Abstract
Pea rust caused by Uromyces fabae (Pers.) de-Bary is a major problem in warm humid regions causing huge economic losses. A mapping population of 136 F6:7 recombinant inbred lines (RILs) derived from the cross between pea genotypes, HUVP 1 (susceptible) and FC 1 (resistant) was evaluated in polyhouse as well as under field conditions during two consecutive years. Infection frequency (IF) and area under disease progress curve (AUDPC) were used for evaluation of rust reaction of the RILs. A linkage map was constructed with 57 polymorphic loci selected from 148 simple sequence repeats (SSRs), 3 sequence tagged sites (STS), and 2 random amplified polymorphic (RAPD) markers covering 634 cM of genetic distance on the seven linkage groups of pea with an average interval length of 11.3 cM. Composite interval mapping (CIM) revealed one major (Qruf) and one minor (Qruf1) QTL for rust resistance on LGVII. The LOD (5.2–15.8) peak for Qruf was flanked by SSR markers, AA505 and AA446 (10.8 cM), explaining 22.2–42.4% and 23.5–58.8% of the total phenotypic variation for IF and AUDPC, respectively. The minor QTL was environment-specific, and it was detected only in the polyhouse (LOD values 4.2 and 4.8). It was flanked by SSR markers, AD146 and AA416 (7.3 cM), and explained 11.2–12.4% of the total phenotypic variation. The major QTL Qruf was consistently identified across all the four environments. Therefore, the SSR markers flanking Qruf would be useful for marker-assisted selection for pea rust (U. fabae) resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pea (Pisum sativum L) rust can be caused by Uromyces fabae (Pers) de-Bary [Syn. Uromyces viciae-fabae (Pers.) J. Schrőt] (Arthur 1934) or by Uromyces pisi (Pers.) Wint. U. fabae is an autoecious fungus, while U. pisi is a heteroecious macrocyclic fungal pathogen that completes its life cycle on Euphorbia cyparrissias (Pfunder and Roy 2000). The two species can be differentiated from each other by morphology of telia and infections structures or by internal transcribed spacer (ITS) markers (Emeran et al. 2005, Barilli et al. 2006; Sillero et al. 2006). Uromyces fabae is reported to infect faba beans (Vicia faba L.), lentil (Lens culinaris Medik.), and vetches (Vicia sativa L.) (Cummins 1987), and is considered as a species complex consisting of several isolates specialized with respect to their hosts (Emeran et al. 2005, 2008). Aeciospores of U. fabae acts as repeating spores and play important role in the outbreak of the disease (Kushwaha et al. 2006), whereas urediospores are the only infecting spores in the case of U. pisi (Xue and Warkentin 2002; Barilli et al. 2009a).
In temperate regions, pea crop is primarily damaged by U. pisi (Emeran et al. 2005; Barilli et al. 2009a, b, c), while in tropical and subtropical regions, U. fabae is the pathogen causing rust. The European and Mediterranean Plant Protection organization (EPPO) reports yield losses of >30% due to U. pisi under favorable environments (EPPO 2009), while U. fabae is known to cause up to 50% reduction in pea yields, particularly in the Northwestern Uttar Pradesh and the Terai regions of India (Upadhayay and Singh 1994; Singh 1998). Yield losses due to U. fabae are particularly high when humidity and temperature are high (Pal et al. 1980; Xue and Warkentin 2002; Kushwaha et al. 2006; Kushwaha et al. 2007), especially during flowering and pod formation stages (Sanger and Singh 1994; Singh et al. 1996; Chand et al. 2006.
The genetic basis of resistance to U. fabae is debatable as different studies indicated either monogenic (Pal et al. 1979; Katiyar and Ram 1987) or polygenic control (Kumar et al. 1994); Vijayalakshmi et al. 2005). Vijayalakshmi et al. (2005) reported a single gene Ruf to govern resistance to U. fabae in the pea cross FC 1 × HUVP 1, but they also presented some evidence suggesting involvement of some polygenes as well. Further, none of the pea genotypes are reported to be free from U. fabae infection (Singh and Srivastava 1985; Chand et al. 2004, 2006) suggesting a polygenic type of resistance. Similarly, resistance to U. pisi in wild pea (Pisum fulvum L.) shows polygenic inheritance (Barilli et al. 2010), quantitative non-hypersensitive type of resistance to rust has been reported in other legumes, such as, chickpea (Cicer arietinum L.) against U. ciceris-arietini (Madrid et al. 2008), and in groundnut (Arachis hypogaea L.) against Puccinia arachidis (Khedikar et al. 2010). However, hypersensitive reaction has been reported in French bean (Phaseolus vulgaris L.) against rust pathogen U. appendiculatus (Stavely et al. 1989), in faba bean (Sillero et al. 2000) and lentil (Saha et al. 2010) against U. viciae-fabae, and in soybean [Glycine max (L.) Marr] against Phakopsora pachyrhizi (Li 2009).
Chemical control for pea rust has been suggested, but it is considered uneconomic as well as hazardous to human health and the environment (Chand et al. 2006). Breeding for rust resistance is hampered by the difficulty in identification of resistant plants since disease expression is markedly influenced by the environment. Therefore, molecular markers linked to resistance genes could facilitate selection of rust resistant segregants and thereby improve breeding efficiency. So far, reports on molecular mapping of resistance to this disease are limited and robust markers are not known. Few years ago, two RAPD markers, SC10-82 360 (10.8 cM) and SCRI-71 1000 (24.5 cM) flanking the rust resistance gene, Ruf, were identified by employing a relatively small (31) backcross (BC1F1) mapping population (Vijayalakshmi et al. 2005). In a recent report (Barilli et al. 2010), a major QTL was detected for resistance to U. pisi in wild pea (P. fulvum L.). A large number of SSR markers are now publicly available from pea microsatellite consortium (Burstin et al. 2001, Loridon et al. 2005) and can be used for identification of markers linked to rust resistance. Therefore, the objective of the present study was to study inheritance and mapping of QTLs controlling resistance of pea to U. fabae using SSR markers (Burstin et al. 2001, Loridon et al. 2005).
Materials and methods
Plant material
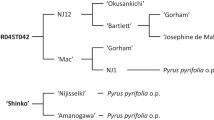
The mapping population consisted of 136 F6:7 RILs derived from individual F2 plants of the cross between pea genotypes HUVP 1 (susceptible) and FC 1 (resistant) handled as per single seed descent (SSD) method. HUVP 1 was developed at Banaras Hindu University, Varanasi, from the cross (Bonneville × 5064) × S 143, and FC 1 was derived from the cross (Bonneville × HR209) × IIHR 656 at Indian Institute of Horticultural Research, Bangalore.
Field evaluation
One hundred thirty-six RILs were evaluated for rust resistance in the Agricultural Research Farm of Banaras Hindu University, Varanasi, India (Northeastern Plains Zone, 25.2ºN and 83.0ºE) during crop seasons 2006–2007 and 2007–2008. Planting was carried out on 30th November in year 2006–2007 and 4th December in 2007–2008; this allowed plants to develop fully when warm and humid conditions favorable for disease occur during late February to early March. Throughout the experiment, plants were maintained in vegetative phase by bud clipping. The maximum and minimum temperature and humidity ranged between 26 and 14.5°C and 70–40.5%, respectively, in 2006–2007 and 30–17°C, 63–31.5% in 2007–2008.
The RILs were planted in randomized block design with two replications in each environment. Each RIL was planted in a single row of 1.5 m, and row-to-row and plant-to-plant distances were maintained at 40 and 10 cm, respectively. A single row of the parents and five rows of spreader line were sown after every 10 rows of RILs. In addition, five rows of spreader line were planted all around the experimental plot.
Polyhouse evaluation
The RILs were also evaluated in a polyhouse at the Institute of Agricultural Sciences, Banaras Hindu University, Varanasi, India. Planting was carried out on 1st December in year 2006–2007 and 5th December in 2007–2008. Five seeds of RILs were sown in pots of 30 cm diameter. Each line was planted in two pots with each pot representing a replication. Seeds of both the parents were also grown in pots and placed after every five pots of RILs. Rust susceptible spreader line (HFP4) was sown in the ground between two pots of each block. The disease appears in first week of February and the plants begin to die in end of March. During these 2 months, the maximum and minimum temperatures ranged from 17.8 to 39.4°C and 9.9 to 19.5°C, respectively, during 2006– 2007, and from 20.6 to 39.2°C, and 4.7 to 21.1°C during 2007–2008.
Artificial inoculation in the field and polyhouse
Aecidial stage of U. fabae dominated on pea in India (Kushwaha et al. 2006) and is characterized by papillated teliospores. Aeciospores derived from a single pustule were multiplied on highly susceptible cultivar HFP4, sown 22 days prior to planting of the main experiment. The rusted leaves were soaked in water and agitated vigorously to prepare a spore suspension of 104 spores per ml. The plants were inoculated at pre-flowering stage by spraying the spore suspension during evening hours (Chand et al. 2004). Field and pots were irrigated immediately after inoculation to provide favorable environment for disease development. Field was irrigated five times in the entire crop period. In polyhouse, high humidity was maintained by misting the plants every 3 h for 48 h after inoculation. Subsequently, high humidity was maintained throughout the experimental period by regular misting.
Assessment of disease
Rust severity (%) displayed by all the plants of each RIL was recorded six times at 3 days interval, in the polyhouse, and four times at 6 days interval in the field. The area under disease progress curves (AUDPC), considered a more practical approach for disease assessment (Jeger and Viljanen-Rollinson 2001), was calculated using following formula given by Shaner and Finney (1977).
Infection frequency scoring as a component of rust resistance
Infection frequency (IF) was scored in terms of number of pustules per cm2 on the tagged leaves when the disease severity % (DS%) of resistant parent was less than 30 and that of susceptible parent was more than 75. Leaves were tagged in the mid vegetation zone since it is the most favorable for disease development (Chand et al. 2006; Kushwaha et al. 2006). Both sporulating and non-sporulating pustules were scored and plants were maintained in vegetative phase by bud clipping which delayed the aging of plants.
Statistical analysis
Analysis of variance (ANOVA) was performed using PROC GLM of SAS software (v 10; SAS Institute Inc., Cary, NC, USA 1997) to determine the variation in disease response in terms of AUDPC and IF among the RILs within and across the environments (year and location). Heritability (h 2) of traits within and across the environments was estimated from mean squares by applying the formulas: h 2 = σ 2 G /(σ 2 G + σ 2 e /r), and h 2 = σ 2 G /(σ 2 G + σ 2 GE /E + σ 2 e /rE), respectively (Hallauer and Miranda Filho 1981). Here, σ 2 G , σ 2 GE , σ 2 e , E and r represent the genetic variance, genotypic × environment interaction variance, residual variance, number of environments, and the number of replicates per line, respectively. Estimates of correlation coefficient among the rust severity, AUDPC and IF across the years and locations were obtained using SPSS software v 12.0 (SPSS, Chicago, IL, USA).
QTL analysis and molecular mapping
We selected SSR primers from Loridon et al. (2005) that detected polymorphism between the two parents and were, as far as possible about 20 cM apart covering the seven linkage groups of pea. The 136 RILs were screened with these markers to create a framework of molecular map. Single marker regression analysis was performed to test each marker for the significance to disease severity. Later on more SSR and other markers, including two RAPD markers viz SC10-82 360 and SCRI-71 1000 , (Vijayalakshmi et al. 2005) and three STS markers, viz, ACCox, Ga20X, and Gs3B (Aubert et al. 2006) were selected and tested to enrich the neighboring region found to be significant in the initial analysis. Finally, Mapmaker v 3.0 (Lincoln et al. 1993) was used to construct the map using 136 RILs with all polymorphic primers. Linkage map was built using a minimum LOD (likelihood of odd ratio) threshold of 2.5 and a maximum recombination fraction of 0.40. Haldane’s function was applied to estimate map unit distances (Haldane 1919). The ‘Order’ and ‘Rip’ commands were used to assign the order of markers, while ‘Try’ and ‘Compare’ commands were used to add additional markers on the map.
The chromosome regions found significant in initial single marker regression analysis, enriched with more markers were then tested with the QTL Cartographer version 2.5 (Wang et al. 2005) to confirm QTL location and effect for both the components by interval mapping (IM) and composite interval mapping (CIM). The standard model (Model 6) of the Zmapqtl procedure of QTL Cartographer was used with window size 10 and walking speed 2 cM along the linkage groups. The cofactors were selected by the forward and backward regression with threshold P < 0.01. Only those QTLs were considered significant that had LOD score higher than the threshold calculated from 1,000 permutation test for P < 0.01 to control Type-I error (Churchill and Doerge 1994). The LOD score and coefficient of determination (R 2) of detected significant QTLs was estimated by CIM through ZmapQTL. The QTLNetwork software (http://ibi.zju.edu.cn/software/qtlnetwork/) was used to estimate QTL × QTL and QTL × environment interactions. This is based on QTL mapper 2.0, which amalgamates the effect of multiple QTLs, and QTL by environment interactions into a single mapping system, called mixed-model-based composite interval mapping.
Results
Phenotypic variation
The continuous distribution of disease in terms of AUDPC and IF (Fig. 1a, b) and the test of normality using Shapiro–Wilk test (P = 0.073, 0.225 for AUDPC and IF, respectively) revealed that RIL data fit normal distribution with transgressive segregates at both the ends of the distribution, except for the absence of lower pustule number per cm2 than the resistant parent. The parents exhibited contrasting phenotype for mean AUDPC and IF in all the four environments.
Analysis of variance for AUDPC (Table 1) revealed highly significant differences for genotypes and genotypes by environment (year and location) interaction. The analysis of variance for IF also revealed a significant variation for genotypes and environment but not for genotype by environment interaction. The mean AUDPC for RILs varied among the four environments from 300 (06–07, field data) to 489 (07–08, field data), being intermediate between the two parental means (Table 2). Similarly, mean IF for RILs showed some variation among the environments (from 9.09 to 10.28) and was intermediate between those for the two parents. Heritability estimates were consistently higher for AUDPC (0.78–0.86) than those for IF (0.43–0.67), and were comparable over the environments, except for IF during 06–07 in polyhouse (Table 2).
Trait correlations
High phenotypic correlation was observed between DS% and AUDPC; the estimates of correlation coefficients were generally higher in the field than in the polyhouse (Table 3). Similarly, IF showed significant (P < 0.01) positive correlation with AUDPC (r = 0.45–0.69) (Table 4); in general, these estimates were lower than those for correlation between AUDPC and DS%. Thus AUDPC was positively correlated with DS% and IF across environments (locations and years). In addition AUDPC estimated in one environment showed significant, high positive correlation (r = 0.65–0.82) with AUDPC estimates from other environments, a similar result was evident for IF as well (Table 4).
Markers polymorphism and segregation
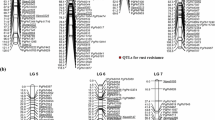
We tested 148 SSR, 2 RAPD and 3 STS markers for detecting polymorphism between the parents HUVP 1 and FC 1. Out of these, 55 SSR, 2 RAPD and 2 STS markers (38.5%) were polymorphic and amplified 21 dominant and 51 co-dominant polymorphic bands. The amplification of multiple bands by some SSR primers could be due to the presence of cryptic sites, up and downstream or between the two specific primer-binding sites (Winter et al. 1999). Fifteen of the polymorphic loci were found to be unlinked, while 57 were distributed over the seven linkage groups, covering 634 cM; the average distance between adjacent markers was approximately 11 cM. LG VII was of 174.8 cM length, and it contained both the QTLs detected in this study (Fig. 2). All the linkage groups were assigned according to the linkage map published by Loridon et al. (2005) based on common markers. LG VII (Fig. 2) had five markers (AA19, AA416, AD146, AD237 and Gs3B) in common with the LGVII of composite linkage map of pea published by Aubert et al. (2006); these markers covered the genomic region containing the two QTLs, Qruf and Qruf1. The population was in advanced stage (F 6:7) and the alleles were expected to segregate in equal proportion. However, 10 polymorphic loci did not fit 1:1 ratio (P < 0.05); this included both dominant and co-dominant markers.
Genetic linkage map of LG VII and LOD profiles of Qruf and Qruf1 on LGVII detected in a population of 136 F6:7 RILs derived from the cross HUVP 1 × FC 1 evaluated for rust reaction in polyhouse as well as field for 2 years (2006–2007 and 2007–2008) using a IF and b AUDPC as disease reaction criteria. The size of linkage group is in Haldane centiMorgans (cM) and thick bands on LGVII represent QTL positions
QTL mapping
Using CIM, one major QTL was detected for IF (Fig. 2a); this QTL had LOD values of 5.2 (polyhouse, 06–07) to 9.8 (field, 07–08), and explained 22.2–42.4% of the total phenotypic variance for IF (Table 5). The peak of LOD for this QTL was between markers interval AA446–AA505 (10.8 cM) and the closest marker was AA505 at about 1 cM from the peak. RAPD markers SC10-82 360 and SCRI-71 1000 reported to be linked with the gene Ruf governing number of rust pustules per cm2 (Vijayalakshmi et al. 2005) were tested for their linkage with the SSR markers surrounding the above major QTL; the marker SC10-82 360 mapped between SSR marker AD237 and AB27, while the other RAPD marker (SCRI-71 1000 ) did not show linkage with the SSR markers (Fig. 2).
Similarly, one major and one minor QTL were detected for AUDPC for pea rust on linkage group VII. The major QTL seems to cover the region between the markers, AA446 and SC10-82 360 , but its peak LOD value was always between the markers, AA446 and AA505 (Fig. 2b). Therefore, this major QTL is considered to be the same as that detected for IF; it consistently expressed in all the environments and the LOD value ranged from 7.5 (polyhouse, 07–08) to 15.8 (field, 06–07) with R 2 ranging from 23.5 to 58.8%, respectively (Table 5). The minor QTL, had peak LOD values between the markers, AD146 and AA416 (7.3 cM), at 0.6 cM from the marker AD146; it was expressed only in polyhouse, the LOD values being 4.8 in 2006–2007, and 4.2 in 2007–2008, and the R 2 values being 12.4 and 11.2%, respectively. The major and minor QTLs accounted for 40.9 and 12.8%, respectively, of the total phenotypic variance in mean over year analysis. There was no significant QTL × QTL interaction between the two identified QTLs. Similarly, QTL effect versus environment (year, location) interaction was also non-significant for both the detected QTLs.
Discussion
A continuous frequency distribution of 136 RILs for rust AUDPC and IF (Fig. 1) suggested rust resistance in the pea cross ‘HUVP 1 × FC 1’ to be a quantitative trait. An earlier study reported continuous distribution and considerable variation for number of pustules per leaf, in BC1F2 population of the same cross, suggestive of polygenic nature of inheritance (Vijayalakshmi et al. 2005).
AUDPC showed significant (P < 0.01) high positive correlation with DS% and IF; therefore AUDPC values were used for QTL analysis. This also suggests that IF is an important component for rust resistance in pea. The genetic variance for AUDPC and IF varied across environments, but the high positive correlation among AUDPC and IF values across environments suggested the reproducibility and validity of experimental conditions and scoring method. The difference in the level of disease infestation between environments could be due to difference in the prevailing temperature and/or humidity that influenced growth and spread of the pathogen. In polyhouse, the environmental effect was minimized by maintaining moisture and temperature, although these conditions were not exactly similar across years. Broad sense heritability estimates for disease severity components were high (for AUDPC, 0.83, for IF, 0.91) representing good reproducibility of the phenotypic data. The h 2 for AUDPC was high in each individual trial ranging from 0.79 to 0.86. Comparatively, h 2 for IF was moderate within individual trials ranging from 0.42 to 0.67, but was very high (0.91) across environments because the G × E interaction was low (non-significant) as compared to the magnitude of genetic variance.
Several molecular-marker-based linkage maps have been developed for pea. The SSR markers selected from composite microsatellite marker map of pea developed by Loridon et al. (2005) facilitated molecular mapping of rust resistance in the present study. Our analysis of the selected SSR markers generated seven linkage groups, which is equal to the haploid number of chromosomes (2n = 2x = 14) for pea. Linkage with SSR markers revealed two QTLs for rust resistance located on linkage group VII of Loridon et al. (2005). Only CIM detected QTLs are presented since CIM takes selected markers as cofactors along with interval markers in QTL detection by analytical model, which reduces background effect of other QTLs, and enables detection of small effect QTLs that is usually not possible by simple interval mapping (Zeng 1994; Melchinger et al. 1998). When IF was used for mapping, only the major QTL, was detected. This agrees well with the earlier proposed Ruf gene for number of rust pustules per cm2 in the same cross of pea. The two RAPD markers (SC10-82 360 and SCRI-71 1000 ) reported to be linked to the Ruf gene (Vijayalakshmi et al. 2005), but only marker SC10-82 360 was found to map in the same region that has Qruf. It seems that the RAPD marker SCRI-71 1000 was incorrectly identified to be linked to gene Ruf most likely due to the rather small BC1F2 population used for mapping, and possibly some incorrect genotyping for the marker. We suggest that the Ruf gene proposed by Vijayalakshmi et al. (2005) be now redesignated as Qruf to signify the quantitative nature of its action Qruf was also detected for AUDPC along with another minor QTL, which is designated as Qruf1. Barilli et al. (2010) reported a single QTL, UP1, located between markers OPY11_1316 and OPV17_1078 to govern P. fulvum accession IFPI3260 resistance to U. pisi, although there was hint of a second QTL between markers OPAB12_125 and OPY11_1361 (in Figure 3 of Barilli et al. 2010). The disease severity in their study was assayed using only 94 F 3 families grown in a single environment. It would be useful to evaluate a large number of RILs in multiple environments.
Rust resistance in lentil (Lens culinaris Medik.) is governed by one (Kumar et al. 2001), two (Chahota et al. 2002), or even four (Kumar et al. 1997) dominant genes. Saha et al. (2010) identified a sequence related amplified polymorphism (SRAP) marker, F7XEM4a, estimated to be 7.9 cM from the lentil rust resistance gene. In faba bean (Vicia faba L.), a single gene (Sillero et al. 2000), three race-specific genes (Conner and Bernier 1982) and even several genes (Rashid and Bernier 1986) were reported to specify rust (U. viciae-fabae) resistance. Avila et al. (2003) identified three RAPD markers, OPD13736, OPL181032, and OPI20900, linked in coupling phase and two RAPD markers, OPP021172 and OPR07930 linked in repulsion phase with the gene Uvf-1 that specifies resistance of faba bean to race 1 of U. viciae-fabae. Rust resistance in groundnut appeared to be polygenic (Motagi 2001; Dwivedi et al. 2002), and a major QTL, QTLrust01, contributing 6.90–55.20% to the phenotypic variation was identified in a mapping population of 268 RILs from the cross TAG 24 × GPBD 4. The QTLrust01 was located between marker interval IPAHM103-pPGSseq19D6 (position, 0–12 cM on LG VI); SSR marker IPAHM103 seems to be associated with the QTLrust01 itself and this marker was validated in a wide range of resistant/susceptible breeding lines as well as in progeny lines of another mapping population, TG 26 × GPBD 4 (Khedikar et al. 2010). A quantitative trait locus for rust resistance in chickpea was located between STMS makers TA18–TA180, on LG7 of C. arietinum genetic linkage map (Madrid et al. 2008).
The QTLs Qruf and Qruf1 explained 40.9 and 12.8% of phenotypic variance, respectively. These values are lower than the R 2 value (63%) reported for the single QTL (Up1) for rust resistance against U. pisi (Pers.) Wint. in wild pea using a mapping population of 94 F2 individuals (Barilli et al. 2010). It is possible that in the present study some minor QTL may have remained undetected. Involvement of one or few major QTLs with or without supporting minor QTL/QTLs is a common feature for disease resistance (Young 1996; Kover and Caicedo 2001; Burnham et al. 2003; Cobos et al. 2009). Major QTLs are usually consistent but minor QTLs are highly fluctuating according to the environment (Paterson et al. 1991).
This is the first report of QTL mapping for resistance to U. fabae in cultivated pea since the earlier mapping of Ruf gene by Vijayalakshmi et al. (2005) was based on oligogenic inheritance of infection frequency. The major QTL (Qruf) detected for AUDPC was also detected for IF; this may be expected since pustule density is reported as an important component of rust resistance in legumes (Statler and McVey 1987; Negussie et al. 2005; Chand et al. 2006). The expression of Qruf was consistent across the environments, which is of paramount significance for MAS. Further, the distance between the closest marker and QTL peak was very small for both Qruf (1.0 cM from AA505) and Qruf1 (0.6 cM from AD146). Therefore, the chance of recombination between the closest markers and the two QTLs is 1% or lower. Thus, these markers are quite promising for MAS, and they could even facilitate cloning and sequencing of these QTLs. If the two flanking markers were used for MAS, the chance of both the markers becoming unlinked to the concerned QTLs, viz, Qruf and Qruf1 would be only 0.1 and 0.04%, respectively. The usefulness of these SSR markers for marker-assisted selection would require their validation in wider germplasm of cultivated pea.
SSR markers developed with Medicago truncatula show considerable cross-amplification in other pulses, e.g., 40% in faba bean, 36.6% in chickpea and 37.6% in pea. In addition, EST-SSR (expressed sequence tagged-short sequence repeats) markers showed about twofold higher cross-amplification in the three pulses (39–43%) than the genomic region SSR markers (21–24%; Gutie′rrez et al. 2005). Aubert et al. (2006) have developed a functional map of pea spanning 1,458 cM and depicting locations of 363 loci, including 111 markers anchored to specific genes having known functions. They also mapped 21 of these genes in M. truncatula, and added 41 new links between pea and M. truncatula. It would be useful to analyze the present RIL population using appropriate markers from Aubert et al. (2006) in an attempt to search for candidate genes corresponding/located close to the rust resistance QTLs reported in this study.
Mixed-model-based composite interval mapping method has been used in legumes (soybean) for detection of interaction between disease resistance QTLs and their interaction with the environment (Wu et al. 2009). The major and minor QTLs (Qruf and Qruf1) were located on the same linkage group (LGVII) and in close proximity separated by <25 cM, but interaction between them was non-significant, although the minor QTL was not expressed in all the environments. Whenever the minor QTL was not expressed, the contribution of the major QTL was overestimated. This could be the reason why percentage of total phenotypic variance explained by the major QTL was always higher in the field than in the polyhouse, where the minor QTL Qruf1 was expressed.
Some SSR markers showed deviation from 1:1 ratio, but their inclusion in the analysis did not affect the linkage map. Skewed segregation of molecular markers is a common feature in different mapping populations concerning disease resistance in pea (Pilet-Nayel et al. 2002; Valderrama et al. 2004; Fondevilla et al. 2008). This study has identified two genomic regions, Qruf and Qruf1 flanked by SSR markers AA446-AA505 and AD146-AA416, respectively, on LG VII, that are involved in resistance of P. sativum against U. fabae. The usefulness of these markers in marker-assisted selection and for map-based cloning of the major QTL will crucially depend on their validation in unrelated germplasm.
References
Arthur JC (1934) Manual of the rusts in United States and Canada. Purdue Research Foundation, Lafayette
Aubert G, Morin J, Jacquin F, Loridon K, Quillet MC, Petit A, Rameau C, Lejeune-He′naut I, Huguet T, Burstin J (2006) Functional mapping in pea, as an aid to the candidate gene selection and for investigating synteny with the model legume Medicago truncatula. Theor Appl Genet 112:1024–1041
Avila CM, Sillero JC, Rubiales DM, Moreno T, Torres AM (2003) Identification of RAPD markers linked to the Ufv-1 genen conferring hypersensitive resistance against rust (Uromyces viciae-fabae) in Vicia faba L. Theor Appl Genet 107:353–358
Barilli, E, Satovic, Z, Sillero, JC, Rubiales, D, Torres, AM (2006) Estudio filogenetico de royas de leguminosas (Uromyces spp.) mediante ITS (Internal Transcribed Spacer). In: de los Mozos Pascual M, Gime′nez Alvear MJ, Rodrı′guez Conde MF, Sa′nchez Vioque R (eds) Proceedings of the 4th conference of AEL. Consejerı′a de Agricultura de Castilla-La Mancha, Toledo, Spain, pp 331–338
Barilli E, Sillero JC, Fernandez-Aparicio M, Rubiales D, Torres AM (2009a) Identification of resistance to Uromyces pisi (Pers.) Wint. In Pisum spp. germplasm. Field Crop Res 114:198–203
Barilli E, Sillero JC, Moral A, Rubiales D (2009b) Characterization of resistance response of pea (Pisum spp.) against rust (Uromyces pisi). Plant Breed 128:665–670
Barilli E, Sillero JC, Serrano A, Rubiales D (2009c) Differential response of pea (Pisum sativum) to rusts incited by Uromyces fabae and U. Pisi. Crop Prot 28:980–986
Barilli E, Satovic Z, Rubiales D, Torres AM (2010) Mapping of quantitative trait loci controlling partial resistance against rust incited by Uromyces pisi (Pers.) Wint. In a Pisum fulvum L. intraspecific cross. Euphytica 175:151–159
Burnham KD, Dorrance AE, VanToai TT, Martin SK (2003) Quantitative trait loci for partial resistance to Phytophthora sojae in soybean. Crop Sci 43:1610–1617
Burstin J, Deniot G, Potier J, Weinachter C, Aubert G, Baranger A (2001) Microsatellite polymorphism in Pisum sativum. Plant Breed 120:311–317
Chahota PK, Gupta VP, Sharma SK (2002) Inheritance of rust resistance in lentil. Indian J Genet Plant Breed 62:226–227
Chand R, Srivastava CP, Sarode SB (2004) Screening technique for pea (Pisum sativum) genotypes against rust disease (Uromyces fabae). Indian J Agric Sci 74:166–167
Chand R, Srivastava CP, Singh BD, Sarode SB (2006) Identification and characterization of slow resistance components in pea (Pisum sativum). Genet Resourc Crop Evol 53:219–224
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Cobos MJ, Winter P, Kharrat M, Cubero JI, Gil J, Millan T, Rubio J (2009) Genetic analysis of agronomic traits in a wide cross of chickpea. Field Crops Res 111:130–136
Conner RL, Bernier CC (1982) Host range of Uromyces viciae-fabae. Phytopathology 72:687–689
Cummins GB (1987) Rust fungi on legumes and composites in North America. The University of Arizona Press, Tucson
Dwivedi SL, Pande S, Rao JN, Nigam SN (2002) Components of resistance to late leaf spot and rust among interspecific derivatives and their significance in a foliar disease resistance breeding in groundnut (Arachis hypogaea L.). Euphytica 125:81–88
Emeran AA, Sillero JC, Niks RE, Rubiales D (2005) Infection structures of host specialized isolates of Uromyces viciae-fabae and of other Uromyces infecting leguminous crops. Plant Dis 89:17–22
Emeran AA, Roma′ n B, Sillero JC, Satovic Z, Rubiales D (2008) Genetic variation among and within Uromyces species infecting legumes. J Phytopathol 156:419–424
EPPO (2009) EPPO Standards Pea http://archives.eppo.org/EPPOStandards/PP2GPP/pp2-14-e.doc. Accessed 08 December 2009
Fondevilla S, Satovic Z, Rubiales D, Moreno MT, Torres AM (2008) Mapping of quantitative trait loci for resistance to Mycosphaerella pinodes in Pisum sativum subsp. syriacum. Mol Breed 21:439–454
Gutiérrez MV, Vaz Patto MC, Huguet T, Cubero JI, Moreno MT, Torres AM (2005) Cross-species amplification of Medicago truncatula microsatellites across three major pulse crops. Theor Appl Genet 110:1210–1217
Haldane JBS (1919) The combination of linkage values, and the calculation of distance between the loci of linked factors. J Genet 8:299–309
Hallauer AR, Miranda Filho JB (1981) Quantitative genetics in maize breeding. Iowa State University Press, Ames
Jeger MJ, Viljanen-Rollinson SLH (2001) The use of area under disease progress curve (AUDPC) to assess quantitative disease resistance in crop cultivars. Theor Appl Genet 102:32–40
Katiyar RP, Ram RS (1987) Genetics of rust resistance in pea. Indian J Genet 47:46–48
Khedikar YP, Gowda MVC, Sarvamangala C, Patgar KV, Upadhyaya HD, Varshney RK (2010) A QTL study on late leaf spot and rust revealed one major QTL for molecular breeding for rust resistance in groundnut (Arachis hypogaea L.). Theor Appl Genet 121:971–984
Kover PX, Caicedo AL (2001) The genetic architecture of disease resistance in plants and the maintenance of recombination by parasites. Mol Ecol 10:1–16
Kumar TBA, Rangaswamy KT, Ravi K (1994) Assessment of tall field pea genotypes for slow rusting resistance. Legume Res 17:79–82
Kumar V, Singh BM, Singh S (1997) Genetics of lentil resistance to rust. Lens Newslett 24:21–22
Kumar R, Mishra SK, Sharma B (2001) Genetics of rust resistance in lentil. Indian J Genet. 61:238–241
Kushwaha C, Chand R, Srivastava CP (2006) Role of aeciospores in outbreaks of pea (Pisum sativum) rust (Uromyces fabae). Eur J Plant Pathol 115:323–330
Kushwaha C, Srivastava CP, Chand R, Singh BD (2007) Identification and evaluation of a critical time for assessment of slow rusting in pea against Uromyces fabae. Field Crops Res 103:1–4
Li S (2009) Reaction of soybean rust-resistant lines identified in Paraguay to Mississippi isolates of Phakopsora pachyrhizi. Crop Sci 49:887–894
Lincoln SE, Daly MJ, Lander ES (1993) Constructing genetic linkage maps with MAPMAKER/EXP version 3.0: a tutorial and reference manual. A Whitehead Institute for Biomedical Research Technical Report. http://www.mapmaker@genome.wi.mit.edu
Loridon K, McPhee K, Morin J, Dubreuil P, Pilet-Nayel ML, Aubert G, Rameau C, Baranger A, Coyne C, Lejeune-He`naut I, Burstin J (2005) Microsatellite marker polymorphism and mapping in pea (Pisum sativum L.). Theor Appl Genet 111:1022–1031
Madrid E, Rubiales D, Moral A, Moreno MT, Millán T, Gil J, Rubio J (2008) Mechanism and molecular markers associated with rust resistance in a chickpea interspecific cross (Cicer arietinum × Cicer reticulatum). Eur J Plant Pathol 121:43–53
Melchinger AE, Utz HF, Schon CC (1998) Quantitative trait locus (QTL) mapping using different tester and independent population samples in maize revels low power of QTL detection and large bias in estimates of QTL effects. Genetics 149:383–403
Motagi (2001) Genetic analysis of resistance to late leaf spot and rust vis-à-vis productivity in groundnut (Arachis hypogaea L.). Dissertation, University of Agriculture Sciences, Dharwad, India
Negussie T, Pretorius ZA, Bender CM (2005) Components of rust resistance in lentil. Euphytica 142:55–64
Pal AB, Sohi HS, Rawal RD (1979) Studies on resistance to rust (Uromyces fabae (Pers.) de-Bary) on peas. SABRAO J 11:101–103
Pal AB, Brahmappa HS, Rawal RD, Ullasa BA (1980) Field resistance of pea germplasm to powdery mildew (Erysiphe polygoni) and rust (Uromyces fabae). Plant Dis 64:1085–1086
Paterson AH, Damon S, Hewitt JD, Zamir D, Rabinowitch HD, Lincoln SE, Lander ES, Tanksley SD (1991) Mendelian factors underlying quantitative traits in tomato: Comparison across species, generations, and environments. Genetics 127:181–197
Pfunder M, Roy B (2000) Pollinator-mediated interactions between a pathogenic fungus, Uromyces pisi (Pucciniaceae), and its host plant, Euphorbia cyparrissias (Euphorbiaceae). Am J Bot 87:48–55
Pilet-Nayel ML, Muehlbauer FJ, McGee RJ, Kraft JM, Baranger A, Coyne CJ (2002) Quantitative trait loci for partial resistance to Aphanomyces root rot in pea. Theor Appl Genet 106:28–39
Rashid KY, Bernier CC (1986) Selection for slow rusting in faba bean (Vicia faba L.) to Uromyces viciae-fabae. Crop Prot 5:218–224
Saha GC, Sarker A, Chen W, Vandemark GJ, Muehlbauer FJ (2010) Identification of markers associated with genes for rust resistance in Lens culinaris Medik. Euphytica 175:261–265
Sanger RBS, Singh VK (1994) Effect of showing dates and pea variety on severity of rust and powdery mildew. Indian J Agric Sci 74:166–167
Shaner G, Finney RE (1977) The effect of nitrogen fertilization on the expression of slow-mildewing resistance in Knox wheat. Phytopathology 67:1051–1056
Sillero JC, Moreno MT, Rubiales D (2000) Charecterization of new sources of resistance to Uromyces viciae-fabae in a germplasm collection of Vicia faba L. Theor Appl Genet 107:226–227
Sillero JC, Fondevilla S, Davidson J, Vaz Patto MC, Warketin TD, Thomas J, Rubiales D (2006) Screening techniques and sources of resistance to rusts and mildews in grain legumes. Euphytica 147:255–272
Singh RS (1998) Plant Diseases (7th edn) Oxford and IBH, New Delhi
Singh RM, Srivastava CP (1985) Evaluation, classification and usefulness of pea germplasm for quantitative characters. Legume Res 8:68–73
Singh VK, Sanger RBS, Singh RN (1996) Effect of varieties and showing dates on disease incidence and productivity of field pea (Pisum sativum L). Indian J Agron 4:451–453
Statler GD, McVey MA (1987) Partial resistance to Uromyces appendiculatus in dry edible beans. Phytopathology 77:1101–1103
Stavely JR, Steadman JR, McMillan RT (1989) New pathogenic variability in Uromyces appendiculatus in North America. Plant Dis 73:428–432
Upadhayay AL, Singh VK (1994) Performance of pea varieties/lines against powdery mildew and rust. Indian J Pulses Res 7:92–93
Valderrama MR, Roman B, Satovic Z, Rubiales D, Cubero JI, Torres AM (2004) Locating quantitative trait loci associated with Orobanche crenata resistance in pea. Weed Res 44:323–328
Vijayalakshmi S, Yadav K, Kushwaha C, Sarode SB, Srivastava CP, Chand R, Singh BD (2005) Identification of RAPD markers linked to the rust (Uromyces fabae) resistance gene in pea (Pisum sativum). Euphytica 144:265–274
Wang, S, Basten, CJ, Gaffney, P, Zeng, ZB (2005) Windows QTL Cartographer version 2.5. Statistical Genetics, North Carolina State University, Raleigh, NC http://statgen9.ncsu.edu/qtlcart/WQTLcart.htm
Winter P, Pfaff T, Udupa SM, Hüttel B, Sharma PC, Sahi S, Arreguin-Espinoza R, Weigand F, Muehlbauer FJ, Kahl G (1999) Characterization and mapping of sequence-tagged microsatellite sites in the chickpea (Cicer arietinum L.) genome. Mol Gen Genet 262:90–101
Wu X, Blake S, Sleper DA, Shannon JG, Cregan P, Nguyen HT (2009) QTL, additive and epistatic for effects for SCN resistance in PI 437654. Theor Appl Genet 118:1093–1105
Xue AG, Warkentin TD (2002) Reaction of field pea varieties to three isolates of Uromyces fabae. Can J Plant Sci 82:253–255
Young ND (1996) QTL mapping and quantitative disease resistance in plants Annu Rev Phytopathol 34:479–501
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136:1457–1468
Acknowledgments
We thank Dr. Sundeep Sharma, National Bureau of Plant Genetic Resources, Indian Council of Agricultural Research, New Delhi 110012, India, and Dr. Uttam Kumar, The Energy and Resources Institute, Darbari Seth Block, IHC Complex, Lodhi Road, New Delhi 110003, India, for their help during the development of this manuscript. The first author (RR) is thankful to University Grants Commission (UGC) for financial support in the form of Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Varshney.
Rights and permissions
About this article
Cite this article
Rai, R., Singh, A.K., Singh, B.D. et al. Molecular mapping for resistance to pea rust caused by Uromyces fabae (Pers.) de-Bary. Theor Appl Genet 123, 803–813 (2011). https://doi.org/10.1007/s00122-011-1628-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-011-1628-2