Abstract
Key message
This study explored the genetic constitutions of several wheat- A. cristatum translocation lines and determined the effects of A. cristatum 6P chromosome segments on fertile tiller number in wheat.
Abstract
Progress in wheat breeding is hampered by a relatively narrow range of genetic variation. To overcome this hurdle, wild relatives of common wheat with superior agronomic traits are often used as donors of desirable genes in wheat-breeding programs. One of the successfully utilized wheat wild relatives is Agropyron cristatum (L.) Gaertn (2n = 4x = 28; genomes PPPP). We previously reported that WAT31-13 was a wheat-A. cristatum 5A-6P reciprocal translocation line with higher fertile tiller number and grain number per spike compared to common wheat. However, WAT31-13 was genetically unstable. In this study, we analyzed the 43 genetically stable progenies from WAT31-13 using genomic in situ hybridization, dual-color fluorescence in situ hybridization, and molecular markers. We classified them into three translocation types (TrS, TrL and TrA) and seven subtypes, and also pinpointed the translocation breakpoint. The genotypic data, combined with the phenotypes of each translocation type, enabled us to physically map agronomic traits to specific A. cristatum 6P chromosome arms or segments. Our results indicated that A. cristatum chromosome 6P played an important role in regulating fertile tiller number, and that positive and negative regulators of fertile tiller number existed on the A. cristatum chromosome arm 6PS and 6PL, respectively. By exploring the relationship between fertile tiller number and A. cristatum chromosome segment, this study presented a number of feasible approaches for creation, analysis, and utilization of wheat-alien chromosome translocation lines in genetic improvement of wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L., 2n = 6x = 42, genomes AABBDD) is one of the most important crops worldwide. A major challenge in modern agriculture is to breed elite wheat varieties with enhanced agronomic traits to meet the growing demands for food. However, common wheat exhibits a relatively narrow range of genetic variation, which has become a bottleneck for yield improvement (Dubcovsky and Dvorak 2007; Haudry et al. 2007; Reif et al. 2005). To broaden the genetic basis of common wheat, its wild relatives harboring superior agronomic traits have often been used in wheat breeding (Wang 2011). The tribe Triticeae contains around 400 species and 25 genera, conferring ample genetic diversity for wheat improvement (Mujeeb-Kazi et al. 2013). To date, more than 100 alien genes/QTLs conferring superior traits have been transferred into cultivated wheat (Bao et al. 2009; Cao et al. 2011; Faris et al. 2008; Friebe et al. 1990, 1996a; Fu et al. 2012; He et al. 2009; Hua et al. 2009; Jiang et al. 1994; Kang et al. 2011; Luo et al. 2009; Monneveux et al. 2003; Sharma et al. 1995; Singh et al. 1998; Wang et al. 2011). Producing wheat-alien species translocation lines and elucidating their genetic constitutions are key steps for effective transfer of desirable genes into common wheat (Chen et al. 1995, 2005; Friebe et al. 1992, 1996a; Gill et al. 2011; Klindworth et al. 2012; Larkin et al. 1995; Larson et al. 2012; Niu et al. 2011, 2014; Qi et al. 2011; Singh et al. 1998; Wang and Zhang 1996; Yu et al. 2009). Some translocation lines, especially 1BL∙1RS, 6VS∙6AL, and 7DL∙7Ag have played important roles in wheat improvement. A 1BL∙1RS translocation line was widely used in bread wheat-breeding programs throughout the world due to the presence of powdery mildew resistance gene Pm8 and rust resistance genes Sr31, Lr26, and Yr9 in 1RS (Hsam and Zeller 1997; Lukaszewski 2000; Mago et al. 2002; Singh et al. 1998). A 6VS∙6AL translocation line (Chen et al. 1995, 2005) has been widely used throughout the world (Duan et al. 1998; Liu et al. 1999; Qi et al. 1995). The 7DL∙7Ag translocation lines produced from Lophopyrum elongatum were reported to be valuable reservoirs of desirable genes conferring resistance to leaf rust, salinity, and waterlogging (Deal et al. 1999; Ma et al. 2000; McDonald et al. 2001; Niu et al. 2014).
Agropyron cristatum (L.) Gaertn (2n = 4x = 28; genomes PPPP) is a perennial wheatgrass that possesses many desirable traits such as enhanced fertile tiller number, high grain number per spike, and resistance to numerous diseases (Dewey 1984; Dong et al. 1992). It has long been considered a useful genetic resource for wheat improvement. The F1 hybrids were successfully obtained between common wheat cv. Fukuhokomugi (Fukuho) and A. cristatum accession Z559, followed by backcrossing with Fukuho for several generations (Li et al. 1995, 1998b; Li and Dong 1991, 1993). A series of disomic addition lines was obtained, and the 6P disomic addition line 4844-12 was one of them. This addition line had significantly higher grain number per spike and floret number per spikelet, as well as enhanced resistance to powdery mildew, compared to its wheat parent (Han et al. 2014; Li et al. 1997, 1998a, b; Luan et al. 2010; Wu et al. 2006). These results indicated the existence of desirable genes on the A. cristatum 6P chromosome. To produce germplasm useful in wheat-breeding programs, wheat-A. cristatum 6P translocation lines were then produced by both gametocidal chromosomes and ionizing radiation (Luan et al. 2010). WAT31-13 (M2 generation), produced from irradiated hybrid seeds (wheat-A. cristatum addition line/Gaocheng 8901), is a 5A-6P reciprocal translocation line with 44 chromosomes. Although it displays superior agronomic traits, it is genetically unstable (Luan et al. 2010).
To further localize these desirable genes on the A. cristatum 6P chromosome and acquire translocation materials with genetic stability, strict self-pollination was carried out over further generations of WAT31-13. Forty-three stable translocation lines in the M8 generation were selected. Surprisingly, most of them contained 6P translocation chromosome segments that were different from those in WAT31-13, although the various elite agronomic traits were retained. In this study, we not only examined the genomic constitutions of the 43 translocation lines by GISH and FISH, but also ascribed superior agronomic traits (especially the high fertile tiller number) to specific A. cristatum 6P chromosomal segments.
Materials and methods
Materials
WAT31-13 is a 5A-6P reciprocal translocation line (2n = 44) produced from irradiated hybrid seeds using the wheat-A. cristatum 6P disomic addition line 4844-12 as the female parent and common wheat Gaocheng 8901 (2n = 6x = 42, genomes AABBDD) as the male parent (Luan et al. 2010). In this study, 43 stable M8 translocation lines were obtained from the progenies of WAT31-13. Wheat-A. cristatum 6P disomic addition line 4844-12, common wheat Gaocheng 8901, and Fukuho were used as contrasting parents in this study. Wheat-A. cristatum 6P disomic addition line 4844-12 and translocation line WAT31-13 were originally produced in our laboratory. Seeds of Gaocheng 8901 and Fukuho were provided by the Institute of Crop Sciences, Chinese Academy of Agricultural Sciences.
GISH and FISH analysis
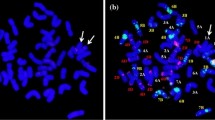
Genomic in situ hybridization (GISH) and dual-color fluorescence in situ hybridization (FISH) were carried out as previously described (Han et al. 2003). Genomic DNA was isolated using the CTAB method (Allen et al. 2006). A. cristatum genomic DNA (labeled with Dig-Nick-Translation Mix) and Fukuho genomic DNA were used as probe and blocker, respectively. Wheat and A. cristatum chromosomes were pseudo-colored as blue and red, respectively. Dual-color FISH was performed using pAs1 and pHvG39 plasmid DNAs as probes in all translocation subtypes, while probes CRW (centromeric retrotransposon of wheat) and pAcCR1 (centromeric retrotransposon of A. cristatum) were applied to analyze the centromere of the arm translocation chromosome (TrA) in the Subtype II. Probe pAs1 from Aegilops tauschii preferentially hybridizes to most D-genome chromosomes, and the probe pHvG39 could hybridize with A, B, and D genome chromosomes (Pedersen and Langridge 1997). Probe CRW was identified from a Triticum boeoticum library and closely associated with the centromeres (Liu et al. 2008). The A. cristatum centromere-specific probe pAcCR1 was developed using the wheat-A. cristatum 6PS ditelosomic lines in our laboratory. To develop the probe pAcCR1, microdissection and degenerate oligonucleotide-primed PCR (DOP-PCR) were conducted as previously described (Vega et al. 1994). All cytological images were taken under a Nikon Eclipse E600 fluorescence microscope and captured with a CCD camera.
Translocation breakpoint analyses
To determine the translocation breakpoints, a total of 25 wheat SSR markers physically or genetically mapped on wheat chromosome 5A were chosen (Somers et al. 2004; Sourdille et al. 2004). Common wheat ‘Chinese Spring’ (CS) nulli-tetrasomic lines (N5AT5B and N5AT5D), CS ditelosomic lines (Dt5AL and Dt5AS), and CS 5AL deletion lines (5AS10-0.98-1.00, 5AS3-0.75-0.98, 5AS1-0.40-0.75, C-5AS1-0.40, C-5AL12-0.35, 5AL12-0.35-0.57, 5AL10-0.57-0.78, 5AL17-0.78-0.87, and 5AL23-0.87-1.00) were applied to verify the chromosomal localization of wheat SSR markers. Nine A. cristatum-specific markers developed from EST sequences of A. cristatum mRNA transcriptome and seven A. cristatum-specific markers (For3-G02, For5-E08, For22-B10, For15-D06, For8-G11, For14-B02, and For22-E03) obtained from Luan et al. (2010) were chosen to distinguish different chromosomal regions of A. cristatum chromosome 6P. All wheat SSR primers were obtained from the Graingenes website (http://wheat.pw.usda.gov/ggpages/maps.shtml).
Evaluation of agronomic traits
Evaluation of traits was conducted in a field trial in Beijing with three replications in each of 2 years (2012 and 2013). For each replication, 20 grains of each line were evenly planted in 2.0 m rows, spaced 0.3 m apart. All the translocation lines were evaluated for a number of agronomic traits, including fertile tiller number, grain number per spike, spikelet number per spike, kernel number per spikelet, and thousand-kernel weight. Traits were measured on ten plants randomly selected from each line of each translocation subtype. Statistical analyses were conducted using the Statistical Analysis System version 9.2 (SAS Institute Inc., Cary, NC, USA), and the t-test was used to test the difference of the agronomic traits between each of the translocation subtypes and two common wheat parents (Gaocheng 8901 and Fukuho).
Results
Distinct translocation types were identified in the progenies of WAT31-13
We previously reported that the 5A-6P reciprocal translocation-addition line WAT31-13 (2n = 44) (M2 generation) harbored a number of superior agronomic traits but displayed genetic instability (Luan et al. 2010). By strict self-pollination by spike bagging for six generations, 43 translocation lines were acquired at the M8 generation. GISH was carried out at each generation to determine the genetic constitutions of individual plants. Plants that lost the A. cristatum chromosome segments were excluded from further investigation. Because plants at the M8 generation were all identical in genetic constitution relative to the M6 and M7 generations, we concluded that these 43 M8 lines were genetically stable. GISH indicated that there were three types of translocation chromosomes in these various lines (Table 1), with each line harboring a single or combinations of three translocation types (Table 2). As shown in Table 1, the small translocation chromosome (TrS) contained a small segment of A. cristatum chromosome (shown in red) of approximately 60 % of the distal portion of one chromosome arm, whereas the remainder of the chromosome including the centromere was from wheat (shown in blue). In contrast, the large translocation chromosome (TrL) contained a small segment of wheat chromosome (shown in blue) at the end of one arm with the rest of the chromosome from A. cristatum (shown in red). The arm translocation chromosome (TrA) seemed to include only the translocated arm found in the TrL type.
According to the translocation chromosomes they harbored, these 43 translocation lines were categorized into seven translocation subtypes (Table 2; Fig. 1). In Subtype I, there were 42 wheat chromosomes as well as one pair of TrSs and one pair of TrLs (Fig. 1a; Table 2). Subtype II contained 42 wheat chromosomes, a pair of TrSs and a pair of TrAs (Fig. 1b; Table 2). Subtype III contained 42 wheat chromosomes and a pair of TrLs (Fig. 1c; Table 2). Subtype IV contained 42 wheat chromosomes and a pair of TrSs (Fig. 1d; Table 2). In Subtype V, there were 40 wheat chromosomes and a pair of TrSs (Fig. 1e; Table 2). Both Subtype VI and VII possessed 42 chromosomes in total, consisting of all wheat chromosomes except one chromosome 5A but add one TrS and TrL, respectively (Fig. 1f, g; Table 2). In Subtype II, we hypothesized that TrA might have derived from TrL by misdivision at the centromere. If this was the case, the centromere of TrA should come from A. cristatum rather than wheat. Indeed, the wheat centromere-specific probe CRW successfully stained all chromosome centromeres except for those of the TrAs, but the A. cristatum centromere-specific probe pAcCR1 stained only the TrA centromeres (Fig. 2). These results supported our assumption that TrA was a telocentric derivative of TrL.
GISH discrimination of seven subtypes of wheat-A. cristatum translocation lines. a Subtype I contained 22 pairs of chromosomes including 20 pairs of wheat chromosomes, one pair of TrSs and one pair of TrLs. b Subtype II consisted of 20 pairs of wheat chromosomes, one pair of TrSs and one pair of TrAs. c, d Both Subtype III and Subtype IV consisted of 44 chromosomes including 42 wheat chromosomes, but including one pair of TrLs and one pair of TrSs, respectively. e Subtype V possessed 42 chromosomes consisting of 40 wheat chromosomes and one pair of TrSs. f, g Both Subtype VI and Subtype VII possessed 42 chromosomes in total, but harbored one TrS and one TrL, respectively. Arrows indicate wheat-A. cristatum translocation chromosomes. Chromosomes or chromosomal segments painted in blu e and red belong to wheat and A. cristatum, respectively (color figure online)
Identification of the centromere of TrA in Subtype II using the CRW and pAcCR1 probes. a Green signals appeared on all 21 pairs of chromosomes except one pair of TrAs when the probe CRW (centromeric retrotransposon of wheat) was used; b Green signals are present on one pair of TrAs with the probe pAcCR1 (centromeric retrotransposon of A. cristatum). Chromosomes in blue and red are wheat and A. cristatum chromosomes, respectively. Arrows indicate the TrA chromosomes in Subtype II (color figure online)
Translocation occurred between the wheat chromosome 5A and A. cristatum chromosome 6P
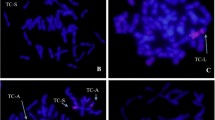
To further determine the identity of the wheat chromosomes involved in the translocation, dual-color FISH was performed on each of the seven translocation subtypes using pAs1 and pHvG39 probes, which were used to distinguish all 21 pairs of wheat chromosomes. As shown in Fig. 3, A. cristatum 6P chromosome segments were translocated onto wheat chromosome 5A in all seven subtypes. In the TrSs that were present in Subtype I, II, IV, V, and VI, A. cristatum 6P chromosome segments were translocated onto wheat chromosome arm 5AL, replacing 60 % of the distal portion of 5AL. In the TrLs that were present in Subtype I, III, and VII, A. cristatum 6P chromosome segments replaced wheat chromosome 5AS and a 40 % portion of 5AL proximal to the centromere.
FISH identification of seven subtypes of wheat-A. cristatum translocation lines using pHvG39 and pAs1 as probes. Images on the right of each panel show FISH results, and those on the left showed corresponding GISH patterns. Chromosomes in blue and red are wheat and A. cristatum chromosomes in the GISH patterns, respectively; whereas pHvG39 and pAs1 signals were pseudo-colored as green and red in the FISH patterns, respectively. a–g, FISH patterns of Subtype I (a), Subtype II (b), Subtype III (c), Subtype IV (d), Subtype V (e), Subtype VI (f), and Subtype VII (g). Note that A. cristatum 6P chromosome segments were translocated to wheat chromosome 5A in all seven subtypes. Arrows indicate wheat-A. cristatum translocation chromosomes (color figure online)
Identification of the breakpoint in the Subtype V translocation line
To pinpoint the location of the breakpoint in the TrS and TrL translocation chromosomes, Subtype V was the only subtype that could be used, since it lacked an intact wheat chromosome 5A. Twenty-five wheat SSR markers physically mapped on wheat chromosome 5A were chosen to analyze the breakpoint in Subtype V. CS nulli-tetrasomic lines (N5AT5B and N5AT5D), CS ditelosomic lines (Dt5AL and Dt5AS), and CS 5AL deletion lines were used to confirm the physical locations of these markers. As shown in Fig. 4a, seven and 18 markers were located to the wheat chromosome arms 5AS and 5AL, respectively. Among them, five markers (Xgwm186, XbarcM158, XbarcM135, Xbarc155, and Xbarc186) were located in bin C-5AL12-0.35, four markers (Xbarc40, XksuM56, Xbarc1, and XksuM5) in bin 5AL12-0.35-0.57, three markers (Xcfa2163, Xgwm617, and Xgwm666) in bin 5AL10-0.57-0.78, three markers (Xwmc110, Xcfa2155, and Xgpw1086) in bin 5AL17-0.78-0.87, and three markers (Xgpw2120, Xgpw2136, and Xgpw2172) in bin 5AL23-0.87-1.00. PCR results showed that only SSR markers in the 5AS and bin C-5AL12-0.35 were amplified in Subtype V, but SSR markers from other bins were not. Xbarc40 and XksuM56 were present in Subtype V, but Xbarc1 and XksuM5 were absent (Fig. 5a), indicating that the translocation breakpoint occurred on wheat chromosome arm 5AL and the breakpoint was located in bin 5AL12-0.35-0.57. All 25 wheat SSR markers were present in the other six subtypes, suggesting that at least one intact 5A chromosome or combinations of one intact 5A chromosome was present in these subtypes.
Physical map of wheat chromosome 5A and A. cristatum chromosome 6P. The map on the left is wheat chromosome 5A consisting of nine deletion bins, and wheat SSR markers are shown on the corresponding regions. The map on the right is A. cristatum chromosome 6P, including 6PS, C-6PL-0.32, 6PL-0.32-0.69, and 6PL-0.69-1.00 chromosomal segments, and A. cristatum-specific markers are shown on the corresponding regions. Fraction breakpoints on chromosomes 5A and 6P are indicated as dashed lines; The location of gene Q is indicated by the arrow according to the previous reports (Galiba et al. 1995; Sutka et al. 1999)
PCR amplification patterns of wheat SSR markers and A. cristatum-specific markers. a Wheat SSR markers Ksum5 and Ksum56 were shown in bin 5AL12-0.35-0.57 by CS nulli-tetrasomic lines, CS ditelosomic lines, and CS 5AL deletion lines. b A. cristatum-specific markers Agc3413, Agc12567, and Agc8937 located in different bins of A. cristatum 6P chromosome were present or absent in some subtypes. Arrows indicate specific bands from wheat (a) and A. cristatum (b), respectively. Lanes in a: 1 CS, 2 N5AT5B, 3 N5AT5D, 4 DT5AS, 5 DT5AL, 6 5AL23-0.87-1.00, 7 5AL17-0.78-0.87, 8 5AL10-0.57-0.78, 9 5AL12-0.35-0.57, 10 C-5AL12-0.35, 11 Subtype V. Lanes in b: 1 Z559, 2 4844-12, 3 Gaocheng 8901, 4 Fukuho, 5 Subtype I, 6 Subtype II, 7 Subtype III, 8 Subtype IV, 9 Subtype V, I0 Subtype VI, 11 Subtype VII
To determine which arm of A. cristatum chromosome 6P was present in these translocation lines, seven EST makers specific for A. cristatum chromosome arm 6PL or 6PS were used. Three markers (For3-G02, For5-E08 and For22-B10) on chromosome 6PS were absent in subtype V, whereas four markers (For15-D06, For8-G11, For14-B02, and For22-E03) on the A. cristatum chromosome 6PL were present in Subtype V (Figs. 4b, 5b). Besides, another nine A. cristatum-specific EST markers were examined. Three markers (Agc3413, Agc7155, and Agc9322) on A. cristatum chromosome arm 6PS, two markers (Agc6900 and Agc34162) in bin C-6PL-0.32, and Agc12567 in bin 6PL-0.32-0.69 were absent in Subtype V, whereas Agc8937 in bin 6PL-0.32-0.69 and two markers (Agc4543 and Agc24535) in the bin 6PL-0.69-1.00 were present in Subtype V (Fig. 5b). The results indicated that the translocation breakpoint occurred on A. cristatum chromosome arm 6PL and the breakpoint was located in bin 6PL-0.32-0.69. A. cristatum chromosome segments including partial 6PL-0.32-0.69 and whole 6PL-0.69-1.00 were translocated onto wheat chromosome arm 5AL, so the constitution of the translocation chromosome in Subtype V could be described as T5AS∙5AL-6PL (Table 2). Similarly, TrS and TrL could also be described as T5AS∙5AL-6PL and T5AL-6PL∙6PS, respectively, while TrA could be designated as T5AL-6PL (Table 2).
Evaluation of agronomic traits in all seven subtypes
Each of these 43 translocation lines was evaluated for five agronomic traits, including fertile tiller number, grain number per spike, spikelet number per spike, kernel number per spikelet, and thousand-kernel weight. Trait observations over 2 years were very similar showing that year-by-year environmental effects were insignificant. We also observed the lines from the same translocation subtype always displayed similar phenotypes, indicating that the genetic constitution was the major factor controlling these phenotypes. Phenotypes for each subtype were calculated using phenotypic data from all lines within the particular subtypes (Table 3; Fig. 6).
Among all the seven subtypes, Subtype V was the only subtype exhibiting inferior agronomic traits including lower fertile tiller number, grain number, spikelet number per spike, kernel number per spikelet, and thousand-kernel weight. Subtype V displayed speltoid spike morphology, and occurred at a very low frequency (only one line). This was probably because Subtype V was the only translocation subtype deficient in a large portion of wheat chromosome arm 5AL. Our results suggested that the distal end of wheat chromosome arm 5AL was essential for normal plant growth and overall fitness and cannot be fully compensated by the A. cristatum chromosome arm 6PL. We compared the agronomic traits among the other six subtypes and their contrast parents. Grain number per spike in Subtype I, VI, and VII were higher than those of two common wheat parents (Gaocheng 8901 and Fukuho), but lower than that of A. cristatum 6P disomic addition line 4844-12. Kernel number per spikelet in all six subtypes were similar to those of two common wheat parents (Gaocheng 8901 and Fukuho), but significantly lower than that of A. cristatum 6P disomic addition line 4844-12. This was also the case for spikelet number per spike. For thousand-kernel weight, no significant difference was observed among the six subtypes as well as two common wheat parents and A. cristatum 6P disomic addition line 4844-12.
Among all traits analyzed, fertile tiller number showed the highest variance. The translocation lines with TrL (Subtype I, III, and VII produced multiple fertile tiller number, which was much higher than those of their two common wheat parents. However, translocation lines without TrL but with TrS (Subtype II, IV, V and VI) displayed reduced fertile tiller number. According to the FISH patterns, TrL included A. cristatum chromosome arm 6PS, chromosome segment 6PL-0.32, and partial 6PL-0.32-0.69, whereas TrS included A. cristatum chromosome segments 6PL-0.69-1.00 and partial 6PL-0.32-0.69. These results indicated that there were positive and negative regulators of fertile tiller number on A. cristatum chromosome 6PS and 6PL, respectively. But when the positive and negative regulators of the fertile tiller number both existed on the same chromosome, as in the case of the A. cristatum 6P disomic addition line 4844-12, the fertile tiller number was not significantly different from those of two common wheat parents, probably because positive and negative effects regulators neutralized each other.
Discussion
Stabilization of the genetic constitution of translocation lines can be achieved spontaneously
Alien genetic resources are important for improving agronomic traits in wheat. Ionizing radiation treatment of alien addition lines, substitution lines, and translocation lines carrying desirable genes is one method to induce chromosome translocation (Chen et al. 1996; Friebe et al. 1996b; Sears and Gustafson 1993; Zhang et al. 2012). In our study, all the translocation lines were identified from the progenies of WAT31-13, which came from an irradiated hybrid. The translocation lines were classified into seven subtypes (I–VII), containing three translocation chromosome types (TrS, TrL, and TrA). TrS and TrL, but not TrA, were present in WAT31-13, indicating that TrA was a new chromosome type which occurred spontaneously. Cytobiological evidences indicated that TrA was derived from TrL as a consequence of a chromosome breakage at the centromere causing loss of the chromosome arm 6PS but retention of the A. cristatum centromere. Such chromosomal breakage happened at a high frequency, since the number of Subtype II individuals (30 lines) was much higher than that of Subtype I (4 lines). The exact biological mechanism of such high frequency of chromosome breakage is currently unknown, but TrA was probably more genetically stable than TrL. Similar spontaneous chromosomal rearrangements have also been reported in wheat-Haynaldia villosa and wheat-Leymus racemosus translocation lines (Cao et al. 2009; Wang et al. 2010).
Non-homoeologous translocations between chromosome 5A and 4A in hexaploid wheat are well known (Devos et al. 1995), and translocations involving 4L/5L also exist in several other species within the tribe Triticeae (King et al. 1994). In our study, all translocations occurred between wheat chromosome arm 5AL and A. cristatum chromosome arm 6PL. Among all the translocation subtypes, Subtype V was the only translocation subtype that exhibited inferior agronomic traits, and also occurred at a very low frequency. This was probably caused by a low overall fitness of the resulting translocation line due to incomplete compensation of the missing segments of chromosome arm 5AL. Indeed, chromosome arm 5AL contains important genes such as gene Q controlling free-threshing and square spike morphology, Vrn-A1 determining winter/spring growth-habit, and Fr1 conferring frost-resistance (Galiba et al. 1995; Sarma et al. 1998; Sutka et al. 1999). Gene Q was reported to locate in bin 5AL23-0.87-1.00 of chromosome arm 5AL, and its location is indicated on the physical map of the wheat chromosome arm 5A (Fig. 4a). Consistent with their lack of gene Q, Subtype V plants displayed speltoid spike morphology, as well as lower seed-set and thousand-kernel weight. Similar phenotypes were observed in the progenies of the cultivar Biscay, which lacks chromosomes 5A carrying Q gene (Forster et al. 2013). Therefore, our results indicated that the distal end of wheat chromosome arm 5AL is essential for normal plant growth and overall fitness, and could not be fully compensated by the A. cristatum chromosome arm 6PL.
The probe pAcCR1 is sufficient to distinguish the A. cristatum and wheat centromere
The plant centromeres are mainly composed of various types of repetitive DNA elements, including transposons, retrotransposons, and telomere-like repeats, most of which are species- or genome-specific (Galasso et al. 1995; Iwabuchi et al. 1991). Thus, centromeric DNA is often a perfect target to distinguish chromosomes of different species (Jiang et al. 2003). A number of repetitive DNA elements were found in wheat centromeres (Cheng and Murata 2003; Kishii et al. 2001; Ito et al. 2004; Zhang et al. 2004), upon which the wheat centromere-specific probe CRW was developed (Liu et al. 2008). In this study, we developed the probe pAcCR1 by a combination of microdissection and DOP-PCR methods. The probe pAcCR1 specifically hybridizes with A. cristatum centromeres, and could not hybridize with wheat centromeres. The probe pAcCR1 could not hybridize with centromeres from several other genomes including genome H, E, R, St, and U (unpublished data).
The translocation lines developed in this study could be used to mechanistically explore the A. cristatum chromosome segments controlling fertile tiller number
Tillering is a key component of yield for most cereals such as wheat, rice, and barley (Sakamoto and Matsuoka 2004; Sreenivasulu and Schnurbusch 2012). While the molecular mechanism of tillering has been extensively studied in dicots and a few monocots (Aguilar-Martinez et al. 2007; Kebrom et al. 2013; Li et al. 2003; Minakuchi et al. 2010; Schmitz et al. 2002; Wang and Li 2008), it has not been fully explained in wheat, probably due to the lack of genetic materials suitable for dissecting the genetic architecture of tillering. A number of quantitative trait loci (QTLs) for tiller number have been identified (Huang et al. 2004; Kato et al. 2000; Kim et al. 1993; Li et al. 2002; Narasimhamoorthy et al. 2006; Naruoka et al. 2011; Snape et al. 1985), but the underlying genes have not been cloned. In those studies, a large number of QTLs with minor effects were identified, indicating that tillering is a complex trait coordinately controlled by many loci.
To successfully clone QTLs, the parents used to construct the cloning population have to be carefully chosen, since QTLs with large effects can only be revealed when alleles from the parents have dramatic effects on tillering. Few cases have been reported where large-effect QTLs controlling tillering have been fine-mapped in wheat, the most prominent example was the tiller inhibition gene (tin1) detected in the uniculm wheat mutant Line 492 and located on the wheat chromosome arm 1AS (Spielmeyer and Richards 2004). Other examples included tin2 and tin3 (Kuraparthy et al. 2007; Peng et al. 1998). The ftin (fertile tiller inhibition) gene was located on wheat chromosome arm 1AS in Pubing 3558 in our previous report (Zhang et al. 2013). The translocation lines reported here have dramatic differences in fertile tiller number, so there should exist gene(s) controlling fertile tiller number. In this study, we determined that the A. cristatum chromosome arm 6PS and 6PL have positive and negative effects on fertile tiller number, respectively. However, the underlying genes controlling fertile tiller number cannot be cloned from these materials using a traditional map-based cloning approach. To overcome this hurdle, the A. cristatum 6P chromosome segments can first be reduced to smaller segments and transferred into a known elite cultivated wheat background, and then the strategy employed in the cloning of Pm21 can be used (Cao et al. 2011).
Some translocation lines showed potential applications for breeding high-yielding wheat
Wheat cultivars can be classified into large-spike and multi-spike types. Compared to large-spike types, multi-spike type cultivars are considered to be more stable in agronomic performance, and it is easier to achieve higher yields per unit field area (Deng et al. 2011). This effect becomes more pronounced when wheat plants are confronted with environmental stresses such as drought and salinity (Reynolds et al. 2007; Tian et al. 2006a, b). In this study, all translocation subtypes except Subtype V showed higher grain number per spike, thus they are potentially useful for breeding wheat lines with large spikes. However, what attracted our attention more were the subtypes with high numbers of fertile tiller, such as Subtype I, III, and VII, which also showed potential applications for breeding of multi-spike types. Although these subtypes showed potential applications for breeding high-yielding wheat, it would be a challenge to introduce the desirable genes conferring multiple fertile tiller number from A. cristatum chromosome arm 6PS into common wheat without the introduced segments also conferring deleterious effects. To overcome this problem, chromosome translocation induced by ionizing radiation or Ph1-deficient genetic stocks can be used.
The Ph1 system is advantageous in that it can significantly promote the frequency of homoeologous chromosome pairing and recombination, thereby producing genetically compensating translocations (Qi et al. 2008; Niu et al. 2011). However, homoeologous recombination is affected by the structurally rearranged segments of alien chromosomes, the genetic relationship between wheat and its related species as well as the genetic distance between the target gene and the centromere (Qi et al. 2007; Monte et al. 1993; Nasuda et al. 1998). In our previous research, wheat-A. cristatum 6P disomic addition lines displayed obvious chromosome rearrangements (Han et al. 2014). Therefore, ionizing radiation might be the preferred choice in our case. Ionizing radiation treatment can cause alien chromosome breakage at different positions, and then the alien chromosome will be transferred onto different regions of wheat chromosomes, as exemplified in Chen et al. (2013). Therefore, it is possible that the desirable genes conferring multiple fertile tiller number from A. cristatum chromosome arm 6PS would be transferred onto other chromosomes rather than wheat chromosome 5A during further cycles of ionizing radiation.
In conclusion, by studying the chromosomal constitution, behavior, and agronomic traits of wheat-A. cristatum translocation lines, we pinpointed the chromosomal segments of A. cristatum 6P positively and negatively regulating fertile tiller number in wheat, respectively. Our work not only laid the foundation for further research on wheat tillering, but also provided the starting materials for high-yield wheat breeding.
Author contribution statement
Li LH conceived the project. Lu YQ and Ye XL analyzed the data and wrote the manuscript. Ye XL, Han HM, and Chen GY performed cytological experiments. Liu WH created the translocation lines. Zhang JP and Gao AN contributed to the development of molecular markers. Yang XM and Li XQ performed the phenotyping.
References
Aguilar-Martinez JA, Poza-Carrion C, Cubas P (2007) Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell 19:458–472
Allen GC, Flores-Vergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325
Bao Y, Li X, Liu S, Cui F, Wang H (2009) Molecular cytogenetic characterization of a new wheat-Thinopyrum intermedium partial amphiploid resistant to powdery mildew and stripe rust. Cytogenet and Genome Res 126:390–395
Cao YP, Bie TD, Wang XE, Chen PD (2009) Induction and transmission of wheat-Haynaldia villosa chromosomal translocations. J Genet Genomics 36:313–320
Cao AH, Xing LP, Wang XY, Yang XM, Wang W, Sun YL, Qian C, Ni JL, Chen YP, Liu DJ, Wang X, Chen PD (2011) Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc Natl Acad Sci USA 108:7727–7732
Chen PD, Qi LL, Zhou B, Zhang SZ, Liu DJ (1995) Development and molecular cytogenetic analysis of wheat-Haynaldia villosa 6VS/6AL translocation lines specifying resistance to powdery mildew. Theor Appl Genet 91:1125–1128
Chen Q, Conner RL, Laroche A (1996) Molecular characterization of Haynaldia villosa chromatin in wheat lines carrying resistance to wheat curl mite colonization. Theor Appl Genet 93:679–684
Chen PD, Liu WX, Yuan JH, Wang XE, Zhou B, Wang SL, Zhang SZ, Feng YG, Yang BJ, Liu GX, Liu DJ, Qi LL, Zhang P, Friebe B, Gill BS (2005) Development and characterization of wheat-Leymus racemosus translocation lines with resistance to Fusarium head blight. Theor Appl Genet 111:941–948
Chen PD, You CF, Hu Y, Chen SW, Zhou B, Cao AZ, Wang XE (2013) Radiation-induced translocations with reduced Haynaldia villosa chromatin at the Pm21 locus for powdery mildew resistance in wheat. Mol Breed 31:477–484
Cheng ZJ, Murata M (2003) A centromeric tandem repeat family originating from a part of Ty3/gypsy-retroelement in wheat and its relatives. Genetics 164:665–672
Deal KR, Goyal S, Dvorak J (1999) Arm location of Lophopyrum elongatum genes affecting K +/Na + selectivity under salt stress. Euphytica 108:193–198
Deng SM, Wu XR, Wu YY, Zhou RH, Wang HG, Jia JZ, Liu SB (2011) Characterization and precise mapping of a QTL increasing spike number with pleiotropic effects in wheat. Theor Appl Genet 122:281–289
Devos KM, Dubcovsky J, Dvorak J, Chinoy CN, Gale MD (1995) Structural evolution of wheat chromosomes 4A, 5A, and 7B and its impact on recombination. Theor Appl Genet 91:282–288
Dewey DR (1984) The genomic system of classification as a guide to intergeneric hybridization with the perennial Triticeae. In: Gustafson JP (ed) Gene manipulation in plant improvement. Plenum Press, New York, pp 209–279
Dong YS, Zhou RH, Xu SJ, Li LH, Cauderon Y, Wang RRC (1992) Desirable characteristics in perennial Triticeae collected in China for wheat improvement. Hereditas 116:175–178
Duan XY, Sheng BQ, Zhou YL, Xiang QJ (1998) Monitoring of the virulence population of Erysiphe graminis f.sp. tritici. Acta Phytophylac Sin 25:31–36
Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316:1862–1866
Faris JD, Xu SS, Cai XW, Friesen TL, Jin Y (2008) Molecular and cytogenetic characterization of a durum wheat-Aegilops speltoides chromosome translocation conferring resistance to stem rust. Chromosome Res 16:1097–1105
Forster S, Schumann E, Baumann M, Weber WE, Pillen K (2013) Copy number variation of chromosome 5A and its association with Q gene expression, morphological aberrations, and agronomic performance of winter wheat cultivars. Theor Appl Genet 126:3049–3063
Friebe B, Hatchett JH, Sears RG, Gill BS (1990) Transfer of Hessian fly resistance from ‘Chaupon’ rye to hexaploid wheat via a 2BS/2RL wheat-rye chromosome translocation. Theor Appl Genet 79:385–389
Friebe B, Zeller FJ, Mukai Y, Forster BP, Bartos P, McIntosh RA (1992) Characterization of rust-resistant wheat-Agropyron intermedium derivatives by C-banding, in situ hybridization and isozyme analysis. Theor Appl Genet 83:775–782
Friebe B, Gill KS, Tuleen NA, Gill BS (1996a) Transfer of wheat streak mosaic virus resistance from Agropyron intermedium into wheat. Crop Sci 36:857–861
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996b) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87
Fu SL, Lv ZL, Qi B, Guo X, Li J, Liu B, Han FP (2012) Molecular cytogenetic characterization of wheat-Thinopyrum elongatum addition, substitution and translocation lines with a novel source of resistance to wheat Fusarium head blight. J Genet Genomics 39:103–110
Galasso I, Schmidt T, Pignone D, Heslop-Harrison JS (1995) The molecular cytogenetics of Vigna unguiculata (L.) Walp: the physical organization and characterization of 18S-5.8S-25S rRNA genes, 5S rRNA genes, telomere-like sequences, and a family of centromeric repetitive DNA sequences. Theor Appl Genet 91:928–935
Galiba G, Quarrie SA, Sutka J, Morgounov A, Snape JW (1995) RFLP mapping of the vernalization (Vrn1) and frost-resistance (Fr1) genes on chromosome 5A of wheat. Theor Appl Genet 90:1174–1179
Gill BS, Friebe BR, White FF (2011) Alien introgressions represent a rich source of genes for crop improvement. Proc Natl Acad Sci USA 108:7657–7658
Han FP, Fedak G, Benabdelmouna A, Armstrong K, Ouellet T (2003) Characterization of six wheat x Thinopyrum intermedium derivatives by GISH, RFLP, and multicolor GISH. Genome 46:490–495
Han HM, Bai L, Su JJ, Zhang JP, Song LQ, Gao AN, Yang XM, Li XQ, Liu WH, Li LH (2014) Genetic rearrangements of six wheat-Agropyron cristatum 6P addition lines revealed by molecular markers. PLoS ONE 9:e91066
Haudry A, Cenci A, Ravel C, Bataillon T, Brunel D, Poncet C, Hochu I, Poirier S, Santoni S, Glemin S, David J (2007) Grinding up wheat: a massive loss of nucleotide diversity since domestication. Mol Biol Eol 24:1506–1517
He RL, Chang ZJ, Yang ZJ, Yuan ZY, Zhan HX, Zhang XJ, Liu JX (2009) Inheritance and mapping of powdery mildew resistance gene Pm43 introgressed from Thinopyrum intermedium into wheat. Theor Appl Genet 118:1173–1180
Hsam SLK, Zeller FJ (1997) Evidence of allelism between genes Pm8 and Pm17 and chromosomal location of powdery mildew and leaf rust resistance genes in the common wheat cultivar Amigo. Plant Breed 116:119–122
Hua W, Liu ZJ, Zhu J, Xie CJ, Yang TM, Zhou YL, Duan XY, Sun QX, Liu ZY (2009) Identification and genetic mapping of Pm42, a new recessive wheat powdery mildew resistance gene derived from wild emmer (Triticum turgidum var. dicoccoides). Theor Appl Genet 119:223–230
Huang XQ, Kempf H, Ganal MW, Röder MS (2004) Advanced backcross QTL analysis in progenies derived from a cross between a German elite winter wheat variety and a synthetic wheat (Triticum aestivum L.). Theor Appl Genet 109:933–943
Ito H, Nasuda S, Endo TR (2004) A direct repeat sequence associated with the centromeric retrotransposons in wheat. Genome 47:747–756
Iwabuchi M, Itoh K, Shimamoto K (1991) Molecular characterization of repetitive DNA sequences in Brassica. Theor Appl Genet 81:349–355
Jiang J, Morris KL, Gill BS (1994) Introgression of Elymus trachycaulus chromatin into common wheat. Chromosome Res 2:3–13
Jiang JM, Birchler JA, Parrott WA, Dawe RK (2003) A molecular view of plant centromeres. Trends Plant Sci 18:570–575
Kang HY, Wang Y, Fedak G, Cao WG, Zhang HQ, Fan X, Sha LN, Xu LL, Zheng YL, Zhou YH (2011) Introgression of chromosome 3Ns from Psathyrostachys huashanica into wheat specifying resistance to stripe rust. PLoS One 6:e21802
Kato K, Miura H, Sawada S (2000) Mapping QTLs controlling grain yield and its components on chromosome 5A of wheat. Theor Appl Genet 101:1114–1121
Kebrom TH, Spielmeyer W, Finnegan EJ (2013) Grasses provide new insights into regulation of shoot branching. Trends Plant Sci 18:41–48
Kim NS, Armstrong KC, Fedak G, Fominaya A, Whelan EW (1993) Cytological and molecular characterization of a chromosome interchange and addition lines in Cadet involving chromosome 5B of wheat and 6Ag of Lophopyrum ponticum. Theor Appl Genet 86:827–832
King IP, Purdie KA, Liu CJ, Reader SM, Pittaway TS, Orford SE, Miller TE (1994) Detection of interchromosomal translocations within the Triticeae by RFLP analysis. Genome 37:882–887
Kishii M, Nagaki K, Tsujimoto H (2001) A tandem repetitive sequence located in the centromeric region of common wheat (Triticum aestivum) chromosomes. Chromosome Res 9:417–428
Klindworth DL, Niu Z, Chao S, Friesen TL, Jin Y, Faris JD, Cai X, Xu SS (2012) Introgression and characterization of a goatgrass gene for a high level of resistance to Ug99 stem rust in tetraploid wheat. Genes Genom Genet 2:665–673
Kuraparthy V, Sood S, Dhaliwal HS, Chhuneja P, Gill BS (2007) Identification and mapping of a tiller inhibition gene (tin3) in wheat. Theor Appl Genet 114:285–294
Larkin PJ, Banks PM, Lagudah ES, Appels R, Xiao C, Zhiyong X, Ohm HW, McIntosh RA (1995) Disomic Thinopyrum intermedium addition lines in wheat with barley yellow dwarf virus resistance and with rust resistances. Genome 38:385–394
Larson SR, Kishii M, Tsujimoto H, Qi LL, Chen PD, Lazo GR, Jensen KB, Wang RRC (2012) Leymus EST linkage maps identify 4NsL-5NsL reciprocal translocation, wheat-Leymus chromosome introgressions, and functionally important gene loci. Theor Appl Genet 124:189–206
Li LH, Dong YS (1991) Hybridization between Triticum aestivum L. and Agropyron michnoi Roshev. Theor Appl Genet 81:312–316
Li LH, Dong YS (1993) A self-fertile trigeneric hybrid, Triticum aestivum x Agropyron michnoi x Secale cereale. Theor Appl Genet 87:361–368
Li LH, Dong YS, Zhou RH, Li XQ, Li P (1995) Cytogenetics and self-fertility of hybrids between Triticum aestivum L. and Agropyron cristatum (L.) Gaertn. Chin J Genet 22:105–112
Li LH, Li XQ, Li P, Dong YS, Zhao GS (1997) Establishment of wheat-Agropyron cristatum alien addition lines. I. Cytology of F3, F2BC1, BC4, and BC3F1 progenies. Acta Genetica Sinica 24:154–159
Li LH, Yang XM, Li XQ, Dong YS, Chen XM (1998a) Introduction of desirable genes from Agropyron cristatum into common wheat by intergeneric hybridization. Sci Agric Sin 31:1–6
Li LH, Yang XM, Zhou RH, Li XQ, Dong YS (1998b) Establishment of wheat-Agropyron cristatum alien addition lines II. Identification of alien chromosomes and analysis of development approaches. Acta Genetica Sinica 25:538–544
Li WL, Nelson JC, Chu CY, Shi LH, Huang SH, Liu DJ (2002) Chromosomal locations and genetic relationships of tiller and spike characters in wheat. Euphytica 125:357–366
Li X, Qian Q, Fu Z, Wang Y, Xiong G, Zeng D, Wang X, Liu X, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li J (2003) Control of tillering in rice. Nature 422:618–621
Liu ZY, Sun QX, Ni ZF, Yang TM, Mclntosh RA (1999) Development of SCAR markers linked to the Pm21 gene conferring resistance to powdery mildew in common wheat. Plant Breed 118:215–219
Liu Z, Yue W, Li DY, Wang RRC, Kong XY, Lu K, Wang GX, Dong YS, Jin WW, Zhang XY (2008) Structure and dynamics of retrotransposons at wheat centromeres and pericentromeres. Chromosoma 117:445–456
Luan Y, Wang XG, Liu WH, Li CY, Zhang JP, Gao AN, Wang YM, Yang XM, Li LH (2010) Production and identification of wheat-Agropyron cristatum 6P translocation lines. Planta 232:501–510
Lukaszewski AJ (2000) Manipulation of the 1RS∙1BL translocation in wheat by induced homoeologous recombination. Crop Sci 40:216–225
Luo PG, Hu XY, Chang ZJ, Zhang M, Zhang HQ, Ren ZL (2009) A new stripe rust resistance gene transferred from Thinopyrum intermedium to hexaploid wheat (Triticum aestivum). Phytoprotection 90:57–63
Ma JX, Zhou RH, Dong YS, Jia JZ (2000) Control and inheritance of resistance to yellow rust in Triticum aestivum-Lophopyrum elongatum chromosome substitution lines. Euphytica 111:57–60
Mago R, Spielmeyer W, Lawrence GJ, Lagudah ES, Ellis JG, Pryor A (2002) Identification and mapping of molecular markers linked to rust resistance genes located on chromosome 1RS of rye using wheat-rye translocation lines. Theor Appl Genet 104:1317–1324
McDonald MP, Galwey NW, Ellneskog-Staam P, Colmer TD (2001) Evaluation of Lophopyrum elongatum as a source of genetic diversity to increase the waterlogging tolerance of hexaploid wheat (Triticum aestivum). New Phytol 151:369–380
Minakuchi K, Kameoka H, Yasuno N, Umehara M, Luo L, Kobayashi K, Hanada A, Ueno K, Asami T, Yamaguchi S, Kyozuka J (2010) FINE CULM1 (FC1) works downstream of strigolactones to inhibit the outgrowth of axillary buds in rice. Plant Cell Physiol 51:1127–1135
Monneveux P, Reynolds MP, Aguilar JG, Singh RP (2003) Effects of the 7DL∙7Ag translocation from Lophopyrum elongatum on wheat yield and related morphophysiological traits under different environments. Plant Breed 122:379–384
Monte JV, Mclintyre CL, Gustafson JP (1993) Analysis of phylogenetic relationships in the Triticeae tribe using RFLPs. Theor Appl Genet 86:649–665
Mujeeb-Kazi A, Kazi AG, Dundas I, Rasheed A, Ogbonnaya F, Kishi M, Bonnett D, Wang RRC, Xu S, Chen P, Mahmood T, Bux H, Farrakh S (2013) Genetic diversity for wheat improvement as a conduit for food security. In: Sparks D (ed) Advances in Agronomy, vol 122. Academic Press, Burlington, pp 179–257
Narasimhamoorthy B, Gill BS, Fritz AK, Nelson JC, Brown-Guedira GL (2006) Advanced backcross QTL analysis of a hard winter wheat x synthetic wheat population. Theor Appl Genet 112:787–796
Naruoka Y, Talbert LE, Lanning SP, Blake NK, Martin JM, Sherman JD (2011) Identification of quantitative trait loci for productive tiller number and its relationship to agronomic traits in spring wheat. Theor Appl Genet 123:1043–1053
Nasuda S, Friebe B, Busch W, Kynast RG, Gill BS (1998) Structural rearrangement in chromosome 2M of Aegilops comosa has prevented the utilization of the compair and related wheat-Ae. comosa translocations in wheat improvement. Theor Appl Genet 96:780–785
Niu Z, Klindworth DL, Friesen TL, Chao S, Jin Y, Cai X, Xu SS (2011) Targeted introgression of a wheat stem rust resistance gene by DNA marker-assisted chromosome engineering. Genetics 187:1011–1021
Niu Z, Klindworth DL, Yu G, Friesen TL, Chao S, Jin Y, Cai X, Ohm JB, Rasmussen JB, Xu SS (2014) Development and characterization of wheat lines carrying stem rust resistance gene Sr43 derived from Thinopyrum ponticum. Theor Appl Genet 127:969–980
Pedersen C, Langridge P (1997) Identification of the entire chromosome complement of bread wheat by two-colour FISH. Genome 40:589–593
Peng ZS, Yen C, Yang JL (1998) Genetic control of oligo-culms character in common wheat. Wheat Inf Serv 86:19–24
Qi LL, Chen PD, Liu DJ, Zhou B, Zhang SZ, Sheng QB, Xiang QJ, Duan XY, Zhou YL (1995) The gene Pm21-a new source of resistance to wheat powdery mildew. Acta Agro Sin 21:257–260
Qi LL, Friebe B, Zhang P, Gill BS (2007) Homoeologous recombination, chromosome engineering and crop improvement. Chromosome Res 15:3–19
Qi LL, Pumphrey MO, Friebe B, Chen PD, Gill BS (2008) Molecular cytogenetic characterization of alien introgressions with gene Fhb3 for resistance to Fusarium head blight disease of wheat. Theor Appl Genet 117:1155–1166
Qi LL, Pumphrey MO, Friebe B, Zhang P, Qian C, Bowden RL, Rouse MN, Jin Y, Gill BS (2011) A novel Robertsonian translocation event leads to transfer of a stem rust resistance gene (Sr52) effective against race Ug99 from Dasypyrum villosum into bread wheat. Theor Appl Genet 123:159–167
Reif JC, Zhang P, Dreisigacker S, Warburton ML, van Ginkel M, Hoisington D, Bohn M, Melchinger AE (2005) Wheat genetic diversity trends during domestication and breeding. Theor Appl Genet 110:859–864
Reynolds M, Dreccer F, Trethowan R (2007) Drought adaptive traits derived from wheat wild relatives and landraces. J Exp Bot 58:177–186
Sakamoto T, Matsuoka M (2004) Generating high-yielding varieties by genetic manipulation of plant architecture. Current Opin Biotechnol 15:144–147
Sarma RN, Gill BS, Sasaki T, Galiba G, Sutka J, Laurie DA, Snape JW (1998) Comparative mapping of the wheat chromosome 5A Vrn-A1 region with rice and its relationship to QTL for flowering time. Theor Appl Genet 97:103–109
Schmitz G, Tillmann E, Carriero F, Fiore C, Cellini F, Theres K (2002) The tomato Blind gene encodes a MYB transcription factor that controls the formation of lateral meristems. Proc Natl Acad Sci USA 99:1064–1069
Sears ER, Gustafson JP (1993) Use of radiation to transfer alien chromosome segments to wheat. Crop Sci 33:897–901
Sharma H, Ohm H, Goulart L, Lister R, Appels R, Benlhabib O (1995) Introgression and characterization of barley yellow dwarf virus-resistance from Thinopyrum-intermedium into Wheat. Genome 38:406–413
Singh RP, Huerta-Espino J, Rajaram S, Crossa J (1998) Agronomic effects from chromosome translocations 7DL∙7Ag and 1BL∙1RS in spring wheat. Crop Sci 38:27–33
Snape JW, Law CN, Parker BB, Worland AJ (1985) Genetical analysis of chromosome 5A of wheat and its influence on important agronomic characters. Theor Appl Genet 71:518–526
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Sourdille P, Singh S, Cadalen T, Brown-Guedira GL, Gay G, Qi L, Gill BS, Dufour P, Murigneux A, Bernard M (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct Integr Genomics 4:12–25
Spielmeyer W, Richards RA (2004) Comparative mapping of wheat chromosome 1AS which contains the tiller inhibition gene (tin) with rice chromosome 5S. Theor Appl Genet 109:1303–1310
Sreenivasulu N, Schnurbusch T (2012) A genetic playground for enhancing grain number in cereals. Trends Plant Sci 17:91–101
Sutka J, Galiba G, Vágújfalvi A, Gill BS, Snape JW (1999) Physical mapping of the Vrn-A1 and Fr1 genes on chromosome 5A of wheat using deletion lines. Theor Appl Genet 99:199–202
Tian F, Zhu ZF, Zhang BS, Tian LB, Fu YC, Wang XK, Sun CQ (2006a) Fine mapping of a quantitative trait locus for grain number per panicle from wild rice (Oryza rufipogon Griff.). Theor Appl Genet 113:619–629
Tian JC, Deng ZY, Hu RB, Wang YX (2006b) Yield components of super wheat cultivars with different spike types and the path coefficient analysis on grain yield. Acta Agron Sin 32:1699–1705
Vega JM, Abbo S, Feldman M, Levy AA (1994) Chromosome painting in plants: in situ hybridization with a DNA probe from a specific microdissected chromosome arm of common wheat. Proc Natl Acad Sci USA 91:12041–12045
Wang RRC (2011) Agropyron and Psathyrostachys. In: Kole C (ed) Wild Crop Relatives: Genomic and Breeding Resources, Cereals, vol 1, chapter 2. Springer, Berlin and Heidelberg, pp 77–108
Wang YH, Li JY (2008) Molecular basis of plant architecture. Annu Rev Plant Biol 59:253–279
Wang RRC, Zhang XY (1996) Characterization of the translocated chromosome using fluorescence in situ hybridization and random amplified polymorphic DNA on two Triticum aestivum-Thinopyrum intermedium translocation lines resistant to wheat streak mosaic or barley yellow dwarf virus. Chromosome Res 4:583–587
Wang LS, Chen PD, Wang XE (2010) Molecular cytogenetic analysis of Triticum aestivum-Leymus racemosus reciprocal chromosomal translocation T7DS∙5LrL/T5LrS∙7DL. Chin Sci Bull 55:1026–1031
Wang SW, Yin LN, Tanaka H, Tanaka K, Tsujimoto H (2011) Wheat-Aegilops chromosome addition lines showing high iron and zinc contents in grains. Breed Sci 61:189–195
Wu J, Yang X, Wang H, Li H, Li L, Li X, Liu W (2006) The introgression of chromosome 6P specifying for increased numbers of florets and kernels from Agropyron cristatum into wheat. Theor Appl Genet 114:13–20
Yu GT, Cai X, Harris MO, Gu YQ, Luo MC, Xu SS (2009) Saturation and comparative mapping of the genomic region harboring Hessian fly resistance gene H26 in wheat. Theor Appl Genet 118:1589–1599
Zhang P, Li W, Friebe B, Gill BS (2004) Simultaneous painting of three genomes in hexaploid wheat by BAC-FISH. Genome 47:979–987
Zhang RQ, Wang XE, Chen PD (2012) Molecular and cytogenetic characterization of a small alien-segment translocation line carrying the softness genes of Haynaldia villosa. Genome 55:639–646
Zhang JP, Wu J, Liu WH, Lu X, Yang XM, Gao AN, Li XQ, Lu YQ, Li LH (2013) Genetic mapping of a fertile tiller inhibition gene, ftin, in wheat. Mol Breed 31:441–449
Acknowledgments
This work was funded by the National High Technology Research and Development Program of China (863 program, No. 2011AA100102), the National Natural Science Foundation of China (No. 31271714), and the National Basic Research Program of China (973 program, No. 2011CB100104). We thank Dr. Xueyong Zhang (Institute of Crop Sciences, Chinese Academy of Agricultural Sciences) for generously providing the CRW probe used in this study.
Conflict of interest
The authors declare that there are no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. S. Xu.
X. Ye and Y. Lu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Ye, X., Lu, Y., Liu, W. et al. The effects of chromosome 6P on fertile tiller number of wheat as revealed in wheat-Agropyron cristatum chromosome 5A/6P translocation lines. Theor Appl Genet 128, 797–811 (2015). https://doi.org/10.1007/s00122-015-2466-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2466-4