Abstract
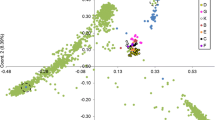
Gossypium species (± 49) represent a vast resource of genetic diversity for the improvement of cultivated cotton. To determine intra- and inter-specific genetic relationships within a diverse collection of Gossypium taxa, we employed 16 AFLP primer combinations on three diploid species, Gossypium herbaceum L. (A1), Gossypium arboreum L. (A2) and Gossypium raimondii Ulbrich (D5), and 26 AD allotetraploid accessions (Gossypium barbadense L. and Gossypium hirsutum L.). A total of 1180 major AFLP bands were observed; 368 of these (31%) were polymorphic. Genetic similarities among all taxa ranged from 0.21 (between the diploid species G. arboreum and G. raimondii) up to 0.89 (within G. barbadense). Phenetic trees based on genetic similarities (UPGMA, N-J) were consistent with known taxonomic relationships. In some cases, well-supported phylogenetic relationships, as well as evidence of genetic reticulation, could also be inferred. UPGMA trees and principal coordinate analysis based on genetic similarity matrices were used to identify genetically distinct cultivars that are potentially important sources of germplasm for cotton improvement, particularly of fiber quality traits. We show that AFLP is useful for estimating genetic relationships across a wide range of taxonomic levels, and for analyzing the evolutionary and historical development of cotton cultivars at the genomic level.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received: 17 January 2000 / Accepted: 4 May 2000
Rights and permissions
About this article
Cite this article
Abdalla, A., Reddy, O., El-Zik, K. et al. Genetic diversity and relationships of diploid and tetraploid cottons revealed using AFLP. Theor Appl Genet 102, 222–229 (2001). https://doi.org/10.1007/s001220051639
Issue Date:
DOI: https://doi.org/10.1007/s001220051639