Abstract
Key message
Genome-wide QTL analysis of potato tuber carotenoid content was investigated in populations of Solanum tuberosum Group Phureja that segregate for flesh colour, revealing a novel major QTL on chromosome 9.
Abstract
The carotenoid content of edible plant storage organs is a key nutritional and quality trait. Although the structural genes that encode the biosynthetic enzymes are well characterised, much less is known about the factors that determine overall storage organ content. In this study, genome-wide QTL mapping, in concert with an efficient ‘genetical genomics’ analysis using bulked samples, has been employed to investigate the genetic architecture of potato tuber carotenoid content. Two diploid populations of Solanum tuberosum Group Phureja were genotyped (AFLP, SSR and DArT markers) and analysed for their tuber carotenoid content over two growing seasons. Common to both populations were QTL that explained relatively small proportions of the variation in constituent carotenoids and a major QTL on chromosome 3 explaining up to 71 % of the variation in carotenoid content. In one of the populations (01H15), a second major carotenoid QTL was identified on chromosome 9, explaining up to 20 % of the phenotypic variation. Whereas the major chromosome 3 QTL was likely to be due to an allele of a gene encoding β-carotene hydroxylase, no known carotenoid biosynthetic genes are located in the vicinity of the chromosome 9 QTL. A unique expression profiling strategy using phenotypically distinct bulks comprised individuals with similar carotenoid content provided further support for the QTL mapping to chromosome 9. This study shows the potential of using the potato genome sequence to link genetic maps to data arising from eQTL approaches to enhance the discovery of candidate genes underlying QTLs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carotenoids play fundamental roles in human nutrition and increasingly their consumption is associated with protection from a range of diseases (reviewed in Fraser and Bramley 2004). Consequently, research focusing on enhancing carotenoid content particularly in fruit, grains and tubers that form the major part of the human diet has gained momentum. In recent years, considerable detail of the plant carotenoid biosynthetic pathway has been elucidated (for reviews see Hirschberg. 2001; Taylor and Ramsay 2005; DellaPenna and Pogson 2006). Most of the genes encoding the structural enzymes have been characterised from multiple sources and genetic engineering of pathways has led to notable enhancement of carotenoid levels in a range of crop species, including tomato, potato and rice (reviewed in Giuliano et al. 2008). Despite these successes, there are still major gaps in our understanding of how carotenoid accumulation in different plant organs is regulated. In addition to biosynthesis, other functions such as carotenoid turnover appear to be important, at least in some cases (Ohmiya et al. 2006; Campbell et al. 2010; Zhou et al. 2011). Genes that impact on plastid development, such as the Or gene of cauliflower, also influence the accumulation of carotenoids in storage tissues (Li et al. 2001; Van Eck et al. 2007). Additionally, factors that control the expression levels of the biosynthetic genes remain to be determined, with few recent exceptions (for example, Welsch et al. 2008; Cazzonelli et al. 2009), a situation that contrasts with other biosynthetic pathways, such as those involved in flavonoid biosynthesis (Tian et al. 2008).
In potato there is wide variation in tuber carotenoid accumulation levels (Brown et al. 2006). White-fleshed tubers contain levels of total carotenoid less than 100 µg/100 g−1 FW (Breithaupt and Bamedi 2002). However, several studies have shown considerably higher levels of carotenoid accumulation in yellow/orange-fleshed tuber varieties (for example Morris et al. 2004). In diploid progeny of crosses between Solanum tuberosum Group Stenotomum (Stenotomum) and S. tuberosum Group Phureja (hereafter referred to as Phureja), Brown et al. (1993) correlated the tuber orange flesh trait in potato with the content of zeaxanthin. In a different Phureja × Stenotomum hybrid population, progeny clones with yellow-fleshed tubers were obtained. In these tubers, lutein-5, 6-epoxide and lutein were the major carotenoids, with much lower levels of zeaxanthin (Lu et al. 2001). Although significant genotype × environment interactions have been reported for potato tuber carotenoid content, heritabilities of carotenoid content are generally high (Haynes 2010). In fact, it has long been mooted that tuber yellow flesh is primarily controlled by a single dominant allele at the Y (Yellow) locus (Bonierbale et al. 1988; Thorup et al. 2000). Collective evidence from several studies has shown that the Y locus maps on chromosome 3 and is likely to encode an isoform of β-carotene hydroxylase (Chy, Brown et al. 2006; Kloosterman et al. 2010). Other enzyme activities were implicated in tuber carotenogenesis by transgenic studies which demonstrated elevated tuber carotenoid content on down-regulation of the gene encoding zeaxanthin epoxidase (Zep, Römer et al. 2002). An inverse relationship between Zep expression level and tuber carotenoid content reinforced this suggestion (Morris et al. 2004) and recently it was shown that within a range of potato diploid genotypes with orange tuber flesh, all were homozygous for one specific Zep allele (Wolters et al. 2010). Whilst alleles for Chy and Zep clearly have a major impact on tuber carotenoid levels, a comprehensive survey of loci affecting tuber carotenoid content has not yet been reported. A further recent suggestion is that potato tuber sugar content can regulate the transcription of Chy (Zhou et al. 2011). In view of our limited knowledge of how carotenoid accumulation is regulated in potato tubers, there remains the possibility that novel regulatory mechanisms exist, as has been suggested for other carotenoid accumulating storage organs. For example, in tomato fruit 16 quantitative trait loci (QTL) were identified that modified fruit red colour (lycopene content) (Liu et al. 2003). Candidate genes associated with the carotenoid biosynthesis pathway were mapped to 23 distinct loci, however, only five carotenoid QTL co-segregated with the same ‘bins’ that contained candidate genes. A further example comes from melon fruit where eight significant QTL for fruit β-carotene content were identified (Cuevas et al. 2008), none of which could be associated with putative carotenoid pathway genes. Clearly, this demonstrates the complexity of mechanisms that regulate storage organ carotenoid content.
Previously, a “genetical genomics” approach was used to investigate gene expression patterns associated with high versus low tuber carotenoid accumulation (Kloosterman et al. 2010). Here, pools of individuals from a segregating population with high and low tuber carotenoid content were compared at the level of the transcriptome. Gene expression in tuber samples extracted from the pools was analysed using a 44,000 feature microarray. A gene encoding β-carotene hydroxylase 2 (Chy2) was shown to be expressed at a much higher level in the high tuber carotenoid pool adding weight to the evidence that this gene underlies the large effect QTL for tuber carotenoid content on chromosome 3.
The aim of this study was to conduct a comprehensive genetic analysis of constituent tuber carotenoids using two distinct diploid populations showing variation in tuber carotenoid content. A conventional QTL analysis of these populations was performed using a range of carotenoid phenotypic data and linkage maps generated using various types of molecular marker. Moreover, an expression QTL (eQTL) approach was used, whereby tuber samples from clones with similar carotenoid traits were bulked and patterns of gene expression were measured for each bulk by microarray analysis. Variation of gene expression between bulks may be due to either expression polymorphisms located near to or within the trait gene (cis-eQTL) or indirectly from a distant location on the genome (trans-eQTL). Differentially expressed genes from bulks with contrasting carotenoid traits were genetically and physically mapped using the potato genome to reinforce the QTL analysis, and provide a rapid means of developing gene markers closely associated with the target trait.
Materials and methods
Development of two F1 diploid Phureja populations segregating for tuber flesh colour
Two diploid F1 S. tuberosum Group Phureja populations, (01H15 and 03TR2) segregating for tuber flesh colour were developed as a resource for tuber carotenoid phenotyping and the genetic mapping of genes controlling tuber carotenoid content. Using long-day adapted Phureja clones selected from the Commonwealth Potato Collection maintained at the James Hutton Institute UK, population 01H15 was generated by crossing parents PHU5010 Yema De Huevo (YDH) and DB378(1) clones. Tuber flesh colour was assessed using a colour reference chart (Online Resource I). Though tubers from both parents have a deep orange flesh colour, indicative of high carotenoid content (Lu et al. 2001), the derived F1 population consisting of 106 progeny clones segregated for deep orange to light yellow tuber flesh colour in an approximately 3:1 ratio. Population 03TR2 was developed from a cross between two Phureja clones, DB271(39) and DB375(2) which displayed cream and deep orange tuber flesh colours, respectively. The resulting population consisted of 177 progeny clones which segregated in an approximately 1:1 ratio for deep yellow/orange to cream tuber flesh colour.
Growth of plant material
All experimental material used in this study was obtained from 01H15 (106 clones) and 03TR2 (177 clones) populations grown in the years 2006 and 2008. Potato plants were grown from seed tubers in 15-cm-diameter pots containing compost. Plants were raised in a glasshouse maintained at 20 °C/15 °C day/night temperature settings and the light intensity varied between 400 and 1,000 µmol m−2 s−1 with a mean day length of 16 h. Developing tubers were harvested at approximately 10 weeks following shoot emergence and mature tubers from fully senesced plants at 20 weeks. For each progeny clone, tubers from three replicated clonal plants were harvested, pooled together then immediately frozen in liquid nitrogen followed by freeze drying then stored at −80 °C.
Analysis of potato tuber carotenoids
Total tuber carotenoids were extracted and both saponified and non-saponified extracts were analysed by reverse-phase high-performance liquid chromatography (RP-HPLC) as detailed in Morris et al. (2004). Peak responses were detected using a variable UV–Vis detector set at 450 nm on a Gynkotek UVD340 detector using Dionex Chromeleon software version 6.6. The remainder of the HPLC equipment consisted of a Dionex UltiMate 3000 autosampler and a Gykotek p580-A LPG pump. Three technical extractions were performed from each bulk replicate and the individual carotenoids identified by co-chromatography with an authentic standard (CarotenNature).
Total RNA extraction from potato tissues
Total RNA was extracted from freeze-dried potato tissues as described by Ducreux et al. (2008). The RNA quality and quantity was assessed using an RNA 6000 Nano Chip on an Agilent 2100 Bioanalyzer (http://www.chem.agilent.com).
Genomic DNA isolation and molecular marker assays
Details of genomic DNA isolation from leaves of the parents and progeny clones and of the Amplified Fragment Length Polymorphism (AFLP) and Simple Sequence Repeats (SSR) assays can be found in Bradshaw et al. (2004). Genomic DNA used for Diversity Array Technology (DArT) analysis was isolated using the procedure recommended by the Diversity Arrays Technology Pty Ltd(http://www.diversityarrays.com/sites/default/files/pub/DArT_DNA_isolation.pdf).
Quantitative RT-PCR
First-strand cDNA was synthesised using an Invitrogen Superscript III reverse transcriptase kit following the manufacturer’s instructions (Invitrogen Life Technologies, Carlsbad, California, USA)., 25 ng was used as a template for real-time PCR using the Universal Probe Library System (http://www.roche-applied-science.com/sis/rtpcr/upl/ezhome.html). Using a conserved region of the Chy2 gene, an assay was designed using the online designer tool. The primer sequences used for the analysis were (forward), CHY2_143F (5′-GCGCTGCCGTAAGTTCCGTT-3′) and (reverse), CHY2_143R (5′-TTATGAAGAAGGTAAAACAGGGT-3′) and probe number 143. Reactions were performed in 25 µl containing 1 X FastStart TaqMan® Probe Master, 0.2 µM gene specific primers and 0.1 µM probe. Samples were amplified using a Perkin Elmer ABI Prism 7500. Thermal cycling conditions were: 10 min denaturation at 95 °C followed by 40 cycles of 15 s at 94 °C, 60 s at 60 °C. Primers were first validated then the sample reactions performed in triplicate alongside elongation factor-1α specific primers as an internal control. Relative expression levels were calculated and the primers validated using the ∆∆C t method (Livak 1997).
Chy2 amplicon assay
Primers were designed to areas of conserved sequence in all 3 Chy2 alleles, (forward), crtRb2HAPLO8F, (5′-TCGAAGGACTTATTAATTGC-3′), (reverse), crtRb2HAPLO1R, (5′-GAAGCATGCCATAGTGCTCT-3′). The forward primer was radiolabelled with ATP[γ 33P] (Perkin Elmer, UK) using methods described by Russell et al. (1997). A standard PCR was performed using the radiolabelled forward primer and genomic DNA from all progeny and parents of populations 01H15 and 03TR2 as a template. The resulting PCR products were denatured to linearise the DNA and separated by electrophoresis using a standard acrylamide gel. Gels were dried and exposed to Kodak BioMax MR film for 2 days prior to development.
Linkage mapping
Genetic linkage analysis for both populations (01H15 and 03TR2) was performed using JoinMap® 4.1 (Van Ooijen 2006). Marker genotyping data were scored as a cross-pollinator (CP; a population resulting from a cross between two heterogeneously heterozygous and homozygous diploid parents, linkage phases unknown) cross-type as specified in JoinMap coding format. A series of quality checks on the data was performed before calculating the linkages. Co-segregating markers including those with ≥10 % missing data excluded before generating linkage groups (LGs). Genetic maps were constructed using the Kosambi mapping function (Kosambi 1944).
QTL analysis
QTL analysis was performed using MapQTL version 5.0 (Van Ooijen 2004). An initial rapid search for associations between markers and traits was performed using non-parametric mapping (Kruskal–Wallis analysis) followed by interval mapping. Logarithm of odds (LOD) thresholds for declaring QTL significance were determined by performing 1,000 permutations for each individual trait’s data (Churchill and Doerge 1994).
Microarray sample bulking
Bulked replicates were constructed by combining equal weights (200 mg) of freeze-dried tuber powder from 10 individual clones selected according to the desired carotenoid phenotype and/or Chy2 genotype. Bulks were constructed from samples harvested at two developing stages, designated the developing tuber and mature tuber stage tuber as defined in Morris et al. (2004). RNA was extracted from the bulked powder as described by Ducreux et al. (2008).
Microarray processing and data analysis
Analysis of gene expression using bulked tuber RNA samples was performed using the 44,000 element Potato Oligo Chip Initiative (POCI) microarray, first described by Kloosterman et al. (2008). Experimental design, array information and complete datasets are available from ArrayExpress (http://www.ebi.ac.uk/arrayexpress/; accession number E-TABM-1200). RNA labelling, array hybridisation, washing and scanning were as detailed in Stushnoff et al. (2010).
Data were extracted using Agilent FE software and imported into Genespring v 7.3 as previously described (Stushnoff et al. 2010). Normalisation of data was performed using the default Lowess algorithm and transformed to account for dye-swaps. Unreliable data flagged as absent in all replicate samples was discarded. Volcano plots were used to identify probes with significant differential expression (Student’s t test p value ≤0.05, fold-change ≥2) between each pool type for all the traits studied.
Zep allele analysis
The determination of Zep alleles was performed using the method and primers described by Wolters et al. (2010).
Tuber soluble sugar analysis
Sugars were extracted from freeze-dried tuber samples and analysed for sucrose, glucose and fructose content as described in Shepherd et al. (2006).
Results
Analysis of tuber carotenoid content in 01H15 and 03TR2 population clones
Carotenoid contents of mature tuber extracts were assayed for each clone in the 01H15 and 03TR2 populations over two seasons (2006 and 2008). The total carotenoid contents of individual clones varied by between 8- and 13-fold in these populations (Fig. 1a, b). Saponified extracts were compared with their non-saponified counterpart to estimate the esterified carotenoid content. Tables 1 and 2 summarises the main carotenoid components of the profiles obtained for the 01H15 and 03TR2 populations. In general, in population 01H15, the major tuber carotenoids were zeaxanthin, antheraxanthin and lutein, and up to 46 % of the carotenoid components were esterified. A small (ca. 3 %) amount of an unidentified carotenoid was also present. For 03TR2 tubers, the carotenoid profiles were more complex. In addition to zeaxanthin, antheraxanthin and lutein, some clones contained significant levels of neoxanthin and violaxanthin. Very unusually for potato, for some clones in both populations and the cream-fleshed parent DB271/39 in population 03TR2, β-carotene comprised a high proportion of tuber carotenoid content (up to 26 % of the total). Up to four unidentified carotenoids were present in the tuber extracts from the 03TR2 clones; however, their cumulative levels were generally less than 10 % of total carotenoid content.
Figure 2a and b shows the correlations between individual tuber carotenoid contents found in the 01H15 and 03TR2 populations. β-carotene, which represents between 0.1 and 18.5 % of the overall tuber carotenoid content in 01H15 clones and between 0.1 and 26.2 % in 03TR2 individuals, is the only tuber carotenoid to show a strong negative correlation with total tuber carotenoid content in samples from both seasons and in both populations. Additionally, β-carotene is strongly negatively correlated with zeaxanthin and antheraxanthin in both 01H15 and 03TR2 clones, in addition to violaxanthin and neoxanthin in population 03RTR2.
Correlation relationships between tuber carotenoid values in a population 01H15 b 03TR2 in data from two growing seasons. Values below the diagonal are from season 1 (2006), values above the diagonal are from season 2 (2008), the correlation comparison between the two seasons is presented along the diagonal. p values are displayed by colour with blue shades representing negative correlations and orange shades representing positive correlations, white boxes have a p value >0.05. S1 2006 samples, S2 2008 samples, NS non-saponified, SAP saponified, Neo neoxanthin, Vio violaxanthin, Unk unidentified carotenoid, Ant antheraxanthin, Lut lutein, Zea zeaxanthin, β-car β-carotene (colour figure online)
Previously, it has been observed that among 11 β-carotene hydroxylase 2 (Chy2) alleles only one dominant allele, designated allele 3, has a major tuber effect, changing white into yellow flesh colour (Wolters et al. 2010). In an attempt to identify different correlations between clones containing or lacking the Chy2 allele 3 both populations were assessed for the presence of the allele. This was achieved by first sequencing the full-length cloned parental PCR products to identify the alleles present, then by developing a radiolabelled PCR assay using primers covering conserved regions of sequence either side of a series of indels located in the second intron which was first described by Brown et al. (2006). A total of 3 Chy2 alleles, including the Wolters et al. allele 3, were identified in the parents and clones from the 01H15 population. The 03TR2 population contained only 2 Chy2 alleles, one of which was the Chy2 allele 3. A nucleotide alignment of the marker assay amplicons for all 3 alleles can be found in Online Resource II. Clones with the Chy2 allele 3 accumulated up to 13-fold more tuber carotenoids in 03TR2 clones and up to 11-fold more in the 01H15 population. Interestingly, no significant increase in tuber carotenoid content was measured in clones from the 01H15 population which contained two copies of the chy2 allele (Table 3). In both populations and in both seasons measured, subsets of clones with at least one copy of the Chy2 allele 3 contain significantly higher levels of antheraxanthin and zeaxanthin and lower relative proportions of lutein and β-carotene content compared with those lacking the allele (Fig. 3). This may be attributed to the increased expression level of the high carotenoid associated β-carotene hydroxylase resulting in increased flux through the carotenoid biosynthetic pathway.
Mean tuber carotenoid composition of 01H15 and 03TR2 clones grouped according to Chy2 genotype. Data is presented from two growing seasons. Genotype groups containing one or more copies of the Chy2 allele 3, designated A, are represented by grey shaded bars in all graphs. Groups of clones lacking the Chy2 allele 3, designated a1 and a2, are represented by the pattern filled bars. Error bars shown are the standard error of the mean. ***p < 0.001, *p < 0.05 for genotypes lacking the Chy2 allele 3 (a1a2, a1a1) versus groups of clones with the allele present (AA, Aa1, Aa2). The number of samples per genotype and the total carotenoid content is presented in Table 3
To test this hypothesis, the expression level of Chy2 was measured in a subset of 12 clones containing 0, 1 or 2 copies of the allele from both populations. The quantitative real-time PCR (RT-PCR) assay was designed to a conserved region of the Chy2 gene to ensure product amplification from all known allelic variants. As expected, Chy2 expression was significantly lower (Student’s t test p value ≤ 0.001) in the subset of clones lacking the Chy2 allele 3 compared with those containing at least one copy in both populations (Fig. 4a, b). Interestingly, no significant increase in Chy2 expression was measured in the subset of 01H15 clones homozygous for the Chy2 allele 3 (Fig. 4a), suggesting a dominant role for this allele and further implying that a single copy of the Chy2 allele 3 is sufficient for the enhanced Chy2 expression and the associated elevated concentrations of tuber carotenoids.
Quantitative RT-PCR analysis of the β-carotene hydroxylase gene Chy2 in a 01H15 clones, b 03TR2 clones. Values shown are the means of triplicate assays ± standard error. Results from each assay were measured from three technical extractions from a bulk powder of two biological replicates. Samples are highlighted according to Chy2 allele copy number, white absent, grey 1, striped 2. ***p < 0.001 for the group lacking the Chy2 allele versus the group with the allele present
Construction of linkage maps
A combination of SSR, AFLP and DArT molecular markers was used to generate linkage maps of the 01H15 and 03TR2 populations. The most numerous marker type were DArT markers with a total of 1,865 and 1,529 polymorphic markers obtained for the 01H15 and 03TR2 populations, respectively. Both datasets contained a significant number (>70 % of total) of duplicate redundant markers and markers which were identical apart from missing values. The physical genomic locations for DArT markers were obtained from the reference potato pseudomolecules and annotated with their chromosomal location where possible prior to map construction (Potato Genome Sequencing Consortium, 2011; Sharma et al. 2013). A small number of SSR markers were used as anchor markers in each population and a single carotenoid biosynthetic gene, Chy2, encoding a plastid associated β-carotene hydroxylase 2 gene, known to map to chromosome 3 in potato (Thorup et al. 2000), was also mapped in both populations.
Linkage analysis of the 01H15 population using JoinMap® 4.1 formed 12 distinct linkage groups at a LOD score between 3.5 and 8.5 with 51 markers forming many fragmented smaller groups. The finalised set of 12 linkage groups contained a total of 397 evenly distributed markers with an overall length of 1,116.2 centiMorgans (cM) (Online Resource IIIa). The 12 linkage groups could be assigned to the 12 potato chromosomes by virtue of ‘known location’ SSR and DArT markers. Assuming that the potato genome contains 840 million base pairs (Mb) (http://www.potatogenome.net), each cM represents on average approximately 752.1 kilobase base pairs (kb) of nucleotide sequence. Key features of each of the linkage maps are summarised in Table 4 and the complete data set presented in Online Resource IIIa.
For linkage analysis of the 03TR2 population, a total of 428 DArT markers were used in the construction of linkage groups. Following annotation of individual DArT markers with their known chromosome location it was apparent that chromosomes 6 and 8 contained far fewer polymorphic markers in comparison with other linkage groups. Linkage analysis using JoinMap® 4.1 detected 12 groups (16–92 markers) segregating at LOD scores between 4.0 and 11.0; however, no groups of markers corresponding to chromosomes 6 and 8 sufficient for linkage were formed although linkage maps for the remaining chromosomes were readily constructed. As results of previous studies (Sharma et al. 2013) and those presented for the 01H15 population in this study have demonstrated a good coverage of DArT markers on both chromosome 6 and 8, the lack of polymorphic markers available in this area for the 03TR2 population suggests both parents have low levels of polymorphism in these linkage groups. Online Resource IIIb and Table 5 detail the finalised linkage maps.
As many of the DArT markers are polymorphic in both populations a comparison of the mapped marker order can be made. The genetic maps from both populations show good correspondence with each other, as illustrated in Fig. 5, depicting a comparison between chromosome 9 maps which share 22 common markers in generally consistent map order positions.
QTL analysis
Chromosomal loci that were associated with modified tuber carotenoid content were identified using the genetic maps and phenotypic data acquired for the 01H15 and 03TR2 populations from two growing seasons. Initial identification of QTL was performed using the non-parametric Kruskal–Wallis (KW) test which tests each locus independently. A significance threshold p value of 0.0005 (KW value > 12.6) was selected as a baseline for identifying significant QTL and the results are summarised in Tables 6 and 7. Efforts were made to include all markers with significant KW values above the stated threshold for various carotenoid traits in the linkage maps for both populations. Interval mapping using MapQTL ® was then carried out on individual linkage groups containing markers possessing significant KW value to estimate the percentage of variance explained and to give a LOD profile for each QTL locus.
As previously reported by Thorup et al. (2000), this study also identified a major QTL for tuber carotenoid content on chromosome 3 in both populations. QTLs for antheraxanthin, β-carotene, carotenoid esters, violaxanthin and zeaxanthin were detected in both populations at the same locus with an additional effect on lutein in the second season in the 03TR2 population (Tables 6, 7). A second smaller QTL for total carotenoid content is present on chromosome 3 in both populations, identified by plotting the KW value against the physical location of each DArT marker in the potato chromosome 3 genome sequence (Fig. 6). This QTL is not identified following restricted interval mapping using the Chy2 marker as a co-factor (data not shown) as the markers involved are not significantly separated on the linkage map. A third major total carotenoid QTL was also detected on chromosome 9 at approximately 70 cM in both populations. This QTL was also present for lutein content in both seasons, as well as antheraxanthin, carotenoid esters, total carotenoid in the 2008 season and zeaxanthin in the 01H15 population only (Table 6).
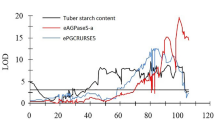
Elucidation of a second QTL for tuber carotenoid content present on chromosome 3 in populations 01H15 and 03TR2 determined by marker physical mapping. Figure shows a scatter plot representation of the physical map location (Mb) of markers present on chromosome 3 in both populations and the corresponding Kruskal–Wallis marker scores for total tuber carotenoid content in two independent growing seasons. The Kruskal–Wallis threshold (12.6) for defining QTL significance is indicated. Markers are annotated according to the genetic information obtained by the parental segregation. Markers annotated by a triangle have genetic information provided by both parents (hkxhk); Square first parent only (lmxll); Rhombus second parent only (nnxnp), Circle is the Chy2 gene marker
Determination of Zep genotype
As alleles at the Zep locus have previously been shown to influence tuber carotenoid content (Zhou et al. 2011), a QTL for carotenoid traits might be expected at the locus on chromosome 2 where the single potato Zep gene maps (Thorup et al. 2000). As no QTL were detected in this region (Tables 4, 5) the parental Zep genotypes were determined for both populations. Using an allelic assay described by Wolters et al. (2010), primers AWZEP9 and AWZEP10 were used to amplify a fragment of the Zep gene from both parental DNA samples from the 01H15 and 03TR2 populations. A single 535 bp amplicon was produced for each parent sample and aligned to sequences published by Wolters et al. (2010), confirming a 100 % match to the recessive Zep allele 1 associated with a reduced level of expression in orange-fleshed diploid genotypes (Wolters et al. 2010; Zhou et al. 2011). The aligned parental sequences from the 01H15 and 03TR2 parents are presented in Online Resource IV.
Bulked transcriptome analysis to identify candidate genes associated with QTL from the 03TR2 population
A pooling strategy was employed to identify candidate genes underlying QTL for the high and low carotenoid trait. RNA bulks were constructed from both developing and mature tubers and analysed using a potato microarray (Kloosterman et al. 2008).
In our initial bulking experiment, high tuber carotenoid bulks where genotypes possess at least one copy of the high carotenoid Chy2 allele 3 were compared with bulks from low tuber carotenoid individuals lacking this allele. A total of four bulks were constructed for each stage of tuber development (two high carotenoid, two low carotenoid at the developing and mature tuber stage), each consisting of 10 genotypes exhibiting the requisite carotenoid content. Thus, for each stage, there were four independent pools of RNA each representing 10 different genotypes. Genes showing consistent differential expression (>twofold) across the four bulk comparisons were considered as candidate genes at each developmental stage. Microarray analysis identified 24 genes that were significantly up-regulated (≥twofold, p ≤ 0.05), the largest fold increase was represented by probe BSK01458FB06.t3m.scf_145, annotated as encoding a miraculin gene which was expressed 376-fold higher in the high tuber carotenoid bulk at the mature stage. A total of 19 probes/genes were down-regulated (≤0.5-fold, p ≤ 0.05) with probe bf_mxlfxxxx_0031f03.t3m.scf_102 which represents chalcone synthase B expressed up to 20-fold lower in the high carotenoid bulks at the mature stage(Online Resource V). Interestingly, Chy2 was the only known carotenoid biosynthetic gene differentially expressed in this experiment and amongst one of the most highly up-regulated genes, reaching levels 176-fold higher in the high tuber carotenoid bulks and thereby corroborating the evidence collected from previous studies (Kloosterman et al. 2010, Wolters et al. 2010). Using the potato genome sequence (The Potato Genome Sequencing Consortium. 2011) the genomic location of >85 % of the genes represented on the POCI array can be identified. A high proportion (16 of 43) of the differentially expressed genes from this experiment mapped to a 7.2 Mb region of chromosome 3 which contains a total of 572 gene models, including the Chy2 gene; and directly underpins the associated QTL (Fig. 7). This proof of principal of our bulking strategy and subsequent microarray analysis proved very effective in defining genomic regions influencing the tuber carotenoid trait, as well as validating the QTL locations.
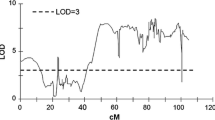
Graphical comparison of a major QTL for total tuber carotenoid content in the 2008 season present on chromosome 3 in the 03TR2 population, presented in cM, and the physical locations (Mb) of markers and differentially expressed microarray probes present within the QTL interval. A LOD threshold of 25 was used to determine the QTL interval and the end markers are indicated by two asterisks. The QTL significance threshold (LOD 3.2) as determined by a permutation test is shown on the LOD graph. The microarray probe ID (POCI ID), physical map position (Mb) and related gene annotation are shown; for further information see Online Resource 6
Further bulks were constructed based on different tuber carotenoid phenotypes, for example, a bulk comprising individuals with at least one copy of the high tuber carotenoid conferring dominant Chy2 allele. Despite this, there was a wide range (threefold) of tuber carotenoid contents within this group, allowing bulks of low and high tuber carotenoid content to be constructed from this group (Online Resource VI). As all individuals contained both the high tuber carotenoid Chy2 allele 3 and Zep allele 1, the intention was to identify genes associated with QTL other than the one on chromosome 3. Microarray analysis revealed 102 differentially expressed genes, 65 with increased expression level and 37 with lower expression level (Online Resource VII) The gene represented by probe cPRO26K2TH_644 annotated as encoding an early light inducible protein (ELIP) gene showed the largest fold-increase in expression No carotenoid biosynthetic genes were present in the generated gene lists. The physical map locations of the differentially expressed genes were investigated, and 20 of the genes mapped to chromosome 9, 11 of these co-localising within an 8.5 Mb section of the chromosome underpinning QTLs for lutein, antheraxanthin, carotenoid esters, and total carotenoid contents (Fig. 8). Examination of the gene content in this genomic region (52.41 – 60.87 Mb) identified a total of 660 gene models. The number of candidate genes can be further reduced to 309 when considering only the QTL area defined by a 1 LOD drop either side of the highest scoring marker (~62 – 80 cM on the genetic map and 53.15 – 56.97 Mb on the physical map). However, both regions were devoid of any known carotenoid biosynthetic genes, indicating that other regulatory or novel yet uncharacterised genes must be present in the identified genomic interval. The map locations of known carotenoid biosynthetic genes are provided in Online Resource VIII.
Graphical comparison of a major QTL for non-saponified lutein content in the 2008 season present on chromosome 9 in the 03TR2 population, presented in cM, and the physical locations (Mb) of markers and differentially expressed microarray probes present within the QTL interval. A LOD threshold of 6 was used to determine the QTL interval and the end markers are indicated by two asterisks. The QTL significance threshold (LOD 3.2) as determined by a permutation test is shown on the LOD graph. The microarray probe ID (POCI ID), physical map position (Mb) and related gene annotation are shown; for further information see Online Resource 8
Tuber soluble sugar analysis
Recently, it has been suggested that the level of Chy2 expression and hence tuber carotenoid content is correlated with the levels of tuber glucose and sucrose (Zhou et al. 2011). This finding was based on comparison of carotenoid and tuber sugar levels in two S. tuberosum cultivars. As Chy2 expression is clearly also an important factor in tuber carotenoid accumulation in the populations described in this study, the opportunity was taken to investigate the potential relationship between tuber soluble sugar and carotenoid contents. Soluble sugar contents (sucrose, glucose and fructose) were compared in 26 genotypes from the 03TR2 population including the parents. Nine genotypes, including the parent DB271(39), were sampled from the low tuber carotenoid group lacking the Chy2 allele 3 associated with high tuber carotenoid content. The remaining 17 samples were taken from genotypes containing one copy of the Chy2 allele 3 which also included the parent, DB375(2). There was no significant correlation between the total tuber soluble sugar content and the levels of tuber carotenoids (p = 0.79), or in the levels of the individual sugars, sucrose (p = 0.69), glucose (p = 0.87) or fructose (p = 0.99), in contrast to the findings for two Tuberosum cultivars (Online Resource IX, Zhou et al. 2011).
Discussion
Although polymorphism in the β-carotene hydroxylase 2 gene may explain some of the variation in potato tuber carotenoid content, variation in other carotenoid pathway genes including Zep may also be important (Brown et al. 2006). Indeed, Wolters et al. (2010) identified a recessive Zep allele that apparently acts in concert with Chy2 to give orange-fleshed tubers. Despite these advances, however, until now there has been no reported genome-wide analysis for loci influencing tuber carotenoid content. As it seems highly likely that less obvious candidate genes also impact on carotenoid biosynthesis (Liu et al. 2003), this study was designed to investigate the genetic structure of the tuber carotenoid trait using two bi-parental populations. A further aim was to use a ‘genetical genomics’ approach in conjunction with QTL analysis to identify genetic loci other than those harbouring known structural genes of the biosynthetic pathway that may have a role in influencing tuber carotenoid content.
A detailed analysis of the carotenoid content of two S. tuberosum Group Phureja F1 populations which segregate in different ways for overall tuber carotenoid concentrations and the individual carotenoid constituents has been carried out. The tuber carotenoid concentrations measured in the two populations were comparable to values published in previous studies for yellow-fleshed potatoes (Breithaupt and Bamedi 2002; Morris et al. 2004). One notable difference between the 01H15 and 03TR2 progeny clones was the extent to which they accumulated the carotenoids zeaxanthin and lutein. The 01H15 clones were typically high in zeaxanthin and low in lutein content (Table 1). In contrast, 03TR2 tuber samples are typically high in lutein and low in zeaxanthin (Table 2). As these carotenoids are on different branch points of the carotenoid biosynthetic pathway, catalysed by a combination of different enzymatic steps, these findings suggest the mechanisms controlling individual carotenoid sequestration are somewhat different in each population. It is also of interest that the parents of both 01H15 and 03TR2 are homozygous for the recessive Zep allele associated in some populations with high tuber zeaxanthin content (Wolters et al. 2010). This explains why there is no detectable QTL for carotenoid traits at the chromosome 2 Zep locus. Clearly, in 03TR2 genotypes other genetic factors over-ride the effects of the recessive Zep allele, resulting in low zeaxanthin content. Interestingly, low tuber carotenoid individuals of both populations contained elevated levels of β-carotene up to 13-fold greater than those previously described in non-transgenic modified potato (Nesterenko and Sink 2003). The low tuber carotenoid individuals also contained higher relative proportions of lutein and lower levels of xanthophylls, produced downstream of β-carotene, compared to individuals containing high levels of carotenoids, suggesting that individuals containing low levels of tuber carotenoids hydroxylate β-carotene less efficiently. This hypothesis is further strengthened by the quantitative real-time expression analysis which showed significantly higher expression of Chy2 in individuals containing at least one copy of the high carotenoid Chy2 allele. In general, there were many positive correlations between individual carotenoids (Fig. 2). However, β-carotene had a negative correlation with several carotenoids, notably with zeaxanthin and antheraxanthin, a lower correlation with neoxanthin and violaxanthin, and a very small association with lutein. These results are fully consistent with presence of a gene acting to block or permit a flow of metabolites through the carotenoid biosynthesis pathway at β-carotene. The most profound effect was on zeaxanthin and antheraxanthin, the products of the reaction catalysed by β-carotene hydroxylase. There may be feedback regulation affecting the flux through the pathway and so altering the tuber total carotenoid content as a result, or differences may be occurring in the control of carotenoid breakdown.
Recent evidence has shown that the Chy2 promoter is responsive to glucose and sucrose and a high tuber carotenoid group Tuberosum cultivar contained a higher content of glucose and sucrose than a low carotenoid cultivar (Zhou et al. 2011). From an examination of 26 genotypes from the 03TR2 population, no relationship was evident between Chy2 expression or tuber carotenoid content with tuber sugar levels. It is possible that there are population dependent differences in the response to soluble sugars and it would be interesting to explore this aspect in a wider range of potato germplasm.
Genetic maps for both populations were constructed using a combination of marker types (AFLP, SSRs and DArTs). The development of DArT markers has provided a low-cost high-throughput method of saturating linkage maps with markers. All potato DArT markers have been sequenced and aligned to the latest version of the potato genome assembly (Sharma et al. 2013), so they are particularly useful for locating the actual genomic locations corresponding to the gene models identified near mapped QTLs. The physical locations of the POCI microarray gene probes on the potato genome were also determined and thereby allowing QTL maps and locations of genes identified by eQTL analysis to be directly compared for the first time in potato, thus facilitating candidate gene identification in this important crop plant.
Genome-wide QTL analysis of tuber carotenoid content was carried out for two bi-parental diploid group Phureja populations and major QTL were identified in both. The major chromosome 3 QTL is most likely due to the action of a dominant Chy2 allele. However, a number of previously undefined carotenoid content QTL, some with smaller effect, were revealed by the current study (Tables 6, 7; Fig. 5). Notable among these is a QTL on chromosome 9 in both populations, associated with carotenoid components including β-carotene, zeaxanthin, violaxanthin and antheraxanthin. This QTL has not previously been observed in potato. The chromosome 9 QTL and the differentially expressed genes identified by bulked transcriptome analysis mapped to an 8.5 Mb covering a total of 660 gene models. Here, we demonstrate that using a novel phenotypic pooling strategy using bulks designed according to phenotypic and genotypic data, a segregating potato population can be rapidly and cost-effectively screened for the association of variance in gene expression with the quantitative trait data. Genetic variations underlying the QTL can influence different aspects of gene transcription either directly or indirectly. Within the list of differentially expressed genes, it is possible, in some cases, to identify candidate genes responsible for the trait. Clearly, for the chromosome 3 QTL, Chy2 is strongly differentially expressed. For the newly identified chromosome 9 QTL, we cannot yet recognise any obvious carotenoid biosynthetic gene within the list of differentially expressed genes or the genes annotated within the genomic region spanned by the QTL interval. It is intriguing, however, that one of the most strongly differentially expressed genes (approximately 16-fold up-regulated in both developing and mature tuber bulks) from this experiment (Fig. 8, Online Resource VII) is annotated as encoding an early light inducible protein. It has been suggested that these proteins have a role in plastid transition processes (Bruno and Wetzel 2004) possibly consistent with their up-regulation in genotypes with elevated tuber carotenoid content. It may well be significant that the chromosomal location of the early light-inducible protein gene (54.45 Mb) coincides with the peak of the chromosome 9 QTL. We believe these examples aptly illustrate the power of combining conventional QTL analysis approach with a novel pooled transcriptomic analysis in increasing the focus and resolution of genetic mapping studies; and in current case, for identifying new candidate genes possibly influencing carotenoid content in potato tubers.
References
Bonierbale MW, Plaisted RL, Tanksley SD (1988) RFLP maps based on a common set of clones reveal modes of chromosomal evolution in potato and tomato. Genetics 120:1095–1103
Bradshaw JE, Pande B, Bryan GJ, Hackett CA, McLean K, Stewart HE, Waugh R (2004) Interval mapping of quantitative trait loci for resistance to late blight [Phytophthora infestans (Mont.) de Bary], height and maturity in a tetraploid population of potato (Solanum tuberosum subsp. tuberosum). Genetics 168:983–995
Breithaupt DE, Bamedi A (2002) Carotenoids and carotenoid esters in potatoes (Solanum tuberosum L.): new insights into an ancient vegetable. J Agric Food Chem 50:7175–7181
Brown CR, Edwards CG, Yang CP, Dean BB (1993) Orange flesh trait in potato: inheritance and carotenoid content. J Am Soc Hort Sci 118:145–150
Brown CR, Kim TS, Ganga Z, Haynes K, De Jong D, Jahn M, Paran I, De Jong W (2006) Segregation of total carotenoid in high level potato germplasm and its relationship to beta-carotene hydroxylase polymorphism. Am J Pot Res 83:365–372
Bruno AK, Wetzel CM (2004) The early light-inducible protein (ELIP) gene is expressed during the chloroplast-to-chromoplast transition in ripening tomato fruit. J Exp Bot 55:2541–2548
Campbell R, Ducreux LJM, Morris WL, Morris JA, Suttle JC, Ramsay G, Bryan GJ, Hedley PE, Taylor MA (2010) The metabolic and developmental roles of carotenoid cleavage dioxygenase 4 from potato (Solanum tuberosum L). Plant Physiol 154:656–664
Cazzonelli CI, Cuttriss AJ, Cossetto SB, Pye W, Crisp P, Whelan J, Finnegan EJ, Turnbull C, Pogson BJ (2009) Regulation of carotenoid composition and shoot branching in arabidopsis by a chromatin modifying histone methyltransferase, SDG8. Plant Cell 21:39–53
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Cuevas HE, Staub JE, Simon PW, Zalapa JE, McCreight JD (2008) Mapping of genetic loci that regulate quantity of beta-carotene in fruit of US Western Shipping melon (Cucumis melo L.). Theor Appl Genet 117:1345–1359
DellaPenna D, Pogson BJ (2006) Vitamin synthesis in plants: tocopherols and carotenoids. Annu Rev Plant Biol 57:711–738
Ducreux LJ, Morris WL, Prosser IM, Morris JA, Beale MH, Wright F, Shepherd T, Bryan GJ, Hedley PE, Taylor MA (2008) Expression profiling of potato germplasm differentiated in quality traits leads to the identification of candidate flavour and texture genes. J Exp Bot 59(15):4219–4231
Fraser PD, Bramley PM (2004) The biosynthesis and nutritional uses of carotenoids. Prog Lipid Res 43:228–265
Giuliano G, Tavazza R, Diretto G, Beyer P, Taylor M (2008) Metabolic engineering of carotenoid biosynthesis in higher plants. Trends Biotechnol 26:139–145
Haynes KG (2010) Genotype-environment interactions for potato tuber carotenoid content. J Amer Soc Hort Sci 135(3):250–258
Hirschberg J (2001) Carotenoid biosynthesis in flowering plants. Curr Opin Plant Biol 4:210–218
Kloosterman B, De Koeyer D, Griffiths R, Flinn B, Steuernagel B, Scholz U, Sonnewald S, Sonnewald U, Bryan GJ, Bánfalvi Z, Hammond JP, Geigenberger P, Nielsen KL, Visser RGF, Bachem CWB (2008) The potato transcriptome: a new look at transcriptional changes during tuber development using the POCI array. Comp Funct Genom 8:329–340
Kloosterman B, Oortwijn M, uit de Willigen J, America T, de Vos R, Visser, RGF Bachem CWB (2010) From QTL to candidate gene: Genetical genomics of simple and complex traits in potato using a pooling strategy. BMC Genomics 11:158
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Li L, Paolillo DJ, Parthasarathy MV, Dimuzio EM, Garvin DF (2001) A novel gene mutation that confers abnormal patterns of beta-carotene accumulation in cauliflower (Brassica oleracea var. botrytis). Plant J 26:59–67
Liu YS, Gur A, Ronen G, Causse M, Damidaux R, Buret M, Hirschberg J, Zamir D (2003) There is more to tomato fruit colour than candidate carotenoid genes. Plant Biotechnol J 1:195–207
Livak KJ (1997) “Relative quantification of gene expression” User Bulletin No.2: ABI PRISM 7700 Sequence detection system. PE Applied Biosystems, Foster City
Lu W, Haynes K, Wiley E, Clevidence B (2001) Carotenoid content and color in diploid potatoes. J Am Soc Hortic Sci 126:722–726
Morris WL, Ducreux L, Griffiths DW, Stewart D, Davies HV, Taylor MA (2004) Carotenogenesis during tuber development and storage in potato. J Exp Bot 55:975–982
Nesterenko S, Sink KC (2003) Carotenoid profiles of potato breeding lines and selected cultivars. HortScience 38(6):1173–1177
Ohmiya A, Kishimoto S, Aida R, Yoshioka S, Sumitomo K (2006) Carotenoid cleavage dioxygenase (CmCCD4a) contributes to white color formation in chrysanthemum petals. Plant Physiol 142:1193–1201
Römer S, Lubeck J, Kauder F, Steiger S, Adomat C, Sandmann G (2002) Genetic engineering of a zeaxanthin-rich potato by antisense inactivation and co-suppression of carotenoid epoxidation. Metab Eng 4:263–272
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor J, Powell W, Waugh R (1997) Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theor Appl Genet 95:714–722
Sharma SK, Bolser D, de Boer J, Sonderkaer M, Amoros W, Federico Carboni M, D’Ambrosio JM, de la Cruz G, De Genova A, Douches DS, Eguiluz M, Guo X, Guzman F, Hackett CA, Hamilton JP, Li G, Li Y, Lozano R, Maass A, Marshall DF, Martinez D, McLean K, Mejia N, Milne L, Munive S, Nagy I, Ponce O, Ramirez M, Simon R, Thomson SJ, Torres Y, Waugh R, Zhang Z, Huang S, Visser RGF, Bachem CWB, Sagredo B, Feingold SE, Orjeda G, Veilleux RE, Bonierbale M, Jacobs JME, Milbourne D, Martin DMA, Bryan GJ (2013) Construction of reference chromosome-scale pseudomolecules for potato: integrating the potato genome with genetic and physical maps. G3 Genes Genomes Genet 3.11:2031–2047
Shepherd LVT, McNicol JW, Razzo R, Taylor MA, Davies HV (2006) Assessing the potential for unintended effects in genetically modified potatoes perturbed in metabolic and developmental processes. Targeted analysis of key nutrients and anti-nutrients. Trans Res 15:409–425
Stushnoff C, Ducreux LJM, Hancock RD, Hedley PE, Holm D, McDougall GJ, McNicol JW, Morris J, Morris WL, Sungurtas J, Verrall SR, Zuber T, Taylor MA (2010) Flavonoid profiling and transcriptome analysis reveals new gene-metabolite correlations in tubers of Solanum tubersoum L. J Exp Bot 61:1225–1238
Taylor M, Ramsay G (2005) Carotenoid biosynthesis in plant storage organs: recent advances and prospects for improving plant food quality. Physiol Plant 124(2):143–151
The Potato Genome Sequencing Consortium (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Thorup TA, Tanyolac B, Livingstone KD, Popovsky S, Paran I, Jahn M (2000) Candidate gene analysis of organ pigentation loci in the Solanaceae. Pro Nat Acad Sci USA 97:11192–11197
Tian L, Pang Y, Dixon RA (2008) Biosynthesis and genetic engineering of proanthocyanidins and (iso)flavonoids. Phytochem Rev 7:445–465
Van Eck J, Conlin B, Garvin DF, Mason H, Navarre DA, Brown CR (2007) Enhancing beta-carotene content in potato by RNAi-mediated silencing of the beta-carotene hydroxylase gene. Amer J Potato Res 84:331–342
Van Ooijen JW (2004) MapQTL® 5, Software for the mapping of quantitative trait loci in experimental populations. Kyazma B.V. Wageningen, Netherlands
Van Ooijen JW (2006) JoinMap® 4, Software for the calculation of genetic linkage maps in experimental populations. Kyazma B.V. Wageningen, Netherlands
Welsch R, Maass D, Voegel T, DellaPenna D, Beyer P (2008) The transcription factor RAP2.2 and its interacting partner SINAT2—stable elements in the carotenogenesis of Arabidopsis leaves. Plant Physiol 145:1073–1085
Wolters AMA, Uitdewilligen JGAML, Kloosterman BA, Hutten RCB, Visser RGF, van Eck HJ (2010) Identification of alleles of carotenoid pathway genes important for zeaxanthin accumulation in potato tubers. Plant Mol Biol 73:659–671
Zhou X, McQuinn R, Fei Z, Wolters AA, Van Eck J, Brown C, Giovannoni JJ, Li L (2011) Regulatory control of high levels of carotenoid accumulation in potato tubers. Plant Cell Environ 34:1020–1030
Acknowledgments
This work was funded by the Scottish Government Rural and Environmental Research and Analysis Directorate, EU-SOL project number PL 016214 and EU-FP7 METAPRO 244348. The authors would also like to thank Christine Hackett of Biomathematics and Statistics, Scotland for assistance and advice on linkage and QTL analysis.
Conflict of interest
The authors declare they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Christiane Gebhardt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2014_2349_MOESM1_ESM.pdf
Online Resource I. The James Hutton Institute in-house visual tuber flesh colour standard scoring chart. The illustration represents the tuber flesh colour attributed individual colour scores. (PDF 272 kb)
122_2014_2349_MOESM2_ESM.pdf
Online Resource II. Chy2 haplotype assay. Figure shows the sequence variations in the second intron of the full length nucleotide sequence within the 3 alleles. Primer sequences are highlighted in red text. Amplicon sizes; allele 1, 213 bp; allele 2, 207 bp; allele 3, 206 bp. (PDF 314 kb)
122_2014_2349_MOESM5_ESM.pdf
Online Resource IV. 01H15 and 03TR2 parental Zep amplicon sequences elucidated using the allele assay and primer sequences described by Wolters et al (2010). (PDF 150 kb)
122_2014_2349_MOESM6_ESM.pdf
Online Resource V. Microarray experiment 1 dataset. Comparison of 03TR2 clones bulked according to tuber carotenoid content. The developing and mature stage expression data are presented compared to wild type (P <0.05). The physical map position and gene annotation are presented for each individual microarray probe. (PDF 101 kb)
122_2014_2349_MOESM7_ESM.pdf
Online Resource VI. Tuber carotenoid contents (µg g-1 DW) of individual 03TR2 clones in two seasons selected for bulked microarray analysis. Rhombus, low carotenoid bulk replicate 1; Square, low carotenoid bulk replicate 2; Triangle, high carotenoid bulk replicate 1; Circle, high bulk replicate 2. (PDF 124 kb)
122_2014_2349_MOESM8_ESM.pdf
Online Resource VII. Physical map locations of the known carotenoid biosynthetic genes. The transcript ID, gene ID and superscaffold location are annotated according to The Potato Genome Browser version 4.03. (PDF 39 kb)
122_2014_2349_MOESM9_ESM.pdf
Online Resource VIII. Microarray experiment 2 dataset. Comparison of 03TR2 clones bulked according to Chy2 allele dosage and tuber carotenoid content. The developing and mature stage expression data are presented compared to wild type (P <0.05). The physical map position and gene annotation are presented for each individual microarray probe. (PDF 169 kb)
122_2014_2349_MOESM10_ESM.pdf
Online Resource IX. Sugar content of select 03TR2 clones containing varying levels of tuber carotenoid contents. (a) Total tuber sugar content, (b) tuber glucose content, (c) tuber fructose content, (d) tuber sucrose content. Values shown are the means of three technical extractions of a two sample bulk powder. Error bars indicate the standard error. (PDF 43 kb)
Rights and permissions
About this article
Cite this article
Campbell, R., Pont, S.D.A., Morris, J.A. et al. Genome-wide QTL and bulked transcriptomic analysis reveals new candidate genes for the control of tuber carotenoid content in potato (Solanum tuberosum L.). Theor Appl Genet 127, 1917–1933 (2014). https://doi.org/10.1007/s00122-014-2349-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-014-2349-0