Abstract
Key message
Stripe rust resistance transferred from Thinopyrum intermedium into common wheat was controlled by a single dominant gene, which mapped to chromosome 1B near Yr26 and was designated YrL693.
Abstract
Stripe rust caused by Puccinia striiformis f. sp. tritici (Pst) is a highly destructive disease of wheat (Triticum aestivum). Stripe rust resistance was transferred from Thinopyrum intermedium to common wheat, and the resulting introgression line (L693) exhibited all-stage resistance to the widely virulent and predominant Chinese pathotypes CYR32 and CYR33 and to the new virulent pathotype V26. There was no cytological evidence that L693 had alien chromosomal segments from Th. intermedium. Genetic analysis of stripe rust resistance was performed by crossing L693 with the susceptible line L661. F1, F2, and F2:3 populations from reciprocal crosses showed that resistance was controlled by a single dominant gene. A total 479 F2:3 lines and 781 pairs of genomic simple sequence repeat (SSR) primers were employed to determine the chromosomal location of the resistance gene. The gene was linked to six publicly available and three recently developed wheat genomic SSR markers. The linked markers were localized to wheat chromosome 1B using Chinese Spring nulli-tetrasomic lines, and the resistance gene was localized to chromosome 1B based on SSR and wheat genomic information. A high-density genetic map was also produced. The pedigree, molecular marker data, and resistance response indicated that the stripe rust resistance gene in L693 is a novel gene, which was temporarily designated YrL693. The SSR markers that co-segregate with this gene (Xbarc187-1B, Xbarc187-1B-1, Xgwm18-1B, and Xgwm11-1B) have potential application in marker-assisted breeding of wheat, and YrL693 will be useful for broadening the genetic basis of stripe rust resistance in wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe rust caused by Puccinia striiformis f. sp. tritici (Pst) continues to be one of the most devastating diseases threatening wheat (Triticum aestivum L.) yields worldwide, especially in wheat-growing areas with cool, moist climates. Based on survey data collected over the last 10 years, China is the largest region for wheat stripe rust in the world, with an average of 4 million hectares potentially affected each year (Li et al. 2006b; Liu et al. 2013). For example, 6.6, 4.9, and 4.08 million hectares of wheat in China were affected by stripe rust in 2002, 2003, and 2009, respectively (Kang et al. 2010).
Stripe rust is most destructive to wheat production in northwest and southwest China (Luo et al. 2005; Wan et al. 2004). With the prevalence of Pst races CYR32 and CYR33, stripe rust has become existing major threat to wheat production because only a few of the known resistance genes (Yr5, Yr10, Yr15, Yr18, Yr24/Yr26, Yr36, Yr39, and Yr41) are effective against these races (Chen et al. 2009; Kang et al. 2010; Li et al. 2006a, b, c; Luo et al. 2009a; Wan et al. 2004). Additionally, a recently identified race, V26, overcomes the resistance gene Yr24/Yr26/YrChuanmai42 present in a number of varieties grown in the region (Han et al. 2012; Liu et al. 2010).
The development of host resistance by selective breeding is the most effective, economical, environmentally sound, and consistently used method of controlling stripe rust in wheat (Chen 2005). To date, more than 60 designated resistance genes or alleles and more than 40 temporarily assigned genes have been identified on all wheat chromosomes except 1A and 7A (http://wheat.pw.usda.gov/; Xu et al. 2013; McIntosh pers. comm.). Among these genes, only Yr9 and Yr24/Yr26 have been widely used to develop stripe rust-resistant cultivars in wheat breeding programs in China. In hexaploid wheat (AABBDD), microsatellites or simple sequence repeats (SSRs) have been used to locate and map many stripe rust resistance genes within specific wheat chromosomes or chromosomal arms; these genes include Yr5 (Sun et al. 2002), Yr17 (Jia et al. 2011), Yr18 and Yr29 (Lillemo et al. 2008), Yr24/Yr26 (Li et al. 2006a), Yr36 (Uauy et al. 2005), Yr41 (Luo et al. 2008), Yr43 and Yr44 (Cheng and Chen 2010), Yr45 (Li et al. 2011), Yr46 (Herrera-Foessel et al. 2011), Yr47 (Bansal et al. 2011), Yr48 (Lowe et al. 2011), Yr50 (Liu et al. 2013), Yr52 (Ren et al. 2012), and Yr53 (Xu et al. 2013).
Alien gene transfer is an important means of increasing the genetic diversity of disease resistance in wheat. Of the permanently named Yr genes, 14 were transferred from wild relatives, including Aegilops comosa Sm., Ae. geniculata Host, Ae. kotschyi Boiss, Ae. neglecta Bertol, Ae. sharonensis Eig., T. dicoccoides Körn., T. tauschii Coss., T. ventricosum Ces., Dasypyrum villosum L., and cereal rye (Secale cereale L.) (http://wheat.pw.usda.gov/; Chen 2005; Xu et al. 2013). Thinopyrum intermedium (Host) Barkworth and D.R. Dewey (2n = 6x = 42; JJJsJsSS) (syn. Elytrigia intermedia [Host] Nevski) has been hybridized extensively with wheat and has also proven to be a useful source of disease resistance in hexaploid wheat (T. aestivum L.) (2n = 42; AABBDD) (Liu et al. 2013; Luo et al. 2009a).
A considerable body of evidence supports the hypothesis that Th. intermedium is a valuable gene pool for wheat disease resistance because of its resistance to wheat streak mosaic virus (Friebe et al. 1996), Fusarium head blight (FHB) (Fedak and Han 2005), leaf rust (Autrique et al. 1995), stem rust (Fedak 1999), and powdery mildew (Liu and Wang 2005). Two powdery mildew resistance genes, Pm40 and Pm43, and one stripe rust resistance gene, Yr50, were transferred from Th. intermedium to common wheat (He et al. 2009; Liu et al. 2013; Luo et al. 2009b). The wheat line L693, which is resistant to stripe rust, powdery mildew, and FHB, was developed from F6:7 families of a cross between Mianyang 11 (MY11) and YU25. The latter was derived from Th. intermedium by an interspecific cross (Zhang et al. 2011). The stripe rust resistance in L693 is conferred by the temporarily designated gene YrYU25 (Luo et al. 2009a). The objectives of the present study were to study the inheritance of stripe rust resistance in L693, to identify the chromosomal location of the resistance gene, and to map the gene with the eventual objective of marker-assisted selection, fine mapping, and map-based cloning.
Materials and methods
Plant materials
Wheat line L693, which is resistant to stripe rust, and susceptible line L661 were selected from F6:7 families of a cross between the susceptible line MY11 and the resistant line YU25. Stripe rust resistance in YU25 is derived from Th. intermedium (Luo et al. 2009a, b; Zhang et al. 2011). The sister lines were used as parents to study the inheritance of stripe rust resistance. The wheat cultivars (or lines) YU25, MY11, Chuanmai 42, L693, and L661 and susceptible control SY95-71 were included in comparative response tests. Th. intermedium and common wheat line Chinese Spring were used as controls for the detection of alien chromatin in YU25 and L693. Chinese Spring nulli-tetrasomic (NT) lines were used to identify the chromosomal location of the resistance gene and linked markers. F1, F2, and F2:3 populations from the reciprocal crosses L661/L693 and L693/L661 were used for genetic analysis of stripe rust response. A total 479 F2:3 lines were used in genetic mapping.
Evaluation of stripe rust reactions
Nine wheat cultivars or lines, including four resistant lines (L658, L693, L696, and L699) and one susceptible line (L661), were derived from the same F6:7 families of MY11/YU25. The response of wheat cultivar Chuanmai 42 (CM42), which carries Yr24/Yr26 (Li et al. 2006a; Liu et al. 2010), was compared with that of YU25 and its derived lines using six representative Chinese Pst races (CYR31, CYR32, CYR33, SY11-4, SY11-7, and V26). Among these Pst races, V26 differs from others in China because of its virulence on Yr24/Yr26 genotypes. Wheat plants at the three-leaf stage were inoculated with urediniospores of a single race, and their reactions were evaluated according to previously described methods (Luo et al. 2005). Inoculated seedlings were kept sufficiently moist during incubation at 10 °C with 100 % relative humidity in the dark for 24 h, after which they were subjected to a 24 h cycle of 16 h of light (at 18 °C) and 8 h of darkness (at 10 °C). When the pustules on susceptible checks were fully developed (14–21 days after inoculation), infection types (IT) were classified using a 0–4 rating scale (Luo et al. 2008).
To determine the genetics of resistance in YU25, race CYR32, which is avirulent on YU25 and virulent on SY95-71, was employed to test F1, F2, and F2:3 populations of L661/L693 and L693/L661. All seeds used for genetic analysis were planted in a greenhouse; 20–30 plants of each parental line and F1, 396 F2 plants and 207 F2:3 lines derived from L661/L693, and 582 F2 plants and 272 F2:3 lines derived from L693/L661 (Table 1) were planted in a randomized design with 20–30 plants in 2.5 m rows with 25 cm spacing. Susceptible control plants (SY95-71) used to spread disease were planted in every third row to ensure that all plants had the same opportunity for infection. SY95-71 was inoculated with the predominant race CYR32 at the seedling stage according to previously described methods (Liu et al. 2013). The reactions of adult plants were scored at the milk stage using the previously described rating scale (Luo et al. 2008).
DNA extraction and bulked segregant analysis
Total DNA was extracted from 5-week-old seedling leaves using a previously described method (Tai and Tanksley 1990). Equal amounts of DNA at a concentration of 60 ng/μl from 10 homozygous resistant and 10 homozygous susceptible F2 individuals (genotypes based on the reactions of F2:3 lines) were mixed to construct resistant (BR) and susceptible (Bs) bulks, respectively, for bulked segregant analysis (BSA) (Michelmore et al. 1991). Markers that were polymorphic between the resistant and susceptible parents and bulks were used to genotype the F2:3 lines and in linkage analysis.
Microsatellite marker analysis
Genomic DNA of parents and individual F2 plants derived from the L661/L693 and L693/L661 crosses was used for molecular analyses. For the initial polymorphic marker survey, selected gwm (Röder et al. 1998) and wmc (Gupta et al. 2002) SSR markers spaced at intervals of 3–4 cM along the chromosome according to the consensus map of Somers et al. (2004) were used in BSA to screen for markers linked to the resistance gene. PCRs (25 μl volume) were performed in a PTC-200 thermocycler (MJ Research, Watertown, MA, USA). SSR analysis was performed following a previously described procedure (Röder et al. 1998) with minor modifications. For each PCR, the 25 μl mixture consisted of each SSR primer at a concentration of 200 nmol/l, 0.2 mmol/l dNTPs, 1.5 mmol/l MgCl2, 1 unit of Taq polymerase, and 60 ng of template DNA. PCR was performed following a previously described program (Luo et al. 2008). Each PCR product was mixed with 3 μl of loading buffer (98 % formamide, 10 mM EDTA [pH 8.0], 0.25 % bromophenol blue, and 0.25 mg/ml xylene cyanol), denatured at 95 °C for 5 min, and chilled on ice. Subsequently, a 6 μl aliquot of each sample was loaded onto a 6 % polyacrylamide gel (19:1 acrylamide:bis-acrylamide, 8 M urea and 1 × TBE [90 mM tris–borate (pH 8.3), 2 mM EDTA]) prior to separation at 80 W for approximately 1.5 h and visualization by silver staining (Bassam et al. 1991).
Development of novel SSR markers
To increase the marker density of the map, we chose various Xcfd, Xbarc, and Xgdm SSR markers situated close to two markers that co-segregated with the resistance locus in BSA. The contig sequences carrying the markers that mapped to chromosome 1B can be found in the draft wheat genome sequence (Brenchley et al. 2012; Jia et al. 2013; Ling et al. 2013) using BLAST. This contig was used to search for additional SSR loci. Using primer3 (Rozen and Skaletsky 2000), we developed 19 novel genomic SSR markers based on 8 contigs carrying 8 different public SSR loci (Table 2). The novel markers were named after previously identified public SSR markers by adding a second number. Because the public markers and newly developed markers are located within the same contig, they are physically close and, therefore, useful for fine genetic mapping. Thus, these markers were employed to screen additional polymorphic markers and subsequently used to construct the high-density genetic map.
Chromosomal location
To further ensure that the chromosomal locations of the linked microsatellite markers used in this study were accurate, the following lines were used: Chinese Spring nullisomic 1A tetrasomic 1B (N1AT1B), nullisomic 1A tetrasomic 1D (N1AT1D), nullisomic 1B tetrasomic 1A (N1BT1A), nullisomic 1B tetrasomic 1D (N1BT1D), nullisomic 1D tetrasomic 1A (N1DT1A), and nullisomic 1D tetrasomic 1B (N1DT1B). All lines were kindly provided by Prof. D.C. Liu, Triticeae Research Institute, Sichuan Agricultural University, Chengdu, Sichuan.
Statistical analysis and linkage mapping
Chi-squared tests were performed to determine the goodness-of-fit of the segregation data with hypothesized 1:2:1 ratios for F2:3 lines using Sigmaplot 2001 software (SPSS Inc., Chicago, IL, USA). Recombination fractions were converted to map distances (cM) between loci using the Kosambi mapping function (Kosambi 1944). Loci showing no significant deviations (P > 0.05) were used in the linkage analysis. The order of the linked SSR markers and the resistance gene was determined using JoinMap 4 (Wageningen, Netherlands). A total of 207 F2:3 lines derived from L661/L693 and 272 F2:3 lines derived from L693/L661 (Table 1) were used to construct two different genetic maps using JoinMap 4 with an LOD threshold of 3.0. An integrated genetic linkage map of the resistance gene was constructed by integrating the two different genetic maps from L661/L693 and L693/L661 using the map integration function of JoinMap 4.
Results
Stripe rust response
The low ITs on nine wheat lines inoculated with six Pst races (CYR31, CYR32, CYR33, SY11-4, SY11-7, and V26) showed that line YU25 and its derived lines L658, L693, L696, and L699 were resistant (IT 0-0) to all six races, whereas MY11 and SY95-71 were susceptible (IT 3-4). CM42, which carries Yr24/Yr26, was susceptible (IT 4) only to the V26 race. Typical adult plant responses of the parents and resistant segregants to race CYR32 are shown in Fig. 1. Thus, the resistance gene in YU25 was likely present in wheat lines L658, L693, L696, and L699.
Inheritance of resistance in L693
To map the resistance gene in L693 using molecular markers, F1 plants, 978 F2 individuals and 479 F2:3 lines from the reciprocal L661/L693 crosses (Table 1) were infected from the susceptible spreader line SY95-71, which was inoculated with race CYR32. F1 plants were resistant, and their response was similar to that of L693, indicating that resistance was dominant. The F2 and F2:3 lines segregated for a single dominant gene (Table 1), which was provisionally designated YrL693.
Identification of microsatellite markers linked with YrL693
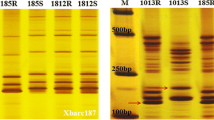
Only 41 (5.2 %) of 781 microsatellites (from the gwm and wmc series) were polymorphic between the susceptible line L661 and resistant line L693. Primer pairs Xgwm273-1B, Xgwm18-1B, Xgwm11-1B, and Xwmc269-1B produced identical bands in the resistant (B R ) F2 bulks (and in L693) and polymorphic bands in the susceptible (B S ) F2 bulks (and in L661), demonstrating that these markers were linked with YrL693. Of the five Xbarc, Xcfd, and Xwmc serial primer pairs close to YrL693, Xbarc137-1B and Xbarc187-1B were linked with YrL693 (Fig. 2). In addition, three of 19 newly developed SSR primer pairs (Table 2), Xbarc187-1B-1, Xcfd65-1B-1 and Xwmc626-1B-1, were closely linked to YrL693. Because six of the nine linked markers were previously shown to be located on wheat chromosome 1B and the contigs containing the other three linked markers must also be located on wheat chromosome 1B, we hypothesize that YrL693 is located on this chromosome.
Silver-stained polyacrylamide gels showing simple sequence repeat (SSR) markers: Xgwm273-1B (a), Xbarc137-1B (b), Xgwm11-1B (c), Xgwm18-1B (d), Xbarc187-1B (e), Xwmc269-1B (f), Xbarc187-1B-1 (g), Xcfd65-1B-1 (h), and Xwmc626-1B-1 (i) linked with YrL693. L661susceptible parent; L693 resistant parent; R1 and R2 resistant F2 individuals; B R resistant F2 DNA pool; H1, H2, and H3 resistant F2 individuals; S1 and S2 susceptible F2 individuals; B S susceptible F2 DNA pool; Marker, 50 bp DNA ladder
Chromosomal assignment and genetic map of YrL693
Based on the published chromosomal locations of the six linked microsatellite markers (Röder et al. 1998; Somers et al. 2004; Song et al. 2002) and the reported chromosomal locations of the contigs containing the other three linked microsatellite markers (http://www.wheatgenome.org/), YrL693 was localized to the centromeric region of wheat chromosome 1B, and the order of SSR loci agreed well with the established SSR maps on chromosome 1B (http://wheat.pw.usda.gov/cgi-bin/graingenes). However, microsatellite markers are not always chromosome-specific because some wheat chromosomes share partial homology (Plaschke et al. 1996). Eight of the nine linked markers were verified on chromosome 1B using Chinese Spring nulli-tetrasomic lines, and only Xwmc626-1B-1 was not assigned to chromosome 1B (Fig. 3).
Chromosomal localization of the microsatellite markers Xgwm273-1B (a), Xbarc137-1B (b), Xgwm11-1B (c), Xgwm18-1B (d), Xbarc187-1B (e), Xwmc269-1B (f), Xbarc187-1B-1 (g), Xcfd65-1B-1 (h), and Xwmc626-1B-1 (i) linked with YrL693 in L661 (S), L693 (R), and a nulli-tetrasomic line of homoeologous group 1. With the exception of Xwmc626-1B-1, no PCR products were generated in nullisomic 1B (N1BT1A and N1B1D)
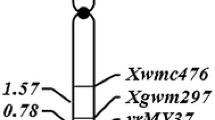
The relationship between the stripe rust resistance gene and the marker genotypes is shown in Supplemental Table 1. Each marker locus exhibited a 1:2:1 segregation ratio. No crossovers were found between YrL693 and the markers Xgwm18-1B, Xgwm11-1B, Xbarc187-1B, and Xbarc187-1B-1 in the forward cross (L661/L693) (Supplemental Table 1; Fig. 4a); YrL693 was, therefore, narrowed to a 0.39 cM interval flanked by markers Xbarc137-1B and Xwmc269-1B/Xcfd65-1B-1 (Fig. 4a). Additionally, no crossovers were found between YrL693 and the markers Xbarc137-1B, Xgwm273-1B, Xgwm18-1B, Xgwm11-1B, Xbarc187-1B, Xbarc187-1B-1, and Xwmc626-1B-1 in the reverse cross (L693/L661); however, the markers Xcfd65-1B-1 and Xwmc269-1B were distally linked to YrL693 with genetic distances of 1.3 and 1.5 cM, respectively (Fig. 4b). The locus order in the forward cross agreed well with that in the reverse cross (Fig. 4a, b). An integrated map was produced from the two genetic maps using the regression mapping algorithm in JoinMap 4 (Fig. 4c). In the integrated map, YrL693 co-segregated with four microsatellite markers (Xgwm18-1B, Xgwm11-1B, Xbarc187-1B, and Xbarc187-1B-1), and it was narrowly flanked by the markers Xgwm273-1B and Xwmc626-1B-1, with distances of 0.08 and 0.07 cM, respectively.
Genetic mapping of YrL693 on chromosome 1B in reciprocal crosses L661/L693 (a), and L693/L661 (b), the integrated map (c), the order of SSR markers in the reference map (d), and the wheat deletion map (e) (http://wheat.pw.usda.gov/cmap/)
Discussion
Origin and mode of inheritance for YrL693
The discovery of novel stripe rust resistance genes and development of new resistant cultivars is the most effective method of controlling stripe rust in wheat. Th. intermedium is an important perennial Triticeae species and is a valuable source of resistance to stripe rust (Liu et al. 2013; Luo et al. 2009a). The wheat line L693 and its resistant parent YU25 are derivatives of Th. intermedium, and because all common wheat parents in the pedigree of these lines are highly susceptible, the resistance in L693 was likely derived from Th. intermedium. Genetic segregation data clearly indicate the presence of a single dominant resistance gene in L693 (Table 1) and its resistant parent YU25 (Luo et al. 2009a). Of the formally named wheat genes for stripe rust resistance, only Yr53 is derived from this donor species. Because of its different chromosomal location the resistance gene in L693 is likely to be novel.
Chromosomal location of YrL693
The microsatellite markers Xgwm273-1B, Xgwm18-1B, Xgwm11-1B, Xwmc269-1B, Xbarc137-1B, and Xbarc187-1B, which were previously assigned to chromosome 1B (Gupta et al. 2002; Röder et al. 1998; Somers et al. 2004; Song et al. 2002), were closely linked with the stripe rust resistance gene in L693. Additionally, the contigs containing the other three linked microsatellite markers were also on chromosome 1B (http://www.wheatgenome.org/). More importantly, eight of the linked markers (except for Xwmc626-1B-1) were verified to be located on chromosome 1B using the Chinese Spring nulli-tetrasomic line (Fig. 3). Furthermore, the order of these SSR loci agreed well with the established SSR maps of chromosome 1B (http://wheat.pw.usda.gov/cgi-bin/graingenes), and the genetic distances between these markers and YrL693 were <1 cM (Fig. 4c). These data provide solid evidence that YrL693 is also on chromosome 1B and is located in the centromeric region, where it is narrowly flanked by the markers Xgwm273-1B and Xwmc626-1B-1 at distances of 0.08 and 0.07 cM, respectively. Xgwm273-1B mapped to the wheat chromosome deletion bin C-1BS-10-0.50, whereas Xwmc626-1B-1 is not mapped to a chromosome. However, Xcfd65-1B, which is located in the same contig as Xcfd65-1B-1 is distal to Xwmc626-1B-1 on the opposite side of Xgwm273-1B, proximal to the centromere in deletion bin C-1BL-6-0.32 (http://wheat.pw.usda.gov/cmap/). The co-segregating marker Xgwm18-1B was also mapped to wheat chromosome deletion bin C-1BS-10-0.50, indicating that YrL693 may be located within wheat chromosome deletion bin C-1BS-10-0.50.
Of the permanently named stripe rust resistance genes, Yr26 is located in the centromeric region of chromosome 1B (Zhang et al. 2013). Yr26 was mapped to chromosome 1B using microsatellite markers, and a 1.9 cM genetic distance between Yr26 and Xgwm11-1B/Xgwm18-1B was estimated using an F2 population of 109 individuals (Ma et al. 2001). The genetic distance between Yr26 and Xgwm11-1B/Xgwm18-1B was reported to be 3.2 cM in a study utilizing 500 F2:3 families (Wang et al. 2008). However, another study using 787 F2 plants and 165 F3 lines reported that Xgwm11-1B, Xgwm18-1B, and Xbarc187-1B were closely linked with Yr26, with genetic distances of 4.2, 3.9, and 2.5 cM, respectively (Wen et al. 2008). The differences among the reported distances between a single marker and Yr26 may result from errors in genotyping and differences in population size and genetic backgrounds of the materials studied because similar mapping software and LOD thresholds were used in the different experiments. Thus, although variable genetic distances were reported between Yr26 and markers Xgwm11-1B, Xgwm18-1B, and Xbarc187-1B in independent experiments, no crossovers were detected between YrL693 and Xgwm11-1B, Xgwm18-1B, or Xbarc187-1B in the present reciprocal crosses (Fig. 4c). More importantly, recent studies have confirmed that Yr26 maps to wheat chromosome deletion bin C-1BL-6-0.32 (Wen et al. 2008; Zhang et al. 2013), whereas the present work indicates that YrL693 might be located within wheat chromosome deletion bin C-1BS-10-0.50. A comparison of the available mapping information regarding Yr26 and YrL693 tends to support the hypothesis that the YrL693 locus is not an allele of Yr26. The new Pst race V26 is virulent to Yr24/Yr26 (Liu et al. 2010; Han et al. 2012) but avirulent to YrL693. The pedigree of L693 indicates that YrL693 should be derived from Th. intermedium, whereas Yr26 is derived from Triticum turgidum (Ma et al. 2001).
Of the permanently named stripe rust resistance genes, Yr15 is also located on wheat chromosome 1BS and genetically close to the centromere, but physically could be some distance from it (McIntosh et al. 1996; Sun et al. 1997; Peng et al. 2000). Several lines of evidence indicate that YrL693 is different from Yr15 and YrH52. First, Yr15 and YrH52 are from Triticum dicoccoides (McIntosh et al. 1996; Peng et al. 2000), whereas YrL693 is potentially from Th. intermedium. Moreover, the presence of YrL693 in L693, together with resistances to powdery mildew and FHB (Luo et al. 2009b; Zhang et al. 2011), further excludes the possibility that YrL693 is by Yr15 or YrH52. Moreover, Yr15 is located on the short arm within 5.7 cM of Xgwm273-1B in the high-density molecular map (Peng et al. 2000), whereas YrL693 is on the long arm within 0.08 cM of Xgwm273-1B based on genetic mapping (Fig. 4c). These data suggest that the locus of YrL693 is different from that of Yr15. Among the formally named stripe rust resistance genes, only Yr9 and Yr17 confer resistant to Pst race V26, whereas YrL693 was effectively resistant to V26 (Han et al. 2012).
Thus, based on pedigree, inheritance, results of molecular marker experiments, gene location, and response specificity, it can be concluded that YrL693 differs from Yr26 and Yr15 and is apparently a new gene.
Transfer of resistance by cryptic translocation is possible
Alien chromosomal translocation is a classic and useful method for transferring genes from wild relatives to common wheat. Despite the potential to carry valuable genes, many alien translocations have questionable value in wheat breeding because the large transferred chromosome segments often carry additional genes for undesirable traits or do not adequately compensate for the wheat genes they replace, resulting in ‘linkage drag’ (e.g., Young and Tanksley 1989). However, traits of interest have been occasionally transferred to recipient genotypes by cryptic translocation without detectable cytological or genetic changes (Kuraparthy et al. 2007). Previous studies have indicated that the wheat genotype YU25 the resistant parent of L693, does not have a cytologically detectable alien chromosome segment and, therefore, may have a cryptic translocation (Luo et al. 2009a, b). Alien chromosomal segments resulting from such small translocations are not easily detected by standard cytogenetic methods other than high-resolution FISH.
In the present study, nearly all of the 781 SSR primer pairs amplified wheat-specific products that were evenly distributed over all the chromosomal arms with 3–4 cM spacing in L661 and L693; however, only 41 (5.2 %) of the 781 SSR primer pairs revealed polymorphisms between L693 and L661. These polymorphisms were located on all chromosomes except 7A, 7B, 1D, 3D, and 6D. This result, which was not unexpected because L661 and L693 are sister lines (Zhang et al. 2011), but they also indicate that a large foreign chromosomal segment is not present in L693. This observation is similar to what was observed in YU25 (Luo et al. 2009a, b). Furthermore, the amplified products of all linked markers in L693 were the same size as those in YU25. Additional evidence for the absence of a large alien chromosomal segment in line L693 includes the following: First, wheat primers linked with the resistance gene produced wheat-specific PCR products in genotype L693, and the SSR loci closely flanking YrL693 (with distances of 0.07 and 0.8 cM) did not show significant alterations in order or between-marker distances compared with the consensus genetic map (Fig. 4); second, L693 is genetically and agronomically uniform based on several years of observations; and third, the resistance gene in L693 behaved as a discrete Mendelian unit (Table 1). Moreover, we could not detect in situ hybridization signals in the resistant parent YU25 using Th. intermedium genomic DNA as a probe (data not shown, personal communication with Prof. F.P. Han, Institute of Genetics and Developmental Biology, Chinese Academy of Science, Beijing). Taken together, these data indicate that L693 does not possess a large alien chromosomal segment and may instead contain a cryptic translocation. The pedigree provides the only evidence that L693 carries a resistance gene from Th. intermedium. Hence, one of the goals for further study is to seek new information concerning the source of stripe rust resistance at the DNA level.
Potential role of YrL693 in the improvement of wheat resistance
Although many wheat stripe rust resistance genes have been identified and incorporated into commercial cultivars, most of them have been overcome by virulent races. In recent years, several physiological races of Pst, including CYR30, CYR31, and CYR32, have become prevalent in southwestern China. These races are virulent to almost all the cultivars developed in this region (Luo et al. 2005). In addition, a new race referred to as V26 is virulent to Yr24/Yr26, and varieties in the region possessing Yr24/Yr26 are now vulnerable to stripe rust epidemics (Liu et al. 2010; Han et al. 2012). The present study identified a novel stripe rust resistance gene located on chromosome 1B in L693 that displayed normal inheritance. Approximately, 66.7 % of the alien genes that confer resistance to wheat stripe rust are located on B chromosomes (http://wheat.pw.usda.gov/), indicating that wheat B chromosomes may be more tolerant to the presence of alien chromatin. Hence, loss, gain, or replacement of genetic material in this genome is likely to have minimal detrimental effects. L693 displays effective resistance to all predominant Pst races in the region. In addition, this line exhibits excellent resistances to powdery mildew (afforded by Pm40 [Luo et al. 2009b]) and FHB (Zhang et al. 2011), but those resistances are not linked to YrL693. Markers that co-segregate with or are closely linked to YrL693 can be used to accelerate the incorporation of this gene into commercial cultivars by marker-assisted selection. Therefore, there is a high likelihood that the material described in the present study can be utilized by breeders, especially in southwestern China.
References
Autrique E, Singh R, Tanksley S, Sorrells M (1995) Molecular markers for four leaf rust resistance genes introgressed into wheat from wild relatives. Genome 38:75–83
Bansal U, Forrest K, Hayden M, Miah H, Singh D, Bariana H (2011) Characterisation of a new stripe rust resistance gene Yr47 and its genetic association with the leaf rust resistance gene Lr52. Theor Appl Genet 122:1461–1466
Bassam BJ, Caetano-Anolles G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Brenchley R, Spannagel M, Pfeifer M et al (2012) Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491:705–710
Chen X (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp. tritici] on wheat. Can J Plant Pathol 27:314–337
Chen W, Wu L, Liu T, Xu S, Jin S, Peng Y, Wang B (2009) Race dynamics, diversity, and virulence evolution in Puccinia stiiformis f. sp. tritici, the causal agent of wheat stripe rust in China from 2003 to 2007. Plant Dis 93:1093–1101
Cheng P, Chen X (2010) Molecular mapping of a gene for stripe rust resistance in spring wheat cultivar IDO377s. Theor Appl Genet 121:195–204
Fedak G (1999) Molecular aids for integration of alien chromatin through wide crosses. Genome 42:584–591
Fedak G, Han F (2005) Characterization of derivatives from wheat—Thinopyrum wide crosses. Cytogenet Genome Res 109:350–359
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87
Gupta PK, Balyan HS, Edwards KJ et al (2002) Genetic mapping of 66 new microsatellite (SSR) in bread wheat. Theor Appl Genet 105:413–422
Han DJ, Wang N, Jiang Z, Wang QL, Wang XJ, Kang ZS (2012) Characterization and inheritance of resistance to stripe rust in the wheat line Guinong775. Hereditas (Beijing) 34:1607–1613
He R, Chang Z, Yang Z, Yuan Z, Zhan H, Zhang X, Liu J (2009) Inheritance and mapping of powdery mildew resistance gene Pm43 introgressed from Thinopyrum intermedium into wheat. Theor Appl Genet 118:1173–1180
Herrera-Foessel SA, Lagudah ES, Huerta-Epino J, Hayden M, Bariana H, Singh D, Singh RP (2011) New slow-rusting leaf rust and stripe rust resistance genes Lr67 and Yr46 in wheat are pleiotropic or closely linked. Theor Appl Genet 122:239–249
Jia J, Li G, Liu C, Lei M, Yang Z (2011) Characterization of wheat yellow rust resistance gene Yr17 using EST-SSR and rice syntenic region. Cereal Res Commun 39:88–99
Jia JZ, Zhang SC, Kong XY et al (2013) Aegilops tauschii draft genome sequence reveals a gene repertoire of wheat adaptation. Nature 496:91–95
Kang Z, Zhao J, Han D, Zhang H, Wang X, Wang C, Han Q, Guo J, Huang L (2010) Status of wheat rust research and control in China. BGRI 2010, Technical Workshop, St Petersburg, 30–31 May 2010
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kuraparthy V, Sood S, Chhuneja P, Dhaliwal HS, Kaur S, Bowder RL, Gill BS (2007) A cryptic wheat—Aegilops triuncialis translocation with leaf rust resistance gene Lr58. Crop Sci 47:1995–2003
Li G, Li Z, Yang W, Zhang Y, He Z, Xu S, Singh R, Qu T, Xia X (2006a) Molecular mapping of stripe rust resistance gene YrCH42 in Chinese wheat cultivar Chuanmai 42 and its allelism with Yr24 and Yr26. Theor Appl Genet 112:1434–1440
Li Z, Xia X, Zhou X, Niu Y, He Z, Zhang Y, Li G, Wan A, Wang D, Chen X, Lu Q, Singh R (2006b) Seedling and slow rusting resistance to stripe rust in Chinese common wheats. Plant Dis 90:1302–1312
Li ZF, Zheng TC, He ZH, Li GQ, Xu SC, Li XP, Yang GY, Singh RP, Xia XC (2006c) Molecular tagging of stripe rust resistance gene YrZH84 in Chinese wheat line Zhou 8425B. Theor Appl Genet 112:1098–1103
Li Q, Chen X, Wang M, Jing J (2011) Yr45, a new wheat gene for stripe rust resistance on the long arm of chromosome 3D. Theor Appl Genet 122:189–197
Lillemo M, Asalf B, Singh R, Huerta-Espino J, Chen X, He Z, Bjørnstad A (2008) The adult plant rust resistance loci Lr34/Yr18 and Lr46/Yr29 are important determinants of partial resistance to powdery mildew in bread wheat line Saar. Theor Appl Genet 116:1155–1166
Ling HQ, Zhao SC, Liu DC, Wang J et al (2013) Draft genome of the wheat A-genome progenitor Triticum urartu. Nature 496:87–90
Liu SB, Wang HG (2005) Characterization of wheat-Thinopyron intermedium substitution line with resistance to powdery mildew. Euphytica 143:229–233
Liu TG, Peng YL, Chen WQ, Zhang ZY (2010) First detection of virulence in Puccinia striiformis f. sp. tritici in China to resistance genes Yr24 (= Yr26) present in wheat cultivars Chuanmai 42. Plant Dis 94:1163
Liu J, Chang ZJ, Zhang XJ, Yan ZJ, Li X, Ji JQ, Zhang HX, Guo HJ, Wang JM (2013) Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor Appl Genet 126:265–274
Lowe I, Jankuloski L, Chao S, Chen X, See D, Dubcovsky J (2011) Mapping and validation of Yr48 and other QTL conferring partial resistance to broadly virulent post-2000 North American races of stripe rust in hexaploid wheat. Theor Appl Genet 123:143–157
Luo PG, Ren ZL, Zhang HQ, Zhang HY (2005) Identification, chromosome location, and diagnostic markers for a new gene (YrCN19) for resistance to wheat stripe rust. Phytopathology 95:1266–1270
Luo PG, Hu XY, Ren ZL, Zhang HY, Shu K, Yang ZJ (2008) Allelic analysis of stripe rust resistance genes on wheat chromosome 2BS. Genome 51:922–927
Luo PG, Hu XY, Chang ZJ, Zhang M, Zhang HQ, Ren ZL (2009a) A new stripe rust resistance gene transferred from Thinopyrum intermedium to hexaploid wheat (Triticum aestivum). Phytoprotection 90:57–63
Luo PG, Luo HY, Chang ZJ, Zhang HY, Zhang M, Ren ZL (2009b) Characterization and chromosomal location of Pm40 in common wheat: a new gene for resistance to powdery mildew derived from Elytrigia intermedium. Theor Appl Genet 118:1059–1064
Ma JX, Zhou RH, Dong YS, Wang LF, Wang XM, Jia JZ (2001) Molecular mapping detection of the yellow resistance gene Yr26 in wheat transferred from Triticum turgidum L. using microsatellite markers. Euphytica 120:219–226
McIntosh RA, Silk J, The TT (1996) Cytogenetic studies in wheat XVII. Monosomic analysis and linkage relationships of gene Yr15 for resistance to stripe rust. Euphytica 89:395–399
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregation populations. Proc Natl Acad Sci USA 88:9828–9832
Peng JH, Fahima T, Röder MS, Huang QY, Dahan A, Li YC, Grama A, Nevo E (2000) High-density molecular map of chromosome region harboring stripe rust resistance genes YrH52 and Yr15 derived from wild emmer wheat, Triticum dicoccoides. Genetica 109:199–210
Plaschke J, Börner A, Wendehake K, Ganal MW, Röder MS (1996) The use of aneuploids for the chromosomal assignment of microsatellite loci. Euphytica 89:33–40
Ren RS, Wang MN, Chen XM, Zhang ZJ (2012) Characterization and molecular mapping of Yr52 for high-temperature adult-plant resistance to stripe rust in spring wheat germplasm PI 183527. Theor Appl Genet 125:847–857
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Song QJ, Fickus EW, Cregan PB (2002) Characterization of trinucleotide SSR motifs in wheat. Theor Appl Genet 104:286–293
Sun GL, Fahima T, Korol AB, Turpeinen T, Grama A, Ronin YI, Nevo E (1997) Identification of molecular markers linked to the Yr15 stripe rust resistance gene of wheat originated in wild emmer wheat Triticum dicoccoides. Theor Appl Genet 95:622–628
Sun Q, Wei Y, Ni C, Xie C, Yang T (2002) Microsatellite marker for yellow rust resistance gene Yr5 introgressed from spelt wheat. Plant Breed 121:539–541
Tai TH, Tanksley SD (1990) A rapid and inexpensive method for isolation of total DNA from dehydrated plant tissue. Plant Mol Biol Rep 8:297–303
Uauy C, Brevis J, Chen X, Khan I, Jackson L, Chicaiza O, Distenfeld A, Fahima T, Dubcovsky J (2005) High-temperature adult-plant stripe rust resistance gene Yr36 from Triticum turgidum ssp. dicoccoides is closely linked to the grain protein content locus Gpc-B1. Theor Appl Genet 112:97–105
Wan AM, Zhao ZH, Chen XM, He ZH, Jin SL, Jia QZ, Yao G, Yang JX, Wang BT, Li GB, Bi YQ, Yuan ZY (2004) Wheat stripe rust epidemic and virulence of Puccinia striiformis f. sp. tritici. Plant Dis 88:896–904
Wang CM, Zhang YP, Han DJ, Kang ZS, Li GP, Cao AZ, Chen PD (2008) SSR and STS markers for wheat stripe rust resistance gene Yr26. Euphytica 159:359–366
Wen WE, Li GQ, He ZH, Yang WY, Xu ML, Xia XC (2008) Development of an STS marker tightly linked to Yr26 against wheat stripe rust using the resistance gene-analog polymorphism (RGAP) technique. Mol Breed 22:507–515
Xu LS, Wang MN, Cheng P, Kang ZS, Hulbert SH, Chen XM (2013) Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor Appl Genet 126:523–533
Young ND, Tanksley SD (1989) RFLP analysis of the size chromosomal segments retained around Tm-2 locus of tomato during backcross breeding. Theor Appl Genet 92:1923–1932
Zhang L, Chang ZJ, Li X, Zhang HY, Ren ZL, Luo PG (2011) Screen and identification of wheat new resistant germplasms to Fusarium head blight. Acta Phytophylacica Sinica 38:569–570
Zhang XJ, Han DJ, Zeng QD, Duan YH, Yuan FP, Shi JD, Wang QL, Wu JH, Huang LL, Kang ZS (2013) Fine mapping of wheat stripe rust resistance gene Yr26 based on collinearity of wheat with Brachypodium distachyon and rice. PLoS One 8:e57885
Acknowledgments
Financial support was provided by the National Natural Science Foundation of China (30971787, 31271721 and 31171557), the Provincial Science and Technology Foundation for Young Scientists of Sichuan China (2010JQ0042), and the Specific Foundation of Agronomy (No. nyhyzx200903035). We are grateful to Dr. R.A. McIntosh of the University of Sydney, Australia; Dr. X.M. Chen of the U.S. Department of Agriculture-Agricultural Research Service (USDA-ARS) in Pullman, WA, USA; and Dr. X.C. Xia of the Institute of Crop Science at the National Wheat Improvement Centre of The National Key Facility for Crop Gene Resources and Genetic Improvement of the Chinese Academy of Agricultural Sciences (CAAS) in Beijing, China, for critically reviewing drafts of this paper. We are also grateful to Prof Q.X. Sun and Prof Z.Y. Liu of the College of Agriculture and Biotechnology, China Agricultural University, Beijing, for providing many useful suggestions and discussing the manuscript.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by A. Graner.
Q. Huang and X. Li contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, Q., Li, X., Chen, W.Q. et al. Genetic mapping of a putative Thinopyrum intermedium-derived stripe rust resistance gene on wheat chromosome 1B. Theor Appl Genet 127, 843–853 (2014). https://doi.org/10.1007/s00122-014-2261-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-014-2261-7