Abstract
Amplified fragment length polymorphism fingerprinting was applied to survey the genetic diversity of primitive South American Gossypium barbadense cotton for establishing a possible link to its pre-Columbian expansion. New germplasm was collected along coastal Peru and over an Andean transect in areas where most of the archaeological evidence relating to cotton domestication has been recorded. Gene bank material of three diploid (G. raimondii, G. arboreum, and G. herbaceum) and four allotetraploid cotton species (G. hirsutum, G. mustelinum, G. tomentosum and additional G. barbadense) was added for inter- and intra-specific comparison. Eight primer combinations yielded 340 polymorphic bands among the 131 accessions. The obtained neighbor joining and unweighted pair-group method with arithmetic means are in full agreement with the known cytogenetics of the tetraploid cottons and their diploid genome donors. The four tetraploid species are clearly distinct based on taxonomic classification. The genetic diversity within G. barbadense reveals geographic patterns. The locally maintained cottons from coastal Peru display a distinct genetic diversity that mirrors their primitive agro-morphological traits. Accessions from the northernmost coast of Peru and from southwestern (SW) Ecuador cluster basal to the east-of-Andes accessions. The remaining accessions from Bolivia, Brazil, Columbia, Venezuela, and the Caribbean and Pacific islands cluster with the east-of-Andes accessions. Northwestern Peru/SW Ecuador (the area flanking the Guayaquil gulf) appears to be the center of the primitive domesticated G. barbadense cotton from where it spread over the Andes and expanded into its pre-Columbian range.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The cotton genus Gossypium L. consists of about 45 diploid and five tetraploid species forming a monophyletic group (Fryxell 1992; Wendel and Cronn 2003). Eight diploid genomes (A–G and K), each comprising 13 chromosome pairs, have been identified (Endrizzi et al. 1985). The native distribution of all five tetraploid cottons (2n=4x=52, AADD) is restricted to the New World, although its emergence involves a combination of an Old World A-genome (derived from an ancestor of G. arboreum L. and G. herbaceum L.) and a New World D-genome (derived from an ancestor of G. raimondii Ulbrich) (Stephens 1944; Phillips 1963, 1964; Seelanan et al. 1997; Cronn et al. 1999; Liu et al. 2001). The underlying trans-oceanic dispersal of the A-genome donor and the timing of the polyploidization event has been controversial, but the current view is that a single Mid-Pleistocene (1–2 mya) polyploidization event has occurred (Phillips 1963; Wendel and Albert 1992; Senchina et al. 2003). It is further assumed, that the emergence of tetraploid cotton was followed by long-distance separation into five species, three of which are truly wild species—G. mustelinum Miers ex Watt (limited distribution in northeast Brazil; Wendel et al. 1994), G. darwinii Watt (endemic to the Galapagos Islands; Wendel and Percy 1990), and G. tomentosum Nutall ex Seemann (endemic to the Hawaiian Islands; DeJoode and Wendel 1992). The other two species have undergone domestication—G. hirsutum L. (predominantly distributed in Meso America and the Caribbean) and G. barbadense L.(main distribution in South America and the Caribbean). Although molecular diversity within and among the tetraploid cottons is limited (Small et al. 1999; Wendel and Cronn 2003), data indicate that G. mustelinum forms a basal branch in the phylogram of the tetraploids, while the remaining four species form two sister groups—one constituted by G. hirsutum and G. tomentosum and the other by G. barbadense and G. darwinii (Small et al. 1998; Liu et al. 2001).
The elongated epidermal seed trichomes of G. herbaceum, G. arboreum, G. hirsutum, and G. barbadense were recognized by humans as useful spinnable fibers and led to four independent domestication events (Wendel 1995; Brubaker et al. 1999). Archaeological evidence of cotton remains in association with human settlements dates back to the sixth millennium BC for the Greater Indus area in the Old World (Moulherat et al. 2002) and to layers dated to 6,400–5,000 years BP for the Zaña river valley in Peru in the New World (Rossen et al. 1996; Piperno and Pearsall 1998). The cotton G. hirsutum has received much attention in this matter due to its prime economic importance in modern cotton production (e.g., Brubaker and Wendel 1994). The South American G. barbadense domestication has also been studied, but mainly by means of archaeological approaches. Cotton remains along the coast of Peru and Ecuador are abundant and show a gradient in traits from wild to domesticated (Stephens 1973; Stephens and Moseley 1973, 1974). Purely wild G. barbadense forms are mentioned in the literature (e.g., Percy and Wendel 1990) but are not clearly defined in terms of fitness-related traits for wild survival. The wild-to-domesticated continuum is, according to Percy and Wendel (1990), delimited in four general categories in gene bank collections: (1) wild; (2) ‘dooryard cottons’ or ‘commensals’, meaning single plants found near habitations thought to be derived directly from local wild populations; (3) landraces; (4) improved modern cultivars. The maritime subsistence for the Andean civilizations depending in part on cotton fishing-nets has led to the perception that the domestication of G. barbadense took place along the coastline, but cotton is also found in more inland located sites like Caral (Solis et al. 2001). An allozyme study (Percy and Wendel 1990) yielded geographic clusters largely congruent with this maritime-based scenario, but only an unspecified geographic domestication region ‘north-western South America west of the Andes’ including Colombia, Ecuador, Peru, and Bolivia was reported.
Until now no DNA marker-based study has assessed the intra-specific genetic diversity of G. barbadense with the objective of describing a geographic pattern that can help in understanding the pre-Columbian domestication events of this species. To this end we conducted a new field collection of dooryard/feral cottons of G. barbadense in Peru and added material from the USDA Cotton Collection in order to have a representative sample from its pre-Columbian range. We included the two wild tetraploid species G. mustelinum and G. tomentosum and the domesticated G. hirsutum to get an inter-specific comparison at the polyploid species level. We also added the three diploid species involved in the polyploidization event so that we could directly compare the diploid and the derived polyploid cottons. We applied the widely used AFLP (amplified fragment length polymorphism, Vos et al. 1995) fingerprinting method, which has been used both to pinpoint domestication events (Heun et al. 1997) and to reveal relationships among diploid and tetraploid cottons (Abdalla et al. 2001). In addition, our study provides information on the genetic diversity of G. barbadense, which at the present time is grown in the ‘Andean’ countries in general and in Peru in particular, with a potential value for conservation strategies.
Material and methods
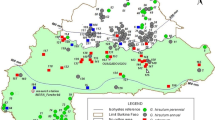
The cotton accessions analyzed in this investigation were obtained in two different ways. (1) Seed of 77 accessions was obtained from the Cotton Collection of the USDA-ARS, College Station, Texas, via Dr. E. Percival, who selected the accessions based on our wishes on which area to cover. (2) New material from Peru was collected by the authors. A collecting permit was obtained from the Instituto Nacional de Recursos Naturales (INRENA) to collect Gossypium spp. in the departments of Ancash, La Libertad, Lambayeque, Piura, Tumbes, Cajamarca, Amazonas, San Martin, Loreto, Ucayali and Huanuco. A Material Transfer Agreement (MTA) was signed to send collected material to Norway. Voucher specimens were deposited both at the Herbario Weberbauer at the Universidad Nacional Agraria La Molina and at the Herbarium Truxillense at the Universidad Nacional de Trujillo. Seed samples were deposited in the Instituto Nacional de Investigacion Agraria (INIA) Lima, Peru. Collecting data included passport information on each locality: longitude, latitude and altitude data were obtained using a global positioning system (GPS) receiver (Garmin, eTrex Summit), and phenotypic observations were added. For the DNA extraction, one small, fresh leaf was taken from each cotton plant and dried in sealed plastic bags containing dry silica gel. Seed was collected from those plants having mature bolls. Branches were cut off from a few representative plants as herbarium vouchers. In total, 100 cotton accessions were collected in this way, six of which are G. raimondii while the rest are G. barbadense.
DNA extraction
From those accessions with seed, three kernels were sown in separate pots under controlled conditions [25°C, 14/10-h (light/dark) photoperiod, pot soil kept constantly humid] at the Agricultural University of Norway (NLH), Ås, Norway, as requested by the Norwegian authorities (import permit obtained from Landbrukstilsynet, Ås, Norway). The seeds germinated between 3 days and 3 weeks after sowing. One plant from every accession was used for DNA extraction. The cotyledons (2–6 cm2 in size) of the plant selected were cut just above the meristem and air-brushed to avoid possible impurities. The tissue was ground in liquid nitrogen and transferred for storage to a −80°C freezer. A total of 131 accessions representing seven Gossypium species were included in the final analysis: 64 samples obtained from the USDA-ARS gene bank (Table 1) and 67 from the material collected in Peru (Table 2). Five samples originate from silica gel-dried material, whereas the remaining ones were obtained from freshly harvested tissue obtained from seed. DNA was extracted using a DNeasy Plant Mini kit from QIAGEN (Valencia, Calif.). The quality and quantity of the undigested DNA obtained was checked and quantified against undigested λ-DNA (Fermentas, Vilnius, Lithuania).
AFLP fingerprinting
AFLP fingerprinting was performed according to the original protocol of Vos et al. (1995) omitting the use of streptavidin beads for selecting EcoRI-biotinylated DNA fragments. Briefly, 500 ng DNA per sample was digested for 2 h at 37°C by 5 U EcoRI (Fermentas) and 5 U MseI (NEB) in 1× RL buffer (10 mM Tris-acetate pH 7.5, 10 mM Mg acetate, 50 mM potassium acetate, 5 mM DTT). The ligation of adapters fitting the cutting sites was done by adding T4 DNA ligase (Fermentas), 10 mM ATP, 10× RL buffer and by incubating the mixture for 3 h at 37°C. Thereafter, the selective pre-amplification with primers complementary to adapter sequence with a one-base extension was performed with E01 and M02. This primer combination was chosen based on the results of Abdalla et al. (2001) and Lacape et al. (2003). The PCR reactions [5.0 μl of the above-mentioned DNA digestion/ligation mix, 1.5 μl of each +1 primer (75 ng), 1.0 μl 10 mM dNTP mix, 0.25 μl Taq polymerase (5 U/μl) (Fermentas), 5.0 μl 10× PCR-buffer, 5.0 μl 25 mM MgCl2, in a total volume of 50 μl] were carried out on a PTC-200 thermocycler (MJ Research, Waltham, Mass.) at the temperature-time profile given by Vos et al. (1995). The resulting pre-amplification products were diluted 1:19 prior to the selective +3/+3 amplification. Primer labeling was done by phosphorylating the 5′-end of the E-primers with radioactive γ-[33P] and T4 kinase (Fermentas). The selective amplification mix with +3/+3 primers [5 μl of the diluted pre-amplified DNA, 0.5 μl *E primer, 0.6 μl M primer (see below for primer sequences), 0.4 μl dNTP (10 mM), 2.0 μl PCR buffer (10×), 1.2 μl MgCl2 (25 mM), 0.1 μl Taq polymerase (5 U/μl), in a total volume of 20 μl] was performed on a GeneAmp thermocycler (Applied Biosystems, Foster City, Calif.) according to Vos et al. (1995). Loading buffer (99% formamide, 10 mM EDTA, pH 8, 1.0 mg/ml xylene cyanol FF, 1.0 mg/ml bromophenol blue) was added to the final PCR products (50%), and the DNA was denatured by heat prior to running the samples (3 μl) on 5% polyacrylamide (acrylamide:bis, 19:1) gels under denaturing conditions. The electrophoresis apparatus (model S2/S2001, Gibco BRL Life Technologies, Gaithersburg, Md.) was filled with 2× TBE in the lower chamber and with 1× TBE in the upper chamber. For comparison 1.5 μl of a labeled 30- to 330-bp AFLP ladder from Invitrogen (Carlsbad, Calif.) was loaded on the polyacrylamide gel electrophoresis (PAGE). The gels were run at 80 W for 90 min and thereafter fixed, dried, and exposed to X-ray film (Kodak BioMax MR) for 8–48 h, depending on radiation intensity. X-ray films were developed with a AGFA Curix 60. The resulting images were scored by the naked eye for the presence/absence of AFLP bands using a light box. Overlapping samples as well as overlapping PCR reactions guaranteed that only reproducible AFLPs were scored and that the different PAGEs were well aligned. Based on a screening of 40 E01/M02 primer combinations with four samples (results not presented here), the following primer combinations were used to fingerprint all 131 cottons: E35/M48, E38/M48, E38/M49, E40/M50, E40/M59, E40/M60, E41/M47, E41/M60 (Invitrogen). The core sequence for the EcoRI primers is 5′-GACTGCGTACCAATTCNNN-3′ and 5′-GATGAGTCCTGAGTAANNN-3′ for MseI. The −3′ end sequences for the primers used in this study are as follows: E01-A, E35-ACA, E38-ACT, E40-AGC, E41-AGG, M02-C, M47-CAA, M48-CAC, M49-CAG, M50-CAT, M59-CTA, M60-CTC.
Data analysis
The X-ray films were scored in a binary manner using “1” to indicate the presence of a polymorphic AFLP band and ‘0’ to indicate its absence. Only unambiguous bands within an 80- to 800-bp range were scored. The data were assembled in a matrix and analyzed for pairwise genetic similarity (Gs) using the Dice similarity coefficient (Dice 1945; Nei and Li 1979) in the simqual option of ntsys-pc v.2.11f (Rohlf 2000). The Dice similarity is given by the equation Gs ij = 2a/(2a+b+c) where Gs ij expresses the genetic similarity between line i and j, a is the number of bands present in both accessions, b is the number of bands present in i and absent in j, and c is the number of bands absent in i and present in j. Genetic distance (Gd) measures were obtained for all Gs coefficients using the equation Gd ij =1−Gs ij . The Dice-based Gd matrix was used for the neighbor joining (NJ) method of Saitou and Nei (1987). The Gs data were used for calculating unweighted pair-group method with arithmetic means (UPGMA) (Sokal and Sneath 1963). The trees were calculated with the program ntsys and paup (ver. 4.0b10 for Macintosh; Swofford 1998). The robustness of the obtained trees was evaluated by comparing the different data analyses [NJ, UPGMA, and principal coordinate analysis (PCA)] and by bootstrapping (Felsenstein 1985) with 1,000 re-samplings (paup).
Results
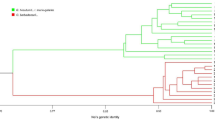
A total of 340 unambiguous, polymorphic bands were scored with eight AFLP primer combinations. Of these polymorphic AFLP bands, 185 were polymorphic among the tetraploid species, and 93 were polymorphic within the final (see below) G. barbadense sample. The level of intra-specific variation in G. barbadense ranged from seven polymorphic bands out of a total of 44 (16%) in primer combination E38/M48 to 16 out of 60 (27%) in E35/M48. Figure 1 displays the NJ tree of the full data set. The intermediate position of the AD-tetraploids relative to the A- and D-diploids is clearly visible. G. herbaceum, G. arboreum, and G. raimondii are well separated and so are the four tetraploids. The average genetic distances between species (see inlet table of Fig. 1, which also contains the within-species distances) range from 0.16 (G. barbadense versus G. tomentosum) to 0.82 (G. herbaceum versus G. raimondii).
To address the tree topology of the tetraploid species, we discarded all diploid species (and 12 tetraploid accessions that had missing data in more than one primer combination) from the data matrix. The resulting Dice similarity-based UPGMA is shown in Fig. 2a. The robustness of the clustering was evaluated by bootstrapping. All species-defining branches are well supported and equal those obtained in Fig. 1. The G. barbadense sub-cluster in Fig. 2a displays a first split distinguishing a first subset comprising 33 coastal Peruvian accessions. The remaining accessions from northernmost coastal Peru (the departments of Piura and Tumbes) and accessions from coastal Ecuador are clustered basal to the east-of-Andes accessions and accessions from Bolivia, Brazil, Columbia, Venezuela, and the Caribbean and Pacific Islands.
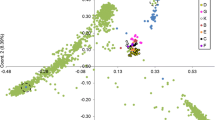
To further investigate these geographic patterns a G. barbadense NJ tree was computed on the basis of the 96 accessions displaying no missing data in any of the primer combinations (Fig. 2b). This NJ intraspecific analysis ‘zooms in’ on the topology of the G. barbadense cluster of Fig. 2a and displays the same general pattern in an unrooted way. The highest distance between two accessions of G. barbadense within this subset is found between the coastal Peruvian P1 and the Hawaiian Gb480 (0.13), while the average distance is limited to 0.06. A principal coordinate analysis [not shown] revealed the same overall pattern for these 96 accessions and the two first principal coordinates accounted for 23.9% of the total variation.
Discussion
Diploid versus tetraploid relationships
The intermediate position of the four allotetraploid AD-cotton species relative to the A- and D-diploids in the NJ analysis (Fig. 1) is in full agreement with the established cytogenetic theory (Endrizzi et al. 1985). The genetic distances between the two different diploid species (A ver. D) and the derived tetraploids are large (0.45–0.59) compared to the distance among the tetraploids (0.16–0.25), as is expected by the bottleneck effect of a single polyploidization event (Cronn et al. 1999; Wendel and Cronn 2003). It is also apparent that G. raimondii is less related to the tetraploids than the two A-genome species (G. arboreum and G. herbaceum), again in agreement with cytogenetic and molecular studies (Endrizzi et al. 1985; Wendel 1989; Cronn et al. 1999; Small et al. 1999; Abdalla et al. 2001; Senchina et al. 2003). This consistency of the results of AFLP analyses with cytogenetic studies is important as caution is recommended when using AFLP markers in inter-specific comparisons (e.g., El-Rabey et al. 2002), since the homology of the bands will be less well preserved the more evolutionary distant the analyzed material is. Abdalla et al. (2001) identify AFLP bands that are shared between a subset of the A-diploids and tetraploids and call these ‘A-related’ markers; they do the same for identifying ‘D-related’ markers. Applying this classification to our data, we count 136 A-related markers and 75 D-related ones, which is a proportion similar to that obtained by Abdalla et al. (2001). These numbers were subsequently used by Wendel and Cronn (2003) in supporting the relative contribution of the two diploids for the emergence of the tetraploid cottons. Similar inter-specific comparisons have shown that Aegilops genomes, for example, are arranged by AFLP/NJ in accordance with known cytogenetics (Sasanuma et al. 2004); this also holds true for Avena (Drossou et al. 2004), Solanum (Kardolus et al. 1998) Musa (Ude et al. 2002), and Cicer (Sudupak et al. 2004). Even in the investigation of El-Rabey et al. (2002), where the problem surrounding the homology assumption of co-migrating AFLP bands was studied, the AFLP clustering of Hordeum genomes was ‘in full concordance with geographic origin, cytology, and taxonomic status’.
Inter-specific relations among AD-tetraploid cottons
Extensive effort has been expended on studying the molecular phylogeny of Gossypium (Wendel 1989; Wendel and Albert 1992; Cronn et al. 1996; Seelanan et al. 1997; Small et al. 1998; 1999; Seelanan et al. 1999; Small and Wendel 2000; Liu et al. 2001; Cronn et al. 2002). However, the phylogenetic relationship among the tetraploids has remained elusive due to their short evolutionary separation and low levels of nucleotide diversity (Small et al. 1998; 1999). The radiation of the tetraploids has only had an estimated 1.5 million years (Senchina et al. 2003), and the separation into ‘good species’ is confounded, as can be concluded from the detection of gene flow across the taxonomic species borders (Percy and Wendel 1990; Wendel and Percy 1990; Brubaker and Wendel 1994) and from examples of testcrosses yielding fertile F2 progenies (e.g., Hutchinson et al. 1947; Endrizzi et al. 1985). Nevertheless, Small et al. (1998) described two main branches within the tetraploid group, with one branch leading to G. mustelinum and one branch leading to the remaining four species of which G. barbadense and G. darwinii form a sister group to G. hirsutum and G. tomentosum. Later studies have either not addressed the relationship between the tetraploids (Small et al. 2000; Cronn et al. 2002) or failed to resolve the relative position of G. tomentosum within the group (Liu et al. 2001). Our data support the basal branching of G. mustelinum but indicate a closer clustering of G. tomentosum towards G. barbadense than towards G. hirsutum. The difference in genetic distance is low (0.16 between G. barbadense and G. tomentosum versus 0.22 between G. hirsutum and G. tomentosum) and can not be taken as a final evidence for a phylogenetic revision. Yet Hutchinson et al. (1947) reported on chromosome pairing and testcross progeny studies among those tetraploids and concluded that G. tomentosum is closer to G. barbadense than to G. hirsutum, as also suggested by our data. Another interesting feature was observed when five G. barbadense accessions from the Galapagos Islands were included—two of the accessions (Gb624/Gb625) clustered separate from all other G. barbadense. This could have been caused by the fact that some of the Galapagos accessions in the USDA-ARS Cotton Collection contain introgressions from G. darwinii growing in close proximity (E. Percival, personal communication). Due to that uncertainty, we eliminated all Galapagos accessions from our analyses. The observed bias possibly caused by the indirect involvements of the fifth tetraploid cotton (i.e., G. darwinii) would make future analyses of defined Galapagos material (G. barbadense and G. darwinii) important.
Domestication and genetic diversity
G. barbadense is widely distributed, covering the whole range of tropical South America including an overlapping distribution with G. hirsutum in the northern part of the continent and in the Caribbean (Brubaker et al. 1999; Wendel and Cronn 2003). Present-day indigenous G. barbadense is grown in gardens and local farmers’ fields in northwestern (NW) South America and is referred to as ‘dooryard’ cotton or ‘commensals’. The plants occur as perennial shrubs with thick basal stems up to a tree size, 4 m tall, and they produce lint in an array of brown colors. The historical record concerning the native distribution of wild G. barbadense is rather anecdotal, yet extant wild populations were reported from Guayas and Los Rios in Ecuador and Tumbes, Peru (Stephens and Moseley 1973, 1974; Schwendiman et al. 1986; Percival and Kohel 1990). The search for truly wild accessions is even more complicated since the wild-to-domesticated continuum in G. barbadense hardly allows categorical distinctions. Cotton remains obtained from archeological excavation sites from central- and northern coastal Peru show this wild-to-domesticated continuum for seed size, boll size, and fiber width. It has also been proposed that a selection for greater differentiation between fuzz and lint led to morphs with a strongly reduced fuzz layer, called ‘tufted’ seeds, and the so-called ‘kidney-seeded’ type of G. barbadense, as they were more easily ginned by hand (Turcotte and Percy 1990; Brubaker et al. 1999). Stephens (1975) found that the remains from the archaeological site Huaca Prieta (from about 2,500 BC) display chocolate-colored fibers and that the oldest excavation layers contain only fuzzy seeds while tufted seeds appear in more recent layers. Selection also eliminated the hard seed coat and delayed germination (Hutchinson et al. 1947). In addition, a low selection pressure was probably induced on traits like percentage lint, lint length and strength, and eventually also on color differences for artistic purposes in weaving and improved catching abilities of fishing nets (Hutchinson et al. 1947; Vreeland 1999). More specialized accessions have undergone selection for photoperiod neutrality, early maturing and fine white lint, etc., but these later domestication traits were selected in colonial times and thereafter (Hutchinson et al. 1947; Brubaker et al. 1999) and should not be considered here. In summary, for primitive cottons there exists no clear separation between wild and present-day ‘dooryard’ cottons. In fact, these dooryards are envisioned to be derived directly from local wild progenitors (Hutchinson et al. 1947; Percy and Wendel 1990; Brubaker et al. 1999). Despite changing climate, intermixing, and other variables, the geographic pattern of present-day dooryards might provide an insight into the pre-Columbian distribution of this plant. Thus, we focused our re-sampling on dooryard cottons to obtain (in combination with gene bank material) a representative sample of G. barbadense across its assumed pre-Colombian range. The most striking observation we obtained is the unique diversity in the large majority of accessions from coastal Peru. A few accessions from the NW Peru cluster with southwestern (SW) Ecuadorian accessions, and together these accessions are basal to the remaining G. barbadense. This indicates that primitive domesticated G. barbadense has its domestication center in the NW Peru/SW Ecuador region, as proposed (mainly derived from archaeological evidence) by Piperno and Pearsall (1998). Further, we also suggest, that cotton from this core area was transported across the Andes, and spread thereafter south into Bolivia, east into Brazil, and north into Colombia, Venezuela, and the Caribbean and Pacific Islands. This scenario is proposed with caution as the data do not yield high bootstrap-values on the most basal nodes of the G. barbadense cluster, but the pattern is evident in all the different analyses.
Our field observations (Fernandez et al. 2003) support the DNA data as they indicate a high diversity of ‘primitive’ types in the coastal departments of Peru. We encountered dooryard and feral plants with strongly arborescent growth habit, small bolls, and lint colors varying from reddish-brown to light yellow-brown, and purplish-brown to deep chocolate-brown. It was also apparent that the cottons east of the Andes displayed only a subset of this variation, with generally larger bolls (long and slender) and lint that varied in color from lighter shades of brown to white. In the ‘Selva Alta’ of San Martin, a region where the Amazonian rainforest meets the Andes, plants in the next stage of domestication, i.e., landraces, are grown in small-scale commercial productions of organic cotton. The domestication trait ‘kidney-seeded’ is known only from east of the Andes (Turcotte and Percy 1990) and was present in some of our accessions from Bolivia and Brazil, indicating that our geographic interpretation of the data follows an expansion towards increased domestication level.
Conclusion
Our DNA analyses confirm the relationship among diploid and tetraploid cottons. They also confirm the archeology/allozyme-derived assumption of South American cotton domestication having its origin in the coastal zone of NW Peru and SW Ecuador. We add that cotton spread out from this center (probably within the framework of domestication) over the Andes and from there into other parts of the continent. We also observed a tremendous diversity in G. barbadense found along the remaining northern Peruvian coast. This diversity is facing serious threats due to habitat destruction and the replacement of local types, and both ex-situ and in-situ conservation strategies should be implemented.
References
Abdalla AM, Reddy OUK, El-Zik KM, Pepper AE (2001) Genetic diversity and relationships of diploid and tetraploid cottons revealed using AFLP. Theor Appl Genet 102:222–229
Brubaker CL, Wendel JF (1994) Reevaluating the origin of domesticated cotton (Gossypium hirsutum Malvaceae) using nuclear restriction fragment length polymorphisms (RFLP). Am J Bot 81:1309–1326
Brubaker CL, Bourland FM, Wendel JF (1999) The origin and domestication of cotton. In: Smith CW, Cothren JT (eds) Cotton: origin history, technology, and production, 1st edn. Wiley, New York, pp 3–31
Cronn RC, Zhao XP, Paterson AH, Wendel JF (1996) Polymorphism and concerted evolution in a tandemly repeated gene family: 5S ribosomal DNA in diploid and allopolyploid cottons. J Mol Evol 42:685–705
Cronn RC, Small RL, Wendel JF (1999) Duplicated genes evolve independently after polyploid formation in cotton. Proc Natl Acad Sci USA 96:14406–14411
Cronn RC, Small RL, Haselkorn T, Wendel JF (2002) Rapid diversification of the cotton genus (Gossypium: Malvaceae) revealed by analysis of sixteen nuclear and chloroplast genes. Am J Bot 89:707–725
Dejoode DR, Wendel JF (1992) Genetic diversity and origin of the Hawaiian-Islands cotton, Gossypium tomentosum. Am J Bot 79:1311–1319
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–308
Drossou A, Katsiotis A, Leggett JM, Loukas M, Tsakas S (2004) Genome and species relationships in genus Avena based on RAPD and AFLP molecular markers. Theor Appl Genet 109:48–54
El-Rabey HA, Badr A, Schafer-Pregl R, Martin W, Salamini F (2002) Speciation and species separation in Hordeum L. (Poaceae) resolved by discontinuous molecular markers. Plant Biol 4:567–575
Endrizzi JE, Turcotte EL, Kohel RJ (1985) Genetics, cytology, and evolution of Gossypium. Adv Genet 23:271–375
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fernandez HAM, Rodriguez REF, Westengen O (2003) Biología y Etnobotánica del Algodón Nativo Peruano (Gossypium barbadense L., Malvaceae). Arnaldoa 10:93–108
Fryxell PA (1992) A revised taxonomic interpretation of Gossypium L., (Malvacea). Rheedea 2:108–165
Heun M, SchaferPregl R, Klawan D, Castagna R, Accerbi M, Borghi B, Salamini F (1997) Site of einkorn wheat domestication identified by DNA fingerprinting. Science 278:1312–1314
Hutchinson JB, Silow RA, Stephens SG (1947) The evolution of Gossypium and the differentiation of the cultivated cottons, 1st edn. Oxford University Press, London
Kardolus JP, van Eck HJ, van den Berg RG (1998) The potential of AFLPs in biosystematics: a first application in Solanum taxonomy (Solanaceae). Plant Syst Evol 210:87–103
Lacape JM, Nguyen TB, Thibivilliers S, Bojinov B, Courtois B, Cantrell RG, Burr B, Hau B (2003) A combined RFLP-SSR-AFLP map of tetraploid cotton based on a Gossypium hirsutum×Gossypium barbadense backcross population. Genome 46:612–626
Liu Q, Brubaker CL, Green AG, Marshall DR, Sharp PJ, Singh SP (2001) Evolution of the FAD2-1 fatty acid desaturase 5′ UTR intron and the molecular systematics of Gossypium (Malvaceae). Am J Bot 88:92–102
First evidence of cotton at Neolithic Mehrgarh, Pakistan: analysis of mineralized fibres from a copper bead. J Archaeol Sci 29:1393–1401
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Percival AE, Kohel RJ (1990) Distribution, collection, and evaluation of Gossypium. Adv Agron 44:225–256
Percy RG, Wendel JF (1990) Allozyme evidence for the origin and diversification of Gossypium barbadense L. Theor Appl Genet 79:529–542
Phillips LL (1963) The cytogenetics of Gossypium and the origin of New World cottons. Evolution 17:460–469
Phillips LL (1964) Segregation in new allopolyploids of Gossypium. V. Multivalent formation in New World × Asiatic and New World × wild American Hexaploid. Am J Bot 51:324–329
Piperno DR, Pearsall DM (1998) The origins of agriculture in the lowland Neotropics. Academic, San Diego
Rohlf FJ (2000) Numerical taxonomy and multivariate analysis system, ver. 2.11. Applied Biostatistics, New York
Rossen J, Dillehay TD, Ugent D (1996) Ancient cultigens or modern intrusions? Evaluating plant remains in an Andean case study. J Archaeol Sci 23:391–407
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasanuma T, Chabane K, Endo TR, Valkoun J (2004) Characterization of genetic variation in and phylogenetic relationships among diploid Aegilops species by AFLP: incongruity of chloroplast and nuclear data. Theor Appl Genet 108:612–618
Schwendiman J, Ano G, Percival AE (1986) Cotton collecting in continental Ecuador and Galapagos Islands. FAO/IPBGR Plant Genet Resources Newsl 64:33–37
Seelanan T, Schnabel A, Wendel JF (1997) Congruence and consensus in the cotton tribe (Malvaceae). Syst Bot 22:259–290
Seelanan T, Brubaker CL, Stewart JM, Craven LA, Wendel JF (1999) Molecular systematics of Australian Gossypium section Grandicalyx (Malvaceae). Syst Bot 24:183–208
Senchina DS, Alvarez I, Cronn RC, Liu B, Rong JK, Noyes RD, Paterson AH, Wing RA, Wilkins TA, Wendel JF (2003) Rate variation among nuclear genes and the age of polyploidy in Gossypium. Mol Biol Evol 20:633–643
Small RL, Wendel JF (2000) Phylogeny, duplication, and intraspecific variation of Adh sequences in new world diploid cottons (Gossypium L., Malvaceae). Mol Phylogenet Evol 16:73–84
Small RL, Ryburn JA, Cronn RC, Seelanan T, Wendel JF (1998) The tortoise and the hare: choosing between noncoding plastome and nuclear ADH sequences for phylogeny reconstruction in a recently diverged plant group. Am J Bot 85:1301–1315
Small RL, Ryburn JA, Wendel JF (1999) Low levels of nucleotide diversity at homoeologous Adh loci in allotetraploid cotton (Gossypium L.). Mol Biol Evol 16:491–501
Sokal RR, Sneath PHA (1963) Principles of numerical taxonomy. Freeman, San Francisco
Solis RS, Haas J, Creamer W (2001) Dating caral, a preceramic site in the Supe Valley on the central coast of Peru. Science 292:723–726
Stephens SG (1944) Phenogenetic evidence for the amphidiploid origin of New World cottons. Nature 153:53–54
Stephens SG (1975) A reexamination of the cotton remains from Huaca Prieta, North Coastal Peru. Am Antiquity 40:406–419
Stephens SG, Moseley EM (1973) Cotton Remains from archeological sites in central coastal Peru. Science 180:186–188
Stephens SG, Moseley ME (1974) Early domesticated cottons from archaeological sites in central coastal Peru. Am Antiquity 39:109–122
Sudupak MA, Akkaya MS, Kence A (2004) Genetic relationships among perennial and annual Cicer species growing in Turkey assessed by AFLP fingerprinting. Theor Appl Genet 108:937–944
Swofford DL (1998) paup*: phylogenetic analysis using parsimony (*and other methods) ver. 4.0b10 for Macintosh. Sinauer, Sunderland
Turcotte EL, Percy RG (1990) Genetics of kidney seed in Gossypium barbadense L. Crop Sci 30:384–386
Ude G, Pillay M, Nwakanma D, Tenkouano A (2002) Analysis of genetic diversity and sectional relationships in Musa using AFLP markers. Theor Appl Genet 104:1239–1245
Vos P, Hogers R, Bleeker M, Reijans M, Vandelee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP—a new technique for DNA-fingerprinting. Nucleic Acids Res 23:4407–4414
Vreeland JM (1999) The revival of colored cotton. Sci Am 280:112–118
Wendel JF (1989) New world tetraploid cottons contain Old-World cytoplasm. Proc Natl Acad Sci USA 86:4132–4136
Wendel JF (1995) Cotton. In: Simmonds S, Smartt J (eds) Evolution of crop plants, 1st edn. Longman, London, pp 358–366
Wendel JF, Albert VA (1992) Phylogenetics of the cotton genus (Gossypium)—character-state weighted Parsimony analysis of chloroplast-DNA restriction site data and its systematic and biogeographic implications. Syst Bot 17:115–143
Wendel JF, Cronn RC (2003) Polyploidy and the evolutionary history of cotton. Adv Agron 78:139–186
Wendel JF, Percy RG (1990) Allozyme diversity and introgression in the Galapagos-Islands endemic Gossypium darwinii and its relationship to continental Gossypium barbadense. Biochem Syst Ecol 18:517–528
Wendel JF, Rowley R, Stewart JM (1994) Genetic diversity in and phylogenetic-relationships of the Brazilian endemic cotton, Gossypium mustelinum (Malvaceae). Plant Syst Evol 192:49–59
Acknowledgements
We thank Dr. E. Percival (USDA-ARS, Texas) for providing seed material. We also extend our gratitude to J. Lazo, G. Arevalo, F. Balavarca, and C. Basurto for assistance and information; K. Ramirez and M.R. Pastor at INRENA for obtaining the permission to collect and export the germplasm; M. White, C. Deza, E. Rodriguez, and V. Medina for accompanying the collecting efforts in La Libertad; M.I. Olivos Farro and J. Escurra Puicon for collaboration in Lambayeque; P.M. Reyes More for information; E. Manco for access to PRONIRGEB germplasm; P. Azang Huaman for collaboration in San Martin (all in Peru). Thanks are also due to K. Vollan and J.H. Sønstebø for their advice and technical assistance during laboratory work at AUN, Norway.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H.C. Becker
This publication is dedicated to Prof. Dr. Drs.h.c. Gerhard Röbbelen on the occasion of his 75th birthday
Rights and permissions
About this article
Cite this article
Westengen, O.T., Huamán, Z. & Heun, M. Genetic diversity and geographic pattern in early South American cotton domestication. Theor Appl Genet 110, 392–402 (2005). https://doi.org/10.1007/s00122-004-1850-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-004-1850-2