Abstract
Key message
Mapping-by-sequencing and SNP marker analysis were used to fine map the Ligon-lintless-1 ( Li 1 ) short fiber mutation in tetraploid cotton to a 255-kb region that contains 16 annotated proteins.
Abstract
The Ligon-lintless-1 (Li 1 ) mutant of cotton (Gossypium hirsutum L.) has been studied as a model for cotton fiber development since its identification in 1929; however, the causative mutation has not been identified yet. Here we report the fine genetic mapping of the mutation to a 255-kb region that contains only 16 annotated genes in the reference Gossypium raimondii genome. We took advantage of the incompletely dominant dwarf vegetative phenotype to identify 100 mutants (Li 1 /Li 1 ) and 100 wild-type (li 1 /li 1 ) homozygotes from a mapping population of 2567 F2 plants, which we bulked and deep sequenced. Since only homozygotes were sequenced, we were able to use a high stringency in SNP calling to rapidly narrow down the region harboring the Li 1 locus, and designed subgenome-specific SNP markers to test the population. We characterized the expression of all sixteen genes in the region by RNA sequencing of elongating fibers and by RT-qPCR at seven time points spanning fiber development. One of the most highly expressed genes found in this interval in wild-type fiber cells is 40-fold under-expressed at the day of anthesis (DOA) in the mutant fiber cells. This gene is a major facilitator superfamily protein, part of the large family of proteins that includes auxin and sugar transporters. Interestingly, nearly all genes in this region were most highly expressed at DOA and showed a high degree of co-expression. Further characterization is required to determine if transport of hormones or carbohydrates is involved in both the dwarf and lintless phenotypes of Li 1 plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
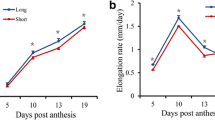
The Ligon-lintless-1 (Li 1 ) short fiber mutant of cultivated Upland cotton (Gossypium hirsutum L.) was originally identified in 1929 and was among the phenotypic markers used to establish the first linkage groups in this important fiber crop (Kohel 1972; Narbuth and Kohel 1990). Beyond its role in the foundation of cotton genetics and genomics, Li 1 has long been used as a model for understanding fiber cell elongation, a trait of great interest to both cotton breeders and cell biologists (Bolton et al. 2010; Kim and Triplett 2001; Triplett et al. 1989; Wang et al. 2013). While wild-type cotton fiber cells of G. hirsutum cv. DP5690 impressively elongate up to 30 mm in 4 weeks of development with fastest elongation around 8 days post anthesis (DPA), the Li 1 fiber cells only extend about 3 mm when fully mature (Gilbert et al. 2013). The single dominant gene that confers this qualitative short fiber phenotype has previously been mapped to a region of chromosome 22 that has also been repeatedly identified in studies of quantitative cotton fiber traits, including quantitative trail loci (QTLs) for fiber length, fiber uniformity and yield of seed cotton (Chee et al. 2005; Fang et al. 2014; Jiang et al. 1998; Karaca et al. 2002; Rong et al. 2005; Yu et al. 2013). Despite many approaches including identification of differentially expressed proteins and transcripts, the number of candidate genes for the Li 1 mutation remains very high (Ding et al. 2014; Gilbert et al. 2013; Liu et al. 2012; Zhao et al. 2009).
Although a reference genome sequence for the cultivated tetraploid cotton has not yet been published, reference genomes for two related diploids, Gossypium raimondii Ulbr. and Gossypium arboreum L. are available (Li et al. 2014; Paterson et al. 2012). Allotetraploid cotton contains A and D subgenomes which are closely related to the genome of the extant diploid G. arboreum (A2 genome) and the genome of G. raimondii (D5 genome), respectively (Wendel and Cronn 2003). We were previously able to significantly narrow the list of candidate genes for a different fiber mutant, Ligon-lintless-2 (Li 2 ), by using the G. raimondii reference sequence, super bulked segregant sequencing (sBSAseq), bioinformatics and traditional fine mapping approaches (Thyssen et al. 2014a). Our current objective was to make similar progress towards the identification of the causative Li 1 mutation using a pseudo-tetraploid reference genome consisting of the reference sequences of the extant diploids.
We took advantage of the long known incomplete dominance of the pleiotropic vegetative phenotypes of Li 1 plants to select and sequence bulks of Li 1 mutant and wild-type homozygotes from a large segregating population (Kohel 1972). This enabled us to confidently identify linked single nucleotide polymorphisms (SNPs) at a high allele frequency, which defined the genomic region containing the Li 1 locus and provided us with markers to test on the F2 mapping population. Ultimately, we were able to define a 255-kb region with only 16 candidate genes, which include a cluster of co-expressed genes, including several mitochondria-targeted genes, and a dramatically under-expressed small molecule transport protein from the major facilitator superfamily.
Materials and methods
Plant materials
The plants and populations used in this study were all described previously (Gilbert et al. 2013). Briefly, near isogenic lines (NIL) of Li 1 mutant and its wild-type G. hirsutum cv. DP5690 were generated by five generations of backcrossing and nine generations of self-pollination with single seed descent to introgress the Li 1 mutation into the DP5690 background. These parental plant lines were grown for mRNA isolation in New Orleans, LA in 2013. A segregating population of 2567 F2 progeny was grown in Stoneville, MS in 2012. Standard conventional field practices were followed at both locations.
RNA isolation, Illumina sequencing and RT-qPCR
Three biological replicates of fibers from different developmental time points were collected from the field in New Orleans, LA. Total RNA from 8-DPA fiber cells were Illumina sequenced by Data2Bio LLC. (Ames, IA) with paired 101-bp reads which are available in the SRA database at NCBI with BioProject accession PRJNA273732. Total RNA from 0, 3, 5, 8, 12, 16, and 20-DPA fiber cells were converted to cDNA and subjected to reverse transcription quantitative polymerase chain reaction (RT-qPCR) as described elsewhere (Naoumkina et al. 2014). Primer sequences are included as Table S1.
Super bulked segregant analysis sequencing (sBSAseq)
The incomplete dominance of the dwarf phenotype of Li 1 plants (Fig. 1 and Fig. S1) allowed us to score homozygosity of the segregating F2 progeny at the Li 1 locus. Based on this phenotype, 100 Li 1 /Li 1 and 100 li 1 /li 1 (wild-type) plants were randomly selected from the F2 population of 2567 individuals to be bulked and sequenced according to a sBSAseq approach (Michelmore et al. 1991; Takagi et al. 2013). Total genomic DNA from each bulk was Illumina sequenced by Data2Bio LLC with paired 101-bp reads.
Identification of diverse genomic regions
We aligned the sBSAseq total genomic reads, using GSNAP software, to a pseudo-reference genome for G. hirsutum that consisted of the 13 reference chromosomes of G. arboreum and 13 chromosomes of the G. raimondii genome which we present as the A and D subgenomes of G. hirsutum, respectively (Jiang et al. 1998; Paterson et al. 2012; Wu and Nacu 2010). We used InterSNP software at three different minor allele frequency (MAF) thresholds, 0.1, 0.2 and 0.3, to call SNPs between the wild-type and mutant bulk sequences (Page et al. 2014). We generated histograms by counting the number of SNPs in 1-Mb and 10-kb intervals.
Differential gene expression
We carried out differential gene expression of RNAseq reads as described elsewhere (Naoumkina et al. 2014, 2015). All reads were aligned to the reference G. raimondii genome following the PolyCat pipeline (Page et al. 2013). PolyCat software uses a database of homeoSNPs to categorize tetraploid reads into subgenomes. We made two adjustments to the PolyCat pipeline, as reported previously: (1) we only counted exonic reads; (2) we used the ratio of A-assigned to D-assigned reads to proportionately divide the total number of mapped reads for each gene to ensure that unassigned reads contribute to the total expression of genes (Thyssen et al. 2014a). The data normalization and ANOVA process were conducted as previously described (Naoumkina et al. 2014). In this paper, we only present RNAseq expression for candidate genes. All the significantly differentially expressed genes are reported elsewhere (Naoumkina et al. 2015).
Subgenome specific primer design
Manual inspection of read alignments in sBSAseq data was used to identify true SNPs and nearby homeoSNPs. We designed subgenome-specific SNP primers pairs essentially as described previously (Thyssen et al. 2014a). Each forward primer ends with the mutant allele SNP, while each reverse primer ends with the D-subgenome homeoSNP. Both primers contain an additional mismatch at the third base from the 3’ end, which increases annealing temperature stringency (Drenkard et al. 2000). Primer sequences are included as Table S2.
Mapping population
The 2567 F2 plants used in this study were previously scored for phenotypes and by simple sequence repeat (SSR) markers (Gilbert et al. 2013). The newly developed SNP markers were validated by running qPCR reactions on parental NILs and F1 plants as described previously (Thyssen et al. 2014a). The flanking SSR markers, C2-034C and DPL0489, were used to identify 85 informative plants with recombinational break points between the markers (Fig. 3). Since Li 1 is a dominant mutation, only plants where one flanking marker was wt/wt and the other was Li 1 /wt were considered informative. These plants were tested with the new SNP markers and scored as either Li 1 /wt or wt/wt based on the presence or absence of a qPCR product with a Ct value that matched the control parental and F1 plants. SNP marker genotypes for the non-informative plants were not imputed but were simply treated as missing data. The SNP marker CFB5857 was tested on the entire population and scored as a dominant marker. A genetic linkage map was constructed in JoinMap with default parameters and a LOD score of 10 (Van Ooijen 2006).
Results
Identification of the genomic region containing the Li 1 locus from sBSAseq
Since the dwarf, twisted stem and wrinkled leaf vegetative phenotypes of the Li 1 mutation are incompletely dominant in the DP5690 background in both the greenhouse (Fig. 1) and the field (Fig. S1), we were able to select only homozygous plants for sBSAseq. This enabled us to set a high allele frequency threshold for SNP calling. If heterozygotes were present in the dominant pool, we would expect reads from both alleles in that pool (Fig. 2). After aligning the reads to a reference pseudo-G. hirsutum genome composed of the A genome species G. arboreum and the D-genome diploid G. raimondii, we called SNPs at increasing minor allele frequencies (MAF). At a MAF of 0.1, there is a striking single peak of 422 SNPs per Mb in the 14th Mb of Chromosome 12 of the D-genome, which corresponds to Chr. 22 of the tetraploid, the expected location of Li 1 based on earlier reports (Gilbert et al. 2013; Karaca et al. 2002; Rong et al. 2005). We took this density of SNPs to indicate a diverse region closely linked to the Li 1 mutation that accompanied Li 1 during the process of NIL development. We developed qPCR based SNP markers to interrogate these polymorphisms in the segregating F2 population.
High confidence SNPs in mapped sBSAseq reads. The 13 reference G. arboreum chromosomes and 13 reference G. raimondii chromosomes are presented as the A and D subgenomes of tetraploid G. hirsutum. The region around the peak on D Chr 12 is expanded for detail; with the locations of the flanking SNP markers (CFB5856, CFB5857) and the Li 1 locus labeled (see Fig. 3). Black indicates a minor allele frequency (MAF) threshold of 0.1, while dark gray indicates MAF <0.2 and light gray MAF <0.3
Segregation of markers in 2567 F2 progeny
Analysis of SNP markers on the segregating population significantly narrowed the genetic interval that contains the Li 1 locus (Fig. 3). Our new genetic map shows good correspondence with the physical map of the homologous chromosome from G. raimondii, and confines the Li 1 locus to an interval of 255-kb.
Li 1 locus in G. hirsutum based on 2567 F2 progeny aligned with the physical map of G. raimondii. SNP and SSR markers associated with the Li 1 gene are shown on G. hirsutum chromosome 22 and on G. raimondii chromosome 12. Genetic map locations are shown in centiMorgans (cM) and physical locations are shown in base pairs (bp)
Differential expression of genes on the interval of the Li 1 locus
The region of reference G. raimondii sequence that is bound by SNP markers CFB5856 and CFB5857 contains only 16 annotated genes, of which only 8 were detected by RNAseq in 8-DPA fibers (Table 1). Three of these were significantly over-expressed in Li 1 (Gorai.012G086600 “PPR”, Gorai.012G086800 “TOM”, and Gorai.012G086900 “DCD”) and two genes were significantly under-expressed (Gorai.012G086000 “DUF” and Gorai.012G086100 “MFS”).
RT-qPCR of candidate genes during fiber development
We tested all sixteen annotated genes in the Li 1 interval by RT-qPCR across the development of cotton fiber cells (Fig. S2 and Fig. S3). In wild-type fiber cells, most of the 8 highly expressed genes were most highly expressed during lint fiber initiation at DOA. The expression of these genes was low at 3- and 5-DPA, with increased expression at the peak of elongation, 8-DPA (Fig. S2). The 8 genes which were not detected by RNAseq in 8-DPA fiber also mostly had their highest expression at DOA, with somewhat increasing expression later in development, at 16 or 20-DPA, though only to levels approximately ten-fold less than the highly expressed genes (Fig. S3). We computed the simple Pearson correlation coefficients for the expression of the 16 genes in mutant and wild-type fibers across the developmental stages as measured by RT-qPCR (Fig. 4). It is clear that two clusters of highly correlated genes are present in the wild-type fibers, with three of the very low expressed genes correlating well with each other, and the remaining 13 genes forming a second cluster of co-expression (Fig. 4). In mutant fibers, four genes lose their correlation with each other and with their neighboring genes (Fig. 4). Namely, Gorai.012G086100 “MFS”, Gorai.012G086400 “SEC”, Gorai.012G086800 “TOM”, and Gorai.012G086900 “DCD” have aberrant expression, most strikingly a dramatically reduced expression in lint fiber initials at DOA (Fig. S2).
Correlation in gene expression at the Li 1 locus. Data presented in Figures S2 and S3 from RT-qPCR of the sixteen candidate genes across the seven developmental time points were used to calculate pairwise Pearson correlation coefficients for each gene from Li 1 and wild-type fiber cells. Genes are labeled with “wt” or “Li 1 ” and with the G. raimondii accession given in Table 1. Black borders locate the correlation of a single gene between Li 1 mutant and wild-type fiber cells. Genes mentioned in the text are bold
Discussion
Mapping the genetic locus by sequencing of two homozygote pools
As genetic resources for cotton continue to improve and costs for sequencing decrease, mapping-by-sequencing will increasingly become the technique of choice for the identification of genetic loci that control phenotypes. We took advantage of the well annotated G. raimondii reference genome and recently released G. arboreum genome to construct a pseudo-reference tetraploid genome for G. hirsutum (Li et al. 2014; Paterson et al. 2012). We also benefitted from the incomplete dominance of the pleiotropic dwarf, twisted stem and wrinkled leaf phenotypes of the Li 1 short fiber mutation to limit our sequencing to two pools of homozygotes, which significantly simplified the SNP-calling bioinformatic analysis. In the future, when mapping genes with complete dominance, we will consider testing the segregation of the progeny of several dominant F2 plants before selecting plants for total genomic sBSAseq to ensure that only homozygotes are included in the bulks. While progeny testing adds time, it increases the expected allele frequency of the causative polymorphism from 0.67 to 1.0 (Mendel 1941). In the present study, at a MAF of 0.3, the SNP-calling software generates considerable background, while a MAF of 0.1 revealed an unambiguous region (Fig. 2). Even with this advantage, we nevertheless failed to discover any polymorphisms on the interval between our two closest flanking SNP markers, CFB5856 and CFB5857 in our short read sequencing data. This may mean that the causative Li 1 mutation is a structural rearrangement, transposon insertion, copy number variation or resides in a region of low sequence complexity or in part of the region that is missing from the reference sequence. Indeed, this region of the G. raimondii assembly contains several long (up to 2-kb) runs of Ns (Paterson et al. 2012). Although there is good colinearity for most of the tetraploid D-subgenome with the G. raimondii genome, we cannot exclude the possibility of significant differences between genomes in the proposed Li 1 interval, including differences in gene content. We look forward to a true G. hirsutum reference genome and longer sequencing read lengths to help resolve these issues.
Traditional fine genetic mapping
Mapping-by-sequencing allowed us to rapidly identify a region of a few Mb that contained a cluster of diversity that differentiates the progenitor of Li 1 from the DP5690 cultivar that we used to generate the NILs. Since diversity between cultivars is not uniformly distributed, there was no warranty that the causative Li 1 mutation would be within the greatest density of polymorphism, although it should be nearby. For this reason we designed markers in the vicinity, but not exclusively within the peak of diversity (Fig. 2). We tested segregation of our new SNP markers and the Li 1 phenotype in the large F2 population of 2567 individuals to narrow the region to 255-kb and 16 candidate genes. However, once flanking markers are confidently established, only a small subset of plants needs to be tested with the new markers to refine the interval (Blair et al. 2003).
Li 1 candidate gene expression
We tested the expression of all 16 annotated G. raimondii genes in the Li 1 locus during fiber development expecting to pick a candidate based on differences in elongation stage (~8-DPA) expression. However, we were surprised to observe that nearly all the candidate genes are most highly expressed at DOA, a time point that is traditionally associated with lint fiber initiation rather than elongation (Basra and Malik 1984). Classically, it is though that the shorter fuzz fibers initiate later than lint fibers, around 5-DPA, and the Li 1 fiber phenotype was originally described as lintless but fuzzy (Kohel 1972; Lang 1938). Differential gene activity at DOA might therefore explain the lack of lint but presence of fuzz on Li 1 seeds. However, it is not clear whether the fibers on Li 1 are lint fibers, fuzz fibers or a combination of both lint and fuzz fibers (Triplett et al. 1989).
Correlated expression of neighboring genes
Because of the highly correlated expression of many genes in the Li 1 locus interval, and potential unknown differences between the G. raimondii reference genome and the D-subgenome of G. hirsutum, it is so far impossible to pick a single candidate gene. By identifying orthologous genes in Arabidopsis, it is clear that the Li 1 locus preserves some syntenic relationships of gene order (Table 1). Operon-like clusters of eukaryotic genes have been observed for triterpene synthesis in Arabidopsis and oat but were not found in Medicago (Field and Osbourn 2008; Naoumkina et al. 2010). Co-regulated genes in developing Arabidopsis root tissues show chromosomal clustering and recently a global analysis of Arabidopsis gene expression confirmed that neighboring genes are co-expressed (Birnbaum et al. 2003; Williams and Bowles 2004). The most significant contribution to neighboring gene co-expression was found to be orientation of gene pairs. Parallel or divergent orientations resulted in higher levels of co-expression, while the convergent orientation of genes reduced their co-expression, suggesting the effect of shared regulatory elements (Williams and Bowles 2004). However, the orientation of genes in the Li 1 locus does not obviously explain their coordinated expression since DCD and ENY are highly correlated and are in a convergent orientation (Fig. 4; Table 1).
Most intriguing for the present study is the report that AT3G27080, the ortholog of “TOM” Gorai.012G086800, a subunit of the mitochondrial outer membrane translocase, is part of a chromosomal region in Arabidopsis that is unusually rich in mitochondrial genes (Elo et al. 2003). This region is also present in rice, and we have observed that the genes adjacent to TOM: Gorai.012G086900 “DCD” and Gorai.012G087000 “ENY” are orthologous to AT3G27090 and AT3G27100 which neighbor AT3G27080 (Elo et al. 2003). Additionally, Gorai.012G086600 “PPR” is predicted to be a mitochondria-targeted RNA binding protein that may affect organellar transcript editing or stability (Barkan and Small 2014). We have recently shown that a different short fiber mutant, Ligon-lintless-2 (Li 2 ), has altered mitochondrial gene activity during fiber elongation, so it is interesting to observe a cluster of co-expressed mitochondrial genes in the Li 1 candidate interval (Thyssen et al. 2014b).
Major facilitator superfamily
We also consider the gene Gorai.012G086100 “MFS” as an attractive candidate for Li 1 . This gene is a member of the major facilitator superfamily of transport proteins, a large family that includes sugar, amino acid and hormone transporters (Pao et al. 1998; Remy et al. 2013). A good match (98 % nucleotide identity) for MFS is NCBI EST DW231799 which has previously been listed as an Li 1 candidate gene based on RNAseq of leaves (Ding et al. 2014). DW231799 was shown to have elevated transcript abundance in leaf and reduced expression in fibers of Li 1 plants (Ding et al. 2014). We observed that MFS is highly expressed at DOA in wild-type fibers but is 40-fold under-expressed in Li 1 fiber initials (Fig. S2). At 8-DPA, wild-type expression of MFS rebounds to about a quarter of the expression level at DOA, but expression in Li 1 fibers remains very low (Fig. S2). The Li 1 vegetative phenotypes mimic the exposure of cotton to low dosages of 2,4-D, a synthetic auxin herbicide, which has long been used to identify mutants of polar auxin transport (Bennett et al. 1996; Sciumbato et al. 2004). Injury to cotton plants by low dosages of auxin-type herbicides is characterized by bending and twisting of stems and wrinkling of leaves, although fiber length is not affected (Sciumbato et al. 2004; Smith and Wiese 1972). However, transgenic over-expression of auxin synthesis during lint fiber initiation resulted in an increase in lint yield (Zhang et al. 2011). Additionally, several important auxin transporters that also belong to the major facilitator superfamily were shown to be under-expressed in developing fibers of Li 1 plants, suggesting a role for defective polar auxin transport in the Li 1 fiber phenotype (Wang et al. 2013). Taken together, it is intriguing to speculate that over-expression of MFS in the leaves of Li 1 plants contributes to the vegetative phenotypes, while under-expression of MFS in the fiber initials results in the short fiber phenotype. It is not possible to determine the substrate specificity of MFS by sequence analysis alone (Pao et al. 1998). However, the action of sugar transporters of the major facilitator superfamily can also produce dwarf plants in Arabidopsis and affect cotton fiber cell elongation (Gottwald et al. 2000; Ruan et al. 2001). Further work is required to determine if the aberrant transport of hormones or metabolites in fact underlies the phenotypes of the Ligon-lintless-1 short fiber mutant.
Author contribution statement
GNT, DDF, and MN conceived and designed the experiment. GT analyzed the sequencing data, designed the SNP markers and wrote the paper. DDF oversaw the project. RT developed the NILs and grew the F2 population. CF and PL conducted the SNP and SSR marker analysis. MN performed and analyzed the RT-qPCR experiments. All authors read and approved the manuscript.
References
Barkan A, Small I (2014) Pentatricopeptide repeat proteins in plants. Annu Rev Plant Biol 65:415–442
Basra AS, Malik C (1984) Development of the cotton fiber. Int Rev Cytol 89:65–113
Bennett MJ, Marchant A, Green HG, May ST, Ward SP, Millner PA, Walker AR, Schulz B, Feldmann KA (1996) Arabidopsis AUX1 gene: a permease-like regulator of root gravitropism. Science 273:948–950
Birnbaum K, Shasha DE, Wang JY, Jung JW, Lambert GM, Galbraith DW, Benfey PN (2003) A gene expression map of the Arabidopsis root. Science 302:1956–1960
Blair MW, Garris AJ, Iyer AS, Chapman B, Kresovich S, McCouch SR (2003) High resolution genetic mapping and candidate gene identification at the xa5 locus for bacterial blight resistance in rice (Oryza sativa L.). Theor Appl Genet 107:62–73
Bolton JJ, Soliman KM, Wilkins TA, Jenkins JN (2010) Aberrant expression of critical genes during secondary cell wall biogenesis in a cotton mutant, Ligon lintless-1 (Li-1). Comp Funct Genomics 2009:659301
Chee PW, Draye X, Jiang C-X, Decanini L, Delmonte TA, Bredhauer R, Smith CW, Paterson AH (2005) Molecular dissection of phenotypic variation between Gossypium hirsutum and Gossypium barbadense (cotton) by a backcross-self approach: III. Fiber length. Theor Appl Genet 111:772–781
Ding M, Jiang Y, Cao Y, Lin L, He S, Zhou W, Rong J (2014) Gene expression profile analysis of Ligon lintless-1 (Li 1 ) mutant reveals important genes and pathways in cotton leaf and fiber development. Gene 535:273–285
Drenkard E, Richter BG, Rozen S, Stutius LM, Angell NA, Mindrinos M, Cho RJ, Oefner PJ, Davis RW, Ausubel FM (2000) A simple procedure for the analysis of single nucleotide polymorphisms facilitates map-based cloning in Arabidopsis. Plant Physiol 124:1483–1492
Elo A, Lyznik A, Gonzalez DO, Kachman SD, Mackenzie SA (2003) Nuclear genes that encode mitochondrial proteins for DNA and RNA metabolism are clustered in the Arabidopsis genome. Plant Cell 15:1619–1631
Fang DD, Jenkins JN, Deng DD, McCarty JC, Li P, Wu J (2014) Quantitative trait loci analysis of fiber quality traits using a random-mated recombinant inbred population in Upland cotton (Gossypium hirsutum L.). BMC Genom 15:397
Field B, Osbourn AE (2008) Metabolic diversification—independent assembly of operon-like gene clusters in different plants. Science 320:543–547
Gilbert MK, Turley RB, Kim HJ, Li P, Thyssen G, Tang Y, Delhom CD, Naoumkina M, Fang DD (2013) Transcript profiling by microarray and marker analysis of the short cotton (Gossypium hirsutum L.) fiber mutant Ligon lintless-1 (Li 1 ). BMC Genom 14:403
Gottwald JR, Krysan PJ, Young JC, Evert RF, Sussman MR (2000) Genetic evidence for the in planta role of phloem-specific plasma membrane sucrose transporters. Proc Natl Acad Sci 97:13979–13984
Jiang C-X, Wright RJ, El-Zik KM, Paterson AH (1998) Polyploid formation created unique avenues for response to selection in Gossypium (cotton). Proc Natl Acad Sci 95:4419–4424
Karaca M, Saha S, Jenkins J, Zipf A, Kohel R, Stelly D (2002) Simple sequence repeat (SSR) markers linked to the Ligon Lintless (Li 1 ) mutant in cotton. J Hered 93:221–224
Kim HJ, Triplett BA (2001) Cotton fiber growth in planta and in vitro. Models for plant cell elongation and cell wall biogenesis. Plant Physiol 127:1361–1366
Kohel R (1972) Linkage tests in Upland cotton, Gossypium hirsutum L. II. Crop Sci 12:66–69
Lang A (1938) The origin of lint and fuzz hairs of cotton. J Agric Res 56:507–521
Li F, Fan G, Wang K, Sun F, Yuan Y, Song G, Li Q, Ma Z, Lu C, Zou C (2014) Genome sequence of the cultivated cotton Gossypium arboreum. Nat Genet 46:567–572
Liu K, Sun J, Yao L, Yuan Y (2012) Transcriptome analysis reveals critical genes and key pathways for early cotton fiber elongation in Ligon lintless-1 mutant. Genomics 100:42–50
Mendel G (1941) Versuche über Pflanzen-Hybriden. Theor Appl Genet 13:221–268
Michelmore RW, Paran I, Kesseli R (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci 88:9828–9832
Naoumkina MA, Modolo LV, Huhman DV, Urbanczyk-Wochniak E, Tang Y, Sumner LW, Dixon RA (2010) Genomic and coexpression analyses predict multiple genes involved in triterpene saponin biosynthesis in Medicago truncatula. Plant Cell 22:850–866
Naoumkina M, Thyssen G, Fang DD, Hinchliffe DJ, Florane C, Yeater KM, Page JT, Udall JA (2014) The Li 2 mutation results in reduced subgenome expression bias in elongating fibers of allotetraploid cotton (Gossypium hirsutum L.). PLoS One 9:e90830
Naoumkina M, Thyssen GN, Fang DD (2015) RNA-seq analysis of short fiber mutants Ligon-lintless -1 (Li 1 ) and – 2 (Li 2 ) revealed important role of aquaporins in cotton (Gossypium hirsutum L.) fiber elongation. BMC Plant Biol 15:65
Narbuth E, Kohel R (1990) Inheritance and linkage analysis of a new fiber mutant in cotton. J Hered 81:131–133
Page JT, Gingle AR, Udall JA (2013) PolyCat: a resource for genome categorization of sequencing reads from allopolyploid organisms. G3 Genes Genom Genet 3:517–525
Page JT, Liechty ZS, Huynh MD, Udall JA (2014) BamBam: genome sequence analysis tools for biologists. BMC Res Notes 7:829
Pao SS, Paulsen IT, Saier MH (1998) Major facilitator superfamily. Microbiol Mol Biol Rev 62:1–34
Paterson AH, Wendel JF, Gundlach H, Guo H, Jenkins J, Jin D, Llewellyn D, Showmaker KC, Shu S, Udall J et al (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492:423–427
Remy E, Cabrito TR, Baster P, Batista RA, Teixeira MC, Friml J, Sá-Correia I, Duque P (2013) A major facilitator superfamily transporter plays a dual role in polar auxin transport and drought stress tolerance in Arabidopsis. Plant Cell 25:901–926
Rong J, Pierce GJ, Waghmare VN, Rogers CJ, Desai A, Chee PW, May OL, Gannaway JR, Wendel JF, Wilkins TA (2005) Genetic mapping and comparative analysis of seven mutants related to seed fiber development in cotton. Theor Appl Genet 111:1137–1146
Ruan Y-L, Llewellyn DJ, Furbank RT (2001) The control of single-celled cotton fiber elongation by developmentally reversible gating of plasmodesmata and coordinated expression of sucrose and K+ transporters and expansin. Plant Cell 13:47–60
Sciumbato AS, Chandler JM, Senseman SA, Bovey RW, Smith KL (2004) Determining exposure to auxin-like herbicides. I. Quantifying injury to cotton and soybean 1. Weed Technol 18:1125–1134
Smith DT, Wiese AF (1972) Cotton response to low rates of 2, 4-D and other herbicides. Tex Agric Exp Stn Bull B-1120
Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74:174–183
Thyssen GN, Fang DD, Turley RB, Florane C, Li P, Naoumkina M (2014a) Next generation genetic mapping of the Ligon-lintless-2 (Li 2 ) locus in upland cotton (Gossypium hirsutum L.). Theor Appl Genet 127:2183–2192
Thyssen GN, Song X, Naoumkina M, Kim H-J, Fang DD (2014b) Independent replication of mitochondrial genes supports the transcriptional program in developing fiber cells of cotton (Gossypium hirsutum L.). Gene 544:41–48
Triplett BA, Busch WH, Goynes WR Jr (1989) Ovule and suspension culture of a cotton fiber development mutant. In Vitro Cell Dev Biol 25:197–200
Van Ooijen J (2006) JoinMap 4 Software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen
Wang M-Y, Zhao P-M, Cheng H-Q, Han L-B, Wu X-M, Gao P, Wang H-Y, Yang C-L, Zhong N-Q, Zuo J-R (2013) The cotton transcription factor TCP14 functions in auxin-mediated epidermal cell differentiation and elongation. Plant Physiol 162:1669–1680
Wendel JF, Cronn RC (2003) Polyploidy and the evolutionary history of cotton. Adv Agron 78:139–186
Williams EJ, Bowles DJ (2004) Coexpression of neighboring genes in the genome of Arabidopsis thaliana. Genome Res 14:1060–1067
Wu TD, Nacu S (2010) Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26:873–881
Yu J, Zhang K, Li S, Yu S, Zhai H, Wu M, Li X, Fan S, Song M, Yang D (2013) Mapping quantitative trait loci for lint yield and fiber quality across environments in a Gossypium hirsutum × Gossypium barbadense backcross inbred line population. Theor Appl Genet 126:275–287
Zhang M, Zheng X, Song S, Zeng Q, Hou L, Li D, Zhao J, Wei Y, Li X, Luo M (2011) Spatiotemporal manipulation of auxin biosynthesis in cotton ovule epidermal cells enhances fiber yield and quality. Nature Biotech 29:453–458
Zhao P-M, Wang L-L, Han L-B, Wang J, Yao Y, Wang H-Y, Du X-M, Luo Y-M, Xia G-X (2009) Proteomic identification of differentially expressed proteins in the Ligon lintless mutant of upland cotton (Gossypium hirsutum L.). J Proteome Res 9:1076–1087
Acknowledgments
This project was financially supported by the USDA-ARS CRIS project #6435-21000-017-0DD and Cotton Incorporated project #12-210. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U. S. Department of Agriculture that is an equal opportunity provider and employer.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Gore.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2015_2539_MOESM1_ESM.tif
Supplementary material 1 (TIFF 1362 kb) Fig. S1 Field grown plants show incomplete dominance of the Li 1 vegetative phenotypes. Homozygous wild-type, heterozygous and homozygous Li 1 plants are labeled
122_2015_2539_MOESM2_ESM.jpg
Supplementary material 2 (JPEG 2153 kb) Fig. S2 RT-qPCR expression of highly expressed Li 1 candidate genes during development of Li 1 and wild-type DP5690 G. hirsutum fiber cells. Error bars indicate standard deviation from three biological replicates. Along the x-axis, DPA indicates the number of days post anthesis, a measure of developmental time for cotton fiber cells
122_2015_2539_MOESM3_ESM.jpg
Supplementary material 3 (JPEG 2119 kb) Fig. S3 RT-qPCR expression of Li 1 candidate genes with no detectable expression at 8-DPA based on RNAseq. Error bars indicate standard deviation from three biological replicates. Along the x-axis, DPA indicated the number of days post anthesis
122_2015_2539_MOESM4_ESM.xlsx
Supplementary material 4 (XLSX 12 kb) Table S1 Primers for RT-qPCR of genes at the Li 1 locus. F—forward, R—reverse, DF—D-subgenome specific forward primer, Table S2 Primers for SNP loci near the Li 1 mutation. mF—mutant-specific forward, DR—D-subgenome-specific reverse primer
Rights and permissions
About this article
Cite this article
Thyssen, G.N., Fang, D.D., Turley, R.B. et al. Mapping-by-sequencing of Ligon-lintless-1 (Li 1 ) reveals a cluster of neighboring genes with correlated expression in developing fibers of Upland cotton (Gossypium hirsutum L.). Theor Appl Genet 128, 1703–1712 (2015). https://doi.org/10.1007/s00122-015-2539-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2539-4