Abstract
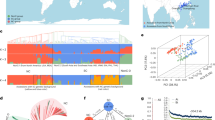
Peas are essential for human nutrition and played a crucial role in the discovery of Mendelian laws of inheritance. In this study, we assembled the genome of the elite vegetable pea cultivar ‘Zhewan No. 1’ at the chromosome level and analyzed resequencing data from 314 accessions, creating a comprehensive map of genetic variation in peas. We identified 235 candidate loci associated with 57 important agronomic traits through genome-wide association studies. Notably, we pinpointed the causal gene haplotypes responsible for four Mendelian traits: stem length (Le/le), flower color (A/a), cotyledon color (I/i) and seed shape (R/r). Additionally, we discovered the genes controlling pod form (Mendelian P/p) and hilum color. Our study also involved constructing a gene expression atlas across 22 tissues, highlighting key gene modules related to pod and seed development. These findings provide valuable pea genomic information and will facilitate the future genome-informed improvement of pea crops.






Similar content being viewed by others
Data availability
The genome sequencing and assembly data of Pisum sativum cultivar Zhewan1 (PeaZW1) have been deposited at National Center for Biotechnology Information under the BioProject PRJNA1042956. The whole-genome sequencing of 237 accessions has also been deposited at NCBI under the BioProject PRJNA1035516. Transcriptome data from different tissues can be found under the BioProject PRJNA1108961. Source data are provided with this paper.
Code availability
All codes and tools used in this study are described in Methods and the Reporting Summary.
References
McCrory, M. A., Hamaker, B. R., Lovejoy, J. C. & Eichelsdoerfer, P. E. Pulse consumption, satiety, and weight management. Adv. Nutr. 1, 17–30 (2010).
Pandey, A. K. et al. Omics resources and omics-enabled approaches for achieving high productivity and improved quality in pea (Pisum sativum L.). Theor. Appl Genet 134, 755–776 (2021).
Yang, T. et al. Improved pea reference genome and pan-genome highlight genomic features and evolutionary characteristics. Nat. Genet. 54, 1553–1563 (2022).
Tayeh, N. et al. Genomic tools in pea breeding programs: status and perspectives. Front. Plant Sci. 6, 1037 (2015).
Liu, N. et al. Comparative transcriptomic analyses of vegetable and grain pea (Pisum sativum L.) seed development. Front. Plant Sci. 6, 1039 (2015).
Smykal, P. et al. From Mendel’s discovery on pea to today’s plant genetics and breeding: commemorating the 150th anniversary of the reading of Mendel’s discovery. Theor. Appl. Genet. 129, 2267–2280 (2016).
Zohary, D. & Hopf, M. Domestication of pulses in the old world: legumes were companions of wheat and barley when agriculture began in the Near East. Science 182, 887–894 (1973).
Smykal, P. et al. Legume crops phylogeny and genetic diversity for science and breeding. Crit. Rev. Plant Sci. 34, 43–104 (2015).
Kreplak, J. et al. A reference genome for pea provides insight into legume genome evolution. Nat. Genet. 51, 1411–1422 (2019).
Makani, J., Nkya, S., Collins, F. & Luzzatto, L. From Mendel to a Mendelian disorder: towards a cure for sickle cell disease. Nat. Rev. Genet. 23, 389–390 (2022).
Charlesworth, B. et al. From Mendel to quantitative genetics in the genome era: the scientific legacy of W. G. Hill. Nat. Genet. 54, 934–939 (2022).
Mendel, G. Versuche über Pflanzen-Hybriden. Brünn, Im Verlage des Vereines, 1822–1884. Biodiversity Heritage Library https://doi.org/10.5962/bhl.title.61004 (1866).
Van Dijk, P. J. & Ellis, T. H. The full breadth of Mendel’s genetics. Genetics 204, 1327–1336 (2016).
Bhattacharyya, M. K., Smith, A. M., Ellis, T. H. N., Hedley, C. & Martin, C. The wrinkled-seed character of pea described by Mendel is caused by a transposon-like insertion in a gene encoding starch-branching enzyme. Cell 60, 115–122 (1990).
Ingram, T. J. et al. Internode length in Pisum: the Le gene controls the 3beta-hydroxylation of gibberellin A20 to gibberellin A 1. Planta 160, 455–463 (1984).
Lester, D. R., Ross, J. J., Davies, P. J. & Reid, J. B. Mendel’s stem length gene (Le) encodes a gibberellin 3 beta-hydroxylase. Plant Cell 9, 1435–1443 (1997).
Weston, D. E. et al. The Pea DELLA proteins LA and CRY are important regulators of gibberellin synthesis and root growth. Plant Physiol. 147, 199–205 (2008).
Lester, D. R., MacKenzie-Hose, A. K., Davies, P. J., Ross, J. J. & Reid, J. B. The influence of the null le-2 mutation on gibberellin levels in developing pea seeds. Plant Growth Regul. 27, 83–89 (1999).
Armstead, I. et al. Cross-species identification of Mendel’s I locus. Science 315, 73 (2007).
Sato, Y., Morita, R., Nishimura, M., Yamaguchi, H. & Kusaba, M. Mendel’s green cotyledon gene encodes a positive regulator of the chlorophyll-degrading pathway. Proc. Natl Acad. Sci. USA 104, 14169–14174 (2007).
Hellens, R. P. et al. Identification of Mendel’s white flower character. PLoS ONE 5, e13230 (2010).
Sussmilch, F. C., Ross, J. J. & Reid, J. B. Mendel: from genes to genome. Plant Physiol. 190, 2103–2114 (2022).
Tayeh, N. et al. Development of two major resources for pea genomics: the GenoPea 13.2K SNP Array and a high-density, high-resolution consensus genetic map. Plant J. 84, 1257–1273 (2015).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Raj, A., Stephens, M. & Pritchard, J. K. fastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197, 573–589 (2014).
Martin, D. N., Proebsting, W. M. & Hedden, P. Mendel’s dwarfing gene: cDNAs from the Le alleles and function of the expressed proteins. Proc. Natl Acad. Sci. USA 94, 8907–8911 (1997).
Ellis, T. H. N. & Poyser, S. J. An integrated and comparative view of pea genetic and cytogenetic maps. New Phytol. 153, 17–25 (2002).
Lamprecht, H. The variation of linkage and the course of crossing over. Agri Hortic. Genet. 6, 10–48 (1948).
Shirasawa, K., Sasaki, K., Hirakawa, H. & Isobe, S. Genomic region associated with pod color variation in pea (Pisum sativum). G3 (Bethesda) 11, jkab081 (2021).
Li, J. A. et al. Mutation of rice BC12/GDD1, which encodes a kinesin-like protein that binds to a GA biosynthesis gene promoter, leads to dwarfism with impaired cell elongation. Plant Cell 23, 628–640 (2011).
Xu, J. et al. HEAT SHOCK PROTEIN 90.6 interacts with carbon and nitrogen metabolism components during seed development. Plant Physiol. 191, 2316–2333 (2023).
Yan, Y. et al. HSP90.2 promotes CO2 assimilation rate, grain weight and yield in wheat. Plant Biotechnol. J. 21, 1229–1239 (2023).
Martinez, C., Pons, E., Prats, G. & Leon, J. Salicylic acid regulates flowering time and links defence responses and reproductive development. Plant J. 37, 209–217 (2004).
Huang, W., Wang, Y., Li, X. & Zhang, Y. Biosynthesis and regulation of salicylic acid and N-hydroxypipecolic acid in plant immunity. Mol. Plant 13, 31–41 (2020).
Tayeh, N. et al. afila, the origin and nature of a major innovation in the history of pea breeding. New Phytol. 243, 1247–1261 (2024).
Bordat, A. et al. Translational genomics in legumes allowed placing in silico 5460 unigenes on the pea functional map and identified candidate genes in Pisum sativum L. G3 (Bethesda) 1, 93–103 (2011).
Weeden, N. F. et al. A consensus linkage map for Pisum sativum. Pisum Genet. 30, 1–3 (1998).
Willoughby, A. C. & Nimchuk, Z. L. WOX going on: CLE peptides in plant development. Curr. Opin. Plant Biol. 63, 102056 (2021).
Balarynova, J. et al. The loss of polyphenol oxidase function is associated with hilum pigmentation and has been selected during pea domestication. N. Phytol. 235, 1807–1821 (2022).
Taylor-Teeples, M. et al. An Arabidopsis gene regulatory network for secondary cell wall synthesis. Nature 517, 571–575 (2015).
Nasmyth, K. The magic and meaning of Mendel’s miracle. Nat. Rev. Genet. 23, 447–452 (2022).
White, O. E. The present state of knowledge of heredity and variation in peas. Proc. Am. Phil. Soc. 56, 487–588 (1917).
Ahmad, I. S., Reid, J. F., Paulsen, M. R. & Sinclair, J. B. Color classifier for symptomatic soybean seeds using image processing. Plant Dis. 83, 320–327 (1999).
Doyle, J. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull. 19, 11–15 (1987).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res. 46, e126 (2018).
Ou, S. & Jiang, N. LTR_retriever: a highly accurate and sensitive program for identification of long terminal repeat retrotransposons. Plant Physiol. 176, 1410–1422 (2018).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinf. 9, 18 (2008).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 46, D41–D47 (2018).
Gremme, G., Steinbiss, S. & Kurtz, S. GenomeTools: a comprehensive software library for efficient processing of structured genome annotations. IEEE/ACM Trans. Comput. Biol. Bioinform. 10, 645–656 (2013).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Keilwagen, J., Hartung, F., Paulini, M., Twardziok, S. O. & Grau, J. Combining RNA-seq data and homology-based gene prediction for plants, animals and fungi. BMC Bioinf. 19, 189 (2018).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res 18, 1979–1990 (2008).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
UniProt, C. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. the gene ontology consortium. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Zheng, Y. et al. iTAK: a program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol. Plant 9, 1667–1670 (2016).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Kalvari, I. et al. Rfam 14: expanded coverage of metagenomic, viral and microRNA families. Nucleic Acids Res. 49, D192–D200 (2021).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl Acad. Sci. USA 117, 9451–9457 (2020).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Bao, W., Kojima, K. K. & Kohany, O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics 4, 4.10.1–4.10.14 (2009).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Marcais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Cingolani, P. Variant annotation and functional prediction: SnpEff. Methods Mol. Biol. 2493, 289–314 (2022).
Retief, J. D. Phylogenetic analysis using PHYLIP. Methods Mol. Biol. 132, 243–258 (2000).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
He, Z. et al. Evolview v2: an online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res. 44, W236–W241 (2016).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Li, M. X., Yeung, J. M., Cherny, S. S. & Sham, P. C. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum. Genet. 131, 747–756 (2012).
Lyu, X. L. et al. A natural mutation of the NST1 gene arrests secondary cell wall biosynthesis in the seed coat of a hull-less pumpkin accession. Hortic. Res. 9, uhac136 (2022).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008).
Kumar, L. & M, E. F. Mfuzz: a software package for soft clustering of microarray data. Bioinformation 2, 5–7 (2007).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Die, J. V., Roman, B., Nadal, S. & Gonzalez-Verdejo, C. I. Evaluation of candidate reference genes for expression studies in Pisum sativum under different experimental conditions. Planta 232, 145–153 (2010).
Acknowledgements
We are grateful to Biomarker Technologies Corporation, Beijing and China National Gene Bank (CNGB), Beijing Novogene Co. Ltd for technical support with PacBio HiFi sequencing, Hi-C sequencing, Iso-seq, RNA-seq and whole genomics sequencing. This work was supported by the Zhejiang Provincial Important Science and Technology Specific Projects (grant no. 2021C02065 to N.L., grant no. 2022C02016 to Y.G.), State Key Laboratory for Managing Biotic and Chemical Threats to the Quality and Safety of Agro-products (grant no. 2021DG700024-ZZ202206 to N.L.), National Natural Science Foundation of China (grant no. 31872114 to N.L.) and Zhejiang Basic Public Welfare Research Project (grant no. LGN20C150006 to N.L., grant no. LGN21C150007 to Z.F.).
Author information
Authors and Affiliations
Contributions
Y.G., N.L., L.Z., T.Z., X.L. and M.Z. conceived the project and designed the study. N.L., T.Z., L.Z., X.L., Z.Z., Y.Z., Z.F., Q.G., K.S., W.S. and Y.D. performed data analyses. N.L. and T.Z. drafted the manuscript. G.Z., X.Z., X.L., X.C., X.Y., Z.F., J.O., B.W. and Y.B. collected samples and performed experiments. N.L. and X.L. wrote the manuscript, and X.G., M.Z., L.Z. and T.Z. revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Aureliano Bombarely, Fanjiang Kong, Petr Smýkal and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–19.
Supplementary Tables
Supplementary Tables 1–21.
Source data
Source Data Fig. 3
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, N., Lyu, X., Zhang, X. et al. Reference genome sequence and population genomic analysis of peas provide insights into the genetic basis of Mendelian and other agronomic traits. Nat Genet 56, 1964–1974 (2024). https://doi.org/10.1038/s41588-024-01867-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01867-8
- Springer Nature America, Inc.