Abstract
The colonization of the neonatal digestive tract provides a microbial stimulus required for an adequate maturation towards the physiological homeostasis of the host. This colonization, which is affected by several factors, begins with facultative anaerobes and continues with anaerobic genera. Accumulating evidence underlines the key role of the early neonatal period for this microbiota-induced maturation, being a key determinant factor for later health. Therefore, understanding the factors that determine the establishment of the microbiota in the infant is of critical importance. Exposure to antibiotics, either prenatally or postnatally, is common in early life mainly due to the use of intrapartum prophylaxis or to the administration of antibiotics in C-section deliveries. However, we are still far from understanding the impact of early antibiotics and their long-term effects. Increased risk of non-communicable diseases, such as allergies or obesity, has been observed in individuals exposed to antibiotics during early infancy. Moreover, the impact of antibiotics on the establishment of the infant gut resistome, and on the role of the microbiota as a reservoir of resistance genes, should be evaluated in the context of the problems associated with the increasing number of antibiotic resistant pathogenic strains. In this article, we review and discuss the above-mentioned issues with the aim of encouraging debate on the actions needed for understanding the impact of early life antibiotics upon human microbiota and health and for developing strategies aimed at minimizing this impact.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gut microbiota is the wide, complex and diverse collection of microorganisms living in the intestine, including bacteria, archaea, viruses and unicellular eukaryotes. This microbiota has adapted to coexist in a commensal relationship with the host and contributes to the programming of gut development and intestinal barrier function, as well as to the metabolic and immune homeostasis, being a key determinant for health [1, 2]. For this reason, the study of the intestinal microbiota has received growing attention during the last years.
The recent advances in this area have been largely a consequence of the development of methods allowing the comprehensive study of complex microbial ecosystems. Until the end of the past century, the intestinal microbial composition had been mainly studied using culture-based methods, but the development of culture-independent techniques and, more recently, the improvement of the massive DNA sequencing platforms have made these tools the current standard in microbiota research [3, 4]. However, despite the efforts for characterizing the human intestinal microbiota composition, the large inter-individual variability makes the interpretation of the results difficult and underlines the need to determine not just the composition but also the microbiota functionality in the interaction with the host. In this sense, the use of germ-free (GF) and gnotobiotic animal models provide an invaluable experimental tool to understand the role of the microbiota establishment on the host health and development. GF animals show anatomical and physiological features that differ from those of specific pathogen-free (SPF) and wild-type animal counterparts [5], underlining the importance of the microbiota for the host.
Accumulating evidence indicates that the early life cross-talk between the microbiota and the host is essential for establishing and maintaining the well-being of the individual. This early microbiota seems to provide a stimulus required for an adequate development of the gut and immune functions [2, 6,7,8,9,10], with effects on distal organs and also at systemic level [11,12,13,14]. Therefore, any disturbance of the neonatal microbiota development process may have important implications for infant health and for the risk of disease later in life.
Establishment of the intestinal microbiota in early life
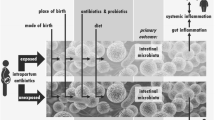
Traditionally the fetus has been considered sterile but recent studies indicate in utero microbial exposure prior to delivery. The presence of low levels of bacteria in placenta, amniotic fluid, umbilical cord blood or meconium has been demonstrated [15,16,17]. This prenatal microbial exposure greatly increases after birth when the newborn is rapidly and densely colonized with a complex myriad of microbes. The microbial colonization process depends on an interplay among different factors including gestational age [18,19,20], mode of delivery [20,21,22,23], use of perinatal antibiotics [19, 24,25,26] or feeding habits [23, 27]. All these factors contribute to the establishment and later development of the microbiota in the infant [28].
During the initial colonization stages, the microbiota is unstable and undergoes microbial succession phenomena. The classical pattern initiates with facultative anaerobes, such as enterobacteria, which reduce the intestinal environment creating suitable conditions for the subsequent proliferation of strict anaerobes such as Bifidobacterium [29]. Then, other anaerobic populations including Bacteroides and different clostridia start to increase, especially after weaning. During the first year of life the infant microbiota maturates at both compositional and functional level, with major changes occurring as a consequence of breastfeeding cessation [23]. During this period, the microbiota of the individual increases in α-diversity whilst the β-diversity is decreased, indicating that the microbiota becomes more complex and homogeneous among individuals along time [23].This progression continues until the age of 2–3 years when the microbiota resembles that of adults, with some bacterial groups already reaching the adult state stability [27]; other bacterial groups, however, may require longer time to reach the adult steady-state [30] with some differences still being present at the preadolescent age [31].
Early life microbiota–host interaction: the window of opportunity
The timing of colonization seems to be very important as underlined by different studies demonstrating that restoration of the microbiota of GF animals in early life, but not during adulthood, is able to normalize some of the functions that are altered in these GF models [6, 8, 32]. Experimental models demonstrate that reduced exposure to microbes early in life is associated with the increased prevalence of diseases such as allergy, IBD or diabetes [33, 34]. Moreover, antibiotic-induced alteration of the early microbiota leads to an increased susceptibility to allergic diseases [35, 36]. Using experimental animals, it has been demonstrated that antibiotic-induced disturbance of the early life microbiota increased the risk of undesirable long-term effects, whereas disturbance during adult life does not [9, 37, 38]. The type of antibiotic administered seems to be relevant since depending on the spectrum of action, different responses can be obtained. This suggests the importance of specific bacterial populations in terms of induction of the host homeostasis [35, 36, 39].
All these studies reveal that the early microbial exposure is important for the development of the host and point to the existence of a “window of opportunity” for the host microbial programming in the neo(peri)natal period. Therefore, any perinatal intervention that modifies the establishment and development of the microbiota may have an influence on later health. However, the exact time of the “window of opportunity” for the establishment of a healthy microbiota is not precisely known, still constituting an area of active and intense research. The development of microbiome-modulating strategies tailored to correct the potential dysbiosis present in early life with the aim of favoring an adequate microbiota development, or to avoid its disturbance by the administration of medication, would represent a good option as a preventive therapy. Among the perinatal factors potentially affecting the microbiota development in the newborn, the use of antibiotics represents the most common threat.
Impact of antibiotics upon the establishing microbiota
Use of antibiotics during pregnancy
Pregnancy is characterized by simultaneous endocrine, metabolic and immune changes aimed at supporting the correct growth and development of the fetus [40]. During pregnancy the maternal microbiota suffers drastic changes [41], with those occurring on the vaginal and intestinal microbiota being of particular relevance [42]. Maternal antibiotic use during pregnancy is known to affect the vaginal microbiota [43], which could hamper the later transfer of microbes to the baby during delivery. Moreover, recent data suggest that during gestation the maternal microbiota plays an important role in programing the future immune system of the offspring [44]. It has been shown that pregnancy complications significantly affect the microbial colonization patterns of the infant gut, with changes persisting during the first year of life [45]. Similarly, antibiotics administered during pregnancy affect the microbial environment of the mother and may impact the child already in utero, disrupting the correct establishment of the nascent microbiota and hence the microbial gut colonization. Studies in animal models showed a reduced microbiota diversity and/or population structure changes in the gut microbiota of offspring from mothers treated with antibiotics during pregnancy [46,47,48,49]. More specifically, the population of lactobacilli seems to be reduced, whereas other populations such as Proteobacteria, Firmicutes or the Clostridium cluster XIVa have been reported to be affected as well [48,49,50].
Use of antibiotics during delivery
Intrapartum antimicrobial prophylaxis for prevention of early onset group B Streptococcus infection
During the 70’s decade group B streptococci (GBS) were the main infectious agents causing morbidity and mortality in the neonatal period in developed countries [51, 52]. At that time the mortality rates in newborns suffering GBS infection reached 50%, a figure that has been now significantly reduced as a result of the advances in neonatal care. However, 25–30% of the affected newborns will still suffer neurologic consequences [53]. During the 1980’s several studies demonstrated that the administration of intravenous antibiotics to the mother during birth was able to prevent neonatal sepsis by GBS. This prompted the American College of Obstetricians and Gynecologists and the Center for Disease Control (CDC, USA) to publish in 1996 the first recommendations for intrapartum antibiotic prophylaxis (IAP) which were recognized by the American Academy of Pediatrics a year later [51]. Prior to the introduction of the IAP the incidence of early neonatal GBS infection was of 2–3 cases per 1000 newborns in the USA and between 0.2 and 4 cases per 1000 in Europe. From the mid-nineties, coinciding with the introduction of the IAP, the incidence of GBS infection dropped to 0.3–0.4 cases per 1000 newborns in most developed countries [52, 53].
The main risk factor for GBS early neonatal infection is the maternal, vaginal or rectal, colonization by GBS, which is present in between 10 and 30% of women [51]. Women colonized by GBS show 25-times higher risk of having a child with early systemic GBS infection than non-colonized mothers [51]. The risk of early onset GBS infection is also increased in preterm neonates, especially in those with low-birth weight (< 2500 g) [54]. In the absence of any preventive measure 1–2% of the newborns colonized during birth will develop infection in the first 7 days of life. There are two approaches for selecting candidates to receive IAP: (1) based on the presence of risk factors (prematurity, prolonged rupture of membranes, intrapartum fever) or (2) based on GBS colonization demonstrated by antepartum screening [51, 52]. Some retrospective studies have compared both strategies and concluded that IAP administration to GBS carriers was twice as effective as the strategy based on risk factors. These prompted the CDC to recommend universal screening between weeks 35 and 37 of pregnancy to optimize the identification of women who should receive IAP [51]. This strategy has been implemented in most European countries, although other countries decided to use the risk-factors based strategy.
Under the strategy of universal screening for GBS, IAP is indicated in all women with a vaginal or rectal positive culture for GBS, those in which GBS is detected in urine during pregnancy, all pregnant women who have had a child with neonatal GBS infection, and in all deliveries for which culture results are not available results and at least one of the following risk factors is present: less than 37 weeks of gestation, membranes ruptured for more than 18 h or intrapartum fever [53]. The antibiotic of choice is intravenous penicillin or, alternatively, vancomycin for women allergic to β-lactam antibiotics.
When the risk factors based protocol is followed, IAP is given to women who have had a previous child with invasive disease by GBS, GBS bacteriuria during pregnancy, preterm labor, ruptured membranes for over 18 h or intrapartum fever, although in some countries preterm labor and premature rupture of membranes have been excluded from the list [53]. The use of the “risk factor” strategy reflects the belief that, given the current low incidence of perinatal GBS infection, the introduction of universal screening would not affect the infection rates but it would increase maternal and fetal exposure to antibiotics, with the subsequent adverse effects [53, 55].
In developed countries IAP is used in over 30% of total deliveries [56] representing the main cause of antibiotic exposure during the perinatal period. In spite of this extended use and although the available studies suggest a reduction of early onset GBS infection, some authors have indicated that these studies present a high risk of methodological bias and that further confirmation on the effectiveness of IAP would be needed [57]. Moreover, the potential adverse effects of antibiotic treatment, and the interest in reducing antibiotics use globally, have raised the attention to alternative approaches to. Several studies have shown that early postnatal antibiotic exposure disturbs the natural establishment of the intestinal microbiota in the newborn with potential negative influence in later health [58, 59]. Similarly, recent studies have reported that IAP affects the later microbiota development in the newborn [19, 60, 61]. Epidemiological evidence points out to a reduced vertical transmission of vaginal lactobacilli from mothers to babies following IAP [60, 62]. Some of the microbial groups that have been reported to be affected include the genera Bacteroides and Clostridium, as well as the family Enterobacteriaceae [19, 60]. These results underline the interest in a rational use of IAP, restricting it to those situations where a beneficial effect has been demonstrated, and point to the need for strategies to minimize the impact of IAP upon the establishing microbiota. This consideration is especially relevant since some studies have suggested incomplete recovery of the gut microbiota after antibiotics administration to infants [63].
Antibiotics in cesarean section delivery
In developed countries another important cause of antibiotic exposure during or immediately after delivery is cesarean section (CS), which occurs in about 20% of deliveries although with a large variability among countries. The WHO indicates that, at the population level, CS rates higher than 10% of total deliveries are not associated with significant reductions in maternal and newborn mortality [64], suggesting that in many cases CS may be unnecessary. Women undergoing CS have a 5- to 20-fold greater risk of infection as compared to those undergoing vaginal delivery, and consequently antibiotic prophylaxis is extensively applied in CS [65]. Nevertheless, some open questions remain about the potential adverse effects of these antibiotics for the woman and the infant. Babies born by C-section display an altered gut microbiota establishment process as compared with babies born by vaginal delivery [21, 22]. The keystone paper by Backhed et al. [23] demonstrates that mother-to-infant transfer of the microbiota is hampered in CS delivered babies, including that of important intestinal anaerobes such as Bifidobacterium or Bacteroides. Bokulich and coworkers [58] demonstrated that the mode of delivery had a stronger effect than repeated infant antibiotic treatment in the establishment of the gut microbiota. However, given their extensive use in the medical practice, the exposure to antibiotics in CS deliveries can be an important confounding factor in the study of the gut microbiota establishment, being difficult to isolate the CS-effects from those attributable to IAP or to postpartum antibiotics. To this regard, several reports have indicated reduced levels of microorganisms such as Bacteroides or Bifidobacterium, and increased levels of certain Firmicutes or Proteobacteria, in CS delivered babies [20, 22, 23, 66] and similar changes on the microbiota composition have been observed as a consequence of perinatal antibiotics [19, 24].
The large evidence substantiating the alteration of the neonatal microbiota in CS delivered babies has prompted researchers to propose the deliberated inoculation of these infants with their mothers’ vaginal microbiota as a way to promote microbiota development [67].
Use of antibiotics during the postnatal period
Antibiotics are the early life medication most frequently administered in neonatal intensive care units (NICUs) [68], with premature babies being the group receiving more empiric antibiotic treatments during the first days of life [69]. Factors like premature rupture of membranes or fears about occult intrauterine infection, that could precipitate spontaneous premature labor or chorioamnionitis, prompt the initiation of empiric antibiotic administration in preterm neonates, even though in practice the incidence of early onset sepsis is low [70, 71]. Premature babies are known to harbor an altered gut microbiota, with higher number of potential pathogens, lower levels of beneficial microorganisms and reduced bacterial diversity than healthy full-term infants [18, 19]. There is not currently enough information to discriminate which part of these alterations is attributable to prematurity itself and which one to concomitant factors such as the antibiotic treatment. However, different studies have demonstrated that exposure to antibiotics during microbiome ontogeny may precede long-term disruptions. Moreover, early empiric antibiotic use has been associated with an increased risk of necrotizing enterocolitis (NEC), sepsis and death in premature babies [72].
The extensive exposure to antibiotics is not restricted to preterm babies and the hospital setting; overall more than half of the infants have been treated with antibiotics during the first months of age [73]. One of the effects of these antibiotic treatments is the perturbation of the correct development of the infant intestinal microbiota and, as a result, the disruption of a proper development of the gut, immune, metabolic and brain systems [9, 36, 38]. Moreover, after the initial antibiotic-induced perturbation the later full recovery of a health-associated microbiota cannot be ensured. In this regard, it has been shown that the caecal microbiota of mice treated with cefoperazone continues to be different from the control group even 6 weeks after finishing the treatment [9, 74]. Different human studies have reported reductions on Bifidobacterium levels and increases in enterobacteria in babies who received antibiotic treatments [19, 24, 75]. Observational studies with babies also revealed incomplete recovery of the gut microbiota some months after cessation of the antibiotic treatment [24, 75].
Breastfeeding constitutes another potential vehicle for antibiotic-mediated effects on the infant microbiota [76]. Antibiotic consumption by the mother may affect the breastfed infant microbiota by two different ways; by altering the microbiota of the milk the infant is receiving and/or by transferring the antibiotic to the infant gut. In this regard, it is well known that maternal antibiotics alter the breast milk microbiota [77] and that some antibiotics may cross the placenta [78].
In spite of the effects stated above, it is important to understand that, in many circumstances, treatment with antibiotics is the most prudent medical option. Mortality due to early life infection has been drastically reduced during the last decades, in good part due to antibiotic treatments. However, given our increasing understanding on the importance of the early life microbiota, it could be wise to avoid the administration of antibiotics when there is not a clear indication of use, to optimize the spectrum of the antibiotics used, the duration of the treatment and the route of administration, as well as to develop strategies for limiting their impact on the microbiota.
Effect of early life antibiotics in disease-risk
As indicated previously the early microbiota establishment process and its further development is determined by several factors. The use of antibiotics influences microbiota–host crosstalk during the critical neonatal period, therefore, being one of the factors that may have profound consequences for later health [25, 79].
In relation with the hygiene hypothesis, different epidemiological studies have reported a link between early antibiotics exposure and allergic disease later in life [80, 81]. It has been repeatedly shown that even prenatal antibiotics exposition may be associated with the occurrence of asthma during childhood [82, 83]. Moreover, not all antibiotics show the same effects; maternal use of penicillin or chloramphenicol, but not of others, shows an association with later infant asthma [83]. However, the presence of potential confounding factors, such as underlying infections, point out to the need for caution when drawing firm conclusions from these epidemiological studies [84].
In recent years, different animal studies, mainly comparing germ-free versus colonized animals, have demonstrated mechanisms linking the early life microbiota with the later development of allergic diseases [8, 85]. The use of germ-free models has provided valuable mechanistic information. However, caution should be taken when extrapolating these data since these animals totally lack the normal microbial antigenic stimulation, whilst in conventional animals, despite the potential differences in microbiota composition, the main antigenic stimuli are still present in every individual. In this context the use of conventional animal models of neonatal exposure to antibiotics provides an alternative tool more closely resembling the normal situation. Using such models Russell and coworkers [35] found that antibiotic-induced changes in the early life microbiota, but not at a later age, increase the susceptibility to asthma. Similar studies have also shown that early life antibiotic treatment hampers the proper development of immune responses [36, 86].
Obesity has also been related to early life antibiotics administration [87]. Animal studies have demonstrated that antibiotic-induced alterations in the early life microbiota, despite the later microbiota restoration, may have long-lasting metabolic consequences with an increase in body fat and weight gain [9, 88]. Epidemiological studies, have also reported an association between exposure to antibiotics during the first months of life and increased body mass during later childhood [89,90,91]. Although the studies focusing on specific antibiotics are still scarce, the available data suggest that broad-spectrum antibiotics have a larger impact [90]. Some data are also available on the effect of prenatal antibiotics exposure as related to childhood obesity [42], indicating an increased risk of obesity in children from mothers that received antibiotics during the second or third trimester. Some evidence on a higher risk for diabetes after early life antibiotic-induced microbiota alteration is also available from animal studies [38, 92].
It is important to underline that, in developed countries, by the end of the first year of life more than half of the infants have been exposed to antibiotics. This is despite the current efforts and measures to restrict antibiotic use, and therefore, this antibiotic exposure has likely been even higher in previous decades when such measures were not in place. Thus, it is reasonable to think that in the 1970’s–1980’s in most developed countries almost all infants were exposed to antibiotics during early life. This suggests that during these years the early life microbiota of almost a whole generation of individuals may have been affected. This opens questions of enormous importance on the true role of early life antibiotic-induced microbiota alterations on the currently observed increased incidence of metabolic and autoimmune disorders.
Antibiotics and the human gut resistome: a pending task
Surprisingly almost no data is available on the impact of early life antibiotics in the harboring of antibiotic resistance genes by the gut microbiota [93]. This may constitute an important issue in view of the increasing problems with antimicrobial resistances. The human gut resistome is defined as the collection of all genes from the gut microbiome that potentially encode for resistance to antibiotics [94, 95]. The enrichment of the reservoir of antibiotic resistance genes (ARG) may increase the risk of transfer towards potential pathogens, compromising the clinical management of infections. Furthermore, it is known from in vitro studies that when microbial communities are exposed to constant antibiotics challenge, they acquire multidrug resistance [96]. Although the way to acquire antibiotic resistance in vivo is less understood, it has been shown that antibiotic therapy helps to select resistant members of the microbial community or those microorganisms capable of acquiring ARG [97]. It is important to consider that ARG may encompass alternative functions in the cell and are not simply for avoiding the effect of antibiotics [95,96,97,, 98]. The recent advance and the cost affordability of DNA sequencing techniques, enabling the exploration of the human microbiome and its resistome, have raised the interest in deciphering the establishment and evolution of the human resistome at the beginning of life and the impact on health later on.
Some available data suggest that the initial acquisition of antibiotic resistance is independent of infant’s exposure and is more likely impacted by maternal and environmental microbes during and after delivery [99,100,101]. The gut resistome begins to develop in the intrauterine environment or at birth even in the absence of selective pressure [102, 103], and the transmission of ARG from mother to infant seems to start before delivery [99, 104]. It is generally accepted that early exposure to some external factors could also contribute to driven the development of the infant evolving resistome. Among these factors are the family environment, hospital and neonatal units, early treatment of infants with antibiotics, intrapartum antibiotic administration, and antibiotic treatment to mothers (before and/or after delivery). However, studies assessing which factors, and to which extent, affect the establishment and prevalence of the infant resistome, are still very scarce. In this way, a recent longitudinal study revealed that chromosomally located ARG increased following antibiotic exposure but decreased afterwards, whereas ARG associated to mobile genetic elements remain high [105].
The prevalence of resistance genes fluctuates over the course of the first few months of life, probably reflecting changes in the establishing intestinal microbial structure and, hence, in the bacterial populations harboring such genes [106]. The most prevalent ARG found in the neonatal microbiota are those encoding resistance to erythromycin, tetracycline, aminoglycosides and beta-lactams [102, 103, 106]. Some differences have been evidenced somehow between the resistome of infants and adults. Versluis and coworkers [98] reported that in infants, but not in adult’s microbiota, the expression of tetracycline resistance genes was predominant over the expression of genes coding for resistance to other antibiotics. In addition, Moore et al. [104] found that chloramphenicol resistance determinants were prevalent in a population of healthy infants during the first year of life and in their mothers; however, whereas multidrug efflux pumps (rarely found in mothers) were the primary effectors of chloramphenicol resistance in infants, acetyltransferases were more common in mothers, appearing also in almost all infants at later times. The proven presence in the microbiota of healthy children of mobile genetic elements close to ARG, or even next to multidrug resistance islands, as well as the apparent persistence of some mobile genetic elements in these microbiotas, point to the potential horizontal transferability and dissemination among the infant gut microbial community of such resistances [99, 107]. Whereas the possibility that the infant gut resistome could constitute a mobile reservoir of ARG is a matter of current concern [102, 107], several studies indicate that antibiotic resistant microbial populations in the early microbiota may be partly replaced later on by more susceptible lineages [104, 106]. Nevertheless, it should be taken into account that ARG carried in mobile elements seems to be more persistent in the resistome than chromosomally encoded ARG [105].
More multi-factor studies are needed to unravel the real contribution of antibiotics administration, including the precise time and way of administration, in the establishment of the infant microbiota and its resistome and the persistence of alterations along time. Long-term observational studies with cohorts composed by a large number of individuals will help to determine to what extent the alterations in the microbiota composition and functionality occurring in the perinatal period are maintained or normalized over time. Whether the nascent gut resistome can represent a threat to human health (with reference to the carrier individual, the community and/or the future offspring) and the alternative roles that such antibiotic resistances may play in the gut microbial community, warrant future research.
Conclusions
The establishment and development of the intestinal microbiota in the early neonatal period constitutes one of the most critical and determinant steps for the later health of the individual. Therefore, it is urgent to decipher and understand the factors determining the microbiota establishment in the neonate, among which the infant exposure to antibiotics may represent an important factor. The evidence discussed in this article clearly shows the impact of early life antibiotics exposure on the developing gut microbiota. These results should open the debate on the need for immediate action to minimize, or limit, the impact of early life antibiotics on the microbiota establishment process, while we decipher and fully understand the role of early microbiota on human health.
References
Sekirov I, Russel SL, Antunes CM, Finlay BB (2010) Gut microbiota in health and disease. Physiol Rev 90:859–904
Sommer F, Bäckhed F (2013) The gut microbiota—masters of host development and physiology. Nat Rev Microbiol 11:227–238
Kuczynski J, Lauber CL, Walters WA et al (2011) Experimental and analytical tools for studying the human microbiome. Nat Rev Genet 13:47–58
Salazar N, Arboleya S, Valdés L et al (2014) The human intestinal microbiome at extreme ages of life. Dietary intervention as a way to counteract alterations. Front Genet 5:406
Al-Asmakh M, Zadjali F (2015) Use of germ-free animal models in microbiota-related research. J Microbiol Biotechnol 25:1583–1588
Hansen CH, Nielsen DS, Kverka M et al (2012) Patterns of early gut colonization shape future immune responses of the host. PLoS One 7:e34043
Renz H, Brandtzaeg P, Hornef M (2012) The impact of perinatal immune development on mucosal homeostasis and chronic inflammation. Nat Rev Immunol 12:9–23
Olszak T, An D, Zeissig S et al (2012) Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336:489–493
Cox LM, Yamanishi S, Sohn J et al (2014) Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158:705–721
Gensollen T, Iyer SS, Kasper DL, Blumberg RS (2016) How colonization by microbiota in early life shapes the immune system. Science 352:539–544
Claus SP, Tsang TM, Wang Y et al (2008) Systemic multicompartmental effects of the gut microbiome on mouse metabolic phenotypes. Mol Syst Biol 4:219
Bercik P, Denou E, Collins J et al (2011) The intestinal microbiota affects central levels of brain-derived neurotrophic factors and behaviours in mice. Gastroenterology 141:599–609
Clarke G, O’Mahony SM, Dinan TG, Cryan JF (2014) Priming for health: gut microbiota acquired in early life regulates physiology, brain and behavior. Acta Paediatr 103:812–819
Neuman H, Debelius JW, Knight R, Koren O (2015) Microbial endocrinology: the interplay between the microbiota and the endocrine system. FEMS Microbiol Rev 39:509–521
Jimenez E, Fernandez L, Marin ML et al (2005) Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Curr Microbiol 51:270–274
Aagaard K, Ma J, Antony KM, Ganu R, Petrosino J, Versalovic J (2014) The placenta harbors a unique microbiome. Sci Transl Med 6:237ra65
Collado MC, Rautava S, Aakko J, Isolauri E, Salminen S (2016) Human gut colonisation may be initiated in utero by distinct microbial communities in the placenta and amniotic fluid. Sci Rep 6:23129
Arboleya S, Binetti A, Salazar N et al (2012) Establishment and development of intestinal microbiota in preterm neonates. FEMS Microbiol Ecol 79:763–772
Arboleya S, Sánchez B, Milani C et al (2015) Intestinal microbiota development in preterm neonates and effect of perinatal antibiotics. J Pediatr 166:538–544
Dogra S, Sakwinska O, Soh S-E et al (2015) Dynamics of infant gut microbiota are influences by delivery mode and gestational duration and are associated with subsequent adiposity. MBio 6:e02419
Dominguez-Bello MG, Costello EK, Contreras M et al (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA 107:11971–11975
Jakobsson HE, Abrahamsson TR, Jenmalm MC et al (2014) Decreased gut microbiota diversity, delayed Bacteroidetes colonization and reduced Th1 responses in infants delivered by caesarean section. Gut 63:559–566
Bäckhed F, Roswall J, Peng Y (2015) Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17:690–703
Fouhy F, Ross RP, Fitzgerald GF, Stanton C, Cotter PD (2012) Composition of the early intestinal microbiota: knowledge, knowledge gaps and the use of high-throughput sequencing to address these gaps. Gut Microbes 3:203–220
Faa G, Gerosa C, Fanni D, Nemolato S, van Eyken P, Fanos V (2013) Factors influencing the development of a personal tailored microbiota in the neonate, with particular emphasis on antibiotic therapy. J Matern Fetal Neonatal Med 26(S2):35–43
Rutten NBMM, Rijkers GT, Meijssen CB et al (2015) Intestinal microbiota composition after antibiotic treatment in early life: the INCA study. BMC Pediatr 15:204
Yatsunenko T, Rey FE, Manary MJ et al (2012) Human gut microbiome viewed across age and geography. Nature 486:222–227
Arrieta MC, Stiemsma LT, Amenyogbe N, Brown EM, Finlay B (2014) The intestinal microbiome in early life: health and disease. Front Immunol 5:427
Fanaro S, Chierici R, Guerrini P, Vigi V (2003) Intestinal microflora in early infancy: composition and development. Acta Paediatr Suppl 91:48–55
Cheng J, Ringel-Kulka T, Heikamp-de Jong I et al (2016) Discordant temporal development of bacterial phyla and the emergence of core in the fecal microbiota of young children. ISME J 10:1002–1014
Hollister EB, Riehle K, Luna RA et al (2015) Structure and function of the healthy preadolescent pediatric gut microbiome. Microbiome 3:36
Sudo N, Sawamura S, Tanaka K, Aiba Y, Kubo C, Koga Y (1997) The requirement of intestinal bacterial flora for the development of an IgE production system fully susceptible to oral tolerance induction. J Immunol 159:1739–1745
Bendtsen KM, Fisker L, Hansen AK, Hansen CH, Nielsen DS (2015) The influence of the young microbiome on inflammatory diseases—lessons from animal studies. Birth Defects Res C Embryo Today 105:278–295
Simonyte Sjodin K, Vidman L, Ryden P, West CE (2016) Emerging evidence of the role of gut microbiota in the development of allergic diseases. Curr Opin Allergy Clin Immunol 16:390–395
Russell SL, Gold MJ, Hartmann M et al (2012) Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep 13:440–447
Russell SL, Gold MJ, Willing BP, Thorson L, Mcnagny KM, Finlay BB (2013) Perinatal antibiotic treatment affects murine microbiota, immune responses and allergic asthma. Gut Microbes 4:158–164
Cho I, Yamanishi S, Cox L et al (2012) Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 488:621–626
Livanos AE, Greiner TU, Vangay P et al (2016) Antibiotic-mediated gut microbiome perturbation accelerates development of type 1 diabetes in mice. Nat Microbiol 1:16140
Watanabe J, Fujiwara R, Sasajima N, Ito S, Sonoyama K (2010) Administration of antibiotics during infancy promoted the development of atopic dermatitis-like skin lesions in NC/Nga mice. Biosci Biotechnol Biochem 74:358–363
Kumar P, Magon N (2012) Hormones in pregnancy. Niger Med J 53:179–183
Nuriel-Ohayon M, Neuman H, Koren O (2016) Microbial changes during pregnancy, birth, and infancy. Front Microbiol 7:1031
Mueller NT, Whyatt R, Hoepner L et al (2015) Prenatal exposure to antibiotics, cesarean section and risk of childhood obesity. Int J Obes 39:665–670
Stokholm J, Schjorring S, Eskildsen CE et al (2014) Antibiotic use during pregnancy alters the commensal vaginal microbiota. Clin Microbiol Infect 20:629–635
Gomez de Agüero M, Ganal-Vonarburg SC, Fuhrer T et al (2016) The maternal microbiota drives early postnatal innate immune development. Science 351(6279):1296–1302
Chernikova DA, Koestler DC, Hoen AG et al (2016) Fetal exposures and perinatal influences on the stool microbiota of premature infants. J Matern Fetal Neonatal Med 29:99–105
Tormo-Badia N, Hakansson A, Vasudevan K, Molin G, Ahrne S, Cilio CM (2014) Antibiotic treatment of pregnant non-obese diabetic mice leads to altered gut microbiota and intestinal immunological changes in the offspring. Scand J Immunol 80:250–260
Munyaka PM, Eissa N, Bernstein CN, Khafipour E, Ghaia JE (2015) Antepartum antibiotic treatment increases offspring susceptibility to experimental colitis: a role of the gut microbiota. PLoS One 10(11):e0142536
Khan I, Azhar EI, Abbas AT et al (2016) Metagenomic analysis of antibiotic-induced changes in gut microbiota in a pregnant rat model. Front Pharmacol 7:104
Tochitani S, Ikeno T, Ito T, Sakurai A, Yamauchi T, Matsuzaki H (2016) Administration of non-absorbable antibiotics to pregnant mice to perturb the maternal gut microbiota is associated with alterations in offspring behavior. PLoS One 11:e0138293
Hu Y, Peng J, Tai N, Hu C, Zhang X, Wong FS (2015) Maternal antibiotic treatment protects offspring from diabetes development in non obese diabetic mice by generation of tolerogenic APCs. J Immunol 195:4176–4184
Verani JR, McGee L, Schrag SF (2010) Division of Bacterial Diseases, National Center for Immunization and Respiratory Diseases, Centers for Disease Control and Prevention (CDC). Prevention of perinatal group B Streptococcal disease. Revised guidelines from CDC. MMWR Recomm Rep 59(RR-10):1–36
Alós Cortés JI, Andreu Domingo A, Arribas Mir L et al (2013) Prevención de la infección perinatal por estreptococo del grupo B. Recomendaciones españolas. Actualización 2012. Documento de consenso SEIMC/SEGO/SEN/SEQ/SEMFYC. Enferm Infecc Microbiol Clin 31:158–172
Di Renzo GC, Melin P, Berandi A et al (2015) Intrapartum GBS screening and antibiotic prophylaxis: a European consensus conference. J Matern Fetal Neonatal Med 28:766–782
Benitz WE, Gould JB, Druzin ML (1999) Risk factors for early-onset group B Streptococcal sepsis: estimation of odd ratios by critical literature review. Pediatrics 103:e77
Brocklehurst P (2015) Screening for Group B Streptococcus should be routine in pregnancy: ACAINST: current evidence does not support the introduction of microbiological screening for identifying carriers of Group B streptococcus. BJOG Int J Obstet Gynecol 122:368
Van Dyke MK, Phares CR, Lynfield R et al (2009) Evaluation of universal antenatal screening for group B streptococcus. N Engl J Med 360:2626–2636
Ohlson A, Shah VS (2014) Intrapartum antibiotics for known maternal group B Streptococcal colonization. Cochrane Database Syst Rev 10(6):CD007467
Bokulich NA, Chung J, Battaglia T et al (2016) Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci Transl Med 8:343ra382
Vangay P, Ward T, Gerber JS, Knights D (2015) Antibiotics, pediatric dysbiosis, and disease. Cell Host Microbe 17:553–564
Azad MB, Konya T, Persaud RR et al (2016) Impact of maternal intrapartum antibiotics, method of birth and breastfeeding on gut microbiota during the first year of life: a prospective cohort study. BJOG Int J Obstet Gynecol 123:983–993
Mazzola G, Murphy K, Ross RP et al (2016) Early gut microbiota perturbations following intrapartum antibiotic prophylaxis to prevent group B Streptococcal disease. PLoS One 11:e0157527
Keski-Nisula L, Kyynarainen HR, Karkkainen U, Karhukorpi J, Heinonen S, Pekkanen J (2013) Maternal intrapartum antibiotics and decreased vertical transmission of Lactobacillus to neonates during birth. Acta Paediatr 102:480–485
Fouhy F, Guinane CM, Hussey S et al (2012) High-throughput sequencing reveals the incomplete, short-term recovery of infant gut microbiota following parenteral antibiotic treatment with ampicillin and gentamicin. Antimicrob Agents Chemother 56:5811–5820
WHO/RHR/15.02 (2015) World Health Organization. Statement on Caesarean Section Rates
Smaill F, Hofmeyr GJ (2002) Antibiotic prophylaxis for cesarean section. Cochrane Database Syst Rev 3:CD000933
Hill CJ, Lynch DB, Murphy K et al (2017) Evolution of gut microbiota composition from birth to 24 weeks in the INFANTMET cohort. Microbiome 5:4
Dominguez-Bello MG, De Jesus-Laboy KM, Shen N et al (2015) Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. Nat Med 22(3):250–253
Clark RH, Bloom BT, Spitzer AR, Gerstmann DR (2006) Reported medication use in the neonatal intensive care unit: data from a large national data set. Pediatrics 117(6):1979–1987. doi:10.1542/peds.2005-1707
Stoll BJ, Hansen NI, Bell EF et al (2010) Neonatal outcomes of extremely preterm infants from the NICHD neonatal research network. Pediatrics 126:443–456
Goldenberg RL, Hauth JC, Andrews WW (2000) Intrauterine infection and preterm delivery. N Engl J Med 342:1500–1507
Stoll BJ, Hansen NI, Sánchez PJ et al (2011) Early onset neonatal sepsis: the burden of group B Streptococcal and E. coli disease continues. Pediatrics 127:817–826
Cotten CM, Taylor S, Stoll B et al (2009) Prolonged duration of initial empirical antibiotic treatment is associated with increased rates of necrotizing enterocolitis and death for extremely low birth weight infants. Pediatrics 123:58–66
Zhang T, Smith MA, Camp PG, Shajari S, MacLeod SM, Carleton BC (2013) Prescription drug dispensing profiles for one million children: a population-based analysis. Eur J Clin Pharmacol 69:581–588
Antonopoulos DA, Huse SM, Morrison HG, Schmidt TM, Sogin ML, Young VB (2009) Reproducible community dynamics of the gastrointestinal microbiota following antibiotic perturbation. Infect Immun 77:2367–2375
Tanaka S, Kobayashi T, Songjinda P et al (2009) Influence of antibiotic exposure in the early postnatal period on the development of intestinal microbiota. FEMS Immunol Med Microbiol 56:80–87
Mathew JL (2004) Effect of maternal antibiotics on breastfeeding infants. Postgrad Med J 80(942):196–200
Soto A, Martín V, Jiménez E, Mader I, Rodríguez JM, Fernández L (2014) Lactobacilli and bifidobacteria in human breast milk: influence of antibiotherapy and other host and clinical factors. J Pediatr Gastroenterol Nutr 59:78–88
Pacifici G (2006) Placental transfer of antibiotics administered to the mother: a review. Int J Clin Pharmacol Ther 44:57
Tamburini S, Shen N, Wu HC, Clemente JC (2016) The microbiome in early life: implications for health outcomes. Nat Med 22:713–722
Ong MS, Umetsu DT, Mandl KD (2014) Consequences of antibiotics of antibiotics and infections in infancy: bugs, drugs, and wheezing. Ann Allergy Asthma Immunol 112:441–445
Johnson CC, Ownby DR, Alford SH et al (2005) Antibiotic exposure in early infancy and risk for childhood atopy. J Allergy Clin Immunol 115:1218–1224
Stensballe LG, Simonsen J, Jensen SM, Bonnelykke K, Bisgaard H (2013) Use of antibiotics during pregnancy increases the risk of asthma in early childhood. J Pediatr 162:832–838
Chu S, Yu H, Chen Y, Chen Q, Wang B, Zhang J (2015) Periconceptional and gestational exposure to antibiotics and childhood asthma. PLOS One 10:e0140443
Wickens K, Ignham T, Epton M et al (2008) The association of early life exposure to antibiotics and the development of asthma, eczema and atopy in a birth cohort: confounding or causality? Clin Exp Allergy 38:1318–1324
Hill DA, Siracusa MC, Abt MC et al (2012) Commensal bacteria-derived signals regulate basophil hematopoiesis and allergic inflammation. Nat Med 18:538–546
Gonzalez-Perez G, Hicks AL, Tekieli TM, Radens CM, Williams BL, Lamouse-Smith ES (2016) Maternal antibiotic treatment impacts development of the neonatal intestinal microbiome and antiviral immunity. J Immunol 169:3768–3779
Cox LM, Blaser MJ (2014) Antibiotics in early life and obesity. Nat Rev Endocrinol 11:182–190
Nobel YR, Cox LM, Kirigin FF et al (2015) Metabolic and metagenomics outcomes from early-life pulsed antibiotic treatment. Nat Commun 6:7486
Trasande L, Blustein J, Liu M, Corwin E, Cox LM, Blaser MJ (2013) Infant antibiotic exposures and early-life body mass. Int J Obes 37:16–23
Bailey LC, Forrest CB, Zhanj P, Richards TM, Livshits A, DeRusso PA (2014) Association of antibiotics in infancy with early childhood obesity. JAMA Pediatr 168:1063–1069
Saari A, Virta LJ, Sankilampi U, Dunkel L, Saxen H (2015) Antibiotic exposure in infancy and risk of being overweight in the first 24 months of life. Pediatrics 133:617–626
Candon S, Perez-Arroyo A, Marquet C et al (2015) Antibiotics in early life alter the gut microbiome and increase disease incidence in a spontaneous model of autoimmune insulin-dependent diabetes. PLOS One 10:e0125448
Gibson MK, Crofts TS, Dantas G (2015) Antibiotics and the developing infant gut microbiota and resistome. Curr Opin Microbiol 27:51–56
D’Costa VM, McGrann KM, Hughes DW, Wright GD (2006) Sampling the antibiotic resistome. Science 311:374–377
Gillings MR (2013) Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Front Microbiol 4:4
Toprak E, Veres A, Michel JB, Chait R, Hartl DL, Kishony R (2011) Evolutionary paths to antibiotic resistance under dynamically sustained drug stress. Nat Genet 44:101–105
Barbosa TM, Levy SB (2000) The impact of antibiotic use on resistance development and persistence. Drug Resist Updat 3:303–311
Versluis D, D’Andrea MM, Garcia JR et al (2015) Mining microbial metatranscriptomes for expression of antibiotic resistance genes under natural conditions. Sci Rep 5:11981
Moore AM, Patel S, Forsberg KJ et al (2013) Pediatric fecal microbiota harbor diverse and novel antibiotic resistance genes. PLoS One 8(11):e78822
Moles L, Gomez M, Jimenez E et al (2015) Preterm infant gut colonization in the neonatal ICU and complete restoration 2 years later. Clin Microbiol Infect 21:936e1–936e10
Zhang L, Kinkelaar D, Huang Y, Li YL, Li XJ, Wang HH (2011) Acquired antibiotic resistance: are we born with it? Appl Environ Microbiol 77:7134–7141
Gosalbes MJ, Valles Y, Jimenez-Hernandez N et al (2016) High frequencies of antibiotic resistance genes in infants’ meconium and early fecal samples. J Dev Orig Health Dis 7:35–44
Fouhy F, Ogilvie LA, Jones BV et al (2014) Identification of aminoglycoside and beta-lactam resistance genes from within an infant gut functional metagenomic library. PLoS One 9:e108016
Moore AM, Ahmadi S, Patel S et al (2015) Gut resistome development in healthy twin pairs in the first year of life. Microbiome 3:27
Yassour M, Vatanen T, Siljander H et al (2016) Natural history of the infant gut microbiome and impact of antibiotic treatment on bacterial strain diversity and stability. Sci Transl Med 8:343ra81
von Wintersdorff CJH, Wolffs PFG, Savelkoul PHM et al (2016) The gut resistome is highly dynamic during the first months of life. Future Microbiol 11:501–510
Ravi A, Avershina E, Foley SL et al (2015) The commensal infant gut meta-mobilome as a potential reservoir for persistent multidrug resistance integrons. Sci Rep 5:15317
Acknowledgements
The work carried out in the authors’ laboratories on the early life microbiota is founded by the EU Joint Programming Initiative—A Healthy Diet for a Healthy Life (JPI HDHL, http://www.healthydietforhealthylife.eu/) and the Spanish Ministry of Economy and Competitiveness (MINECO) (Project EarlyMicroHealth). The Grant GRUPIN14-043 from “Plan Regional de Investigación del Principado de Asturias” is also acknowledged. A. M. N. is the recipient of a JPI predoctoral fellowship and N. S. benefits from a JdC contract, from the Spanish Ministry of Economy and Competitiveness (MINECO).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nogacka, A.M., Salazar, N., Arboleya, S. et al. Early microbiota, antibiotics and health. Cell. Mol. Life Sci. 75, 83–91 (2018). https://doi.org/10.1007/s00018-017-2670-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2670-2