Abstract
Although hereditary hearing loss is highly heterogeneous, only a few loci have been implicated with low-frequency hearing loss. Mutations in one single gene, Wolfram syndrome 1 (WFS1), have been reported to account for most familial cases with this type of hearing impairment. This study was conducted to determine the cause of nonsyndromic low-frequency hereditary hearing impairment in two large families. Two large families from Switzerland and United States with low-frequency hearing loss were identified. Genomewide linkage analysis was performed followed by mutation screening in the candidate gene WFS1 with direct DNA sequencing and restriction fragment analysis. Both families were linked to DFNA6/14/38 with lod scores>3. Two novel heterozygous missense mutations in WFS1 were identified: c.2311G>C leading to p.D771H in the Swiss family and c.2576G>C leading to p.R859P in the US family. The sequence alteration was absent in 100 control chromosomes. Nonsyndromic low-frequency hereditary hearing impairment seems to be predominantly a monogenic disorder due to WFS1. We confirm that most mutations in WFS1 associated with isolated low-frequency hearing loss are clustered in the C-terminal protein domain coded by exon 8.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hearing among mammals ranges from lower frequency limit of 20 Hz to higher frequency limit of 150 kHz. The span is wide enough that some species have nonoverlapping hearing range; further, no single species hears over this entire spectrum. Human hearing ranges from 20 to 20,000 Hz, with the lower frequencies contributing to the sensation of loudness while the higher frequencies are critical for the clarity with which we hear. With ageing, hearing is stereotypically lost in the higher frequencies first (>2000 Hz), followed by slow encroachments into the lower frequencies with progression. Similarly, some dominant forms of nonsyndromic hearing impairment mimic presbycusis or age-related hearing loss; in contrast, recessive form of hearing impairment is usually profound and affects all frequencies at birth. Low-frequency hearing loss, genetic or acquired, is uncommon. Sensitivity to low-frequency sound is localized in the apex of the human cochlea. The factors responsible for the predilection of hearing loss to affect the basal turn of the cochlea that is exquisitely sensitive to higher frequencies or the sparing of the cochlear apex are poorly understood.
Hope for understanding molecular features related to low- and high-frequency hearing loss has been raised by the significant recent advances in genetics of nonsyndromic hereditary hearing impairment (HHI). Of the more than 90 loci for nonsyndromic HHI, only three are characterized by low-frequency hearing loss: DFNA1, DFNA6/14/38, and DFNA54 [1–6]. DFNA1 on 5q31 was the first autosomal dominant nonsyndromic HHI locus to be mapped using the large Monge family from Costa Rica. Subsequently, the causative mutation in diaphanous (DIAPH) was identified; to date, no additional mutations in DIAPH associated with low-frequency HHI have been identified [4]. In contrast, DFNA6/14/38, due to mutation in WFS1, is a common cause of low-frequency hearing loss [7]. Interestingly, while single mutation in WFS1 is associated with isolated low-frequency hearing impairment, recessively inherited double mutations in WFS1 are associated with Wolfram syndrome characterized by diabetes mellitus, diabetes insipidus, optic atrophy, and high-frequency hearing loss [8]. The protein product of WFS1, wolframin, is a membrane glycoprotein that is primarily localized in the endoplasmic reticulum, where it may serve as a calcium channel or regulator of calcium channel [9]; its function in the cochlea is unknown [10]. Functional analysis of mutant wolframin suggests that reduced protein dosage rather than dysfunction leads to the development of the syndrome [11]. Mutations in WFS1 provide a unique opportunity to understand the molecular mechanism underlying low- vs. high-frequency hearing loss as dominant alterations lead to former while the recessive cause the latter. In this study, we report on the genetic studies of two large families with nonsyndromic, low-frequency HHI in whom WFS1 mutations have been implicated.
Materials and methods
Families
Two multigenerational families with nonsyndromic autosomal dominant HHI were identified: a four-generation Swiss family with 31 members and a six-generation US family from Utah with 74 members. The pedigrees were established, and the auditory phenotype investigated with pure tone audiometry. The onset, severity, presence, or absence of progression and the type of hearing loss were noted. Syndromic and environmental causes were excluded by a questionnaire and examination. Blood samples were obtained from every member relevant for linkage analysis. Informed consent was obtained from all participants; the human study protocols were approved by the appropriate institutional review boards of University Hospital of Basel, Switzerland, and University of California–San Francisco, San Francisco, CA.
Genotyping
The ABI prism linkage mapping set MD-10 (Applied Biosystems, Foster City, CA) of fluorescent-labeled PCR markers was used to amplify highly polymorphic dinucleotide repeats for genotyping. PCR amplification reactions were performed following the manufacturer’s recommendation, pooled, and separated on an ABI 3700 DNA analyzer and sized with ABI GeneMapper software. Additional markers, used for fine mapping of the DFNA6/14/38 locus, were derived from the respective publications [2, 3] and synthesized by Applied Biosystems.
Linkage analysis
Two-point linkage analysis and multipoint analysis were performed using the linkage programs (linkage package: MLINK, CILINK) by Ott [12]. Low-frequency hearing loss was assumed to be inherited in a dominant manner and fully penetrant. The disease allele frequency was estimated at 10−4; changing the disease allele frequency to 10−3 only slightly modified the lod score value. The allele frequencies of the polymorphic markers were assumed to be equal. The meiotic recombination frequencies for males and females were taken from database provided by the Genetic Epidemiology Research Group (http://www.cedar.genetics.soton.ac.uk).
Sequencing of Wolfram syndrome gene
Genomic DNA was harvested from whole blood using the Qiagen DNA isolation kit (QIAamp DNA BloodMaxi Kit, Basel, Switzerland). Coding exons 2–7 were amplified using primers and PCR protocol previously described [13]. For the coding sequence in exon 8, three primer pairs were designed yielding overlapping products of around 700 base pairs. The primer pairs are as follows: 5′-AGCAGTGGGGGTCCTGTC-3′ and 5′-GCCATGCGGAAGAAGAGATA-3′, 5′-ACCTGGTCGTCCTCAACG-3′ and 5′-AGGTGGCTGCAGAGGATCT-3′, 5′-AGATCCTCTGCAGCCACCT-3′ and 5′-CACTGGTGCATGCCTGTC-3′. The first and last fragments were amplified using a touchdown program, decreasing Tm from 65 to 55°C. For PCR for the second fragment, Tm was 59°C. After PCR amplification, excess primers and dNTP were removed by Exosap-IT (Amersham Biosciences, Otelfingen, Switzerland) according to the manufacturer’s instructions. DNA sequencing was performed on a 3700 ABI DNA analyzer (Applied Biosystems) using manufacturer’s instructions. Doublepass sequencing was carried out using both the forward and reverse primers. Before loading the amplicons on the gel, they were purified by running them through a filtration block (Edgebiosystems, Gaithersburg, MD). Each fragment was sequenced twice following independent PCR amplification to detect and confirm sequence changes. The sequences were analyzed using the Sequencher 4.0 software (Gene Codes Corporation, Ann Arbor, MI).
Restriction enzyme analysis
Fragment three of the coding sequence of exon 8 was amplified as described above and digested with type II restriction enzyme Bsa0I (5′..CGRY^CG..3′) using the protocol supplied by the manufacturer and resolved on a 10% TBE polyacrylamide gel (Novex, Invitrogen, Carlsbad, CA, USA). The mutation in the Swiss family abolishes one of three restriction sites and therefore yields a different pattern than the wild type.
Results
Swiss family: linkage analysis
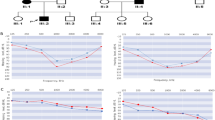
The family from Switzerland is comprised of 31 members and spans four generations (Fig. 1). There are 11 affected members based on the history of hearing loss and type of audiogram in the family with postlingual low-frequency hearing loss (Fig. 2). Additional high-frequency loss was only seen in one 73-year-old member (Fig. 3). The onset is between 5 and 20 years of age; the age of onset was established by audiologic testing in the young affected and by history in the elderly. In a single 34-year-old member, progression to moderate hearing loss was documented (Fig. 2); hearing loss was stable in other affected members. The severity of the hearing loss was moderate to profound. There was no history of psychiatric disorders in the family. A genomewide scan identified evidence for linkage on chromosome 4 with a two-point lod score of 4.8 for marker D4S2366. Recombinants were observed at telomeric marker D4S2375 and centromeric marker D4S2935 defining a 2 cM linkage region containing the WFS1 gene.
Swiss family: WFS1 mutation screening
Mutations screening of the WFS1 gene identified a nonconservative G>C missense mutation at position 2311, changing Asp to His 771 (Fig. 4). The p.D771H mutation was observed in heterozygous form in all affected individuals and was absent in the unaffected individuals by DNA sequence analysis. The mutation was additionally confirmed with restriction enzyme analysis as the sequence alteration leads to a loss of one of three Bsa0I sites. The mutation was absent in 100 control chromosomes.
US family: linkage analysis
Figure 5 shows the extensive pedigree of the Utah family. In this family, the hearing loss is similar to the Swiss family: a postlingual, stable, moderate low-frequency hearing loss (Fig. 6). In two older affected family members, an additional decrease in the high frequencies was noted. The onset is noted between 5 and 30 years. The family does not have a history of psychiatric disorders. A genomewide scan, using 24 members available for genotyping, identified evidence for linkage on chromosome 4 with a multipoint lod score of 4.05 between markers D4S412 and D4S2366.
US family: WFS1 mutation screening
As the linked region contained WFS1, mutation screening was performed. A nonconservative G to C missense mutation at position 2576, changing Arg to Pro was identified (Fig. 7). Sequence analysis of WFS1 in the affected and unaffected confirmed segregation of the p.R859P in all affected members and none of the unaffected members. The mutation was absent in 100 control chromosomes.
WFS polymorphisms
WFS1 is characterized as having numerous polymorphisms. In the two families, eight previously reported polymorphisms were detected with the majority being heterozygous conservative changes and located in exon 8 (Table 1).
Discussion
The auditory system remains mysterious and fascinating in the 21st century. While trailing the visual system and understanding of phototransduction, the molecular mechanisms of mechanotransduction in the auditory system are rapidly being deciphered. However, the predilection for high-frequency hearing loss and the relative hardiness of low-frequency hearing remains poorly understood. Evolutionarily, low-frequency hearing is “older” than high frequency as the former was present in the vertebrates while the latter is only present in mammals [14]. There are likely structural and molecular determinants of frequency gradient within the cochlea. Structural differences in the cochlea from the apex to the base are known to be important contributors to the tonotopic distribution in the cochlea. The basilar membrane that partitions the scala media from the scala tympani provides the scaffold upon which rests the organ of Corti is wider and less stiff in the apex when compared to the base. Further, the hair cells decrease in height from apex to base and likewise the stereocilia decrease in length and numbers [15].
The identification of WFS1 as responsible for nonsyndromic autsomal dominant low-frequency HHI is a significant step in understanding the molecular determinants of frequency distribution in the cochlea. Located across a 33.4 kb of genomic DNA on chromosome 4p16, the gene contains 8 exons and has an mRNA of 3.6 kb coding for an 890 amino acid protein with a predicted molecular mass of 100 kDa. The gene encodes for the protein wolframin that is estimated to have nine helical transmembrane segments. Located primarily in the endoplasmic reticulum, it is believed to be an integral, endoglycosidase H-sensitive membrane glycoprotein [8, 10]. As recessive inactivating mutations cause high-frequency hearing loss as part of Wolfram syndrome and dominant noninactivating missense mutations cause low-frequency hearing loss, the duality of WFS1 in frequency-specific hearing is intriguing and provides a unique genetic tool for dissecting tonotopicity in cochlea [7, 16]. Its putative function in cochlear ion homeostasis may play an important role in determining frequency selectivity.
In this genetic study of two large families with nonsyndromic low-frequency HHI, we provide additional evidence that WFS1 is an important contributor to familial low-frequency hearing loss. Following linkage to WFS1-containing region on 4p16, mutation analysis identified two novel mutations in C-terminal protein domain coded for in exon 8. In the Swiss family, a missense mutation at c.2311G>C results in an amino acid substitution, p.D771H, in a highly conserved area across human, mouse, and rat. Previously, in a single Greek patient with schizophrenia (nonfamilial), Torres et al. [17] reported a missense mutation at c.2312A>G that resulted in substitution of glycine for aspartic acid (p.D771G). The hearing status of his patient was not reported. Our family does not have a history of psychiatric disorders. The variable phenotype associated with spectrum of WFS1 mutations is remarkable but remains poorly understood.
In the Utah family, c.2576G>C missense mutation occurs in an equally conserved region of WFS1 among species. The latter mutation substitutes proline for naturally occurring arginine which is known to disrupt helical chains. It is likely that the mutation significantly alters the secondary structure of wolframin by creating a bending point in the amino acid chain. Both mutations were confirmed in all affected members and were absent in the unaffected. Further, they were not found in 100 control chromosomes. To our knowledge, these mutations have neither been previously described as such nor as polymorphism. The mutations in the WFS1 gene-causing hearing loss share common findings. They are all heterozygous and are all located in exon 8, in the C-terminal domain [5, 7, 16, 18, 19]. In contrast, Hardy et al. [20] found in an analysis of 19 Wolfram syndrome kindred that all patients except two had mutations on both alleles of their WFS1 gene; majority of these mutations were located in the transmembrane domains.
Polymorphisms are common in WFS1 gene. Over 70 such sequence variations have been reported in the gene; while most changes are synonymous, others are nonsynonymous. The exact contribution of the polymorphism to health, to disease, to documented variable expression is unknown. We identified eight different polymporphisms in our two families; of these, six have already been reported [20]. Two nonsynonymous changes were novel and identified in homozygous state in the unaffected members of the family: c.632A>G (p.D211G), c.713C>G (p.S238K). The first polymorphism affects an amino acid that has been previously associated with Wolfram syndrome in the heterozygous state; the responsible mutation was c.631G>A. van den Ouweland et al. [21] reported p.D211N/p.P607R occurring in two individuals of a family with Wolfram syndrome characterized by diabetes mellitus, optic atrophy, deafness in one individual while hypothyroidism and hypertension in the other. The p.S238K has not been previously reported but was not associated with disease in our family. In absence of population studies or functional analysis, the significance of these mutations remains unknown.
At first glance, the phenotype seems to be similar, but there are a few differences among the reported families with WFS-related low-frequency hearing loss. In the original DFNA6 family reported by The Vanderbilt University Hereditary Deafness Study Group, the hearing loss was limited to low frequency, stable, with onset between 5 and 15 years of age [7]. The onset in the Swiss family is before 20 and in the Utah family between 5 and 30 years. The differences in age of onset might be due to delay of diagnosis as low-frequency hearing loss can be asymptomatic. The severity of hearing loss is moderate to profound in the two families. The difference in phenotype across families may reflect influences of a modifier gene similar to DFNB1 and the mitochondrial defect 12SrRNA. Although no such gene has yet been identified, possible loci have been found in mice and man [22–24].
Of note, there are differences in the hearing loss associated with the other low-frequency hearing loss loci DFNA1 and DFNA54. DFNA1, due to mutations in diaphanous, progresses from low-frequency hearing loss to involve all frequencies. The involvement of the high frequencies in two US and one Swiss family members over the age of 60 years is likely due to presbycusis rather than progressive involvement due to WFS1 [25, 26]. Differences in phenotype between the two reported families and DFNA54 also exist [6]. The range of onset in DFNA54 was slightly larger, between 5 and 40 years. Progression from mild to moderate and even profound hearing loss was seen in the majority of affected members, whereas in the Swiss and Utah families, hearing loss was stable. The vestibular symptoms reported in two affected members in DFNA54 were absent in these two families. Demonstration of linkage to DFNA6/14/38 and identification of novel mutations in WFS1 in these two large families lends additional support for the majority of familial autosomal dominant low-frequency hearing loss being due to mutations in WFS1, while the contribution of DFNA1 and DFNA54 to this type of hearing loss remains unknown.
References
Van Camp G, Smith R Hereditary hearing loss homepage. http://dnalab-www.uia.ac.be/dnalab/hhh/
Lesperance MM, Hall JW III, Bess FH et al (1995) A gene for autosomal dominant nonsyndromic hereditary hearing impairment maps to 4p16.3. Hum Mol Genet 4:1967–1972
Van Camp G, Kunst H, Flothmann K et al (1999) A gene for autosomal dominant hearing impairment (DFNA14) maps to a region on chromosome 4p16.3 that does not overlap the DFNA6 locus. J Med Genet 36:532–536
Lynch ED, Lee MK, Morrow JE, Welcsh PL, Leon PE, King MC (1997) Nonsyndromic deafness DFNA1 associated with mutation of a human homolog of the Drosophila gene diaphanous. Science 278:1315–1318
Young TL, Ives E, Lynch E et al (2001) Non-syndromic progressive hearing loss DFNA38 is caused by heterozygous missense mutation in the Wolfram syndrome gene WFS1. Hum Mol Genet 10:2509–2514
Gürtler Nea, Kim Y, Mhatre A, Schlegel C, Mathis A, Lalwani AK (2004) DFNA54, a third locus for low-frequency hearing loss. J Mol Med 82:775–780
Bespalova IN, Van Camp G, Bom SJH et al (2001) Mutations in the Wolfram syndrome 1 gene (WFS1) are a common cause of low frequency sensorineural hearing loss. Hum Mol Genet 10:2501–2508
Inoue H, Tanizawa Y, Wasson J et al (1998) A gene encoding a transmembrane protein is mutated in patients with diabetes mellitus and optic atrophy (Wolfram syndrome). Nat Genet 20:143–148
Osman AA, Saito M, Makepeace C, Permutt MA, Schlesinger P, Mueckler M (2003) Wolframin expression induces novel ion channel activity in endoplasmic reticulum membranes and increases intracellular calcium. J Biol Chem 278:52755–52762
Takeda K, Inoue H, Tanizawa Y et al (2001) WFS1 (Wolfram syndrome 1) gene product: predominant subcellular localization to endoplasmic reticulum in cultured cells and neuronal expression in rat brain. Hum Mol Genet 10:477–484
Hofmann S, Philbrook C, Gerbitz KD, Bauer MF (2003) Wolfram syndrome: structural and functional analyses of mutant and wild-type wolframin, the WFS1 gene product. Hum Mol Genet 12:2003–2012
Ott J. Genetic linkage programs. ftp://linkage.rockefeller.edu/software
Strom TM, Hortnagel K, Hofmann S et al (1998) Diabetes insipidus, diabetes mellitus, optic atrophy and deafness (DIDMOAD) caused by mutations in a novel gene (wolframin) coding for a predicted transmembrane protein. Hum Mol Genet 7:2021–2028
Heffner RS, Koay G, Heffner HE (2001) Audiograms of five species of rodents: implications for the evolution of hearing and the perception of pitch. Hear Res 157:138–152
Raphael Y, Altschuler RA (2003) Structure and innervation of the cochlea. Brain Res Bull 60:397–422
Cryns K, Pfister M, Pennings RJ et al (2002) Mutations in the WFS1 gene that cause low-frequency sensorineural hearing loss are small non-inactivating mutations. Hum Genet 110:389–394
Torres R, Leroy E, Hu X et al (2001) Mutation screening of the Wolfram syndrome gene in psychiatric patients. Mol Psychiatry 6:39–43
Kunz J, Marquez-Klaka B, Uebe S, Volz-Peters A, Berger R, Rausch P (2003) Identification of a novel mutation in WFS1 in a family affected by low-frequency hearing impairment. Mutat Res 525:121–124
Komatsu K, Nakamura N, Ghadami M et al (2002) Confirmation of genetic homogeneity of nonsyndromic low-frequency sensorineural hearing loss by linkage analysis and a DFNA6/14 mutation in a Japanese family. J Hum Genet 47:395–399
Hardy C, Khanim F, Torres R et al (1999) Clinical and molecular genetic analysis of 19 Wolfram syndrome kindreds demonstrating a wide spectrum of mutations in WFS1. Am J Hum Genet 65:1279–1290
van den Ouweland JM, Cryns K, Pennings RJ et al (2003) Molecular characterization of WFS1 in patients with Wolfram syndrome. J Mol Diagn 5:88–95
Bykhovskaya Y, Estivill X, Taylor K et al (2000) Candidate locus for a nuclear modifier gene for maternally inherited deafness. Am J Hum Genet 66:1905–1910
Ikeda A, Zheng QY, Rosenstiel P et al (1999) Genetic modification of hearing in tubby mice: evidence for the existence of a major gene (moth1) which protects tubby mice from hearing loss. Hum Mol Genet 8:1761–1767
Riazuddin S, Castelein CM, Ahmed ZM et al (2000) Dominant modifier DFNM1 suppresses recessive deafness DFNB26. Nat Genet 26:431–434
Lesperance MM, Hall JW III, San Agustin TB, Leal SM (2003) Mutations in the Wolfram syndrome type 1 gene (WFS1) define a clinical entity of dominant low-frequency sensorineural hearing loss. Arch Otolaryngol Head Neck Surg 129:411–420
Pennings RJ, Bom SJ, Cryns K et al (2003) Progression of low-frequency sensorineural hearing loss (DFNA6/14-WFS1). Arch Otolaryngol Head Neck Surg 129:421–426
Acknowledgements
We thank participating family members for their cooperation and the Molecular Genetics Facility at the University Hospital of Basel for technical support. This study was supported in part by grants from the Swiss National Science Foundation, the Novartis Foundation, and “Freiwillige Akademische Gesellschaft”, the latter two from Basel, Switzerland.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gürtler, N., Kim, Y., Mhatre, A. et al. Two families with nonsyndromic low-frequency hearing loss harbor novel mutations in Wolfram syndrome gene 1. J Mol Med 83, 553–560 (2005). https://doi.org/10.1007/s00109-005-0665-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-005-0665-1