ND conditions were determined by qRT-PCR and are shown as natural logarithms. R indicates Pearson’s correlation coefficient; ***p < 0.001
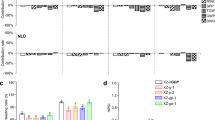
To test the performance stability of the ghd7/Ehd1-Ri lines, we compared the phenotypes of four ghd7/Ehd1-Ri T2 lines that covered the range of heading date variation in Wuhan (ND), a subtropical cultivating region, and in Hainan (natural short-day (NSD)), a tropical cultivating region (Fig. 5a–f). Compared with the control ghd7 mutant, the transgenic lines showed stably extended heading date and increased yield, plant height and yield components under both conditions. However, the corresponding phenotype changes were smaller under NSD conditions than those under ND conditions (Fig. 5a–f).
Comparison of phenotype performances of ghd7/Ehd1-Ri lines under different growth conditions. a–f Comparison of performances of heading date (a), plant height (b), panicle length (c), spikelets on the main panicle (d), number of the primary branches (e), and number of the secondary branches (f) of ghd7, Ghd7-ox and ghd7/Ehd1-Ri lines #2, #17, #15 and #18 between ND conditions in Wuhan and natural short-day (NSD) conditions in Lingshui County, Hainan (110°01′E, 18°30′N)
Enhancing heading date variation in Nipponbare via manipulating Ehd1 expression
To demonstrate the generality of this strategy, we interfered with Ehd1 in another rice variety, Nipponbare (Oryza sativa L. ssp. japonica), that possessed functional Ghd7 and Ehd1 alleles (Doi et al. 2004; Xue et al. 2008) (Fig. 6a). As in the ghd7 background, transgenic T0 plants with differential Ehd1 expression also displayed a range of heading dates but with a smaller span (Fig. 6b, c), and significant correlations between Ehd1 expression and heading date and spikelets on the main panicle were detected in these lines (Fig. 6d–f). These results suggested that this strategy could work not only in the ghd7 mutant but also in varieties harboring functional Ghd7 which is a strong heading date suppressor.
Phenotypes of Ehd1 RNA interference T0 plants in the Nipponbare background (Nip-Ehd1-Ri). a The phenotypes of partial Nip-Ehd1-Ri plants under artificial controlled long-day conditions in the greenhouse. bEhd1 levels in leaves of 35-day-old Nip-Ehd1-Ri T0 plants grown in the greenhouse under artificial long-day (LD, 14-h light/10-h dark) conditions were determined by qRT-PCR and shown as natural logarithms. c Heading dates are distributed in sequentially increasing order. Solid black and hollow bars in b and c represent negative and positive transgenic plants, respectively. d–f Correlations of Ehd1 expression levels with heading date (d), spikelets on the main panicle (e) and plant height (f). R indicates Pearson’s correlation coefficient; ***p < 0.001, **p < 0.01, *p < 0.05
Discussion
New strategy for improvement of regional and seasonal adaptation in rice
Ehd1 plays a central role in the regulation of heading date by integrating signals from multiple upstream regulators and transmitting them to the florigen genes (Shrestha et al. 2014). Combination of different alleles of Ehd1 upstream regulators that possess plenty of natural variations enable the fine tuning of expression of Ehd1 and florigen genes, which provide the most flexibilities of adaptation to different environment conditions (Gómez-Ariza et al. 2015; Zhang et al. 2015). However, the functional strength of different alleles of Ehd1 upstream regulators is not completely elucidated (Gómez-Ariza et al. 2015; Zhang et al. 2015). Thus, how to efficiently select and combine of these alleles is still a challenge during breeding process.
In this study, we demonstrated that Ehd1 acting downstream of Ghd7 regulating not only heading date but also yield potential, suggesting that other pleiotropic genes such as Ghd8 (Wei et al. 2010; Yan et al. 2011), Ghd7.1 (Gao et al. 2014; Yan et al. 2014), Ghd2 (Liu et al. 2016b), OsCOL9 (Liu et al. 2016a) and OsCOL10 (Tan et al. 2016) might increase yield via the same genetic pathway. Enhanced yield or yield components were observed in Ehd1 suppression lines (Fig. 3e), and significant positive correlations were found between yield or yield components and Ehd1 expression (Figs. 4b, d–h, 6e). These results demonstrated the practicability of improving yield potential by modulating Ehd1 expression level.
Besides of great value for understanding of yield contribution of heading date gene Ehd1, the most important impact of our finding is the application potential in breeding varieties by directly regulating heading date. Our approach with direct suppression of downstream common target Ehd1 could attenuate the effects from upstream regulators, such as Ghd7, Hd1, Ghd8, Ghd7.1, which possess complicated variations in different varieties or backgrounds. Like the case of Ghd7, with differential suppression of Ehd1 in genetic backgrounds with either functional or non-functional Ghd7, we successfully generated continuums of heading date variation that previously required of time-consuming combinations of different natural alleles of several genes (Figs. 3c, d, 6b, c). A smaller span of variations, which might be caused by smaller extent (about 1.5-fold in Nipponbare and threefold in ghd7) of suppression of Ehd1, was generated in Nipponbare compared with that in ghd7 mutant. But 22 days’ delay at most in Nipponbare background would be practically useful in the field production. For a specific region of cultivation, we could screen for the heading date best suited for the local environmental conditions from a series of transgenic lines to maximize grain production. Generally, lines with late heading dates can be utilized to increase yield potential in tropical and subtropical regions where the light and temperature resources are sufficient for rice growth. In contrast, lines with early heading dates may be favored in regions where multiple cropping programs are implemented or that have short growing seasons. Furthermore, our approach allows for an efficient phenotype, including heading date and yield, selection and fixation far beyond the reach of traditional breeding methods. Taken together, we proposed an efficient and flexible strategy for breeding high-yield rice varieties by optimizing heading date through RNA interference of Ehd1 regardless of genetic background.
Genetic improvement by transcriptional modulation of key regulators
Compared with mutations in the coding region that alters protein structure, transcriptional diversification with gradual and subtle phenotypic change provides increased plasticity for crop improvement (Wittkopp and Kalay 2011). And multiple transcriptional modulation strategies could be applied. Using an inducible system, Okada et al. (2017) reported the synthetic control of flowering time in rice by inducing Hd3a expression in non-flowering rice via Ghd7 overexpression. Moreover, CRISPR-Cas9 genome editing of promoters has been reported to generate diverse beneficial cis-regulatory alleles for tomato breeding (Rodriguez-Leal et al. 2017), and in this study, we successfully generated continuums of heading date variation through differential suppression of Ehd1 in genetic backgrounds with either functional or non-functional Ghd7 (Figs. 3c, d; 6b, c). Our simple and efficient strategy provides an additional choice for transcriptional crop improvement.
Lower correlation was observed between Ehd1 expression and heading date among 45 varieties from a core germplasm (Fig. 2k–m) compared with that in the ghd7/Ehd1-Ri (Fig. 4a–c) or Nip-Ehd1-Ri (Fig. 6d–f) lines. This might be caused by variations of other genes like DTH2, RID1 and SID1 (Deng et al. 2017; Wu et al. 2013) within the backgrounds, which bypass the Ehd1 pathway and regulated Hd3a/RFT1 directly. In rice, florigen genes Hd3a and RFT1 integrate all flowering signals including that from Ehd1 to switch to flowering. Thus, Hd3a and RFT1 could also be potential targets for genetic improvement of rice cultivars. With increasing knowledge of the mechanisms and pathways underlying plant growth and development, our strategy could be applied in other pathways to facilitate future crop breeding.
Author contribution statement
YH performed the map-based cloning and genetic analysis of qEhd10, the transformation and generation of transgenic materials, and the expression analysis; YH and SL performed the phenotypic analysis and picture preparation; YX directed the project; YX and YH wrote the manuscript.
References
Cho L-H, Yoon J, Pasriga R, An G (2016) Homodimerization of Ehd1 is required to induce flowering in rice. Plant Physiol 01723.02015
Deng L et al (2017) Suppressor of rid1 (SID1) shares common targets with RID1 on florigen genes to initiate floral transition in rice. PLoS Genet 13:e1006642
Doi K et al (2004) Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev 18:926–936
Gao H et al (2013) Ehd4 encodes a novel and Oryza-genus-specific regulator of photoperiodic flowering in rice. PLoS Genet 9:e1003281
Gao H et al (2014) Days to heading 7, a major quantitative locus determining photoperiod sensitivity and regional adaptation in rice. Proc Natl Acad Sci 111:16337–16342
Gibson DG, Young L, Chuang R-Y, Venter JC, Hutchison CA, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345
Gómez-Ariza J et al (2015) Loss of floral repressor function adapts rice to higher latitudes in Europe. J Exp Bot erv004
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Hori K, Matsubara K, Yano M (2016) Genetic control of flowering time in rice: integration of Mendelian genetics and genomics. Theor Appl Genet 1–12
Huang X et al (2012) A map of rice genome variation reveals the origin of cultivated rice. Nature 490:497–501
Itoh H, Nonoue Y, Yano M, Izawa T (2010) A pair of floral regulators sets critical day length for Hd3a florigen expression in rice. Nat Genet 42:635–638
Koo B-H et al (2013) Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes. Mol Plant 6:1877–1888
Kovi MR, Hu Y, Bai X, Xing Y (2015) QTL mapping for thermo-sensitive heading date in rice. Euphytica 205:51–62
Lee YS et al (2010) OsCOL4 is a constitutive flowering repressor upstream of Ehd1 and downstream of OsphyB. Plant J 63:18–30
Lei Y, Lu L, Liu H-Y, Li S, Xing F, Chen L-L (2014) CRISPR-P: a web tool for synthetic single-guide RNA design of CRISPR-system in plants. Mol Plant 7:1494–1496
Liu H, Gu F, Dong S, Liu W, Wang H, Chen Z, Wang J (2016a) CONSTANS-like 9 (COL9) delays the flowering time in Oryza sativa by repressing the Ehd1 pathway. Biochem Biophys Res Commun 479:173–178
Liu J, Shen J, Xu Y, Li X, Xiao J, Xiong L (2016b) Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J Exp Bot 67:5785–5798
Nemoto Y, Nonoue Y, Yano M, Izawa T (2016) Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7. Plant J 86:221–233
Okada R, Nemoto Y, Endo-Higashi N, Izawa T (2017) Synthetic control of flowering in rice independent of the cultivation environment. Nat Plants 3:17039. https://doi.org/10.1038/nplants.2017.39
Rodriguez-Leal D, Lemmon ZH, Man J, Bartlett ME, Lippman ZB (2017) Engineering quantitative trait variation for crop improvement by genome editing. Cell 171:470–480.e478. https://doi.org/10.1016/j.cell.2017.08.030
Sheng P et al (2016) A CONSTANS-like transcriptional activator, OsCOL13, functions as a negative regulator of flowering downstream of OsphyB and upstream of Ehd1 in rice. Plant Mol Biol 92:209–222
Shrestha R, Gómez-Ariza J, Brambilla V, Fornara F (2014) Molecular control of seasonal flowering in rice, arabidopsis and temperate cereals. Ann Bot 114:1445–1458
Sun Y et al (2016) Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of acetolactate synthase. Mol Plant 9:628–631
Takahashi Y, Teshima KM, Yokoi S, Innan H, Shimamoto K (2009) Variations in Hd1 proteins, Hd3a promoters, and Ehd1 expression levels contribute to diversity of flowering time in cultivated rice. Proc Natl Acad Sci 106:4555–4560
Tan J et al (2016) OsCOL10, a CONSTANS-like gene, functions as a flowering time repressor downstream of Ghd7 in rice. Plant Cell Physiol 57:798–812
Wei X et al (2010) DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol 153:1747–1758
Weng X et al (2014) Grain number, plant height, and heading date7 is a central regulator of growth, development, and stress response. Plant Physiol 164:735–747
Wittkopp PJ, Kalay G (2011) Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat Rev Genet 13:59
Wu W et al (2013) Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia. Proc Natl Acad Sci USA 110:2775–2780. https://doi.org/10.1073/pnas.1213962110
Xue W et al (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yan W et al (2014) Natural variation in Ghd7. 1 plays an important role in grain yield and adaptation in rice. Cell Res 23:2013, 2023 (2017): 2969–2971
Yan W-H et al (2011) A major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height, and heading date in rice. Mol Plant 4:319–330
Zhang Y, Luo L, Xu C, Zhang Q, Xing Y (2006) Quantitative trait loci for panicle size, heading date and plant height co-segregating in trait-performance derived near-isogenic lines of rice (Oryza sativa). Theor Appl Genet 113:361–368
Zhang J et al (2015) Combinations of the Ghd7, Ghd8 and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice. New Phytol 208:1056–1066
Zhang Z et al (2017) Alternative functions of Hd1 in repressing or promoting heading are determined by Ghd7 status under long-day conditions. Sci Rep 7:5388
Zhao J et al (2015) Genetic interactions between diverged alleles of early heading date 1 (Ehd1) and Heading date 3a (Hd3a)/RICE FLOWERING LOCUS T1 (RFT1) control differential heading and contribute to regional adaptation in rice (Oryza sativa). New Phytol 208:936–948
Acknowledgements
We are grateful to Dr. Yunde Zhao for the kind donation of vectors for CRISPR. This study was supported by the National Special Program for Transgenic Plant Research of China (2011ZX08009-001-002), the National Key Research and Development Program of China (2016YFD0100301), and the National Natural Science Foundation of China (31571751).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Ian D. Godwin.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, Y., Li, S. & Xing, Y. Lessons from natural variations: artificially induced heading date variations for improvement of regional adaptation in rice. Theor Appl Genet 132, 383–394 (2019). https://doi.org/10.1007/s00122-018-3225-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-018-3225-0