Abstract
To estimate genetic diversity of the residual northern populations of Oryza rufipogon, a total of 232 individuals from six populations were analyzed using microsatellites (SSRs). The O. rufipogon populations with different status included three from Dongxiang (Jiangxi Province) and three from Chaling (Hunan Province) in China. The 23 rice SSR primer pairs selected from the RiceGenes Database detected a total of 115 alleles, indicating that all the SSR loci were polymorphic in this study. The total gene diversity was 0.919 in the six O. rufipogon populations, and the Donxiang populations showed higher diversity than the Chaling populations. More significant genetic differentiation and less gene flow were found among the Dongxiang populations than those from Chaling. The two putative introgressed populations showed relatively high genetic variation. One in situ conserved population from Dongxiang had the lowest level of genetic diversity. The re-introduced population from Chaling restored about 90% of the genetic variation, compared with the original source population. It is concluded from these results that a relatively high level of genetic variation resided in the northern O. rufipogon populations and continued efforts of conservation of these populations are needed; and that the conservation of some Chaling and Dongxiang populations has been effective in preventing gene flow from cultivated rice. Introgression of cultivated rice demonstrated significant impacts on genetic variability of the O. rufipogon populations, and should be carefully considered in conserving this wild rice. This study also suggested that re-introduction to its original habitats is an effective approach to restore O. rufipogon populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The perennial common wild rice, Oryza rufipogon Griff., known as the ancestor of Asian cultivated rice (Oryza sativa L.), is the most important germplasm for rice improvement (Oka 1988). The first male-sterility (MS) gene was found in O. rufipogon and introduced to cultivated rice, which has led to the development of high yielding hybrid rice varieties (Yuan et al. 1989). Other agronomically beneficial traits, such as rice tungro virus resistance, elongation ability and tolerance to acid sulfate soil found in the wild rice, are of great importance for rice breeding (Xiao et al. 1996; Bellon et al. 1998). China is the northern most distribution of O. rufipogon and great genetic diversity has previously been documented in Chinese populations (Wang et al. 1996; Ge et al. 1999; Gao et al. 2000a; Song et al. 2001). However, this species has been under considerable threats in China in the past decades due to changes in farming systems, economic development, urbanization and other human disturbances. Effective conservation of O. rufipogon is urgently needed to preserve the remaining populations (Hong 1995).
O. rufipogon populations in Dongxiang (26°14'N, 116°36'E) of Jiangxi Province and Chaling (26°50'N, 113°40'E) of Hunan Province in China are the residual populations of this species (Zhou 1995; Gao et al. 2000b). The first survey of these areas was carried out independently by the Institute of Rice Research (IRR) belonging to Jiangxi Academy of Agricultural Science (JAAS) and Hunan Academy of Agricultural Science (HAAS) in 1983. During the survey, a total of nine natural O. rufipogon populations were found in Dongxiang, covering an area of about 15 km2. Habitats of O. rufipogon have considerably deteriorated, since then, due to the expansion of rice fields, and currently only three populations remain (Gao et al. 2000b). To avoid complete extinction of O. rufipogon populations in Dongxiang, the IRR of JAAS built a 2-m-high concrete wall in 1995 to surround the Anjashan and Shuitaoshu populations (DX-Pa and DX-Ps) for in situ conservation. The third population (DX-Pz) was left unprotected.
The Chaling O. rufipogon populations are located in the Huli wetland with an area of about 2 ha. Due to changes of water level, some of the Chaling O. rufipogon populations were already extinct before 1994. Zhou (1995) re-introduced O. rufipogon individuals using samples collected in 1983 and were preserved as clonal strains in the wild rice conservation plots of HAAS in Changsha city to restore the extinct population (CL-Pr). In addition, some O. rufipogon plants were germinated from seeds remaining in soil in the same year in the area where O. rufipogon was not re-introduced (Zhou 1995). Around this time farmers started to grow cultivated rice varieties in the area, and the closest distance between an O. rufipogon population (CL-Pi) and rice fields was about 10 m. Individuals in this O. rufipogon population showed considerable morphological variations, possibly due to introgression between the wild and cultivated rice species.
Microsatellites or simple sequence repeats (SSRs) are tandem arrays of short nucleotide repeats with 1–5 bp (Wu and Tanksley 1993) and are widely dispersed in all eukaryotic genomes. Dinucleotide repeats (AT/TA)n and (GA/CT)n are commonly found in vascular plants. SSRs are characterized as co-dominant, highly polymorphic, abundant and randomly distributed markers in genomes. SSR markers can be easily amplified by PCR and are probably selectively neutral (Akagi et al. 1997). Microsatellites have been used for studies of parentage (Roa et al. 2000), genetic mapping and breeding (McCouch et al. 1997), gene flow (Chase et al. 1996; Innan et al. 1997), genetic diversity and population differentiation (Byrne 1996; Cho et al. 2000). To date, more than two thousand SSR markers of cultivated rice are available, which provides a powerful tool for studying O. rufipogon, as SSR markers have good cross-species amplification to the close relatives (Roa et al. 2000).
In order to effectively implement an in situ conservation strategy of threatened northern O. rufipogon populations, several questions need to be addressed. First, what is the genetic variation pattern of the populations? Second, what impact does the current conservation measures, such as isolating populations by a concrete wall, impose on their genetic structure? Third, does gene flow occur between the cultivated and wild rice, and what will be its genetic consequence to wild rice populations? And finally, whether re-introduction will restore genetic diversity of extinct wild rice populations? To answer these questions, all six populations of O. rufipogon from the Dongxiang and Chaling sites were analyzed using 23 SSR markers developed from cultivated rice. The objective of this study was to assess the genetic variation pattern of the O. rufipogon populations, to determine the occurrence and genetic consequence of introgression of cultivated rice to wild rice populations, and to estimate the effectiveness of in situ conservation under different conditions.
Materials and methods
Samples used for SSR analysis
Leaf samples from six O. rufipogon populations were used for SSR analysis. Three populations were collected from Dongxiang, Jiangxi Province. Two populations (DX-Pa and DX-Ps) are in situ conserved sites, and surrounded by a concrete wall, separated over 2 km. One population (DX-Pz) was composed of individuals that were dispersed along the ridges and/or ditches of cultivated rice fields, about 1 km from the DX-Pa population and 3 km from the DX-Ps population. Other three populations were collected from the Chaling O. rufipogon conservation site in Hunan Province and the spatial distance between these populations was about 50–90 m. The CL-Po was an original population conserved since it was found in 1978, the CL-Pr was a re-introduced population to the site where the original population was extinct in 1994, and the CL-Pi population was an original population but adjacent to the farmer's cultivated rice field.
Conservation status and the distance of these O. rufipogon populations from the cultivated rice fields are shown in Table 1. A total number of 232 individuals were sampled from different population for the analysis (Table 1). About 1 g of fresh leaves of each O. rufipogon individual was collected and placed in a plastic bag containing silica gel for fast drying following the description by Xie et al. (1999). To avoid collecting the same clone, samples were collected 5-m apart.
PCR assay
For DNA preparation, total genomic DNA was extracted following the method described by Xie et al. (1999). A total number of 23 SSR primer pairs (one each on different chromosome arms) were selected to assay genetic variation in O. rufipogon (Table 2), based on the RiceGenes Database (http://gramene.org). Primers were chosen based on the number of alleles in each locus with relatively high polymorphism. The PCR reactions were performed in a PTC 10096v thermocycler (MJ Research Inc, Watertown, Mass.) programmed as described by Wu and Tanksley (1993). A denaturation period of 4 min at 94°C was followed by 36 cycles of 40 s at 94°C, 30 s at 55°C and 40 s at 72°C, and then 10 min at 72°C for final extension. Reactions were carried out in a volume of 20 μl containing 1 × buffer, 1 mM each of dATP, dCTP, dGTP and dTTP, 10 μM of SSR primer, 50 ng of genomic DNA and 0.6 units of Taq polymerase (TaKaRa Inc.).
The PCR products were separated in 6% polyacrylamide denaturing gels of 200×125×1 mm (length × width × thickness) in size. Before sample loading, the PCR products mixed with equal amount of buffer were denatured at 95°C for 5 min. After electrophoresis, bands were revealed using the following modified silver-staining procedure. The gel was peeled after electrophoresis and fixed with 10% ethanol containing 0.5% acetic acid solution. The fixed gel was washed twice with ddH2O, then placed into a 200–400 ml 0.1% AgNO3 solution to stain for 10–18 min. The stained gel was transferred into ddH2O and washed twice; then, the gel was placed in 1.5% NaOH solution (about a 200–400 ml vol.) containing 0.4% formaldehyde and 0.019% sodium borate to develop for about 10 min to obtain visible DNA bands. The developed gel was washed with ddH2O twice and placed in 0.75% NaHCO3 to stop developing. The gel was covered with cellophane to dry, and photos were taken using a digital camera (Sony Inc.).
Genotype score
Because SSR markers are co-dominant, the amplified DNA bands represented different alleles and different banding patterns can be scored as different genotypes. To assist in allele scoring, DNA of rice varieties IR36, IR64 and Nipponbare were included in our study for amplification and analysis. Banding patterns identified in the rice varieties IR36, IR64 and Nipponbare, available at the RiceGenes Database, were used as reference materials to help the score of the different alleles in the O. rufipogon samples. As a consequence, the co-dominant SSR banding patterns were scored as AA or BB (for homozygote) and AB (for heterozygote) genotypes corresponding to the alleles identified in the RiceGenes Database.
Statistical analysis
Allele frequencies, effective allele number (Ae), expected (He) and observed heterozygosity (Ho) (Nei 1978), and gene diversity (Shannon's information index = I) were calculated to estimate genetic variation level. The single-locus and mean fixation indices (FI) (Wright 1978) were computed for polymorphic loci to test for departure from Hardy-Weinberg equilibrium values. According to the FI, the outcrossing rate [t=(1−FI)/(1+FI)] was calculated to indirectly estimate the mating pattern of the O. rufipogon populations (Wright 1978). Population differentiation was analyzed for polymorphism between populations, within locations, and between regions by F-statistics (Weir and Cockerham 1984). Gene flow (Nm) was estimated from Nm=(1/4)(1−Fst)/Fst (Nei 1987). All of the above calculations were performed using POPGENE program ver. 1.31 (Yeh et al. 1999). Relationships of the O. rufipogon populations were estimated from the SSR data using the UPGMA clustering method on the basis of Nei's (1978) unbiased genetic distance. The UPGMA tree was constructed using NTsys program ver. 1.8 (Rohlf 1994).
Results
All of the 23 selected rice SSR primer pairs amplified visible DNA fragments for the O. rufipogon populations and no null alleles were found. Each plant within the six populations had a unique genotype, except for two individuals from the DX-Ps population that had identical genotypes. A total of 115 alleles were scored from the 232 wild rice individuals. The highest number of alleles was scored from the locus RM44 (eight alleles) and the lowest was found from the locus RM19 (two alleles) in the O. rufipogon samples. The selected SSR primers generated an average of 4.61 alleles per locus. The allele frequency data (Table 3) indicated one (RM230) to three monomorphic loci, (RM167, RM230 and RM289) in the Chaling populations. The loci, RM14, RM205, RM211 and RM276 showed a monomorphic pattern in the DX-Pa and DL-Pz population. The DX-Ps population showed a low polymorphism with 11 monomorphic loci. An example of the polymorphism patterns generated with the SSR markers RM180, RM219 and RM280 is shown in Fig. 1.
SSR amplification products generated by the primer pairs, RM44, RM180 and RM219. A Products of RM44; lanes 1–13 were samples of the DX-Pa populations, lanes 16–34 were samples of the DX-Ps. B Products of RM180; lanes 1–24 were samples of the DX-Pa population. C Products of RM219; lanes 1–25 were samples of CL-Pi; M, pUC19DNA/MspI(HpaII) markers (MBI)
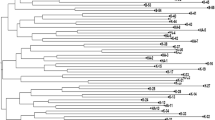
The UPGMA tree based on Nei's (1978) unbiased genetic identity for all populations revealed two distinct groups (Fig. 2). The three populations from the Chaling site were clustered in one group with relatively low differentiation among each other, whereas the three populations from the Dongxiang site were clustered in another group, showing relatively high differentiation among the populations. Within Dongxiang, the two populations, DX-Pa and DX-Pz with relatively close spatial distance were clustered together, and the DX-Ps population that had a longer spatial distance showed considerable differentiation from the other two populations.
A UPGMA tree for the six populations of O. rufipogon based on Nei's (1978) unbiased genetic identity
The average number of effective alleles (Ae), the observed (Ho) and expected heterozygosity (He), and Shannon's index (I) were used to demonstrate the level of population genetic diversity, which are listed in Table 1. In general, the populations included in this study showed a relatively high level of genetic diversity with Ae=2.269, He=0.480, and I=0.919, respectively. The populations from Chaling showed a slightly lower level of genetic variation than that from Dongxiang. Within the Chaling site, the CL-Pi population had the highest level of effective alleles, the expected heterozygosity and Shannon's index, whereas the CL-Pr population exhibited the lowest level of the effective allele, the expected heterozygosity and Shannon's index. For the Dongxiang populations, DX-Pz had the highest level of the effective allele, the expected heterozygosity and Shannon's index, DX-Pa, and had the moderate and lowest level of effective allele number, the expected heterozygosity and Shannon's index, respectively. The DX-Pz and DX-Ps populations were also the highest and lowest in the level of genetic diversity among all the six populations of O. rufipogon (Table 1).
The relatively high mean Fixation indexes (FI) (Table 1) were found in all the six populations and all were positive, indicating significant departure from the H-W equilibrium and heterozygosity deficiency, although these values were different for each population. The outcrossing rate (t) was inferred from the value of FI. The overall outcrossing rate of the studied O. rufipogon populations was about 20%, and the Chaling populations had a relatively lower outcrossing rate compared with that of Dongxiang populations (19.6% vs 28.7%) (Table 1).
The F-statistics showed that genetic variability mainly existed among O. rufipogon individuals rather than among populations in Chaling (Fst=0.066), whereas relatively much genetic variation resided among Dongxiang populations (Fst=0.242), although genetic variability was also mainly presented among individuals (about 76% of total genetic variation); corresponding to the higher gene flow estimated in Chaling than that in Dongxiang (3.532 vs 0.782). About 27.6% of the total genetic variation of the six populations was scored between the two sites of Chaling and Dongxiang (Table 1).
Discussion
Genetic diversity of O. rufipogon and its conservation
Continued in situ conservation of wild rice species is increasingly important due to their tremendous agronomic values and the current critical deterioration of their habitat. One of the important strategies of in situ conservation is to preserve the genetic integrity and diversity of the conserved populations. A relatively high level of genetic variation was detected in six O. rufipogon populations in Jiangxi and Hunan Provinces of China using 23 rice SSR markers, compared with the results obtained from allozyme (Gao et al. 2000a, b, 2001) and RAPD (Ge et al. 1999; Song et al. 2001; Xie et al. 2001) studies of the same populations. This indicates that rice SSR markers have a good cross-amplified motif in its close relatives such as O. rufipogon, and that SSR assay provides a useful tool in estimating genetic variation of wild rice populations. The SSR analysis of the northern most O. rufipogon populations in China under different conditions demonstrates an overall genetic variation pattern that is in agreement with the assumption that most of the genetic diversity of O. rufipogon resided within its populations in the same regions of China (Wang et al.1996; Ge et al. 1999; Song et al. 2001). It is therefore valuable to continue the in situ conservation of O. rufipogon at the population level in China, particularly considering the extensive genetic erosion that has occurred in O. rufipogon populations during recent decades.
Genetic differentiation and spatial isolation
Genetic variation patterns of the Dongxiang and Chaling O. rufipogon populations are considerably different, with a much higher level of differentiation among populations in Dongxiang than those in Chaling (Fig. 2). This is in good accord with the level of estimated gene flow among populations at different sites. Among the Chaling populations, gene flow was scored nearly five-times as high as the Dongxiang populations. The limited gene flow may well explain the reason why considerable differentiation has occurred among the Dongxiang populations, which is in agreement with theoretical expectations (Slatkin 1987). On the contrary, a relatively high level of gene flow reduced genetic differentiation among the Chaling populations. In fact, the spatial distance among the Dongxiang populations is much longer (>1 km) than that among the Chaling populations (approximately 50–90 m). The Chaling populations were within the range of pollen flow, which we have experimentally shown to be approximately 110 m (Song et al., submitted to Biodiversity and Conservation). Based on pollen movement ability, we deduce that limited gene flow through cross-pollination should be responsible for the high level of differentiation among the Dongxiang populations and that the relatively high rate of gene flow led to low differentiation among the Chaling O. rufipogon populations.
Inter-population gene flow in this study was mainly influenced by spatial isolation, although gene flow should also be related to other factors such as mating systems. The outcrossing rate of the Dongxiang populations was higher than that of Chaling populations (mean 28.7% vs 19.6%), showing that the mating systems in Dongxiang populations were much open to gene flow, although they all represent a type of mixed-mating system (Oka and Morishima 1967; Gao et al. 2000a; Xie et al. 2001). The DX-Pa and DX-Ps populations were surrounded by a 2-m high wall, isolated from each other by a distance over 2 km, and the two populations were away from rice fields over 100 m, beyond the range of rice pollen flow. These facts could explain the limited gene flow observed between the two populations, and corresponding differentiation among the Dongxiang populations. These populations are still of special conservation concern as they may loose genetic diversity over time due to the synergistic effects of small population size and genetic drift. Therefore, the fate of the conserved Dongxiang populations inside the wall, especially the DX-Ps population is uncertain, because a relatively small population may encounter relatively stronger genetic drift and too little gene flow to countervail it (Slaktin 1987).
Genetic diversity of O. rufipogon from the Chaling region mainly exists among individuals rather than among populations. In general, the CL-Po population receives limited disturbances by human activities and represents a more pristine population of O. rufipogon at this site. The CL-Pr population is composed of individuals re-introduced from the ex situ conserved materials that were collected from the Chaling populations in 1983. The level of estimated genetic diversity of the CL-Pr population was nearly 90% of the genetic variability scored in the primitive CL-Po population, indicating that the re-introduction of ex situ conserved materials to the natural habitats where O. rufipogon populations are extinct or significantly reduced, is an effective alternative to restore populations of this wild species. In addition, the re-introduction enhanced the gene flow between populations, which promotes long-term maintenance of genetic diversity in O. rufipogon. Therefore, the conservation of the CL-Po and CL-Pr populations in Chaling is effective in comparison with other populations in Chaling or Dongxiang.
Introgression and genetic diversity
The populations DX-Pz and CL-Pi showed higher genetic diversity than the other four populations that are well isolated from rice fields. This can be explained by a higher level of introgression from cultivated rice to the two populations. The individuals of the DX-Pz population were scattered in a rice field, whereas the CL-Pi population grew near-by cultivated rice fields about 10 m away. They were in the range of maximum pollen flow (110 m) of cultivated rice. The overall outcrossing rate of O. rufipogon populations in this study is about 20%, which is nearly the same as the results reported in other studies (Morishima and Barbier 1990; Gao et al. 2000a). The considerable outcrossing rate allows O. rufipogon to receive the genes of cultivated rice varieties through cross-pollination, given that the two species grow sympatrically (Morishima et al. 1984) and high compatibility exists between O. rufipogon and O. sativa (Song et al. 2002). In addition, the two populations showed noticeable morphological variations having characters, such as white stigmas, short lemma awns and anthers, which are obviously from cultivated rice varieties. These confirmed the introgression from cultivated rice into the two populations, and consequently changed their genetic structure. The DX-Pz population had a significant higher level of genetic diversity than the CL-Pi population, suggesting the history of introgression and extent of mixed-growing with cultivated rice that may attribute to the differences. The DX-Pz is highly mixed-grown with cultivars and has a long history of gene exchange with cultivated rice, compared with the CL-Pi population, where farmers started to grow cultivated rice in 1995 and a distance of 10 m was observed from the rice field.
The results support the hypothesis that relatively high genetic diversity in Chinese O. rufipogon populations is probably due to a high level of introgression resulting from gene flow from Oryza sativa to O. rufipogon (Morishima et al. 1984; Wang et al. 1996; Ge et al. 1999; Gao et al. 2000a; Song et al. 2001). Therefore, gene exchange between cultivated crops and their wild relatives could increase detectable genetic diversity in the wild species, except for its opposite consequence, i.e. leading to the extinction of wild populations (Ellstrand et al. 1999). The above results are helpful for our decision-making with design in situ conservation strategies for O. rufipogon populations. Gene flow from cultivated rice (particularly when transgenic rice varieties are involved) should be carefully considered by having sufficient isolation distance to avoid gene flow from cultivated rice and to maintain genetic integrity of the conserved wild rice populations.
The lowest level of genetic diversity was observed in the DX-Ps population, which is most likely attributed to its small population size. Individuals in the DX-Ps population occurred densely within the area of about 60 m2. The samples from 23 individuals in this study possibly included almost all genotypes in this population. However, of the total number of 23 loci analyzed 11 were found to be monomorphic, indicating that most of the samples had the same genotypes. The occurrence of intra-clonal reproduction is probably due to the limited spatial extent (Xie et al. 2001) or the limited number of initiating genotypes at the onset of its in situ conservation. On the contrary, the least disturbed DX-Pa and CL-Po populations showed the highest levels of genetic diversity with only a small set of samples. This further confirms the above conclusion that population size may have a significant effect on the level of genetic diversity in O. rufipogon, provided that no introgression has taken place.
The reproductive pattern in O. rufipogon
It is noteworthy that all the studied O. rufipogon populations showed heterozygosity deficiency and their observed heterozygosity significantly deviated from the H-W equilibrium. This is primarily due to the mixed mating system of O. rufipogon. A mixed mating system with an outcrossing rate of 20% indicates that the non-random mating may result in a large number of self-pollinated seeds, i.e. inbreeding. This process would certainly increase the percentage of homozygosity. Secondly, clonal reproduction by ratoons or rhizomatic stems is an important complement to seed reproduction for the population development of O. rufipogon (Oka and Morishima 1967; Xie et al. 2001). The so-called "intra-clone outcrossing" or cross-pollination within the same clones (Morishima and Barbier 1990; Gao et al. 2000a) will have the same effect as self-pollination. In addition, a relatively narrow spatial range and a small population size would increase the inbreeding rate, and as a consequence resulted in heterozygosity deficiency.
In summary, our present genetic study reveals that: (1) considerable genetic diversity exists in the northernmost populations of O. rufipogon, although the diversity level varied among populations, which is most likely due to different conditions, under which the populations are grown and conserved; (2) the current population size limited by the concrete wall has led to considerable genetic drift of the populations, which does not guarantee their effective in situ conservation; (3) significant introgression from cultivated rice varieties grown in the vicinity of some O. rufipogon populations was detected, which might impose a destructive effect on the genetic integrity of these populations, therefore it is recommended to establish a safe isolation distance between in situ conservation sites of O. rufipogon and cultivated rice to avoid possible contamination of the wild species through gene flow; and (4) for implementation of in situ conservation; re-introduction of O. rufipogon individuals with reasonable genetic diversity is an effective approach to restore the populations that have already undergone extinction.
References
Akagi H, Yokozaki Y, Inagaki A, Fujimura T (1997) Highly polymorphic microsatellites of rice consist of AT repeats, and a classification of closely related cultivars with these microsatellite loci. Theor Appl Genet 94:61–67
Bellon MR, Brar DS, Lu BR, Pham JL (1998) Rice genetic resources. In: Dwoling NG, Greenfield SM, Fischer KS (eds) Sustainability of rice in the global food system. Chapter 16, Davis, California (USA). Pacific Basin Study Center and IRRI, Manila, pp 251–283
Byrne M, Marquez-Garcia MI, Uren T, Smith DS, Moran GF (1996) Conservation and genetic diversity of microsatellite loci in the genus Eucalyptus. Aus J Bot 44:331–341
Chase MR, Moller C, Kessell R, Bawa KS (1996) Distant gene flow in tropical trees. Nature 383:397–398
Cho YG, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch SR, Park WD, Ayres N, Cartinhour S (2000) Diversity of microsatellites derived from genomic libraries and GeneBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100:713–722
Ellstrand NC, Prentice HC, Hancock JF (1999) Gene flow and introgression from domesticated plants into their wild relatives. Annu Rev Ecol Syst 30:539–564
Gao LZ, Hong DY, Ge S (2000a) Allozyme variation and population genetic structure of common wild rice Oryza rufipogon Griff. in China. Theor Appl Genet 101:494–502
Gao LZ, Chen W, Jiang WZ, Ge S, Hong DY, Wang XQ (2000b) Genetic erosion in the Northern marginal population of the common wild rice Oryza rufipogon Griff., and its conservation, revealed by the change of population genetic structure. Hereditas 133:47–53
Gao LZ, Wei CL, Yang QJ (2001) Intra-population genetic structure of Oryza rufipogon Griff. in Yunnan, China. J Plant Res 114:107–113
Ge S, Oliveira GCX, Schaal BA, Gao LZ, Hong DY (1999) RAPD variation within and between natural populations of the wild rice Oryza rufipogon from China and Brazil. Heredity 82:638-644
Hong DY (1995) Rescue the wild rice resources (in Chinese). Bull Chinese Acad Sciences 4:325–326
Innan H, Terauchi, Miyashita NT (1997) Microsatellite polymorphism in natural populations of the wild plant Arabidopsis thaliana. Genetics 146:1441–1452
Morishima H, Barbier P (1990) Mating system and genetic structure of natural populations in wild rice, Oryza rufipogon. Plant Species Biol 5:31–39
Morishima H, Sano Y, Oka HI (1984) Differentiation of perennial and annual types due to habitat conditions in the wild rice Oryza perennis. Plant Syst Evol 144:119–135
McCouch SR, Chen X, Panaud O, Temnykh S, Xu Y, Cho YG, Huang N, Ishii T, Blair M (1997) Microsatellite marker development, mapping and applications in rice genetics and breeding. Plant Mol Biol 35:89–99
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Oka HI (1988) Origin of cultivated rice. Japanese Scientific Societies Press, Tokyo
Oka HI, Morishima H (1967) Variation in the breeding systems of a wild rice, Oryza perennis. Evolution 21:249–258
Roa AC, Chavarriaga-Aguirre P, Duque MC, Maya MM, Bonierbale MW, Iglesias C, Tohme J (2000) Cross-species amplification of Cassava (Manihot esculenta) (Euphorbicaceae) microsatellites: allelic poplymorphism and degree of relationship. Am J Bot 87:1647–1655
Rohlf FJ (1994) NTSYS-pc, numerical taxonomy and multivariate analysis system, ver. 1.80. Exeter Software, New York
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236:787–792
Song ZP, Lu BR, Zhu YG, Chen JK (2001) Changes of genetic structure in common wild rice (Oryza rufipogon) populations as referred to by RAPD markers. J Genet Mol Biol 12:78–84
Song ZP, Lu BR, Zhu YG, Chen JK (2002) Pollen competition between cultivated and wild rice species (Oryza sativa and O. rufipogon). New Phytol 153:289–296
Wang ZS, Zhu LH, Liu ZY, Wang XK (1996) Genetic diversity of natural wild rice populations detected by RFLP markers (in Chinese). Agric Biotechnol 4:111-117
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358-1370
Wu KS, Tanksley SD (1993) Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet 241:225–235
Xiao JH, Grandillo S, Ahn SN, McCouch SR, Tanksley SD, Li JM, Yuan LP (1996) Genes from wild rice improve yield. Nature 384:223–224.
Xie ZW, Ge S, Hong DY (1999) Preparation of DNA from silica gel a dried mini-amount of leaves of Oryza rufipogon for RAPD study and total DNA bank construction. Acta Bot Sinica 41:807–812
Xie ZW, Lu YQ, Ge S, Hong DY, Li FZ (2001) Clonality in wild rice (Oryza rufipogon, Poaceae) and its implications for conservation management. Am J Bot 88:1058–1064
Wright S (1978) Variability within and among natural populations. Vol. 4. The University Of Chicago Press, Chicago
Yeh FC, Yang RC, Boyle T (1999) Microsoft Window-based freeware for population genetic analysis (POPGENE), ver.1.31. ftp://ftp.microsoft.com/softlib/MSLFILES/HPGL.EXE
Yuan LP, Virmani SS, Mao CX (1989) Hybrid rice: achievements and further outlook. In: Progress in irrigated rice research. International Rice Research Institute, Manila, The Philippines, pp 219–223
Zhou J (1995) Studies on conservation biology of three Northern populations of common wild rice (Oryza rufipogon) (in Chinese). PhD dissertation, Wuhan University, Wuhan, China
Acknowledgements
We thank Mr. Guihua Liu and Ji Qian for their assistants in sample collection, and to Mr. M. Reagon of the Ohio State University for his comments on this manuscript. This research was supported by the National Natural Science Foundation of China for Distinguished Young Scholars (Grant No. 30125029), the 863 Program of Ministry of Science and Technology of China (Grant No. 2001AA227141), and Shanghai Commission of Science and Technology (Grant No. 02JC140022).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J.S. Heslop-Harrison
Rights and permissions
About this article
Cite this article
Song, Z.P., Xu, X., Wang, B. et al. Genetic diversity in the northernmost Oryza rufipogon populations estimated by SSR markers. Theor Appl Genet 107, 1492–1499 (2003). https://doi.org/10.1007/s00122-003-1380-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-003-1380-3