Abstract
Key message
Three QTL regions controlling three fiber quality traits were validated and further fine-mapped with 27 new single nucleotide polymorphism (SNP) markers. Transcriptome analysis suggests that receptor-like kinases found within the validated QTLs are potential candidate genes responsible for superior fiber strength in cotton line MD52ne.
Abstract
Fiber strength, length, maturity and fineness determine the market value of cotton fibers and the quality of spun yarn. Cotton fiber strength has been recognized as a critical quality attribute in the modern textile industry. Fine mapping along with quantitative trait loci (QTL) validation and candidate gene prediction can uncover the genetic and molecular basis of fiber quality traits. Four previously-identified QTLs (qFBS-c3, qSFI-c14, qUHML-c14 and qUHML-c24) related to fiber bundle strength, short fiber index and fiber length, respectively, were validated using an F3 population that originated from a cross of MD90ne × MD52ne. A group of 27 new SNP markers generated from mapping-by-sequencing (MBS) were placed in QTL regions to improve and validate earlier maps. Our refined QTL regions spanned 4.4, 1.8 and 3.7 Mb of physical distance in the Gossypium raimondii reference genome. We performed RNA sequencing (RNA-seq) of 15 and 20 days post-anthesis fiber cells from MD52ne and MD90ne and aligned reads to the G. raimondii genome. The QTL regions contained 21 significantly differentially expressed genes (DEGs) between the two near-isogenic parental lines. SNPs that result in non-synonymous substitutions to amino acid sequences of annotated genes were identified within these DEGs, and mapped. Taken together, transcriptome and amino acid mutation analysis indicate that receptor-like kinase pathway genes are likely candidates for superior fiber strength and length in MD52ne. MBS along with RNA-seq demonstrated a powerful strategy to elucidate candidate genes for the QTLs that control complex traits in a complex genome like tetraploid upland cotton.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton fibers are trichomes that initiate from the ovule surface approximately −3 to +3 days post-anthesis (DPA) (Fang et al. 2014b; Gilbert et al. 2014). Over the next ~50 days the cotton fiber undergoes four overlapping developing stages: rapid elongation and primary cell wall (PCW) formation (6–22 DPA), cell wall thickening and PCW remodeling (12–22 DPA), secondary cell wall (SCW) formation via deposition of cellulose between PCW and plasmalemma (16–40 DPA), and finally maturation and cell death before boll opening (40–50 DPA). The duration and rate of each stage varies among genotypes and environment (Haigler 2010; Hinchliffe et al. 2011). During the transition stage from elongation to secondary cell wall thickening, expression of genes related to PCW formation and elongation starts to decline while genes related to SCW biosynthesis are being up-regulated (Hinchliffe et al. 2010). Fiber bundle strength (FBS) increases significantly during this period although cell wall thickening and cellulose deposition are minimal (Hinchliffe et al. 2011; Hsieh 1999).
The demand for cotton fiber with higher quality has increased with the introduction of high-speed spinning technologies. Fiber quality attributes include length, strength, fineness, maturity, and elongation, etc. These fiber properties have been linked to cotton fiber cell wall formation and composition (Wakelyn et al. 2010). FBS is an important indicator of commercial fiber quality and is affected by other properties such as fiber maturity, fineness and length (Kelly et al. 2015). Fiber strength also contributes to yarn quality (Wakelyn et al. 2010) and is suggested to be affected by the thin “winding” cell wall layer and SCW cellulose (Hinchliffe et al. 2011; Hsieh et al. 1995). The genetic mechanisms of fiber traits are complex and often involve multiple genes (Zhang et al. 2013). The most effective way to improve fiber quality is through breeding, although there is a negative correlation between fiber quality and lint yield. The use of molecular marker technologies in cotton breeding through marker-assisted selection (MAS) makes simultaneous improvement of both fiber quality and yield possible (Yu et al. 2013). As suggested by meta-QTL analysis, 1075 quantitative trait loci (QTLs) in intraspecific G. hirutum L. and 1059 QTLs in interspecific (G. hirsutum × G. barbadense) populations have been detected for yield, fiber, seed quality, and stress tolerance (Said et al. 2013). After the report published by Said et al. (2013), there were a few reports also published related to cotton fiber quality QTL. A total of 131 fiber QTLs and 37 QTL clusters were identified using a random mated population originated from crosses between 11 upland cotton cultivars (Fang et al. 2014a). Very recently, Shang et al. (2015) identified 20 QTLs and validated four QTLs related to fiber quality. However, very few QTLs have actually been used in MAS breeding since they are localized in very large genomic regions and are often not stable across wide genetic backgrounds. Thus, it is essential to identify and validate a stable QTL in a specific genomic region in order to improve the trait in practical breeding.
In addition to traditional DNA marker technology, high-throughput next generation sequencing (NGS) offers a new tool to explore the genetic control of complex traits. By using NGS technology, scientists are able to add new sequence-based markers to their genetic maps and narrow down the interval until candidate genes are discovered. One such technology is mapping-by-sequencing (MBS) which has already been used by many researchers to accelerate gene discovery. The increasing availability of draft genome sequences for species in the Gossypium genus (Li et al. 2014; Paterson et al. 2012; Zhang et al. 2015) have enabled scientists to fine map and predict candidate genes. Thyssen et al. (2014, 2015) fine mapped the candidate genes for the qualitative short fiber Li 2 and Li 1 mutants by using the MBS technique and transcriptome analysis. So far, QTL fine mapping in cotton has been mainly conducted with SSR markers and has been unable to identify candidate genes (Chen et al. 2009; Shen et al. 2010; Su et al. 2013). Very few candidate genes for fiber strength have been identified through transcriptome and gene expression analysis alone. Fang et al. (2014b) reported that the carbonhydrate metabolic pathway and SCW biosynthesis can affect fiber strength. So far, no report has been published to identify candidate genes for QTLs by combining MBS with gene expression data in upland cotton. However, similar approaches were used to identify candidate genes in QTLs in other organisms (Bolon et al. 2010; Street et al. 2006).
The upland cotton lines MD52ne and MD90ne are near-isogenic lines (NILs) with significant differences in FBS. Earlier studies showed that MD52ne has approximately 10–25 % higher FBS than MD90ne (Hinchliffe et al. 2010, 2011; Islam et al. 2014; Meredith 2005). These two cotton NILs provide an opportunity to dissect the genetic mechanisms of complex fiber traits especially FBS. Previously, we mapped a stable FBS QTL from MD52ne on chromosome (Chr.) 3 along with other fiber quality QTLs using two separate F2 populations (Islam et al. 2014). In this study, we used an F3 population to validate and fine map the identified QTLs by adding single nucleotide polymorphism (SNP) markers in the QTL regions, anchoring the loci to the Gossypium raimondii physical map. RNA-seq was also utilized to analyze transcriptome and gene expression in order to identify candidate genes and better understand gene function related to fiber quality traits.
Materials and methods
Plant materials and field experiments
The two upland cotton lines (MD52ne and MD90ne) used in this study were reported earlier (Islam et al. 2014). The development scheme of the NILs and related pedigree are presented in Fig. S1. The F2 plants were grown in 2012 in a field in Stoneville, Mississippi, and F3 seeds were collected. Twenty-six plants from each of 375 F3 lines along with the parents were planted in a field in Stoneville, Mississippi, in 2014. The soil type in Stoneville is Bosket fine sandy loam. Each line was grown in single-row plots 12.2 m × 1.0 m with 45.7 cm space between plants. Standard conventional field practices were followed during the growing season. In fall 2014, all naturally opened bolls were harvested by hand from each of the F3 lines. The parental lines were grown in New Orleans, Louisiana in 2013 for mRNA and DNA isolation for sequencing.
Fiber quality measurements
Cotton bolls were ginned using a laboratory saw gin. Fiber properties of all F3 lines were measured using a High-Volume Instrument 1000 (USTER Technologies Inc., Charlotte, NC, USA) in the Cotton Fiber Testing Lab in USDA ARS-SRRC, New Orleans, Louisiana. Among the fiber quality traits, FBS, short fiber index (SFI) and upper-half mean fiber length (UHML) were measured. Fibers were pre-equilibrated in a room (65 % humidity at 21 °C) for 48 h prior to measuring the fiber properties.
DNA isolation and simple sequence repeat (SSR) marker analysis
Young leaves were bulked from 10 randomly selected plants of each F3 line along with their parents and stored at −80 °C. The genomic DNA was extracted from frozen leaves following the protocol previously described (Fang et al. 2010). Based on our previous report (Islam et al. 2014), a total of 60 SSR markers that were mapped on Chr. 3, 14, and 24 were used to genotype the 375 F3 progeny along with two parents for validation. The SSR primer sequences are available at CottonGen database (http://www.cottongen.org). SSR marker amplification, separation and scoring were according to the previous report (Islam et al. 2014).
RNA extraction and sequencing
Total RNA was extracted from the developing cotton fibers [10, 15, 20, and 24 days post anthesis (DPA)] using the Sigma Spectrum™ Plant Total RNA Kit (Sigma-Aldrich, St. Louis, MO) with DNaseI digestion according to the manufacturer’s protocol. The quality and quantity of total RNA was determined using a NanoDrop 2000 spectrophotometer (NanoDrop Technologies Inc., Wilmington, DE, USA) and an Agilent Bioanalyzer 2100 (Agilent Technologies Inc., Santa Clara, CA, USA). The RNA samples of three biological replications at two different developmental stages (15 and 20 DPA) from both NIL fibers were sent to Data2Bio LLC (Ames, Iowa) for library preparation and subsequent paired-end Illumina mRNA sequencing according to the methods described previously (Naoumkina et al. 2014).
DNA sequencing
Genomic DNA of MD52ne and MD90ne were sent to BGI Americas (Cambridge, MA, USA) for library construction and sequencing. DNA library construction and sequencing were described by Thyssen et al. (2015). DNA was sequenced with paired-end 100-bp reads on an Illumina HiSeq 2000 platform. Adaptors, contaminants and low quality reads were removed from further analysis.
Analysis of sequencing data for SNP discovery
The methods of both RNA and DNA sequence data processing were described earlier (Thyssen et al. 2014, 2015). Briefly, by using GSNAP software (Wu and Nacu 2010), clean DNA sequence reads were aligned to a 26 chromosome pseudo-tetraploid reference genome for G. hirsutum composed of the chromosomes from both of the diploid cotton reference genomes. The 13 chromosomes of the G. arboreum genome were used as the At-subgenome and the 13 chromosomes of the G. raimondii genome were used as the Dt-subgenome (Li et al. 2014; Paterson et al. 2012). This time, we used vcf tools software to call SNPs between the MD90ne and MD52ne NILs (Danecek et al. 2011). We filtered for SNPs that were called as homozygous in each NIL and had the highest quality (QUAL = 999). Histograms were generated by counting the number of SNPs in 1-Mb and 100-kb intervals. We also determined which SNP resulted in altered amino acids to identify non-synonymous mutations as before (Thyssen et al. 2014).
SNP primer design
Prior to designing SNP primers from whole genome sequence data, we constructed a genetic map based on SSR marker data to roughly locate the putative QTLs. The sequences of SSRs located in QTL regions were aligned to the reference G. raimondii genome. Read alignment in whole genome DNA sequence data on the QTL genomic regions were manually inspected to identify putative true SNPs and nearby homeo-SNPs. Two versions (one for each NIL allele) of forward primers were designed which end with the SNP alleles from MD52ne and MD90ne, while one common reverse primers was synthesized. Both forward primers contain an additional mismatch, usually at the third base from the 3ʹ end, which has been shown to increase stringency (Drenkard et al. 2000). The SNP nomenclature follows previous report (Islam et al. 2015).
SNP genotyping and linkage map construction
The newly synthesized SNP markers were first validated by running allele specific qPCR reactions using parental NILs and F1 DNA template as described earlier (Thyssen et al. 2014). Unlabeled primer, template DNA and a SYBR green PCR mix were run on a C1000 thermal cycler with CFX96 Real Time System (Bio-Rad, Hercules, CA, USA), which recorded fluorescent signal after each cycle. Samples were scored as positive or negative for each reaction by a Ct value threshold determined for each primer pair. After both allele-specific reactions were run for each individual F3 progeny, a SNP marker was scored as either homozygous for either allele or heterozygous. The SSR and SNP marker genotypes were used to construct a linkage map using the JoinMap 4.0 software with default parameters and LOD score of 10 (Van Ooijen 2006).
QTL and statistical analysis
The phenotypic data of 375 F3 lines were used to validate QTLs associated with fiber quality traits FBS, SFI and UHML reported earlier (Islam et al. 2014). Tests of normality were conducted prior to QTL analyses using PROC UNIVARIATE NORMAL PLOT (SAS Institute Inc., Cary, NC, USA). Multiple QTL modeling (MQM) was performed to localize and detect fiber quality QTL using MapQTL version 6.0 (Van Ooijen 2009). For MQM, forward and backward stepwise regressions were performed to select markers as cofactors with 10 cM window size and 1 cM walking speed. At each interval, the significance of the QTL trait association was tested by likelihood ratio statistics. For each trait, a critical LOD threshold level was estimated by 1000 permutations at p < 0.05 using the MapQTL6.0 program. However, a LOD score of 3.5 was used as a threshold to declare the presence of a putative QTL. R 2 values, representing the percentage of the phenotypic variance explained by the QTL, were determined from the MQM genetic models. QTL and chromosome maps were created using Mapchart 2.2 (Voorrips 2002). The confidence interval (95 %) for QTL location was set as a mapping distance interval corresponding to one LOD decline on either side of the peak. The QTL nomenclature was according to McCouch et al. (1997).
Detection of differentially expressed genes (DEG)
RNA-seq reads were aligned to the G. raimondii reference genome and assigned to At and Dt-subgenomes by PolyCat software as before (Page et al. 2013; Thyssen et al. 2014). Reads per Kilobase per Million reads (RPKM) was used to estimate gene expression levels. DEGs were identified by the negative binomial method of the EdgeR software using the tagwise estimation of dispersion (Robinson et al. 2010). The accuracy of the test result was corrected by false discovery rate (FDR). In this study, FDR < 0.05 and the absolute value of the log2 ratio of 1 (in which ‘ratio’ refers to the fold change in expression of the target unigene among libraries) were used to select DEGs. In this report, we only present RNA-seq expression results for the purpose of candidate genes prediction. All the DEGs were reported elsewhere (Islam et al. 2015, 2016).
Validation of RNA-seq results with reverse transcription quantitative PCR (RT-qPCR)
The experimental procedures and data analysis related to RT-qPCR were performed according to the minimum information for publication of quantitative real-time PCR experiments guidelines (Bustin et al. 2009). Four fiber development stages (10, 15, 20 and 24 DPA) were used for RT-qPCR analyses for validating the RNA-seq results of selected genes. The detailed descriptions of cDNA preparation, qPCR and calculation were previously reported (Kim et al. 2013). Three biological replications and three technical replications for each time-point were used for RT-qPCR.
Results
Analysis of fiber quality traits
The frequency distributions of fiber quality traits including FBS, SFI and UHML displayed continuous and unimodal variation in the F3 population of 375 lines (Fig. 1). Similar trends were also observed in the F2 populations that were reported earlier (Islam et al. 2014). Since all tested traits were normally distributed, they are suitable for QTL analysis. FBS, SFI and UHML data for the F3 progeny ranged from 30.64 to 39.10 g/tex, 6.08 to 9.14 % and 1.05 to 1.25 inches averaging 34.45 g/tex, 7.28 % and 1.17 inches, respectively.
Alignment of sequence reads against reference genome and SNP detection
Mapping-by-sequencing reads were aligned to a pseudo-reference genome of G. hirsutum that consisted of the reference chromosomes of both the extant A and D diploid Gossypium species which we treated as the At and Dt-subgenomes, respectively (Li et al. 2014; Paterson et al. 2012). A total of 29,125 high quality SNPs were detected between the two NILs. Of the 29,125 SNPs, 16,984 and 12,141 were detected in the At-subgenome and Dt-subgenome, respectively. Three large peaks of SNPs were observed on Chr. 4 (15–22 Mb) and Chr. 5 (15–22 Mb, and 50–55 Mb) in the histogram of Dt-subgenome reads. Chr. 4 and 5 of G. raimondii are homeologs of Chr. 24 and 14, respectively, of G. hirsutum (Fig. 2). A total of 902 SNPs per Mb were observed in the highest peak on Chr. 4 of Dt-subgenome reads, and the two peaks on Chr. 5 contained 609 and 715 SNPs per Mb, respectively. No peaks above 600 were observed on the At-subgenome, with the highest at 576 SNPs per Mb on Chr. 13 (Fig. 2). At each of the three largest peaks in the Dt-subgenome, 160, 119 and 153 SNPs per 100-kb were observed at 18.6, 19.9 and 52.9 Mb of Chrs. 4, 5 and 5, respectively (Fig. 2). Since MD52ne and MD90ne are highly near isogenic BC6F6 with a major difference in fiber strength, this result suggested that the fiber strength QTL might reside in those genomic regions with greatest SNPs diversity. Since, very recently a draft upland cotton (G. hirsutum) tetraploid genome (TM-1) was published (Zhang et al. 2015), we also aligned our sequence reads to the TM-1 draft genome sequence following similar methods. SNP distribution histograms of At and Dt subgenomes are presented in Fig. S2. We found four large clusters of SNPs on the TM-1 reference genome. However, we did not observe obvious SNP diversity in one of the regions (D08, or Chr. 24) where fiber QTLs reside. Manual inspection of the TM-1 regions homologous to Chr. 4 (15–22 Mb) and Chr. 5 (50–55 Mb) of the G. raimondii genome suggested an incomplete sorting of homeologous reads in the alignment to the TM-1 draft genome in these regions, which may have interfered with SNP calling. Since we started this project and discovered genetically linked SNPs before the TM-1 genome sequence was published, we decided to proceed with the SNPs detected using the pseudo-tetraploid reference genome, which the G. raimondii reference genome (Paterson et al. 2012) is partially based on.
SNPs per Mb in whole genome sequence data. The 13 reference G. raimondii chromosomes are presented as the Dt-subgenome and the 13 G. arboreum chromosomes represent the At-subgenome. The three most diverse genomic region were located on Chr. 4 (15–22 Mb, a), Chr. 5 (15–22 Mb, b) and Chr. 5 (47–60 Mb, c) are presented as SNPs per 100 kb. The locations of markers that flank the QTLs controlling bundle fiber strength are shown. G.r. = Gossypium raimondii; G.h. = Gossypium hirsutum
Initial linkage map construction and QTL detection
At first, 60 SSR markers anchored on G. hirsutum Chrs. 3, 14 and 24 associated with fiber QTLs identified in our previous report (Islam et al. 2014) were selected for genotyping 375 F3 lines. Finally, all tested SSR loci were placed on three respective chromosomes using JoinMap 4.0 (Fig. S3). A total of 40, 16 and 4 SSR markers were mapped on upland cotton Chrs. 3, 14 and 24, respectively. Marker orders for each respective chromosome were essentially identical to the map reported earlier based on F2 populations (Islam et al. 2014) and to the upland cotton consensus map (Blenda et al. 2012). QTL analysis of these three selected chromosomes was conducted using MapQTL 6.0. One of each QTL associated with FBS, SFI and UHML were confirmed on Chrs. 3, 14 and 24, respectively whereas two additional QTL for FBS were detected on Chrs. 14 and 24 which co-localized with both UHML and SFI QTL. To identify the physical locations of SSRs of these QTL regions, sequences of SSR markers were aligned to the G. raimondii reference genome. BLAST results revealed that the SSR markers of QTL region Chr. 3, 14 and 24 were physically located on G. raimondii Chr. 5 (49–58th Mb), Chr. 5 (17–21th Mb) and Chr. 4 (18–22nd Mb) which agrees with the expected correspondence of homeologous chromosomes in G. hirsutum (Fig. 2) (Altschul et al. 1990; Blenda et al. 2012; Paterson et al. 2012).
SNP marker analysis, linkage map construction and QTL detection
As expected, all three initial QTL regions were physically located in the three most highly diverse genomic regions, so we decided to select SNPs from near the QTL locations to interrogate in the segregating F3 population. Primers for a total of 27 SNP loci were designed using NCBI Primer-BLAST tools and validated (Ye et al. 2012), and all showed expected genotypes for the NILs and F1. The allele specific primers for these 27 loci (Table S1) were used to genotype the 375 F3 progeny.
Finally, all tested SNP and SSR loci were placed on three respective chromosomes using JoinMap 4.0. A total of 49 (40 SSRs and 9 SNPs), 24 (16 SSRs and 8 SNPs) and 13 (3 SSRs and 10 SNPs) markers were mapped on upland cotton Chrs. 3, 14 and 24, respectively (Fig. 3a–c). Anchored SSR marker orders for each respective chromosome were identical to the map constructed initially (Fig. S3). Most of the SNP markers were placed inside the QTL region of the respective chromosome. The average distances between two adjacent loci were 0.96, 0.89 and 0.75 cM in Chrs. 3, 14 and 24, respectively. We also compared the genetic and physical orders of mapped markers which showed significant co-linearity (Fig. 3).
QTLs for bundle fiber strength (FBS), short fiber index (SFI) and upper-half mean length (UHML) identified in F3 population derived from a cross of MD90ne × MD52ne. QTLs were detected by multiple QTL modeling using MapQTL6 software. The LOD scores were plotted against centiMorgans (cM) on the chromosome. SNPs markers (in red) start with CFB followed by number. SNPs and SSRs were also mapped on physical locations as base pairs (bp) of G. raimondii reference genome. a Chr. 03; b Chr. 14, and c Chr. 24
QTL analysis of these three chromosomes was conducted using fiber trait values of 375 F3 lines. One of each QTL associated with FBS and SFI were re-identified on Chrs. 3 and 14, respectively. While two QTLs for UHML were confirmed on Chrs. 14 and 24. Additionally, two QTLs associated with FBS were identified on Chrs. 14 and 24. One additional SFI QTL was also identified on Chr. 24 (Table 1; Fig. 3c). Additive values of the respective QTLs indicated that the beneficial alleles for the respective traits originated from MD52ne (Table 1).
Three QTLs associated with FBS, SFI and UHML (qFBS-c24, qSFI-c24 and qUHML-c24, respectively) were co-localized on G. hirsutum Chr. 24, which is homeologous to G. raimondii Chr. 4 and accounted for 15.7, 9.8 and 11.6 % of the total phenotypic variation. The FBS QTL LOD score peaked near the SSR marker GH454, while two other QTLs were linked with SNP marker CFB5879. However, all three QTLs were flanked by CFB5878 and GH222. The physical and genetic distances between these two flanking markers were 3.7 Mb and 8.1 cM, respectively (Fig. 3c). Similar to Chr. 24, three QTLs associated with all three tested fiber traits (qFBS-c14, qSFI-c14 and qUHML-c14) were co-localized on G. hirsutum Chr. 14, which is homeologous to G. raimondii Chr. 5. The phenotypic variations associated with these QTLs were 8.4, 9.3 and 11.5 % for FBS, SFI and UHML, respectively. The FBS QTL linked with SSR marker CGR6683 and was flanked by markers BNL3502 and BNL3033 that are 1.8 Mb and 4.4 cM apart physically and genetically, respectively. The QTL related to SFI was linked with SSR marker JESPR006 and was flanked by CGR6217-CFB5875 (10.8 cM apart), while the UHML QTL LOD score peaked near the SNP marker CFB5870 and was flanked by JESPR006-CFB5875 (0.61 Mb and 9.0 cM apart physically and genetically, respectively). The third QTL for FBS (qFBS-c3) is linked to SNP markers that reside in a cluster of diversity identified on G. raimondii Chr. 05 of the pseudo-tetraploid genome (Figs. 2, 3a). The SSR markers at this locus were previously reported to reside on G. hirsutum Chr. 03 (Blenda et al. 2012; Islam et al. 2014) although we found suitable alignments close to our linked SNP markers in the Dt-subgenome of the pseudo-tetraploid sequences (Fig. 2). This QTL region was bordered by BNL4034-NAU3541 (4.4 Mb and 6.8 cM apart physically and genetically, respectively) and contributed 4.4 % of the total phenotypic variation.
DEGs near/within a fiber QTL region
Based on the fiber crystallinity data obtained from developing fibers (Fig. S4; Islam et al. 2016) and our previous results (Hinchliffe et al. 2011), transition from fiber elongation to secondary cell wall thickening occurs about 16–17 DPA in MD52ne. Thus, we selected fibers at 15 and 20 DPA for RNA-seq analysis. We analyzed the RNA-seq reads with EdgeR in order to investigate the expression pattern of genes near the identified QTL regions (Robinson et al. 2010). The genes were considered as DEGs, when at-least two fold difference in their expression level between MD52ne and MD90ne at either developing time point was found. In Table 2, we presented all three QTL regions which are physically located on two chromosomes (Chr.04 and Chr.05) of the G. raimondii genome. We included only the differentially expressed genes in Table 2; that are physically located between or are very near to the two flanking markers. A total of 20, 8, and 10 genes were differentially expressed between NILs in 15 and/or 20 DPA developing fiber in QTL regions on Chrs. 3, 14 and 24, respectively. Of the 38 DEGs, only 21 were significantly different (adjusted p value <0.05) between MD52ne and MD90ne at either developing time point. Among those 21 DEGs, interestingly Goraii.005G178700, a ribosomal protein was highly up-regulated (1618- and 568-fold at 15 and 20 DPA, respectively) in MD52ne. This gene was physically located just 2.7 kb away from left flanking marker (BNL4034) of QTL qFBS-c3. Another gene, Gorai.005G108500 (Protein kinase super family protein) was also highly abundant in MD52ne at both DPA and located near QTLs qFBS-c14, qSFI-c14, qUHML-c14. One gene (Gorai.005G102600) also located near QTLs qFBS-c14, qSFI-c14, qUHML-c14 was found down regulated in MD52ne at both the fiber developing stages. Several other genes that are responsible for cell wall growth and development were also up-regulated in MD52ne at 15 DPA.
RT-qPCR of the selected genes during fiber growth
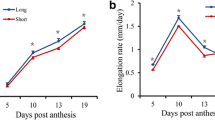
A total of twelve genes across three QTL regions were selected for RT-qPCR in order to validate the RNA-seq expression results using fiber tissue from four developmental stages (Fig. 4; Table S2). All of the selected genes control cell wall growth and development either directly or indirectly. Of the 12 tested genes, the expression level of five genes were significantly higher in MD52ne developing fiber at all four tested time points (Fig. 4a–e) while two genes had lower expression in MD52ne fibers (Fig. 4f, g). Two genes, Gorai.004G102000 and Gorai.004G103500 showed significant differences between MD52ne and MD90ne expression during late stages of fiber development (15, 20 and 24 DPA) (Fig. 4h, i). A higher level of expression was observed in MD52ne with gene MATE efflux family protein (DTX1, Gorai.005G178900) (Fig. 4j) during early stages of fiver development while NAC domain protein 83 (NAC83, Gorai.005G195300) showed opposite direction of expression (Fig. 4k). The gene Gorai.005G198900, showed zigzag expression pattern between two NILs across three fiber development stages while it is indistinguishable at 10 DPA.
Quantitative RT-PCR validation of select genes related to cell wall activity during fiber development. a Protein kinase super family protein (Gorai.005G108500); b Ribosomal L18e/L15 protein (Gorai.005G178700); c homeobox 1 (Gorai.005G189800); d Leucine rich repeat (LRR) transmembrane protein kinase family protein (Gorai.005G193500); e Oxidoreductase, zinc-binding dehydrogenase family protein (Gorai.005G182400); f Calcium-binding EF-hand family protein (Gorai.005G102600); g thaumatin-like protein 3 (Gorai.005G106400); h CONSTANS like 5 (Gorai.004G102000); i trehalose phosphatase/synthase 11 (TPP/TPS 11, Gorai.004G103500); j MATE efflux family protein (DTX1, Gorai.005G178900); k NAC domain protein 83 (NAC83, Gorai.005G195300); l C2H2 and C2HC zinc fingers super family protein (Gorai.005G198900)
Identification of non-synonymous SNPs within genes near/within fiber QTL regions
To find out whether a SNP resulted in amino acid change, we first identified SNPs in genes adjacent to fiber QTL, and then obtained annotated proteins. Of the 27 SNPs, 18 were located in exons. Of these 18 SNPs, eight produced silent amino acid substitution and the remaining 10 SNPs formed mis-sense (9) and nonsense (1) substitutions of amino acid sequence (Table 3). The only nonsense mutation was observed in Gorai.005G106400 (Thaumatin-like protein 3). The NIL with shorter and weaker fibers, MD90ne contains a C-to-T transition resulting in a truncated protein (Table 3). To investigate which missense amino acid substitutions might have significant effects on fiber cell development, we compared the chemical nature of the substituted to original amino acids and searched Genbank for orthologous proteins with identical mutations. We found that three (CONSTANS-like 5, Photosystem II reaction center protein G, Poor homologous synapsis 1) of the nine missense mutations were conversions to chemically similar amino acids, while the other six were synonymous to orthologous proteins (Table 3).
Discussion
Increasing marker density via MBS
In our previous report (Islam et al. 2014), the coverage of SSR markers throughout the cotton genome was low since MD52ne and MD90ne are highly near isogenic BC6F6 lines. We used all the available SSR markers in the three QTL regions to screen for polymorphism between the two NILs. However, we were not able to saturate the QTL regions with SSRs. Recently we explored the MBS technique to identify SNP markers in the QTL regions in order to increase marker density. MBS has already been proven to rapidly identify a genomic region of few Mb that contained a cluster of introgressed diversity that differentiates NILs (Thyssen et al. 2014, 2015). In this study, we were able to place 27 new SNP markers in the three QTL regions using MBS. Consequently, the average distance between two adjacent loci was substantially decreased (0.96, 0.89 and 0.75 cM in Chrs. 3, 14 and 24, respectively; Fig. 3) compared to our initial map (1.25, 1.27 and 3.05 cM in Chrs. 3, 14 and 24, respectively; Fig. S3) and previously reported linkage maps (1.71, 5.16 and 4.50 cM in Chrs. 3, 14 and 24, respectively) (Islam et al. 2014).
QTL identification and validation
In order to use DNA markers in practical breeding, an identified QTL needs to be validated using different genetic backgrounds, different generations and/or environments. QTLs which can be repeatedly identified are stable and useful in MAS breeding (Su et al. 2010). So far, very few identified fiber QTL have been used in cotton MAS breeding due to a lack of stability. The effects of some QTLs are minimized or lost under different environments or in different genetic backgrounds. In this study, we were able to validate four QTLs (qFBS-c3, qSFI-c14, qUHML-c14 and qUHML-c24) related to FBS, SFI and UHML identified on upland cotton Chrs. 3, 14 and 24, respectively (Islam et al. 2014) using individuals of a different generation (F3) planted in a different year. The qFBS-c3 region identified in this study shared many common markers (e.g. BNL4034, CGR6164, BNL3463, NAU3541 and SHIN-0463) with the FBS QTL region detected in our earlier report. The right flanking marker NAU3541 was also identical. However, the left flanking marker BNL4034 was the peak marker in our previous report. According to the high-density consensus map (Blenda et al. 2012), the FBS QTL intervals identified in this study overlapped with our earlier identified FBS QTL (Islam et al. 2014). It is not uncommon that a QTL interval varies slightly between populations or between environments (Shen et al. 2005). Hence, it is very reasonable to believe that these two QTLs are the same. The qSFI-c14 was confirmed since both had an identical peak marker (CGR6683) in the two studies. Finally, qUHML-c24 also shared a common peak marker (GH454) and right flanking markers (GH222) in the two studies. By including SNPs, we were able to reduce the QTL regions substantially. We conclude that these three QTLs from MD52ne are validated and ready to use in MAS.
In this study, we also identified two, one and one new QTLs for FBS, SFI and UHML, respectively. For F2 population, the fiber samples were from only one individual plant. However, fiber samples were from a progeny row of 26 plants for F3 population. Consequently, the phenotypic data may be more accurate in F3 than in F2. It is reasonable to believe that we missed some fiber QTLs in our previous study (Islam et al. 2014). The combined effects of the three FBS QTLs explained 28.5 % phenotypic variation in this study. As mentioned earlier, MD52ne has 10–25 % higher FBS than MD90ne. It is conceivable that we identified most, if not all, QTL responsible for FBS difference between MD52ne and MD90ne.
Eight and nine QTLs associated with fiber length and strength, respectively, have been reported on Chr. 14 in earlier studies, but no QTL was related to SFI (He et al. 2007; Lacape et al. 2005; Qin et al. 2008; Said et al. 2013; Wang et al. 2006) and none of those were positioned in the same genomic region reported in this study. A large number (26) of FBS QTLs along with five fiber length QTLs have been reported by several researchers (Said et al. 2013). One QTL associated with both FBS and fiber length was reported earlier on Chr. 24 that possibly overlaps with the QTL region identified in this study (Zhang et al. 2012). Among the newly identified QTLs, qFBS-c14 and qUHML-c14 appear unique. The possibility that qFBS-c24 and qSFI-c24 may be the same as the QTLs identified in prior studies merits further investigation.
It is well-known that FBS values are affected by individual fiber strength and other fiber quality traits which affect fiber-to-fiber interactions within a bundle (Bradow and Davidonis 2000; Haigler 2010; Hsieh 1999; Hsieh et al. 1995; Wakelyn et al. 2010). Fiber-to-fiber interactions are affected by fiber length within a bundle and impact the FBS values (Naylor et al. 2014). Our results revealed that FBS, SFI and UHML are inter-related, since QTLs associated with those traits were co-localized on Chrs. 14 and 24. Thus, we believe that same genomic regions of MD52ne regulate these three fiber traits.
Candidate genes
To uncover the key regulating genes related to the superior fiber traits in MD52ne, we conducted a combination of three steps of structural and functional genomic analysis. First we fine mapped the QTL regions using SNPs. Second, we identified significant DEGs related to fiber cell development inside the QTL regions. Then we determined the alteration of amino acids due to SNPs in genes. Of the 38 DEGs, only 21 were significantly different (adjusted p value <0.05) between MD52ne and MD90ne at 15 and/or 20 DPA. Interestingly, eight of these were related to multiple receptor-like kinases (RLKs) localized in the plasma membrane, which were differentially expressed in the developing fibers of MD52ne as compared to MD90ne. The leucine-rich repeat (LRR) RLKs comprising a transmembrane region and a kinase domain have recently been reported as a novel signaling pathway that regulates plant cell wall integrity maintenance (Hamann 2015). Two LRR RLKs named FEI1 and FEI2 have been reported to play a vital role in cellulose deposition during elongation of root tips and seeds in Arabidopsis (Harpaz-Saad et al. 2011; Xu et al. 2008). In another report, other LRR RLKs are suggested as regulators of SCW formation in Arabidopsis (Wang et al. 2013) and poplar trees (Song et al. 2011). A plasma membrane bound cotton LRR RLK named GhRLK1 was found to be induced during the active SCW synthesis stage (Li et al. 2005). Thus, RLKs signaling pathways suggested being engaged in mediating a combination of cell elongation and SCW biosynthesis during cotton fiber development in MD52ne. Members of several transcription factor families that are related to cotton fiber cell development including NAC domain protein (Fang et al. 2014b, c), CONSTANS-like 5 (Islam et al. 2015, 2016), and zinc finger family protein (Thyssen et al. 2014) were also located inside the QTL regions and are highly transcribed in developing MD52ne fibers (Table 2). Both RNA-seq and RT-qPCR indicated that two trehalose-phosphatase/synthase (TPP/TPS) genes located in QTL regions on Chrs.14 and 24 were also highly up-regulated in MD52ne developing fibers. Literature suggests that a class II TPS/TPP enzyme involved in sugar catabolism has a role in the control of cell morphogenesis and acts as an important regulator of plant development via a role in generating a sugar-based signal or more directly by participating in transcriptional regulation (Chary et al. 2008).
Protein change due to SNPs within QTLs
To characterize key genes involved in cotton fiber development, we determined which mapped SNP resulted in altered amino acids between NILs (Table 3). Out of nine missense changes, three alterations cause the substitution of chemically similar amino acids, thus they may not have any effect on protein structure and function (Griffiths et al. 2000). The other six missense mutations result in amino acid substitutions that are synonymous to orthologous proteins. While is often considered evidence that a missence mutation is not deleterious (Ng and Henikoff 2003), we cannot exclude the possibility that these missense mutations contribute to the different fiber properties of MD52ne and MD90ne.
Interestingly, one RLK gene (thaumatin-like protein 3) has a premature stop codon in MD90ne that has inferior fiber. We observed significantly higher expression (3.3- and 5.8-fold in 15 and 20 DPA, respectively) of this transcript in MD90ne, which may indicate feedback regulation that is attempting to compensate for the lack of a functional protein (Table 2). A thaumatin like protein (GbTLP1) from sea island cotton (G. barbadense L.) has been reported to be involved in secondary cell wall thickening (Munis et al. 2010). Several other RLK genes are up-regulated in MD52ne relative to MD90ne and may therefore be downstream of the mutated protein in a signaling cascade (Islam et al. 2015, 2016). Further work is required to confirm that RLK signaling pathways are involved in mediating cell elongation and SCW biosynthesis and in the development of superior fiber length and strength in upland cotton cultivar MD52ne.
Conclusions
We confirmed four previously-identified QTLs (qFBS-c3, qSFI-c14, qUHML-c14 and qUHML-c24) related to fiber bundle strength, short fiber index and fiber length, respectively, using an F3 population that originated from a cross of MD90ne × MD52ne. These QTL regions were further fine-mapped by placing a group of 27 additional SNP markers developed through MBS. Our refined QTL regions spanned 4.4, 1.8 and 3.7 Mb of physical distance in the Gossypium raimondii reference genome. The QTL regions also contained 21 significant DEGs between MD52ne and MD90ne. SNPs that result in non-synonymous substitutions to amino acid sequences of annotated genes were identified within these DEGs, and mapped. Transcriptome and amino acid mutation analysis suggest that receptor-like kinase pathway genes are likely candidates for superior fiber strength and length in MD52ne.
Author contribution statement
DDF conceived the project and revised the manuscript. DDF and MSI designed the experiment. MSI design SNP markers, conducted marker analysis, prepared fiber samples, analyzed data and wrote the manuscript. LZ made crosses and conducted field tests. GT analyzed the sequencing data; CDD measured fiber properties using HVI. HJK assisted in designing the project and editing the manuscript. PL assisted in SSR genotyping. All authors read and approved the final manuscript.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Blenda A, Fang DD, Rami JF, Garsmeur O, Luo F, Lacape JM (2012) A high density consensus genetic map of tetraploid cotton that integrates multiple component maps through molecular marker redundancy check. PLoS One 7:e45739
Bolon YT, Joseph B, Cannon SB, Graham MA, Diers BW, Farmer AD, May GD, Muehlbauer GJ, Specht JE, Tu ZJ, Weeks N, Xu WW, Shoemaker RC, Vance CP (2010) Complementary genetic and genomic approaches help characterize the linkage group I seed protein QTL in soybean. BMC Plant Biol 10:41
Bradow JM, Davidonis GH (2000) Quantitation of fiber quality and the cotton production-processing interface: a physiologist’s perspective. J Cotton Sci 4:34–64
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Chary SN, Hicks GR, Choi YG, Carter D, Raikhel NV (2008) Trehalose-6-phosphate synthase/phosphatase regulates cell shape and plant architecture in Arabidopsis. Plant Physiol 146:97–107
Chen H, Qian N, Guo W, Song Q, Li B, Deng F, Dong C, Zhang T (2009) Using three overlapped RILs to dissect genetically clustered QTL for fiber strength on Chro. D8 in Upland cotton. Theor Appl Genet 119:605–612
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R, Genomes Project Analysis G (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Drenkard E, Richter BG, Rozen S, Stutius LM, Angell NA, Mindrinos M, Cho RJ, Oefner PJ, Davis RW, Ausubel FM (2000) A simple procedure for the analysis of single nucleotide polymorphisms facilitates map-based cloning in Arabidopsis. Plant Physiol 124:1483–1492
Fang DD, Xiao J, Canci PC, Cantrell RG (2010) A new SNP haplotype associated with blue disease resistance gene in cotton (Gossypium hirsutum L.). Theor Appl Genet 120:943–953
Fang DD, Jenkins JN, Deng DD, McCarty JC, Li P, Wu J (2014a) Quantitative trait loci analysis of fiber quality traits using a random-mated recombinant inbred population in Upland cotton (Gossypium hirsutum L.). BMC Genom 15:397
Fang L, Tian R, Chen J, Wang S, Li X, Wang P, Zhang T (2014b) Transcriptomic analysis of fiber strength in upland cotton chromosome introgression lines carrying different Gossypium barbadense chromosomal segments. PLoS One 9:e94642
Fang L, Tian R, Li X, Chen J, Wang S, Wang P, Zhang T (2014c) Cotton fiber elongation network revealed by expression profiling of longer fiber lines introgressed with different Gossypium barbadense chromosome segments. BMC Genom 15:838
Gilbert MK, Kim HJ, Tang Y, Naoumkina M, Fang DD (2014) Comparative transcriptome analysis of short fiber mutants Ligon-lintless 1 and 2 reveals common mechanisms pertinent to fiber elongation in cotton (Gossypium hirsutum L.). PLoS One 9:e95554
Griffiths A, Miller J, Suzuki D, Lewontin R, Gelbart W (2000) How DNA changes affect phenotype. In: Freeman WH (ed) Introduction to genetic analysis, 7th edn, New York. http://www.ncbi.nlm.nih.gov/books/NBK21955/
Haigler C (2010) Physiological and anatomical factors determining fiber structure and utility. In: Stewart JM, Oosterhuis DM, Heitholt JJ, Mauney JR (eds) Physiology of cotton. Springer, New York, pp 33–47
Hamann T (2015) The plant cell wall integrity maintenance mechanism—a case study of a cell wall plasma membrane signaling network. Phytochem 112:100–109
Harpaz-Saad S, McFarlane HE, Xu S, Divi UK, Forward B, Western TL, Kieber JJ (2011) Cellulose synthesis via the FEI2 RLK/SOS5 pathway and cellulose synthase 5 is required for the structure of seed coat mucilage in Arabidopsis. Plant J 68:941–953
He D, Lin Z, Zhang X, Nie Y, Guo X, Zhang Y, Li W (2007) QTL mapping for economic traits based on a dense genetic map of cotton with PCR-based markers using the interspecific cross of Gossypium hirsutum × Gossypium barbadense. Euphytica 153:181–193
Hinchliffe DJ, Meredith WR, Yeater KM, Kim HJ, Woodward AW, Chen ZJ, Triplett BA (2010) Near-isogenic cotton germplasm lines that differ in fiber-bundle strength have temporal differences in fiber gene expression patterns as revealed by comparative high-throughput profiling. Theor Appl Genet 120:1347–1366
Hinchliffe DJ, Meredith WR, Delhom CD, Thibodeaux DP, Fang DD (2011) Elevated growing degree days influence transition stage timing during cotton fiber development resulting in increased fiber-bundle strength. Crop Sci 51:1683–1692
Hsieh YL (1999) Structural development of cotton fibers and linkages to fiber quality. In: Basra AS (ed) Cotton fibers developmental biology, quality improvement, and textile processing. Haworth Press Inc, New York, pp 137–165
Hsieh YL, Honik E, Hartzell M (1995) A developmental study of single fiber strength: greenhouse grown SJ-2 Acala cotton. Textile Res J 65:101–112
Islam MS, Zeng L, Delhom CD, Song X, Kim HJ, Li P, Fang DD (2014) Identification of cotton fiber quality quantitative trait loci using intraspecific crosses derived from two near-isogenic lines differing in fiber bundle strength. Mol Breed 34:373–384
Islam MS, Thyssen GN, Jenkins JN, Fang DD (2015) Detection, validation, and application of genotyping-by-sequencing based single nucleotide polymorphisms in upland cotton. Plant Genome. doi:10.3835/plantgenome2014.3807.0034
Islam MS, Fang DD, Thyssen GN, Delhom CD, Liu Y, Kim HJ (2016) Comparative fiber property and transcriptomic analyses reveal key genes potentially related to high fiber strength in cotton (Gossypium hirsutum L.) line MD52ne. BMC Plant Biol 16:36
Kelly B, Abidi N, Ethridge D, Hequet EF (2015) Fiber to fabric. In: Fang DD, Percy RG (eds) Cotton, 2nd edn. ASSA-CSSA-SSSA, Madison, pp 665–744
Kim HJ, Tang Y, Moon HS, Delhom CD, Fang DD (2013) Functional analyses of cotton (Gossypium hirsutum L.) immature fiber (im) mutant infer that fiber cell wall development is associated with stress responses. BMC Genom 14:889
Lacape J, Nguyen T, Courtois B, Belot J, Giband M, Gourlot J, Gawryziak G, Roques S, Hau B (2005) QTL analysis of cotton fiber quality using multiple G. hirsutum × G. barbadense backcross generations. Crop Sci 45:123–140
Li YL, Sun J, Xia GX (2005) Cloning and characterization of a gene for an LRR receptor-like protein kinase associated with cotton fiber development. Mol Genet Genom 273:217–224
Li F, Fan G, Wang K, Sun F, Yuan Y, Song G, Li Q, Ma Z, Lu C, Zou C, Chen W, Liang X, Shang H, Liu W, Shi C, Xiao G, Gou C, Ye W, Xu X, Zhang X, Wei H, Li Z, Zhang G, Wang J, Liu K, Kohel RJ, Percy RG, Yu JZ, Zhu YX, Wang J, Yu S (2014) Genome sequence of the cultivated cotton Gossypium arboreum. Nat Genet 46:567–572
McCouch S, Cho Y, Yano P, Blinstrub M, Morishima H, Kinoshita T (1997) Report on QTL nomenclature. Rice Genet Newsl 14:11–13
Meredith WR (2005) Minimum number of genes controlling cotton fiber strength in a backcross population. Crop Sci 45:1114–1119
Munis MF, Tu L, Deng F, Tan J, Xu L, Xu S, Long L, Zhang X (2010) A thaumatin-like protein gene involved in cotton fiber secondary cell wall development enhances resistance against Verticillium dahliae and other stresses in transgenic tobacco. Biochem Biophys Res Commu 393:38–44
Naoumkina M, Thyssen G, Fang DD, Hinchliffe DJ, Florane C, Yeater KM, Page JT, Udall JA (2014) The Li2 mutation results in reduced subgenome expression bias in elongating fibers of allotetraploid cotton (Gossypium hirsutum L.). PLoS One 9:e90830
Naylor GR, Delhom CD, Cui X, Gourlot J-P, Rodgers J (2014) Understanding the influence of fiber length on the High Volume Instrument™ measurement of cotton fiber strength. Textile Res J 84:979–988
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 31:3812–3814
Page JT, Gingle AR, Udall JA (2013) PolyCat: a resource for genome categorization of sequencing reads from allopolyploid organisms. G3 (Bethesda) 3:517–525
Paterson AH, Wendel JF, Gundlach H, Guo H, Jenkins J, Jin D, Llewellyn D, Showmaker KC, Shu S, Udall J, Yoo MJ, Byers R, Chen W, Doron-Faigenboim A, Duke MV, Gong L, Grimwood J, Grover C, Grupp K, Hu G, Lee TH, Li J, Lin L, Liu T, Marler BS, Page JT, Roberts AW, Romanel E, Sanders WS, Szadkowski E, Tan X, Tang H, Xu C, Wang J, Wang Z, Zhang D, Zhang L, Ashrafi H, Bedon F, Bowers JE, Brubaker CL, Chee PW, Das S, Gingle AR, Haigler CH, Harker D, Hoffmann LV, Hovav R, Jones DC, Lemke C, Mansoor S, ur Rahman M, Rainville LN, Rambani A, Reddy UK, Rong JK, Saranga Y, Scheffler BE, Scheffler JA, Stelly DM, Triplett BA, Van Deynze A, Vaslin MF, Waghmare VN, Walford SA, Wright RJ, Zaki EA, Zhang T, Dennis ES, Mayer KF, Peterson DG, Rokhsar DS, Wang X, Schmutz J (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492:423–427
Qin H, Guo W, Zhang YM, Zhang T (2008) QTL mapping of yield and fiber traits based on a four-way cross population in Gossypium hirsutum L. Theor Appl Genet 117:883–894
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Said JI, Lin Z, Zhang X, Song M, Zhang J (2013) A comprehensive meta QTL analysis for fiber quality, yield, yield related and morphological traits, drought tolerance, and disease resistance in tetraploid cotton. BMC Genom 14:776
Shang L, Liang Q, Wang Y, Wang X, Wang K, Abduweli A, Ma L, Cai S, Hua J (2015) Identification of stable QTLs controlling fiber traits properties in multi-environment using recombinant inbred lines in Upland cotton (Gossypium hirsutum L.). Euphytica 205:877–888
Shen X, Guo W, Zhu X, Yuan Y, Yu J, Kohel R, Zhang T (2005) Molecular mapping of QTLs for fiber qualities in three diverse lines in Upland cotton using SSR markers. Mol Breed 15:169–181
Shen X, He Y, Lubbers EL, Davis RF, Nichols RL, Chee PW (2010) Fine mapping QMi-C11 a major QTL controlling root-knot nematodes resistance in Upland cotton. Theor Appl Genet 121:1623–1631
Song D, Xi W, Shen J, Bi T, Li L (2011) Characterization of the plasma membrane proteins and receptor-like kinases associated with secondary vascular differentiation in poplar. Plant Mol Biol 76:97–115
Street NR, Skogstrom O, Sjodin A, Tucker J, Rodriguez-Acosta M, Nilsson P, Jansson S, Taylor G (2006) The genetics and genomics of the drought response in Populus. Plant J 48:321–341
Su C, Lu W, Zhao T, Gai J (2010) Verification and fine-mapping of QTLs conferring days to flowering in soybean using residual heterozygous lines. Chin Sci Bull 55:499–508
Su CF, Wang W, Qiu XM, Yang L, Li S (2013) Fine-mapping a fibre strength QTL QFS-D11-1 on cotton chromosome 21 using introgressed lines. Plant Breed 132:725–730
Thyssen GN, Fang DD, Turley RB, Florane C, Li P, Naoumkina M (2014) Next generation genetic mapping of the Ligon-lintless-2 (Li(2)) locus in upland cotton (Gossypium hirsutum L.). Theor Appl Genet 127:2183–2192
Thyssen GN, Fang DD, Turley RB, Florane C, Li P, Naoumkina M (2015) Mapping-by-sequencing of Ligon-lintless-1 (Li) reveals a cluster of neighboring genes with correlated expression in developing fibers of Upland cotton (Gossypium hirsutum L.). Theor Appl Genet 128:1703–1712
Van Ooijen JW (2006) JoinMap 4.0: software for the calculation of genetic linkage maps in experimental populations. Wageningen: Kyazma B.V
Van Ooijen J (2009) MapQTL 6, software for the mapping of quantitative trait loci in experimental populations of diploid species. Kyazma BV, Wageningen
Voorrips R (2002) MapChart: software for the graphical presentationof linkage maps and QTLs. J Hered 93:77–78
Wakelyn PJ, Bertoniere NR, French AD, Thibodeaux DP, Triplett BA, Roussell MA, Goynes WRJ, Edward JV, Hunter L, McAlister DD (2010) Cotton fiber chemistry and technology. In: Wakelyn PJ (ed) International fiber chemistry and technology. New York, USA: CRC Press, Taylor and Francis Group
Wang B, Guo W, Zhu X, Wu Y, Huang N, Zhang T (2006) QTL mapping of fiber quality in an elite hybrid derived-RIL population of Upland cotton. Euphytica 152:367–378
Wang J, Kucukoglu M, Zhang L, Chen P, Decker D, Nilsson O, Jones B, Sandberg G, Zheng B (2013) The Arabidopsis LRR-RLK, PXC1, is a regulator of secondary wall formation correlated with the TDIF-PXY/TDR-WOX4 signaling pathway. BMC Plant Biol 13:94
Wu TD, Nacu S (2010) Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26:873–881
Xu SL, Rahman A, Baskin TI, Kieber JJ (2008) Two leucine-rich repeat receptor kinases mediate signaling, linking cell wall biosynthesis and ACC synthase in Arabidopsis. Plant Cell 20:3065–3079
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134
Yu J, Zhang K, Li S, Yu S, Zhai H, Wu M, Li X, Fan S, Song M, Yang D, Li Y, Zhang J (2013) Mapping quantitative trait loci for lint yield and fiber quality across environments in a Gossypium hirsutum × Gossypium barbadense backcross inbred line population. Theor Appl Genet 126:275–287
Zhang K, Zhang J, Ma J, Tang S, Liu D, Teng Z, Liu D, Zhang Z (2012) Genetic mapping and quantitative trait locus analysis of fiber quality traits using a three-parent composite population in upland cotton (Gossypium hirsutum L.). Mol Breed 29:335–348
Zhang T, Qian N, Zhu X, Chen H, Wang S, Mei H, Zhang Y (2013) Variations and transmission of QTL alleles for yield and fiber qualities in upland cotton cultivars developed in China. PLoS One 8:e57220
Zhang T, Hu Y, Jiang W, Fang L, Guan X, Chen J, Zhang J, Saski CA, Scheffler BE, Stelly DM, Hulse-Kemp AM, Wan Q, Liu B, Liu C, Wang S, Pan M, Wang Y, Wang D, Ye W, Chang L, Zhang W, Song Q, Kirkbride RC, Chen X, Dennis E, Llewellyn DJ, Peterson DG, Thaxton P, Jones DC, Wang Q, Xu X, Zhang H, Wu H, Zhou L, Mei G, Chen S, Tian Y, Xiang D, Li X, Ding J, Zuo Q, Tao L, Liu Y, Li J, Lin Y, Hui Y, Cao Z, Cai C, Zhu X, Jiang Z, Zhou B, Guo W, Li R, Chen ZJ (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotech 33:531–537
Acknowledgments
This project was supported by USDA-ARS CRIS project 6054-21000-017-00D. Our appreciation goes to Data2Bio and BGI Americas for their excellent support in library construction, genome wide RNA and DNA sequencing, respectively. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture that is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The authors declare that the experiments comply with the current laws of the country in which they were carried out.
Additional information
Communicated by Y. Xu.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2016_2684_MOESM2_ESM.docx
Figure S2. SNPs per Mb in whole genome sequence data. The 26 chromosomes of Gossypium hirsutum TM-1 draft reference genome are presented (DOCX 198 kb)
122_2016_2684_MOESM3_ESM.docx
Figure S3. QTLs for fiber bundle strength (FBS), short fiber index (SFI) and upper-half mean length (UHML) identified in F3 population derived from cross of MD90ne x MD52ne. QTLs were detected by interval mapping using MapQTL6 software. The LOD scores were plotted against centiMorgans (cM) on the chromosome (DOCX 127 kb)
122_2016_2684_MOESM6_ESM.xlsx
Table S2. Sub-genome (Dt) specific primer names and their sequences used in this study for RT-qPCR validation (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Islam, M.S., Zeng, L., Thyssen, G.N. et al. Mapping by sequencing in cotton (Gossypium hirsutum) line MD52ne identified candidate genes for fiber strength and its related quality attributes. Theor Appl Genet 129, 1071–1086 (2016). https://doi.org/10.1007/s00122-016-2684-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2684-4