Abstract
MicroRNAs have continued to attract enormous interest in the scientific community ever since their discovery. Their allure stems from their unique role in posttranscriptional gene expression control as well as their potential application as therapeutic targets in various disease pathologies. While much is known concerning their general biological function, such as their interaction with RNA-induced silencing complexes, many important questions still remain unanswered, especially regarding their functions in the skin. In this review, we summarize our current knowledge of the role of microRNAs in the skin in order to shine new light on our understanding of cutaneous biology and emphasize the significance of these small, single-stranded RNA molecules in the largest organ of the human body. Key events in epidermal and hair follicle biology, including differentiation, proliferation, and pigmentation, all involve microRNAs. We explore the role of microRNAs in several cutaneous processes, such as appendage formation, wound-healing, epithelial-mesenchymal transition, carcinogenesis, immune response, and aging. In addition, we discuss current trends in research and offer suggestions for future studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The skin in our culture and science: a semi-scientific analysis
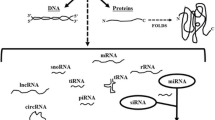
The skin is and always has been one of the most important organs in popular culture. Its implications on the perceptions of beauty, health, and age are paramount [1, 2]. A simple Google search with the term “skin” results in an astonishing 1.74 trillion hits—more hits than in searches for “brain,” “liver,” “lung,” “colon,” “kidney,” “immune system,” and “stomach” combined—falling second only to a search for the term “heart,” which results in 2.63 trillion hits (Table 1). This analysis supports the notion that popular culture is obsessed with the skin and its adnexa (hair, breasts, lips, nails), and they invoke an old adage from the poet, James Joyce: “We might say indeed of modern man that he has an epidermis rather than a soul.” In this review, we share the recent advances in our knowledge of microRNA function in the skin, an important organ as well as an excellent model system to explore the complexities of the relatively young field of microRNA biology (Fig. 1).
Indeed, the skin fulfills many important functions. In addition to its enormous impact on physical attractiveness, the skin most importantly protects the underlying organism from its external surroundings: a cruel environment of extreme temperatures, UV radiation, and parasites, among other potential hazards. Fortunately, the skin of mammals has adapted throughout evolution to master each of these challenges with some assistance from the immune system. Prominent evolutionary innovations of the mammalian lineage include appendages of the skin, such as hair follicles and mammary glands [3]. These complex, novel mini-organs require progressively complex genetic programs for their development and maintenance. In this context, it is interesting to note that, in general, the emergence of novel, increasingly complex body plans—with the appearance of vertebrates and mammals in particular—correlates well with the expansion of the microRNA repertoire [4].
MicroRNAs and mammals: microRNAs as a major tool in evolution
MicroRNAs are small, single-stranded RNA molecules with an average size of about 22 nucleotides [5, 6]. Their prevalence has increased dramatically throughout animal evolution, and this expansion is believed to have had a major role in the formation of progressively complex body plans. In sharp contrast to other types of genes—such as protein-coding genes—the repertoire of microRNAs continues to expand and may even have been critical for our own evolution [7] (Fig. 2).
Evolutionary expansion of microRNAs in deuterostomes. Deuterstomes and protostomes share an ancient set of microRNAs, including let-7 family members, miR-31, and miR-34. Several waves of microRNA expansion occurred during deuterstome diversification, most notably associated with the emergence of vertebrates and placentals. The vast majority of microRNAs discussed in this review are either vertebrate or placental specific. For reasons of space and simplicity, only microRNAs up to number miR-375 were included
Two major expansions of the microRNA repertoire took place in evolutionary history: one associated with the appearance of placentals and the other corresponding to vertebrate evolution. Of approximately 11,000 eukaryotic genes present in humans, over 90 % existed prior to the emergence of mammals [8]. Mammals have added approximately 17 new microRNA families to their genome, and, with a few exceptions, we know hardly anything about their role in mammalian biology. Throughout vertebrate evolution, at least 41–72 new microRNA families found their way into the genomes of the most basic vertebrates, lampreys [9]; as time progressed, advanced vertebrates added another 16 microRNA families to the repertoire, many of which have several members each, producing a myriad of microRNAs. In this light, the expansion of the microRNA gene repertoire is remarkable.
Where did all of these new microRNAs come from? Many of them may have developed from CpG islands, repetitive elements within a genome, while others may have developed from pre-existing microRNAs. Such a constant flow of new microRNAs explains how so many microRNAs are species-specific and lineage-specific [10, 11].
The correlation between evolutionary increases in microRNAs and the generation of new cell types is also striking. The Cnidarian Nematostella has approximately 49 microRNAs; the fruit fly has 240 microRNAs; Homo sapiens has more than 1,500 microRNAs. At the same time, Homo sapiens has at least 200 different cell types, while a mere 14 different cell types exist in Cnidaria. Simply contemplate the complex organization of the hair follicle and estimate the number of different cell types involved in the formation and maintenance of such a structure: stem cells, matrix cells, inner root sheath cells with all of their different subclasses, dermal papilla cells with their precursors, melanocytes with their precursors, and cells of the sebaceous glands with all of their differentiation stages. With such a diversity of cell types and lineages, microRNAs seem to be the ideal tool to provide a robust system for swift and clean transitions between various differentiation stages.
As a final piece of supporting evidence, the rapid evolutionary radiation of cichlids in East African lakes serves as a wonderful model to demonstrate the potential power of microRNAs. In the last million years, cichlids have diversified enormously; yet, while minimal alterations have occurred within their genomes, an analysis of microRNA-binding sites in cichlid mRNAs from different species suggests an important contribution from microRNAs in the evolution of the diverse phenotypes of various cichlid species [12].
These facts highlight the potential role of microRNAs on complex gene expression regulation: microRNAs may have been used evolutionarily to fulfill increasingly important functions, and they may eventually contribute to increasingly complex gene and network regulation possibilities in the future. This concept highlights the observation that evolutionarily younger genes harbor more transcription factor and microRNA binding sites than older genes, implying that younger genes retain more potential for additional regulation than older genes [7–12].
How microRNAs work: another plethora of options for noise control (or how working behind the scenes can still make you famous)
MicroRNAs serve as the guides of a protein complex known as RNA-induced silencing complex (RISC), which regulates the translation of mRNAs. Each RISC is associated with one of four different Argonaute proteins (Ago 1–4). While the different functions of each Ago remain for the most part unknown, Ago2 is considered to be the most important of the four in as much as it interacts with the majority of the microRNAs in the skin relative to the others. Recently, Yi and colleagues have demonstrated using the skin as their model system that the loading of microRNAs into individual RISCs is a stochastic process independent of the type of Argonaute protein in each complex [13].
After loading, each RISC utilizes its incorporated microRNA in order to interact with a target mRNA transcript. While the exact mechanisms behind this targeting process are not well understood, several crude principles have emerged through observation of its activity. For example, nucleotides two through seven of the microRNA transcript appear to play a significant role in the interaction of RISC with the target mRNA. These six nucleotides are collectively referred to as the “seed sequence” [14] of the microRNA transcript, and several target prediction algorithms utilize this seed sequence as a major factor in their predictions (for example, Targetscan). Although the error rates of such predictions can be high, a better understanding of the factors influencing the targeting process, such as target abundance and seed pairing stability, may improve target prediction [15].
Once the target mRNA is bound to the microRNA-RISC, translation of the mRNA transcript will not occur. The exact fate of the complex-bound target mRNA remains debatable; however, it is known that target mRNAs can be shuttled to specific cytoplasmic locations in the cell for subsequent storage or degradation. Such loci/foci are known as “P bodies” (“processing bodies”); while their exact significance to proper microRNA function remains unknown, their presence within a cell is tightly associated with microRNA activity [16].
The previously described mechanism is well-accepted among the scientific community. Outside of the popular dogma, Moser and Fritzler [17] raised an important issue regarding microRNA function when they compared microRNA profiles of total intracellular RNA extractions with microRNA profiles of RNA extracted directly from immunoprecipitations of RISC. In their comparison, they found that many microRNAs were differentially represented between the two populations. While it remains to be seen whether such findings are erroneous or whether they truly represent another step in the control of microRNA activity, such findings raise several novel questions: Could there be a potential function of microRNAs outside of the RISC? How valuable are microRNA expression data after all? And what are the validities of microRNA arrays and microRNA qRT-PCR analyses [18]?
Several gaps in knowledge still exist in regard to the exact mechanisms behind microRNA activity; yet, the notion that microRNAs play a direct role in the regulation of gene expression is well established by the literature. If we accept this notion, as well as the notion that “negative feedback through mRNA provides the best control of gene-expression noise” [19], then the next thing we need to know is how to tame such a tool.
How to control the minions
MicroRNAs are powerful tools for the control of gene expression, and their expression within a cell is tightly regulated as well. As an example of this high level of control, we would like to mention the work by Martello et al. [20] that illustrates the role of microRNAs in vertebrate embryonic development. The researchers demonstrate that microRNAs that target the nodal receptor during embryogenesis are expressed in a gradient that opposes that of the nodal gradient itself. In addition to serving as a practical example of how microRNAs actually work during embryogenesis and improve the robustness of biological processes, such findings demonstrate that the expression of microRNAs can be closely regulated. Indeed, microRNAs can be controlled at several points throughout their life cycle: from the level of their transcription down to their degradation. While the exact mechanisms are still ambiguous, we would like to highlight several significant discoveries in the recent literature.
A recent article in Nature by Poliseno et al. [21] indicates that 3′UTRs from different genes may participate in microRNA regulation. The authors found that the 3′UTRs of pseudogenes, which were once considered superfluous pieces of genetic information, may serve as buffers/traps/decoys/sponges for microRNAs and may thus protect functional mRNAs from the microRNAs that target them. This study sheds new light in general on the many non-coding RNAs that may potentially serve as such decoys.
In reality, few pseudogenes are expressed at sufficient levels for microRNA regulation, and the microRNA system should be robust enough to tolerate a few extra 3′UTR sequences that function as decoys; however, the authors extrapolated their findings to reach some intriguing conclusions regarding the astonishing complexities of microRNA regulation [22, 23]. In their work, they propose the existence of several decoy mRNAs, termed competitive endogenous RNAs (ceRNAs), that protect PTEN mRNA from microRNAs and contribute to tumor suppression.
So far, the most convincing evidence of natural competitor mRNAs contributing to gene expression control in animals comes from a study on glioblastomas [24]. The study identifies and implicates an additional set of ceRNAs in protecting PTEN mRNA from microRNA-mediated control. Furthermore, previous studies in plants established such a decoy mechanism several years prior [25].
While the interpretations of these findings remain controversial, results of such research do assure of us of one thing: the regulation of microRNAs is complex; thus, microRNAs should not be underestimated in regard to their impact on gene expression control. Not surprisingly, the questions that such findings pose are manifold. What do they mean regarding the application of microRNAs as gene expression noise buffers and noise reducers? Will we be able to simultaneously account for all of these complexities in the context of cancer and other complex diseases? And if so, will we be able to overcome them in order to apply microRNA technology as cures for such illnesses? It seems unlikely that clear answers to these questions will be obtained in the near future, but new discoveries continue to arise. For example, recent studies indicate that overexpression of the 3′UTR of CD44 can by itself induce metastasis, potentially via interference of microRNA function [26]. Taken together, this wave of relatively new data draws attention to our limited understanding of the role of the “mysterious agents” known as microRNAs in the process of gene expression control [27].
In addition to regulating intracellular microRNA levels at various stages, cells can direct the impact of microRNAs on gene expression via additional tools. As a “simple” example, alteration of the 3′UTRs of mRNA transcripts eliminates microRNA binding sites and thus helps avoid targeting by microRNAs [28, 29]. Although it is not well understood how such a mechanism is carried out and controlled, this process occurs on a global level in proliferating cells and seems to function as a powerful mechanism that enables cells to enhance protein output without the inactivation of a substantial amount of mRNA by microRNAs. Such a process may be disease-associated and, if so, could be exploited to revert cells and tissues to their previous states via restoration of the normal 3′UTR/microRNA balance. However, a greater understanding of 3′UTR-regulation during disease processes is required before potential benefits may be reaped.
As an alternative form of regulation, RNA-binding proteins can directly alter the ability of RISC to interact with the target mRNA transcripts, leaving 3′UTR sequences unchanged. Such mechanisms are utilized by p53, a tumor suppressor, which actually has so many microRNA interactions that it is surprising that we elucidated anything about its mechanism of action prior to the discovery of microRNAs [30].
Lastly, underscoring the importance of microRNAs is the discovery that viruses too possess integrated countermeasures against microRNA-mediated cellular defense mechanisms. A recent finding shows that murine cytomegalovirus specifically targets the antiviral miR-27 for degradation via a novel mechanism that utilizes an mRNA, m169, to inhibit and degrade miR-27 [31]. Interestingly, m169 can be modified at the miR-27-binding site and redirected to target a different microRNA. This exciting discovery is another one of the latest—and unlikely one of the last—examples of how microRNA activity can be controlled.
Indeed, the complexity that microRNAs add to the regulation of gene expression is mind-boggling; yet, such complexity leaves little doubt about their crucial role in the evolution of mammals. Given what we know, it is not irrational to believe that the vast array of microRNAs at the disposal of mammals contributed directly to the large variety of novel structures associated with their skin.
MicroRNAs in the skin: regulators of differentiation, proliferation, and appendage formation
The hair follicle mini-organ is the most apparent novel structure shared by present day mammals. In the fossil record, hair can be traced back an astonishing 164 million years to proto-mammals. Hair follicles share similarities with other vertebrate ectodermal adnexa—such as feathers, scales, and even teeth—and mammals utilize similar genetic programs to those of birds and fish in order to form these novel epidermal structures [32]. Within these genetic programs, microRNAs play major roles.
However, there are neither data on the expression pattern nor on the role of any microRNA in bird skin and feather development. This is unfortunate since feathers are probably the most complex adnexa of vertebrate skin. Feathers are clearly detectable in the fossil record in several dinosaur groups as far back as 130 million years ago, making them valuable objects to study the contribution of microRNAs to the development of novel intricate structures characteristic for vertebrates in general. A comparison of microRNA data from bird feather development and hair folliculogenesis may allow for a better understanding of the basic principles of convergent evolution that governed the invention of hair, feather, teeth, and scales.
The majority of the microRNAs expressed in the skin or epidermis of vertebrates are from a limited number of clusters or families (Table 2). Of the microRNAs that are differentially expressed in skin (with or without hair follicles), three microRNAs stand out as they exist only in mammals: miR-105, miR-127, and miR-224 [33]. In addition, Yi et al. identified five microRNAs that are prevalent in hair follicles compared to their presence in the epidermis alone: miR-199a, miR-214, miR-126, miR-143, and miR-152 [34]. While the majority of these microRNAs are associated with the telogen phase of the hair growth cycle [35], several of these microRNAs may play a crucial role in the formation of this novel appendage.
In mice, the prototypic mammalian genetic model animal, hair follicles are affected by the loss of the key enzymes of microRNA biogenesis: Dicer, Drosha, and DGCR8 [33, 34, 36]. Tissue-specific deletion of either gene results in very similar phenotypes, including the inability of many hair follicles to grow into the dermis and the emergence of dermal papillae in the epidermis. Another phenotype involves a disruption of the normal scale pattern on the tail. The development of such a phenotype supports the idea that microRNAs ensure a high degree of canalization or the ability to produce a consistent phenotype regardless of environmental or genotypic variability during development [37].
MicroRNAs are also required for postnatal hair growth. Through a knockout of the essential microRNA biogenesis enzymes Drosha and Dicer, we recently demonstrated that microRNAs are necessary to maintain the highly proliferative matrix cells of the hair follicle [38]. In addition, recent work by Yi and colleagues supports previous findings of Dicer, Drosha, and DRGC8 knockout models through their Argonaute protein knockouts in keratinocytes [13].
Limitations of Dicer, Drosha, and DGCR8 knockouts
These knockout studies reveal that microRNAs in general are essential for the proper formation and maintenance of hair follicles because of decreased proliferation and increased apoptosis in their absence; however, such Dicer, Drosha, DGCR8, and Argonaute knockout models come with flaws. Most significantly, while these knockout models adequately represent the simultaneous loss of large sets of microRNAs (multi-knockouts), they fail to represent the loss of individual microRNAs (single knockouts). Thus, while they have served to establish the significance of microRNAs in the skin in general, they cannot help elucidate the significance of individual microRNAs.
Although microRNAs are functionally closer to minions than to master regulators of a control system, their allure makes it tempting to treat them as individual master genes. While the vast majority of microRNA knockout studies in worms and mice do not result in obvious phenotypes, there are some exceptions (miR-126 [39], miR-96 (deafness at birth) [40], and miR-9-2/9-3 [41]). From the perspective of skin biology, the most remarkable of these exceptions is miR-205. The “Keck miRKO Knockout Pipeline” (http://rna.keck.ucsf.edu/miRKO-DB) at the University of California, San Francisco (UCSF), has just released its first set of data on several microRNA knockouts, and regarding the skin, the only knockout with an embryonal phenotype was that of miR-205. Overall, their efforts indicate that up to 15 % of microRNA knockout models may exhibit embryonal phenotypes [42].
Indeed, knockouts have become the standard technique for studying the roles of microRNAs in other systems, but regarding the skin, miR-205 is the exception. In fact, it is the only microRNA associated with the skin or squamous epithelia (mir-205, miR-203, or the miR-200 family) that has been evaluated with knockouts as of yet. This is surprising, considering that studies on miR-203 in particular indicate that individual microRNAs may play important roles in skin biology. Furthermore, the number of studies that even address the role of individual microRNAs in the skin is small. In the following paragraphs, we discuss the results from those that have done so.
miR-31: regulation of the hair growth cycle
While the hair follicle phenotypes observed in conditional Dicer knockout mice imply that microRNAs in general are crucial for normal hair growth, the Botchkareva laboratory published the first paper to address the significance of an individual microRNA in such processes [35].
Their analysis demonstrates an association between miR-31 and the anagen phase of the hair growth cycle, with an emphasis on matrix cells [35, 43]. Together, with family members from the miR-17–92 cluster [43], miR-31 seems to function as a crucial regulator of proliferation in matrix cells. Suppression of miR-31 results in an increased or accelerated anagen phase, the active phase of the hair growth cycle.
As implied earlier, microRNAs are far from simple, and miR-31 is no exception. For example, miR-31 is associated with psoriasis [44], and other data that have accumulated on miR-31 merely hint at the complexities of microRNA biology. On one hand, miR-31 is associated with pro-growth and anti-metastatic properties in breast cancer [45]; on the other hand, it is regarded as a tumor suppressor in mesothelioma [46].
miR-203: differentiation and regulation of stemness
In 2008, Pivarcsi and colleagues were the first group to suggest that miR-203 serves a significant function in skin biology [47]; in the subsequent year, a much clearer picture of the role of miR-203 in the skin emerged [48]. Yi et al. implicated miR-203 in mediating, and, perhaps more precisely, contributing to a proper and distinct transition from basal cells to suprabasal cells in the epidermis. Its mechanism of action as a squamous differentiation marker involves p63, one of its keratinocyte targets and the master regulator of squamous cell fate. Through the downregulation of p63 during epidermal stratification, miR-203 appears to ensure the induction of differentiation. While the study lacks a clear loss-of-function model, it manages to present miR-203 as a classic example of a microRNA ensuring the transition between two cellular states: in this case, from a proliferative, undifferentiated state to a post-mitotic, differentiated compartment of a squamous epithelium. By assisting in this decision process and preventing the retention of any undifferentiated cells, miR-203 aids the epidermis in its key functions, such as the prevention of water loss and of penetration by hostile germs.
While miR-203 is frequently likened to a stemness inhibitor, such a comparison may be a bit of a stretch. miR-203 is not used to inhibit stemness, but instead to ensure that the cell continues on the road to terminal differentiation without complications. Therefore, it is more appropriate to consider miR-203 as a sort of “roadblock” to proliferation that is set up after the decision to differentiate has been made.
Beyond miR-203: p63 and its entourage of microRNAs
p63 is a key regulator of epidermal cell fate. Since the discovery of the connections between miR-203 and p63, additional microRNAs have been implicated in the function of p63 as well. However, the data are not quite consistent, most likely because of the differences in platforms used to measure changes in microRNA expression as well as different experimental approaches. Nevertheless, miR-193a appears to be under the direct negative control of p63 [49], as are miR-138, miR-130b, and miR-181 family members [50], in addition to miR-34 family members [51]. On the other hand, miR-200 family members may be upregulated by p63 [52]. The functional evaluations of some of these p63 target genes indicate that they contribute to the ability of p63 to control proliferation and senescence.
As a final note, one of the most convincing studies on the relationship between p63 and microRNAs, by Chikh et al., indicates that PPP1R13L, also known as iASPP, protects p63 from being targeted by microRNAs, miR-720 and miR-574-3p. These microRNAs are suppressed by PPP1R13L, which prevents p63 destabilization and terminal differentiation. This study is significant in as much as it emphasizes the importance of PPP1R13L in epidermal homeostasis and illustrates the intricate interplay between key epidermal regulators and microRNAs [53].
miR-125b: regulation of proliferation
The discoveries of lin-4 and members of the let-7 family were epiphanic events in the fields of gene expression regulation and microRNA biology. Not only were they among the first microRNAs to be discovered and determined to be evolutionary-conserved, but the elucidation of their important roles in non-vertebrate development and stem cell biology have catapulted microRNAs to stardom [54].
The lin-4/miR-125 family is as old as miR-31, and thanks to studies in C. elegans, much is known about the function of lin-4/miR-125 in non-vertebrates. miR-125 is associated with tumorigenic processes in many tissues. For example, in melanoma cells, miR-125 induces senescence [55]. miR-125 is also downregulated in psoriasis [56] and verrucous carcinoma [57], and generally has reduced expression in squamous cancers. In cutaneous squamous cell carcinoma cell lines, miR-125b was found to suppress proliferation, colony formation, and the migratory and invasive capacity of the cells [58].
The Fuchs laboratory provides the clearest data set on the involvement of miR-125 in skin biology [43]. In their publication, the authors present two significant findings: an enrichment of miR-125b in early mouse hair follicle stem cells and the reversible inhibition of hair growth by overexpression of miR-125b. A target of miR-125b in vitro and in vivo within keratinocytes, VDR, functions as a mediator of this phenotype. VDR itself explains many of the differentiation problems in the hair follicle after miR-125b overexpression; however, they identified additional miR-125b target genes that may have even more important functions in the regulation of stem cell behavior. Hopefully, these additional targets will shed light on the complex regulation, maintenance, and proliferation of stem cells in the skin [43].
The analysis of targets in various species also supports the notion that miR-125 is a general regulator of apoptosis and proliferation rather than an isolated controller of stem cell proliferation. Furthermore, miR-125 may be associated with differentiation in normal human skin and inhibition of proliferation in human keratinocytes [56]. The analysis, which linked miR-125b to the p53 network, illuminated an intriguing issue with the unbiased analysis of target genes: while most of the microRNA targets are not conserved across species (human, mice, zebrafish, etc.), the pathways that are affected are conserved [59].
The let-7 family of microRNAs
let-7 is another ancient microRNA family with several extremely conserved target genes that serve various functions, such as the control of proliferation, differentiation, and stemness, among other roles. If there ever was an anti-stemness factor throughout evolution, it would have been let-7. Among these conserved target genes, lin-28 and lin-41/TRIM71 stand tall; however, let-7 family members also appear to control cell cycle genes, including cyclins and CDKs [60], as well as the oncogenes Ras [61] and HMGA2 [62].
All of the data on the let-7 family indicate that its members can function as tumor suppressors. In the skin, the organ with the highest tumor incidence, let-7 members could be critical players in tissue homeostasis via control of the growth and differentiation of stem cells. Indeed, the analysis of lin-41/Trim71 expression in mouse embryonic epidermis reveals a basal cell pattern inverse to let-7 [63]. Such data support the model of microRNAs mediating the transition from an undifferentiated cellular state to a differentiated suprabasal state.
With this in mind, we speculate that let-7 microRNAs function as key regulators of cutaneous differentiation processes. Not only are they, along with miR-203, among the most abundant microRNAs expressed in the epidermis, but they are also preferentially expressed in differentiated cells [64]. Therefore, let-7 microRNAs may complement miR-203 in the suppression of genes associated with the basal layer cell phenotype.
In addition, a new link was recently established between let-7 and lin-28, a let-7-regulating protein [65]. The mRNA AU-rich element binding factor ZFP36 (tristetraproline, TTP), a key regulator of skin inflammation, mediates the degradation of lin-28, thereby increasing the expression of let-7. ZFP36 is regarded as a potential tumor suppressor, specifically in melanoma and squamous cell carcinomas [66, 67].
Depending on the analysis and tissue starting material, data demonstrate that the miRNome of the skin is dominated by let-7 family members along with a small number of other microRNAs: miR-143, miR-203, miR-451, miR-21, and miR-26a [44]. Based on such expression data, these microRNAs seem to occupy the majority of RISC complexes (Table 2). However, despite such discoveries, no efforts have been undertaken to test the hypothesis that let-7 microRNAs are crucial regulators of epithelial differentiation. The fact that the family has 12 members scattered across the genome and often aligned with other microRNAs in clusters may have hampered the analysis of the function of let-7 microRNAs in mice and humans.
Redundancy of the microRNA system
Then perhaps microRNAs function as a palladium of differentiation: gatekeepers of the differentiated phenotype that insulate cells from retrograde routes of dedifferentiation [63]. Given the high expression and domination of microRNA activity within differentiated cells, it is possible that miR-203, let-7, and miR-125 engage in a ménage à trois to prevent the system from drifting back to a basal cell fate (Fig. 3). Thus, instead of each microRNA working alone to fulfill an important task, microRNAs could instead work together.
The palladium of differentiation hypothesis: gatekeepers of the differentiated phenotype. MicroRNAs function as roadblocks to proliferation set up after the decision to differentiate has been made. In the epidermis, miR-203, let-7, and miR-125b work together with other highly expressed microRNAs to prevent the system from drifting back to a basal cell fate (blue cells) by suppressing genes associated with the basal layer cell phenotype: lin-41/TRIM71, FGFR2, P63, and cell cycle-related genes
As mentioned earlier, microRNAs mainly function as effectors—not master regulators—of a control system that participate in gene expression buffering, noise reduction, and fine-tuning of biological processes. Thus, redundancy within the microRNA system is appropriate, and by definition, most microRNA knockouts should only result in minimal to mild phenotypes. Indeed, redundant systems are a recurring theme in biology. For example, organisms often have functionally redundant proteins encoded by completely different portions of the genome in order to ensure survival in case of mutation, loss, or other unforeseen circumstances. The microRNA system need not be an exception to this trend, and in fact, studies in C. elegans support this notion.
In C. elegans, most individual microRNA losses are of no consequence in this animal [68]. However, harm can be accomplished through weakening the entire system via reduction of overall microRNA production followed by deletion of a specific microRNA [69]. In this scenario, not only is the microRNA in question downregulated, but by weakening the entire system, the possibility of any unknown functionally redundant microRNAs is accounted for. Thus, the elimination of the microRNA in question remains unchecked by redundant mechanisms, and a phenotype develops.
As in C. elegans, few microRNA knockouts in mice result in obvious embryonal phenotypes, with most knockout animals surviving to adulthood without any problems. To add insult to injury, within these unsuccessful first waves of microRNA knockout mouse models, researchers focused on popular microRNAs with potentially important functions. As a recent example, it has been demonstrated that a complete inactivation of miR-34, an attractive candidate for a knockout phenotype, is still compatible with normal development in mice. While miR-34 is implicated as an integral modulator of the p53 pathway, p53 function still remains intact in miR-34-deficient tissues; thus, a knockout of miR-34 does not inhibit p53-induced apoptosis [70]. Such results are suprising; however, a redundancy in the microRNA system could account for the lack of phenotypes seen in these knockouts by compensating for individual eliminations and masking phenotypic effects.
If this is this case, then in order to elucidate the role of individual microRNAs in the skin, we must apply novel methods to our research: manipulating individual microRNAs in combination with weakening the entire microRNA system. The latter could be induced via downregulation—not knockout—of proteins essential to microRNA biogenesis: Dicer, Drosha, DRGC8, or RISC-associated proteins. Through doing so, one would reduce possible redundancies that could interfere with and mask phenotypic effects of the original elimination.
To further validate the notion that individual microRNAs can play major roles in the skin, we will spend the remainder of this review highlighting the current knowledge of microRNAs in clinical processes of the epidermis. These highlights also emphasize the significance of microRNAs in therapeutic applications for various pathologies and the direction of microRNA-related research.
MicroRNAs in regeneration and wound healing
The skin is also a place of rare mammalian tissue regeneration: hair follicle neogenesis in wounded mice, post-amputation bone regeneration, and the annual de novo formation of antlers in deer [71], among other examples. However, the classical models of tissue regeneration stem from other vertebrates, including the fish fin and almost everything regarding the newt. Thus, the little amount that we do know regarding the role of microRNAs in regeneration stems solely from non-mammalian models [72].
As an example, zebrafish fin regeneration is accompanied by global changes in microRNA expression involving miR-31, miR-21, and, most notably, miR-133. Downregulation of miR-133 by FGF signaling is essential for proper fin regeneration [73]. In addition, miR-133 is also downregulated in salamander tail regeneration [74]. However, several other microRNAs show more dramatic changes, such as miR-196b, whose upregulation also affects proliferation in the blastema [75].
Since regeneration in these examples follows wound healing, formation of a blastema, and differentiation of lost tissue, questions arise regarding the role of microRNAs in wound healing and their application to overcome obstacles in chronic wound care. Chronic wounds exemplify the challenges that health-care systems face in Western countries with an increased occurrence among aging populations and patients with diabetes and/or cancer.
A study by Biswas et al. [76] reports the first significant contribution to our understanding of microRNAs in poorly healing ischemic wounds. Biswas et al. demonstrated that miR-210, a microRNA well documented to be induced by hypoxia and HIF-1α, inhibits the proliferation of keratinocytes in vitro and, therefore, may contribute to impaired healing of ischemic wounds in vivo.
MicroRNAs may also play a role in the formation of fibroproliferative keloids, which represent a form of abnormal wound healing. The Yamashita laboratory recently identified several microRNAs associated with keloid formation by comparing the expression profiles of keloid-derived fibroblasts with those of normal fibroblasts. Results include 20 downregulated and 7 upregulated microRNAs, with miR-196a in particular exhibiting the greatest change. The researchers found that the level of expression of miR-196a was inversely related to the level of secretion of type I and III collagens [77]. Unfortunately, aside from these isolated reports, little is known regarding the role of microRNAs in cutaneous wound healing. Thus far, the only microRNAs implicated in proliferation and migration control during wound healing are mir-483-3p and miR-21, respectively, in addition to miR-210 [78].
The process often associated with wound healing but not clearly demonstrated to occur until recently [79] has fared better with the microRNA community: epithelial-mesenchymal transitions (EMT) are key to many developmental processes, wound healing, and, of course, tumor invasion and metastasis.
MicroRNAs in cancer
The TGF-β-induced epithelial-mesenchymal transition (EMT) is a prime example of the relevance of microRNAs to cancer biology. The miR-200 family members constitute an integral portion of the network controlling EMT through their suppression of the transcription factors ZEB1 and ZEB2. In turn, ZEB1 and ZEB2 mediate the switch from epithelial to mesenchymal cell type via two processes: control of E-cadherin, the master of epithelial integrity, and suppression of the expression of miR-200 family members. miR-200 is also activated by another set of master regulators, p63 and Notch signaling (Fig. 4) [80, 81].
Unfortunately, this straightforward and mechanistically beautiful miR-200/ZEB network in EMT cannot be extrapolated to other cell types, such as melanocytes/melanoma [82, 83]. This inability highlights the problems that come with oversimplification and ignorance of the complexities of molecular biology. There is no good reason to assume that a given microRNA retains the same target genes across different cell types and that target spectrums remain the same in identical cell types across different species. Nevertheless, in keratinocytes, miR-200 downregulation is indeed associated with EMT and suppression of metastatic behavior in vivo [84].
It remains to be seen whether our enthusiasm about microRNAs will precipitate into therapeutic applications in the fields of wound healing, tissue regeneration, and cancer. If so, dermatology may become the forefront of testing microRNA-based therapies because of easy access to affected tissues and an urgent need for improved care of wounds. Only time will tell.
However, microRNAs continue to demonstrate significant roles within all of the signaling networks implicated in cancer: p53, TGF-β signaling and EMT, cell cycle regulation, apoptotic cell pathways, and invasion and metastasis, among others. We will take a step back and look at the changes in gene expression networks that take place in cancer cells as well as the role of microRNAs in such processes.
Let us assume that signaling networks function as complex adaptive systems that allow “a high degree of resilience and robustness to environmental challenges through their self-adaptation and internal self-organization” [85]. These complex adaptive systems utilize simple networks with feedback and feedforward circuitries in order to stabilize gene expression or promote cell fate decision, and microRNAs make important contributions to the stability of these networks [86]. The well-balanced nature of these networks likely relies upon these microRNAs, which prevent them from falling from the edge of chaos into actual chaos. Thus, in this scenario, microRNAs serve as foot soldiers against chaotic forces in order to support the maintenance of gene expression in its tissue- and cell-appropriate healthy limits.
But how does this impact cancer development in the skin? The microRNA system seems to remain more or less intact in squamous cell cancer (SCC), since only limited changes in microRNA expression occur during SCC formation. However, the perturbations of mRNA expression are profound and consequently raise questions regarding the functions of the microRNA/RISC system in this new environment: How do they confront, buffer, and control gene expression with this altered mRNA expression [21–28]? We still await answers to such questions, but the knowledge base of the role of microRNAs in cancer has grown because of recent discoveries in various skin cancers.
Squamous carcinogenesis from the microRNA perspective: miR-21
As mentioned previously, tumor formation is accompanied by massive changes in gene expression [87]; however, in contrast to the drastic alterations observed in mRNA and protein expression, the miRNome actually appears to remain relatively stable. Does this mean that microRNAs are mere bystanders in carcinogenesis? Hardly.
Although the literature is slightly lacking in this case, crude evidence exists regarding changes in microRNA expression during squamous cell carcinogenesis [88]. Specifically, the microRNA miR-21 has been evaluated for its impact on skin carcinogenesis. While an elimination of miR-21 does not result in any obvious consequences regarding development or day-to-day adult living in mice [89, 90], the absence of this tumor-associated microRNA has an impact on mice challenged with a chemical skin carcinogensis regimen. Mice deficient in miR-21 exhibit reduced incidences of tumors, slightly reduced rates of proliferation, and increased rates of apoptosis. Furthermore, keratinocytes utilize miR-21 to activate the Ras signaling pathway, as miR-21 inhibits PTEN, SPRY1, and other molecules involved in the negative control of Ras signaling [89]. In Ras-transformed keratinocytes, the downregulation of several tumor-related targets by miR-21 is actually enhanced, presumably because of a downregulation of DND1, an RNA-binding protein, during transformation [91]. Such findings establish that the elevated expression of miR-21 observed in all malignancies truly participates in tumor formation in contrast to merely serving as bystander noise.
Like miR-31, miR-21 is regarded as a TGF-β superfamily target gene, but while miR-21 is induced by TGF-β, it is repressed by BMP4 (Fig. 5) [92, 93].
Model of miR-21 function. miR-21, the prototypic oncomiR, promotes tumorigenesis by targeting the mRNA transcripts of molecules involved in the negative control of Ras signaling: PTEN, Spry1, and Grhl3. miR-21 is also upregulated by TGF-β and downregulated by BMP4. Conversely, competitive endogenous RNAs (ceRNAs) suppress tumorigenesis by protecting PTEN mRNA from microRNAs, such as miR-21
A separate set of experiments implicates miR-21 as a regulator of Grhl3, a member of the grainyhead family [94]. In turn, Grhl3 has been demonstrated to bind and repress the miR-21 promoter; thus, the two agents seem to function in a sort of regulatory homeostatic interplay [91]. A lack of Grhl3 results in increased tumor formation in chemically induced skin carcinogenesis studies through loss of PTEN regulation. PTEN and Grhl3 are both targets of miR-21, which is overexpressed in essentially all forms of squamous cell carcinoma. Indeed, while miR-21 is highly expressed in both squamous cell carcinoma and verrucous carcinoma, PTEN is downregulated in both, and the differential expression of p63 in verrucous carcinoma inversely correlates with levels of miR-21 [57]. Thus, miR-21 may play a role in skin carcinogenesis via direct targeting of PTEN or by targeting of one of its prime epidermal inducers, Grhl3 [94].
Melanocytes and melanoma
Like keratinocytes, melanocytes cannot grow in vitro without the microRNA-processing enzyme, Dicer. In turn, Dicer is under the control of MITF [95], the melanocyte lineage master gene, and can also be regulated by TAp63, an isoform of the keratinocytes’ master gene, p63 [96]. This regulation indicates that Dicer and microRNAs may serve as central signals in cutaneous cell lineage maintenance networks.
Unfortunately, the data on microRNAs and their role in melanocytes and melanoma are relatively limited. While an authoritative review of our current knowledge has recently been published [97], we would like to highlight a few significant findings (Fig. 6).
A meta-analysis of several studies on microRNAs in melanoma and melanocytes reveals the upregulation of several microRNAs in melanoma: miR-21 (the ubiquitous tumor marker), the miR-17-93/106 cluster (the prototypic oncomiR cluster), miR-214, and miR-155. The meta-analysis also reveals the downregulation of miR-211, miR-193b, and miR-196a.
However, the data set is difficult to interpret because of the wide variety of platforms and techniques utilized to determine these “melano-miRs.” In addition, studies of cultured melanocytic cells and real tumor samples are difficult to compare to one another, since real tissue samples retain an underlying complexity because of the presence of multiple cell types within a sample. For example, comparisons between in vivo and in vitro sources demonstrate a downregulation of miR-203, miR-205, and miR-23 in melanoma samples. This finding is not surprising considering that the three are epidermal microRNAs that dominate most skin samples (Table 2). Such “contamination” issues have plagued many melanoma microRNA studies. On the other hand, in vitro analyses of microRNAs may completely neglect microRNAs that play crucial roles in metastasis as well as microRNAs that are not expressed under in vitro conditions but play important roles in vivo for melanocytes and melanoma cells.
miR-214 is an example of the latter case. miR-214 exhibits pro-metastatic abilities, but its in vitro and in vivo expression patterns are confusing. While a few studies demonstrate a reduction of miR-214 levels in melanoma, other studies demonstrate an increased expression of miR-214. In vitro, miR-214 is upregulated during melanocyte differentiation but is also expressed at higher levels in melanoma cells than in melanocytes. At the same time, in an in vivo melanoma progression and metastasis model, miR-214 exhibits a correlation with the metastatic phenotype and could confer metastatic potential by interfering with the AP2γ (TFAP2C) gene expression program [98].
The situation is slightly clearer with miR-211, which is primarily expressed in pigmented tissues, making it the best bet for a “melano-miR” [99]. Most reports describe a reduction of its expression in melanoma versus nevi as well as in melanoma cells versus melanocytes. miR-211 is also upregulated during melanocyte differentiation.
Two more microRNAs frequently overexpressed in tumors, miR-221 and miR-222, are also overexpressed in melanoma. In general, miR-221 and miR-222 are associated with more aggressive melanoma cells [100]. One of miR-221’s confirmed target genes is c-KIT [101, 102]. An SNP in a conserved miR-221 binding site in the 3′UTR of c-KIT is associated with an increased risk of acral melanoma. This same variant is also associated with higher levels of c-KIT in melanoma samples as well as conferred “resistance” to the activity of miR-221. Thus, this c-KIT 3′UTR “mutant” allele may serve as a critical contributor to melanomagenesis in this subset of melanoma [103].
In general, however, c-KIT expression levels are reduced in melanoma samples and in melanoma tumor cell lines, with miR-221/222 serving as major contributors to this reduction. In turn, the expression of miR-221/222 depends upon the loss of promyelocytic leukemia zinc finger (PLZF), a strong repressor of miR-221/222 expression.
As a final note, the miR-17-92 cluster is also overexpressed in more aggressive melanoma cells and increases their proliferation rate [100, 101]. The miR-17 family of microRNAs is notorious for its complex distribution throughout the genome in three polycistronic microRNA clusters containing members of three additional microRNA families (miR-19, miR-25, and miR-363) with four different seed sequences. Out of the three clusters, the miR-17–92 cluster serves as the most important in cancer based upon the widespread overexpression of the miR-17–92 cluster in various types of malignancies [104].
Basal Cell Carcinoma
While basal cell carcinoma (BCC) is by far the most common cancer in humans, research regarding the role of microRNAs in BCC has just begun. Sand et al. have been one of the first to document the differential expression of several microRNAs in BCC. Through analyses of microarray and quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR) data, the researchers characterized the BCC miRNome and discovered 16 statistically significant upregulated microRNAs as well as 10 statistically significant downregulated microRNAs in BCC biopsies versus normal skin samples. Several of these microRNAs, including miR-17, miR-20a, and miR-92a, are associated with known tumorigenesis pathways, such as the MAPK/ERK signaling cascade [105]. However, this represents just the tip of the iceberg, and we await additional studies on the role of these microRNAs in BCC.
Cutaneous T cell lymphoma: microRNA signatures as diagnostic tools?
Ralfkiaer et al. also utilized microarray and qRT-PCR technology in order to develop a differential expression profile for cutaneous T cell lymphomas (CTCLs); however, the researchers took their work one step further and actually applied their findings clinically in the form of diagnostic profiling. CTCLs are the most prevalent primary cutaneous lymphomas, and early diagnosis of the disease can be an issue because of its similar appearance (both grossly and histologically) to other pathologies. After identifying several upregulated (miR-326, miR-663b, miR-711, and miR-155) and downregulated (miR-203 and miR-205) microRNAs in CTCLs versus other cutaneous diseases, the researchers developed a qRT-PCR-based classifier consisting of miR-155, miR-203, and miR-205, capable of differentiating CTCLs from other cutaneous pathologies with high accuracy [106]. Their publication is among the first to demonstrate the practical potential of developing and utilizing disease-specific microRNA classifiers in a clinical setting.
Another recent example involves work on microRNA alterations in different subtypes of melanoma. Researchers at the New York University School of Medicine developed a microRNA signature that differentiates between the two most prevalent melanoma histological subtypes: nodular and superficial spreading melanoma [107]. Such examples demonstrate that, in addition to their potential as future therapeutic targets, microRNAs may serve a useful clinical function even today. Inasmuch as different diseases are likely to have very different microRNA expression profiles, such microRNA classifiers could function as relatively simple and practical disease markers with high diagnostic potential.
Alterations of the microRNA machinery in skin disease
In addition to the differences in microRNA expression profiles of various dermatologic diseases, expression levels of the genes responsible for microRNA processing and the RISC have been evaluated in several cutaneous pathologies. Thus far, the data are complex and deviate from the simple notion that a reduction of Dicer levels is beneficial for cancer formation [108, 109].
For example, in actinic keratosis, basal cell carcinoma, and squamous cell carcinoma, the microprocessor complex component DGCR8 and several components of the RISC were found to be significantly upregulated compared to normal skin controls [110]. Drosha too was demonstrated to be upregulated in both basal cell carcinoma and squamous cell carcinoma, while Dicer expression levels were found to be significantly lower in basal cell carcinoma compared to normal skin controls [111].
Regarding melanoma, some components of the RISC (Argonaute-1, TARBP2, and SND1) were downregulated in primary cutaneous melanoma versus benign melanocytic nevi; however, two of these same components (TARBP2 and SND1) were actually upregulated in melanoma metastases versus benign melanocytic nevi [112].
A separate group of researchers, Ma et al., have demonstrated an upregulation of Dicer expression in cutaneous melanoma [113], and Jafarnejad et al. examined such aberrations in closer detail to find that Dicer expression is essential for the inhibition of melanoma cell invasion. The latter researchers found that a knockdown of Dicer enhances melanoma cell invasion, and clinically speaking, Dicer expression has a negative correlation with melanoma disease progression. Interestingly, the researchers also demonstrated that Sox4, which is downregulated in metastatic melanoma, upregulates the expression of Dicer via binding to its promoter sequences [114].
On the other hand, increased expression levels of Dicer have been implicated as a potential molecular marker revealing a negative prognostic influence in some subtypes of cutaneous T cell lymphomas, including mycosis fungoides [115].
These findings highlight once again the complexities of microRNA biology. Although reduced levels of Dicer enhance tumorigenesis in animal models, and many human tumors show lower Dicer levels, the retention of Dicer activity seems important for the tumorigenic process. It should be noted, however, that tumors can be derived from Dicer-deficient cells [116]. Thus, it is still unclear as to what drives this haploinsufficient tumor suppressor pathway. More detailed studies on the status of Dicer in cutaneous malignancies may help better explain the biology of these tumors.
Holding the line: microRNAs contribute to the first line of defense against pathogens
The skin participates in innate immunity by functioning as the first physical barrier to impede the entry of infectious agents. In addition to serving as a mere hydrophobic barrier, the epidermis functions as the first guard of the immunosurveillance system and plays a role in adaptive immunity as well. Langherans cells, lymphocytes, mast cells, and resident dermal macrophages combine forces to make up skin-associated lymphoid tissue (SALT), which contributes to the defensive functions of the skin. Epidermal cells also have their own antibacterial weapons at their disposal, such as defensins and cathelicidins.
While the exact role of microRNAs in the battle against invaders remains unclear, evidence exists of their role in the immune system. MicroRNAs are implicated in the host response to bacteria and other pathogens [117]. MicroRNAs are implicated in managing essential features of inflammatory responses as well [118], such as the modulation of cytokines during skin inflammation [119]. Consequently, they are thought to play a role in inflammatory disorders of the skin, such as atopic dermatitis [120] and allergic contact dermatitis [121]. MicroRNAs are also implicated in the pathogenesis of several other disorders of the skin, such as psoriasis and scleroderma [122, 123]. In psoriasis, levels of miR-424 have been demonstrated to be markedly decreased and associated with keratinocyte hyperproliferation [124]; in scleroderma, miR-92a is associated with pathogenesis of the disease [125].
In Langherans cells of the skin, microRNAs are necessary for proper maturation, function, and maintenance. An absence of Dicer, and thus of microRNAs, results in greater turnover and apoptosis rates of Langherans cells in vivo. These Dicer-deficient cells are inhibited in their ability to stimulate T cells and to induce their proliferation [126].
Regarding dendritic cells in general, microRNAs have been shown to modulate tolerogenic properties. In particular, miR-23b has been shown to initiate tolerogenic dendritic cell activity and to stimulate the differentiation of T-regulatory cells in vitro. These responses are thought to be carried out through a downregulation of the Notch1 and NF-κB signaling pathways. Thus, microRNAs may also have potential as therapeutic targets in allergen immunotherapy [127]. Interestingly, miR-23b has also been demonstrated to be downregulated within inflammatory lesions of patients with rheumatoid arthritis or lupus. In human cell lines and mouse models, IL-17 is responsible for this suppression of miR-23b and thus contributes to the autoimmune pathogenesis of these diseases [128].
Exosomes: microRNAs in intercellular signaling
In order to magnify immune effects, dendritic cells communicate with one another through various mediums, including direct cell-to-cell communication, soluble mediators (proteins), and vesicle exchange. The latter includes the transfer of exosomes, nanovesicles produced from an endocytic mechanism. These exosomes have been found to contain fully functional microRNAs, and their microRNA profiles differ significantly from those of their maternal cells [129]. Following fusion with the target cell membrane, these exogenous exosome-shuttle microRNAs are released into the cytosol and repress target mRNAs in order to regulate functions of the target dendritic cell [130].
Exosomes are generated by other cells of the immune system, such as mast cells, and have been shown to present viral antigens and activate immune cells during cellular responses [131, 132]. It has even been demonstrated that microRNAs exported from malignant cells are packaged differently than those released by normal cells [133]. It is also true that exosomes are present in the circulation and various biological fluids [134].
The origin of circulating microRNAs, however, is controversial. Through various assays, the Burwinkel laboratory previously claimed that the majority of circulating microRNAs are actually associated with the highly stable Ago2 protein, a part of the RNA-induced silencing complex (RISC), and are most likely mere remnants of dead cells [135]. Recently, though, the Illei laboratory demonstrated that the majority of microRNAs in both serum and saliva are in fact of exosomal origin. The researchers attribute the differences in findings to their superior method of exosome isolation and claim that proper exosome isolation is essential for sensitivity of detection [136]. Nevertheless, while the connotation behind “serum-derived microRNAs” remains under debate, the novel exosome-shuttle mechanism supports the notion that microRNAs may be utilized for intercellular signaling and may function as part of a signaling system with resemblance to hormones.
Beauty and microRNAs: the miRNome of aging
It is well known that aging is not skin deep, but in reality, lies far beneath the surface, affecting every system within our bodies and creating serious problems for the health care sectors of modern societies because of increasing average life expectancies. A dire need exists for contributions from the biomedical research community toward treatments of aging-associated diseases that threaten our continuously enlarging elderly population; while the skin is not regarded as a major contributor to the frailty of the aging process, it is definitely the organ that best epitomizes this slow process of decline [137]. Indeed, through hair loss, wrinkles, and pigmentation abnormalities, the skin fully conveys the long story of aging.
In addition to conveying the process, the skin contributes towards research on the subject. Studies on in vitro aging and senescence have been driven forward by the analyses of skin keratinocytes and fibroblasts. In vitro, mimicking aging in senescence assays results in very different sets of senescence-associated microRNAs in fibroblasts and in keratinocytes [50, 138]. Only miR-23b, miR-24, and miR-34 appear to be upregulated during the senescence program in both cell types, while microRNAs of the miR-17–92 and related clusters as well as miR-15/16 and miR-155 family members are negatively affected in fibroblasts. In vivo, the expression of microRNAs of the very same miR-17–92 clusters is reduced in human skin with age [139].
During senescence, the epigenome is altered with the formation of senescence-associated heterochromatin foci (SAHFs). A recent report indicates that microRNAs interact with Ago2 to repress RB1/E2F-target genes and contribute to the silencing of proliferation-associated genes [140]. However, the importance of such microRNA/Ago2-mediated transcriptional gene silencing is difficult to evaluate, since data exist indicating that a lack of Dicer, and thus a lack of microRNAs, induces senescence and upregulates p19(Arf)-p53 signaling [141]. In addition, SAHFs are unlikely universal markers for senescence [142].
As interesting as it may be to establish the role of microRNAs in embryogenesis, the faculty of microRNAs to stabilize cellular phenotypes may make them equally important for the maintenance of the body throughout adult life. The development of an organism or tissue during embryogenesis is a fast and complex process; however, once the organism is fully established, the lifelong maintenance of its health and fitness is the most important task at hand. G.C. Williams eloquently summarized this paradoxical inability of mammals more than 50 years ago: “It is indeed remarkable that after a seemingly miraculous feat of morphogenesis a complex metazoan should be unable to perform the much simpler task of merely maintaining what is already formed” [143].
In a sense, aging is merely the failure of keeping up with such a task. Frailty associated with aging reminisces upon a lack of robustness of the machinery in charge of homeostasis. If, as mentioned previously, microRNAs are believed to contribute to the robust nature of genetic programs, then perhaps a loss of proper microRNA function, and thus a loss of this robustness, results in the failure of carrying out the “much simpler task” (Fig. 7). Recent advances in our understanding of microRNAs in aging and cellular senescence that support this notion have been summarized by Vikos, Slack, Gorospe, and colleagues [144, 145].
As a final note, very few articles exist on the subject, but in C. elegans, several microRNAs have been implicated in aging and perform well as aging markers [146]. Perhaps more detailed studies on the role of microRNAs in aging will help our comprehension of troubles associated with the aging process. microRNAs are regarded as excellent markers for a plethora of cellular states and biological processes; therefore, determining the changes in the miRNome of aging tissues and cells may provide insights into the aging process itself.
Conclusions
Based on the notion that microRNAs function to fine tune and buffer gene expression, it is safe to assume that they are involved in complex biological processes. In the skin, the appendage best demonstrated as susceptible to aberrations in microRNA function is the hair follicle. This makes perfect sense. In addition to requiring a multifarious set of genetic programs involving a variety of cell types and lineages and producing a variety of differentiation states, hair follicle formation demands a high degree of gene expression control—control that could only be achieved with the help of microRNAs.
Mice have four different types of hair follicles, each fulfilling a crucial function in the combined effort to provide a magnificent shield against environmental challenges. Such a diverse and complex set of miniorgans surely requires a high degree of canalization or ability to produce a consistent phenotype, regardless of environmental or genotypic variability, during development. Otherwise, hair follicle types would get mixed up, spacing would be inaccurate, and differentiation would fail to initiate appropriately. As described previously, microRNAs have been implicated in maintaining this high degree of canalization. Thus, even an appendage as small as the hair follicle requires microRNAs—yes, putting hair on one’s chest requires microRNAs.
Indeed, while microRNAs were perhaps of less importance in the formation and maintenance of the ancient, simple epidermis, experimental evidence supports the notion that microRNAs serve as major players in hair follicle biology. Unfortunately, the hair follicle currently stands alone as a decently developed example of the role of microRNAs in the skin. While bits and pieces of the puzzle continue to be discovered, the exact involvement of microRNAs in the biology of the modern epidermis has yet to be determined.
In order to elucidate the role of individual microRNAs in the formation and maintenance of the epidermis, we must move away from our current gold standards and guides: Dicer, Drosha, and DRGC8 squamous epithelial-specific knockout mice. In reality, many factors influence the impact of an individual microRNA on gene expression control: the ratio between target concentration and microRNA concentration, competition with other microRNAs for RISCs, co-expression of functionally redundant microRNAs, and expression of decoys, among other variables. Therefore, Drosha, Dicer, and DRGC8 knockout models may misrepresent, at least to some extent, microRNA functions in the hair follicle and in the epidermis.
Given the diversity of roles that individual microRNAs can play in the skin processes and pathologies described throughout this review, microRNAs should serve major functions in the epidermis. While some progress has been made through analysis of miR-31, miR-203, miR-125b, and let-7, as we search for answers, we await loss-of-function studies involving miR-203, miR-205, other miR-200 family members, let-7, and other highly expressed epidermal microRNAs as well as comparisons of these phenotypes with those of previous Dicer, Drosha, and DRGC8 knockouts.
Just when we thought that we had a decent grip on the genetics of normal and pathological processes, the discovery of microRNAs reopened our eyes and revealed a novel layer of regulation finely interwoven among other transcriptional and posttranscriptional regulation systems. While we managed to clear the rain, the microRNA world, with its undiscovered complexities, continues to fog our view. As we return to our drawing boards, we must realize that our linear thought process in studying disease may be outdated. MicroRNAs are utilized by animals to reach a level of network control that requires careful reevaluation of our current perspective of how things work in our bodies. Given their association with various pathologies, the more we discover about these small, single-stranded RNA molecules, the closer we should come to their use in therapeutic applications for disease.
References
Little AC, Jones BC, DeBruine LM (2011) Facial attractiveness: evolutionary based research. Philos Trans R Soc Lond B Biol Sci 366(1571):1638–1659
Tucker P (2009) Bald is beautiful?: the psychosocial impact of alopecia areata. J Health Psychol 14(1):142–151
Mikkola ML, Millar SE (2006) The mammary bud as a skin appendage: unique and shared aspects of development. J Mammary Gland Biol Neoplasia 11(3–4):187–203
Hertel J, Lindemeyer M, Missal K, Fried C, Tanzer A, Flamm C, Hofacker IL, Stadler PF et al (2006) The expansion of the metazoan microRNA repertoire. BMC Genomics 7:25
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294(5543):853–858
Lau NC, Lim LP, Weinstein EG, Bartel DP (2001) An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294(5543):858–862
Wheeler BM, Heimberg AM, Moy VN, Sperling EA, Holstein TW, Heber S, Peterson KJ (2009) The deep evolution of metazoan microRNAs. Evol Dev 11(1):50–68
Warnefors M, Eyre-Walker A (2011) The accumulation of gene regulation through time. Genome Biol Evol 3:667–673
Wang QH, Zhou M, Sun J, Ning SW, Li Y, Chen L, Zheng Y, Li X, Lv SL, Li X (2011) Systematic analysis of human microRNA divergence based on evolutionary emergence. FEBS Lett 585(1):240–248
Yuan Z, Sun X, Liu H, Xie J (2011) MicroRNA genes derived from repetitive elements and expanded by segmental duplication eventsin mammalian genomes. PLoS ONE 6(3):e17666
Borchert GM, Holton NW, Williams JD, Hernan WL, Bishop IP, Dembosky JA, Elste JE, Gregoire NS, Kim JA, Koehler WW, Lengerich JC, Medema AA, Nguyen MA, Ower GD, Rarick MA, Strong BN, Tardi NJ, Tasker NM, Wozniak DJ, Gatto C, Larson ED (2011) Comprehensive analysis of microRNA genomic loci identifies pervasive repetitive-element origins. Mob Genet Elem 1(1):8–17
Loh YH, Yi SV, Streelman JT (2011) Evolution of microRNAs and the diversification of species. Genome Biol Evol 3:55–65
Wang D, Zhang Z, O’Loughlin E, Lee T, Houel S, O’Carroll D, Tarakhovsky A, Ahn NG, Yi R (2012) Quantitative functions of Argonaute proteins in mammalian development. Genes Dev 26(7):693–704
Li LC, Okino ST, Zhao H, Pookot D, Place RF, Urakami S, Enokida H, Dahiya R (2006) Small dsRNAs induce transcriptional activation in human cells. Proc Natl Acad Sci USA 103(46):17337–17342
Garcia DM, Baek D, Shin C, Bell GW, Grimson A, Bartel DP (2011) Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat Struct Mol Biol 18(10):1139–1146. doi:10.1038/nsmb.2115
Pontes O, Pikaard CS (2008) siRNA and microRNA processing: new functions for Cajal bodies. Curr Opin Genet Dev 18(2):197–203
Moser JJ, Fritzler MJ (2010) The microRNA and messenger RNA profile of the RNA-induced silencing complex in human primary astrocyte and astrocytoma cells. PLoS ONE 5(10):e13445
Dreher A, Rossing M, Kaczkowski B, Nielsen FC, Norrild B (2010) Differential expression of cellular microRNAs in HPV-11 transfected cells. An analysis by three different array platforms and qRT-PCR. Biochem Biophys Res Commun 403(3–4):357–362
Singh A (2011) Negative feedback through mRNA provides the best control of gene-expression noise. IEEE Trans Nanobiosci 10(3):194–200
Martello G, Zacchigna L, Inui M, Montagner M, Adorno M, Mamidi A, Morsut L, Soligo S, Tran U, Dupont S, Cordenonsi M, Wessely O, Piccolo S (2007) MicroRNA control of nodal signalling. Nature 449(7159):183–188
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP (2010) A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465(7301):1033–1038
Karreth FA, Tay Y, Perna D, Ala U, Tan SM, Rust AG, DeNicola G, Webster KA, Weiss D, Perez-Mancera PA, Krauthammer M, Halaban R, Provero P, Adams DJ, Tuveson DA, Pandolfi PP (2011) In vivo identification of tumor- suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell 147(2):382–395
Tay Y, Kats L, Salmena L, Weiss D, Tan SM, Ala U, Karreth F, Poliseno L, Provero P, Di Cunto F, Lieberman J, Rigoutsos I, Pandolfi PP (2011) Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 147(2):344–357
Sumazin P, Yang X, Chiu HS, Chung WJ, Iyer A, Llobet-Navas D, Rajbhandari P, Bansal M, Guarnieri P, Silva J, Califano A (2011) An extensive microRNA-mediated network of RNA–RNA interactions regulates established oncogenic pathways in glioblastoma. Cell 147(2):370–381
Franco-Zorrilla JM, Valli A, Todesco M, Mateos I, Puga MI, Rubio-Somoza I, Leyva A, Weigel D, García JA, Paz-Ares J (2007) Target mimicry provides a new mechanism for regulation of microRNA activity. Nat Genet 39(8):1033–1037
Jeyapalan Z, Yang BB (2012) The non-coding 3′UTR of CD44 induces metastasis by regulating extracellular matrix functions. J Cell Sci [Epub ahead of print]
Ambros V (2010) MicroRNAs: genetically sensitized worms reveal new secrets. Curr Biol 20(14):R598–R600
Di Giammartino DC, Nishida K, Manley JL (2011) Mechanisms and consequences of alternative polyadenylation. Mol Cell 43(6):853–866
Lutz CS, Moreira A (2011) Alternative mRNA polyadenylation in eukaryotes: an effective regulator of gene expression. Wiley Interdiscip Rev RNA 2(1):22–31. doi:10.1002/wrna.47
Léveillé N, Elkon R, Davalos V, Manoharan V, Hollingworth D, Oude Vrielink J, le Sage C, Melo CA, Horlings HM, Wesseling J, Ule J, Esteller M, Ramos A, Agami R (2011) Selective inhibition of microRNA accessibility by RBM38 is required for p53 activity. Nat Commun 2:513. doi:10.1038/ncomms1519
Libri V, Helwak A, Miesen P, Santhakumar D, Borger JG, Kudla G, Grey F, Tollervey D, Buck AH (2012) Murine cytomegalovirus encodes a miR-27 inhibitor disguised as a target. Proc Natl Acad Sci USA 109(1):279–284
Ji Q, Luo ZX, Yuan CX, Tabrum AR (2006) A swimming mammaliaform from the middle Jurassic and ecomorphological diversification of early mammals. Science 311(5764):1123–1127
Andl T, Murchison EP, Liu F, Zhang Y, Yunta-Gonzalez M, Tobias JW, Andl CD, Seykora JT, Hannon GJ, Millar SE (2006) The microRNA-processing enzyme dicer is essential for the morphogenesis and maintenance of hair follicles. Curr Biol 16(10):1041–1049
Yi R, O’Carroll D, Pasolli HA, Zhang Z, Dietrich FS, Tarakhovsky A, Fuchs E (2006) Morphogenesis in skin is governed by discrete sets of differentially expressed microRNAs. Nat Genet 38(3):356–362
Mardaryev AN, Ahmed MI, Vlahov NV, Fessing MY, Gill JH, Sharov AA, Botchkareva NV (2010) Micro-RNA-31 controls hair cycle-associated changes in gene expression programs of the skin and hair follicle. FASEB J 24(10):3869–3881
Yi R, Pasolli HA, Landthaler M, Hafner M, Ojo T, Sheridan R, Sander C, O’Carroll D, Stoffel M, Tuschl T, Fuchs E (2009) DGCR8-dependent microRNA biogenesis is essential for skin development. Proc Natl Acad Sci USA 106(2):498–502
Andl T (2007) microRNAs: miracle or mirage?: the limes against the barbaric floods of leaky and undesired transcripts. Organogenesis 3(1):25–33
Teta M, Choi YS, Okegbe T, Wong G, Tam OH, Chong MM, Seykora JT, Nagy A, Littman DR, Andl T, Millar SE (2012) Inducible deletion of epidermal Dicer and Drosha reveals multiple functions for miRNAs in postnatal skin. Development 139(8):1405–1416
Kuhnert F, Mancuso MR, Hampton J, Stankunas K, Asano T, Chen CZ, Kuo CJ (2008) Attribution of vascular phenotypes of the murine Egfl7 locus to the microRNA miR-126. Development 135(24):3989–3993
Lewis MA, Quint E, Glazier AM, Fuchs H, De Angelis MH, Langford C, van Dongen S, Abreu-Goodger C, Piipari M, Redshaw N, Dalmay T, Moreno-Pelayo MA, Enright AJ, Steel KP (2009) An ENU-induced mutation of miR-96 associated with progressive hearing loss in mice. Nat Genet 41(5):614–618
Shibata M, Nakao H, Kiyonari H, Abe T, Aizawa S (2011) MicroRNA-9 regulates neurogenesis in mouse telencephalon by targeting multiple transcription factors. J Neurosci 31(9):3407–3422
Park CY, Jeker LT, Carver-Moore K, Oh A, Liu HJ, Cameron R, Richards H, Li Z, Adler D, Yoshinaga Y, Martinez M, Nefadov M, Abbas AK, Weiss A, Lanier LL, de Jong PJ, Bluestone JA, Srivastava D, McManus MT (2012) A resource for the conditional ablation of microRNAs in the mouse. Cell Rep 1(4):385–391
Zhang L, Stokes N, Polak L, Fuchs E (2011) Specific microRNAs are preferentially expressed by skin stem cells to balance self-renewal and early lineage commitment. Cell Stem Cell 8(3):294–308
Joyce CE, Zhou X, Xia J, Ryan C, Thrash B, Menter A, Zhang W, Bowcock AM (2011) Deep sequencing of small RNAs from human skin reveals major alterations in the psoriasis microRNAome. Hum Mol Genet 20(20):4025–4040
Valastyan S, Reinhardt F, Benaich N, Calogrias D, Szász AM, Wang ZC, Brock JE, Richardson AL, Weinberg RA (2009) A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137(6):1032–1046
Valastyan S, Reinhardt F, Benaich N, Calogrias D, Szász AM, Wang ZC, Brock JE, Richardson AL, Weinberg RA (2010) Pro-tumorigenic effects of miR-31 loss in mesothelioma. J Biol Chem 285(30):22809–22817
Sonkoly E, Ståhle M, Pivarcsi A (2008) MicroRNAs: novel regulators involved in the pathogenesis of psoriasis? Clin Exp Dermatol 33(3):312–315
Yi R, Poy MN, Stoffel M, Fuchs E (2008) A skin microRNA promotes differentiation by repressing ‘stemness’. Nature 452(7184):225–229
Ory B, Ramsey MR, Wilson C, Vadysirisack DD, Forster N, Rocco JW, Rothenberg SM, Ellisen LW (2011) A microRNA-dependent program controls p53-independent survival and chemosensitivity in human and murine squamous cell carcinoma. J Clin Invest 121(2):809–820
di Val Rivetti, Cervo P, Lena AM, Nicoloso M, Rossi S, Mancini M, Zhou H, Saintigny G, Dellambra E, Odorisio T, Mahé C, Calin GA, Candi E, Melino G (2012) p63-microRNA feedback in keratinocyte senescence. Proc Natl Acad Sci USA 109(4):1133–1138
Antonini D, Russo MT, De Rosa L, Gorrese M, Del Vecchio L, Missero C (2010) Transcriptional repression of miR-34 family contributes to p63-mediated cell cycle progression in epidermal cells. J Invest Dermatol 130(5):1249–1257
Knouf EC, Garg K, Arroyo JD, Correa Y, Sarkar D, Parkin RK, Wurz K, O’Briant KC, Godwin AK, Urban ND, Ruzzo WL, Gentleman R, Drescher CW, Swisher EM, Tewari M (2012) An integrative genomic approach identifies p73 and p63 as activators of miR-200 microRNA family transcription. Nucl Acids Res 40(2):499–510
Chikh A, Matin RN, Senatore V, Hufbauer M, Lavery D, Raimondi C, Ostano P, Mello-Grand M, Ghimenti C, Bahta A, Khalaf S, Akgül B, Braun KM, Chiorino G, Philpott MP, Harwood CA, Bergamaschi D (2011) iASPP/p63 autoregulatory feedback loop is required for the homeostasis of stratified epithelia. EMBO J 30(20):4261–4273. doi:10.1038/emboj.2011.302
Ambros V (2011) microRNAs and developmental timing. Curr Opin Genet Dev 21(4):511–517
Glud M, Manfé V, Biskup E, Holst L, Dirksen AM, Hastrup N, Nielsen FC, Drzewiecki KT, Gniadecki R (2011) microRNA miR-125b induces senescence in human melanoma cells. Melanoma Res 21(3):253–256
Xu N, Brodin P, Wei T, Meisgen F, Eidsmo L, Nagy N, Kemeny L, Ståhle M, Sonkoly E, Pivarcsi A (2011) miR-125b, a microRNA downregulated in psoriasis, modulates keratinocyte proliferation by targeting FGFR2. J Invest Dermatol 131(7):1521–1529. doi:10.1038/jid.2011.55
Odar K, Boštjančič E, Gale N, Glavač D, Zidar N (2012) Differential expression of microRNAs miR-21, miR-31, miR-203, miR-125a-5p and miR-125b and proteins PTEN and p63 in verrucous carcinoma of the head and neck. Histopathology 61(2):257–265. doi:10.1111/j.1365-2559.2012.04242.x
Xu N, Zhang L, Meisgen F, Harada M, Heilborn J, Homey B, Grander D, Stahle M, Sonkoly E, Pivarcsi A (2012) microRNA-125b down-regulates matrix metallopeptidase 13 and inhibits cutaneous squamous cell carcinoma cell proliferation, migration and invasion. J Biol Chem [Epub ahead of print]
Le MT, Shyh-Chang N, Khaw SL, Chin L, Teh C, Tay J, O’Day E, Korzh V, Yang H, Lal A, Lieberman J, Lodish HF, Lim B (2011) Conserved regulation of p53 network dosage by microRNA-125b occurs through evolving miRNA-target gene pairs. PLoS Genet 7(9):e1002242
Johnson CD, Esquela-Kerscher A, Stefani G, Byrom M, Kelnar K, Ovcharenko D, Wilson M, Wang X, Shelton J, Shingara J, Chin L, Brown D, Slack FJ (2007) The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 67(16):7713–7722
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A, Labourier E, Reinert KL, Brown D, Slack FJ (2005) RAS is regulated by the let-7 microRNA family. Cell 120(5):635–647
Park SM, Shell S, Radjabi AR, Schickel R, Feig C, Boyerinas B, Dinulescu DM, Lengyel E, Peter ME (2007) Let-7 prevents early cancer progression by suppressing expression of the embryonic gene HMGA2. Cell Cycle 6(21):2585–2590
Büssing I, Slack FJ (2008) Grosshans H (2008) let-7 microRNAs in development, stem cells and cancer. Trends Mol Med 14(9):400–409
Rybak A, Fuchs H, Hadian K, Smirnova L, Wulczyn EA, Michel G, Nitsch R, Krappmann D, Wulczyn FG (2009) The let-7 target gene mouse lin-41 is a stem cell specific E3 ubiquitin ligase for the microRNA pathway protein Ago2. Nat Cell Biol 11(12):1411–1420
Viswanathan SR, Daley GQ, Gregory RI (2008) Selective blockade of microRNA processing by Lin28. Science 320(5872):97–100
Bourcier C, Griseri P, Grépin R, Bertolotto C, Mazure N, Pagès G (2011) Constitutive ERK activity induces downregulation of tristetraprolin, a major protein controlling interleukin8/CXCL8 mRNA stability in melanoma cells. Am J Physiol Cell Physiol 301(3):C609–C618
Van Tubergen E, Vander Broek R, Lee J, Wolf G, Carey T, Bradford C, Prince M, Kirkwood KL, D’Silva NJ (2011) Tristetraprolin regulates interleukin-6, which is correlated with tumor progression in patients with head and neck squamous cell carcinoma. Cancer 117(12):2677–2689. doi:10.1002/cncr.25859
Alvarez-Saavedra E, Horvitz HR (2010) Many families of C. elegans microRNAs are not essential for development or viability. Curr Biol 20(4):367–373
Brenner JL, Jasiewicz KL, Fahley AF, Kemp BJ, Abbott AL (2010) Loss of individual microRNAs causes mutant phenotypes in sensitized genetic backgrounds in C. elegans. Curr Biol 20(14):1321–1325
Concepcion CP, Han YC, Mu P, Bonetti C, Yao E, D’Andrea A, Vidigal JA, Maughan WP, Ogrodowski P, Ventura A (2012) Intact p53-dependent responses in miR-34-deficient mice. PLoS Genet 8(7):e1002797
Kierdorf U, Li C, Price JS (2009) Improbable appendages: deer antler renewal as a unique case of mammalian regeneration. Semin Cell Dev Biol 20(5):535–542
Thatcher EJ, Patton JG (2010) Small RNAs have a big impact on regeneration. RNA Biol 7(3):333–338
Yin VP, Thomson JM, Thummel R, Hyde DR, Hammond SM, Poss KD (2008) Fgf-dependent depletion of microRNA-133 promotes appendage regeneration in zebrafish. Genes Dev 22(6):728–733
Sehm T, Sachse C, Frenzel C, Echeverri K (2009) miR-196 is an essential early-stage regulator of tail regeneration, upstream of key spinal cord patterning events. Dev Biol 334(2):468–480
Sehm T, Sachse C, Frenzel C, Echeverri K (2009) miR-196 is an essential early-stage regulator of tail regeneration, upstream of key spinal cord patterning events. Dev Biol 334(2):468–480
Biswas S, Roy S, Banerjee J, Hussain SR, Khanna S, Meenakshisundaram G, Kuppusamy P, Friedman A, Sen CK (2010) Hypoxia inducible microRNA 210 attenuates keratinocyte proliferation and impairs closure in a murine model of ischemic wounds. Proc Natl Acad Sci USA 107(15):6976–6981
Kashiyama K, Mitsutake N, Matsuse M, Ogi T, Saenko VA, Ujifuku K, Utani A, Hirano A, Yamashita S (2012) miR-196a downregulation increases the expression of type I and III collagens in keloid fibroblasts. J Invest Dermatol 132(6):1597–1604. doi:10.1038/jid.2012.22
Madhyastha R, Madhyastha H, Nakajima Y, Omura S, Maruyama M (2011) MicroRNA signature in diabetic wound healing: promotive role of miR-21 in fibroblast migration. Int Wound J 9(4):355–361. doi:10.1111/j.1742-481X.2011.00890.x
Yan C, Grimm WA, Garner WL, Qin L, Travis T, Tan N, Han YP (2010) Epithelial to mesenchymal transition in human skin wound healing is induced by tumor necrosis factor-alpha through bone morphogenic protein-2. Am J Pathol 176(5):2247–2258
Knouf EC, Garg K, Arroyo JD, Correa Y, Sarkar D, Parkin RK, Wurz K, O’Briant KC, Godwin AK, Urban ND, Ruzzo WL, Gentleman R, Drescher CW, Swisher EM, Tewari M (2012) An integrative genomic approach identifies p73 and p63 as activators of miR-200 microRNA family transcription. Nucleic Acids Res 40(2):499–510
Ohashi S, Natsuizaka M, Naganuma S, Kagawa S, Kimura S, Itoh H, Kalman RA, Nakagawa M, Darling DS, Basu D, Gimotty PA, Klein-Szanto AJ, Diehl JA, Herlyn M, Nakagawa H (2011) A NOTCH3-mediated squamous cell differentiation program limits expansion of EMT competent cells that express the ZEB transcription factors. Cancer Res 71(21):6836–6847
Elson-Schwab I, Lorentzen A, Marshall CJ (2010) MicroRNA-200 family members differentially regulate morphological plasticity and mode of melanoma cell invasion. PLoS One 5(10). pii: e13176
Dykxhoorn DM, Wu Y, Xie H, Yu F, Lal A, Petrocca F, Martinvalet D, Song E, Lim B, Lieberman J (2009) miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS ONE 4(9):e7181
Sun L, Yao Y, Liu B, Lin Z, Lin L, Yang M, Zhang W, Chen W, Pan C, Liu Q, Song E, Li J (2012) miR-200b and miR-15b regulate chemotherapy-induced epithelial-mesenchymal transition in human tongue cancer cells by targeting BMI1. Oncogene 31(4):432–445. doi:10.1038/onc.2011.263
Janecka IP (2007) Cancer control through principles of systems science, complexity, and chaos theory: a model. Int J Med Sci 4(3):164–173
Herranz H, Cohen SM (2010) MicroRNAs and gene regulatory networks: managing the impact of noise in biological systems. Genes Dev 24(13):1339–1344
Watt SA, Pourreyron C, Purdie K, Hogan C, Cole CL, Foster N, Pratt N, Bourdon JC, Appleyard V, Murray K, Thompson AM, Mao X, Mein C, Bruckner-Tuderman L, Evans A, McGrath JA, Proby CM, Foerster J, Leigh IM, South AP (2011) Integrative mRNA profiling comparing cultured primary cells with clinical samples reveals PLK1 and C20orf20 as therapeutic targets in cutaneous squamous cell carcinoma. Oncogene 30(46):4666–4677. doi:10.1038/onc.2011.180
Dziunycz P, Iotzova-Weiss G, Eloranta JJ, Läuchli S, Hafner J, French LE, Hofbauer GF (2010) Squamous cell carcinoma of the skin shows a distinct microRNA profile modulated by UV radiation. J Invest Dermatol 130(11):2686–2689
Ma X, Kumar M, Choudhury SN, Becker Buscaglia LE, Barker JR, Kanakamedala K, Liu MF, Li Y (2011) Loss of the miR-21 allele elevates the expression of its target genes and reduces tumorigenesis. Proc Natl Acad Sci USA 108(25):10144–10149
Hatley ME, Patrick DM, Garcia MR, Richardson JA, Bassel-Duby R, van Rooij E, Olson EN (2010) Modulation of K-Ras-dependent lung tumorigenesis by microRNA-21. Cancer Cell 18(3):282–293
Bhandari A, Gordon W, Dizon D, Hopkin AS, Gordon E, Yu Z, Andersen B (2012) The Grainyhead transcription factor Grhl3/Get1 suppresses miR-21 expression and tumorigenesisin skin: modulation of the miR-21 target MSH2 by RNA-binding protein DND1. Oncogene. doi:10.1038/onc.2012.168
Ahmed MI, Mardaryev AN, Lewis CJ, Sharov AA, Botchkareva NV (2011) microRNA-21 is an important downstream component of BMP signalling in epidermal keratinocytes. J Cell Sci 124(Pt 20):3399–3404
Zavadil J, Narasimhan M, Blumenberg M, Schneider RJ (2007) Transforming growth factor-beta and microRNA: mRNA regulatory networks in epithelial plasticity. Cells Tissues Organs 185(1–3):157–161
Darido C, Georgy SR, Wilanowski T, Dworkin S, Auden A, Zhao Q, Rank G, Srivastava S, Finlay MJ, Papenfuss AT, Pandolfi PP, Pearson RB, Jane SM (2011) Targeting of the tumor suppressor GRHL3 by a miR-21-dependent proto-oncogenic network results in PTEN loss and tumorigenesis. Cancer Cell 20(5):635–648
Levy C, Khaled M, Robinson KC, Veguilla RA, Chen PH, Yokoyama S, Makino E, Lu J, Larue L, Beermann F, Chin L, Bosenberg M, Song JS, Fisher DE (2010) Lineage-specific transcriptional regulation of DICER by MITF in melanocytes. Cell 141(6):994–1005
Su X, Chakravarti D, Cho MS, Liu L, Gi YJ, Lin YL, Leung ML, El-Naggar A, Creighton CJ, Suraokar MB, Wistuba I, Flores ER (2010) TAp63 suppresses metastasis through coordinate regulation of Dicer and microRNAs. Nature 467(7318):986–990
Bonazzi VF, Stark MS, Hayward NK (2012) microRNA regulation of melanoma progression. Melanoma Res 22(2):101–113
Penna E, Orso F, Cimino D, Tenaglia E, Lembo A, Quaglino E, Poliseno L, Haimovic A, Osella-Abate S, De Pittà C, Pinatel E, Stadler MB, Provero P, Bernengo MG, Osman I, Taverna D (2011) microRNA-214 contributes to melanoma tumour progression through suppression of TFAP2C. EMBO J 30(10):1990–2007
Boyle GM, Woods SL, Bonazzi VF, Stark MS, Hacker E, Aoude LG, Dutton-Regester K, Cook AL, Sturm RA, Hayward NK (2011) Melanoma cell invasiveness is regulated by miR-211 suppression of the BRN2 transcription factor. Pigment Cell Melanoma Res 24(3):525–537. doi:10.1111/j.1755-148X.2011.00849.x
Greenberg E, Hershkovitz L, Itzhaki O, Hajdu S, Nemlich Y, Ortenberg R, Gefen N, Edry L, Modai S, Keisari Y, Besser MJ, Schachter J, Shomron N, Markel G (2011) Regulation of cancer aggressive features in melanoma cells by microRNAs. PLoS ONE 6(4):e18936
Igoucheva O, Alexeev V (2009) MicroRNA-dependent regulation of cKit in cutaneous melanoma. Biochem Biophys Res Commun 379(3):790–794
Felicetti F, Errico MC, Bottero L, Segnalini P, Stoppacciaro A, Biffoni M, Felli N, Mattia G, Petrini M, Colombo MP, Peschle C, Carè A (2008) The promyelocytic leukemia zinc finger–microRNA-221/-222 pathway controls melanoma progression through multiple oncogenic mechanisms. Cancer Res 68(8):2745–2754
Godshalk SE, Paranjape T, Nallur S, Speed W, Chan E, Molinaro AM, Bacchiocchi A, Hoyt K, Tworkoski K, Stern DF, Sznol M, Ariyan S, Lazova R, Halaban R, Kidd KK, Weidhaas JB, Slack FJ (2011) A variant in a microRNA complementary site in the 3′UTR of the KIT oncogene increases risk of acral melanoma. Oncogene 30(13):1542–1550
Volinia S, Galasso M, Costinean S, Tagliavini L, Gamberoni G, Drusco A, Marchesini J, Mascellani N, Sana ME, Abu Jarour R, Desponts C, Teitell M, Baffa R, Aqeilan R, Iorio MV, Taccioli C, Garzon R, Di Leva G, Fabbri M, Catozzi M, Previati M, Ambs S, Palumbo T, Garofalo M, Veronese A, Bottoni A, Gasparini P, Harris CC, Visone R, Pekarsky Y, de la Chapelle A, Bloomston M, Dillhoff M, Rassenti LZ, Kipps TJ, Huebner K, Pichiorri F, Lenze D, Cairo S, Buendia MA, Pineau P, Dejean A, Zanesi N, Rossi S, Calin GA, Liu CG, Palatini J, Negrini M, Vecchione A, Rosenberg A, Croce CM (2010) Reprogramming of microRNA networks in cancer and leukemia. Genome Res 20(5):589–599
Sand M, Skrygan M, Sand D, Georgas D, Hahn S, Gambichler T, Altmeyer P, Bechara FG (2012) Expression of microRNAs in basal cell carcinoma. Br J Dermatol. doi:10.1111/j.1365-2133.2012.11022.x
Ralfkiaer U, Hagedorn PH, Bangsgaard N, Løvendorf MB, Ahler CB, Svensson L, Kopp KL, Vennegaard MT, Lauenborg B, Zibert JR, Krejsgaard T, Bonefeld CM, Søkilde R, Gjerdrum LM, Labuda T, Mathiesen AM, Grønbæk K, Wasik MA, Sokolowska-Wojdylo M, Queille-Roussel C, Gniadecki R, Ralfkiaer E, Geisler C, Litman T, Woetmann A, Glue C, Røpke MA, Skov L, Odum N (2011) Diagnostic microRNA profiling in cutaneous T-cell lymphoma (CTCL). Blood 118(22):5891–5900
Poliseno L, Haimovic A, Segura MF, Hanniford D, Christos PJ, Darvishian F, Wang J, Shapiro RL, Pavlick AC, Berman RS, Hernando E, Zavadil J, Osman I (2012) Histology-Specific microRNA alterations in melanoma. J Invest Dermatol 132(7):1860–1868. doi:10.1038/jid.2011.451
Kumar MS, Pester RE, Chen CY, Lane K, Chin C, Lu J, Kirsch DG, Golub TR, Jacks T (2009) Dicer1 functions as a haploinsufficient tumor suppressor. Genes Dev 23(23):2700–2704
Lambertz I, Nittner D, Mestdagh P, Denecker G, Vandesompele J, Dyer MA, Marine JC (2010) Monoallelic but not biallelic loss of Dicer1 promotes tumorigenesis in vivo. Cell Death Differ 17(4):633–641
Sand M, Skrygan M, Georgas D, Arenz C, Gambichler T, Sand D, Altmeyer P, Bechara FG (2011) Expression levels of the microRNA maturing microprocessor complex component DGCR8 and the RNA-induced silencing complex (RISC) components Argonaute-1, Argonaute-2, PACT, TARBP1, and TARBP2 in epithelial skin cancer. Mol Carcinog. doi:10.1002/mc.20861
Sand M, Gambichler T, Skrygan M, Sand D, Scola N, Altmeyer P, Bechara FG (2010) Expression levels of the microRNA processing enzymes Drosha and dicer in epithelial skin cancer. Cancer Invest 28(6):649–653
Sand M, Skrygan M, Georgas D, Sand D, Gambichler T, Altmeyer P, Bechara FG (2012) The miRNA machinery in primary cutaneous malignant melanoma, cutaneous malignant melanoma metastases and benign melanocytic nevi. Cell Tissue Res [Epub ahead of print]
Ma Z, Swede H, Cassarino D, Fleming E, Fire A, Dadras SS (2011) Up-regulated dicer expression in patients with cutaneous melanoma. PLoS ONE 6(6):e20494
Jafarnejad SM, Ardekani GS, Ghaffari M, Martinka M, Li G (2012) Sox4-mediated dicer expression is critical for suppression of melanoma cell invasion. Oncogene. doi:10.1038/onc.2012.239
Valencak J, Schmid K, Trautinger F, Wallnöfer W, Muellauer L, Soleiman A, Knobler R, Haitel A, Pehamberger H, Raderer M (2011) High expression of dicer reveals a negative prognostic influence in certain subtypes of primary cutaneous T cell lymphomas. J Dermatol Sci 64(3):185–190
Ravi A, Gurtan AM, Kumar MS, Bhutkar A, Chin C, Lu V, Lees JA, Jacks T, Sharp PA (2012) Proliferation and tumorigenesis of a murine sarcoma cell line in the absence of DICER1. Cancer Cell 21(6):848–855
Eulalio A, Schulte L, Vogel J (2012) The mammalian microRNA response to bacterial infections. RNA Biol 9(6) [Epub ahead of print]
O’Connell RM, Rao DS, Baltimore D (2011) microRNA regulation of inflammatory responses. Annu Rev Immunol 30:295–312
Bak RO, Mikkelsen JG (2010) Regulation of cytokines by small RNAs during skin inflammation. J Biomed Sci 17:53
Sonkoly E, Janson P, Majuri ML, Savinko T, Fyhrquist N, Eidsmo L, Xu N, Meisgen F, Wei T, Bradley M, Stenvang J, Kauppinen S, Alenius H, Lauerma A, Homey B, Winqvist O, Ståhle M, Pivarcsi A (2010) miR-155 is overexpressed in patients with atopic dermatitis and modulates T-cell proliferative responses by targeting cytotoxic T lymphocyte-associated antigen 4. J Allergy Clin Immunol 126(3):581–589 e1-20
Vennegaard MT, Bonefeld CM, Hagedorn PH, Bangsgaard N, Løvendorf MB, Odum N, Woetmann A, Geisler C, Skov L (2012) Allergic contact dermatitis induces upregulation of identical microRNAs in humans and mice. Contact Dermatitis. doi:10.1111/j.1600-0536.2012.02083.x
Zibert JR, Løvendorf MB, Litman T, Olsen J, Kaczkowski B, Skov L (2010) MicroRNAs and potential target interactions in psoriasis. J Dermatol Sci 58(3):177–185
Zhu H, Li Y, Qu S, Luo H, Zhou Y, Wang Y, Zhao H, You Y, Xiao X, Zuo X (2012) MicroRNA expression abnormalities in limited cutaneous scleroderma and diffuse cutaneous scleroderma. J Clin Immunol 32(3):514–522
Ichihara A, Jinnin M, Yamane K, Fujisawa A, Sakai K, Masuguchi S, Fukushima S, Maruo K, Ihn H (2011) microRNA-mediated keratinocyte hyperproliferation in psoriasis vulgaris. Br J Dermatol 165(5):1003–1010. doi:10.1111/j.1365-2133.2011.10497.x
Sing T, Jinnin M, Yamane K, Honda N, Makino K, Kajihara I, Makino T, Sakai K, Masuguchi S, Fukushima S, Ihn H (2012) microRNA-92a expression in the sera and dermal fibroblasts increases in patients with scleroderma. Rheumatology (Oxford) [Epub ahead of print]
Kuipers H, Schnorfeil FM, Fehling HJ, Bartels H, Brocker T (2010) Dicer-dependent microRNAs control maturation, function, and maintenance of Langerhans cells in vivo. J Immunol 185(1):400–409
Zheng J, Jiang HY, Li J, Tang HC, Zhang XM, Wang XR, Du JT, Li HB, Xu G (2012) MicroRNA-23b promotes tolerogenic properties of dendritic cells in vitro through inhibiting Notch1/NF-kB signaling pathways. Allergy 67(3):362–370. doi:10.1111/j.1398-9995.2011.02776.x
Zhu S, Pan W, Song X, Liu Y, Shao X, Tang Y, Liang D, He D, Wang H, Liu W, Shi Y, Harley JB, Shen N, Qian Y (2012) The microRNA miR-23b suppresses IL-17-associated autoimmune inflammation by targeting TAB 2, TAB 3 and IKK-α. Nat Med. doi:10.1038/nm.2815
Diehl P, Fricke A, Sander L, Stamm J, Bassler N, Htun N, Ziemann M, Helbing T, El-Osta A, Jowett JB, Peter K (2012) Microparticles: major transport vehicles for distinct microRNAs in circulation. Cardiovasc Res 93(4):633–644
Montecalvo A, Larregina AT, Shufesky WJ, Stolz DB, Sullivan ML, Karlsson JM, Baty CJ, Gibson GA, Erdos G, Wang Z, Milosevic J, Tkacheva OA, Divito SJ, Jordan R, Lyons-Weiler J, Watkins SC, Morelli AE (2012) Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood 119(3):756–766
Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9(6):654–659
Zomer A, Vendrig T, Hopmans ES, van Eijndhoven M, Middeldorp JM, Pegtel DM (2010) Exosomes: fit to deliver small RNA. Commun Integr Biol 3(5):447–450
Palma J, Yaddanapudi SC, Pigati L, Havens MA, Jeong S, Weiner GA, Weimer KM, Stern B, Hastings ML, Duelli DM (2012) MicroRNAs are exported from malignant cells in customized particles. Nucleic Acids Res [Epub ahead of print]
Diehl P, Fricke A, Sander L, Stamm J, Bassler N, Htun N, Ziemann M, Helbing T, El-Osta A, Jowett JB, Peter K (2012) Microparticles: major transport vehicles for distinct microRNAs in circulation. Cardiovasc Res 93(4):633–644
Turchinovich A, Weiz L, Langheinz A, Burwinkel B (2011) Characterization of extracellular circulating microRNA. Nucleic Acids Res 39(16):7223–7233
Gallo A, Tandon M, Alevizos I, Illei GG (2012) The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 7(3):e30679
Matts PJ, Fink B (2010) Chronic sun damage and the perception of age, health and attractiveness. Photochem Photobiol Sci 9(4):421–431
Faraonio R, Salerno P, Passaro F, Sedia C, Iaccio A, Bellelli R, Nappi TC, Comegna M, Romano S, Salvatore G, Santoro M, Cimino F (2012) A set of microRNAs participates in the cellular senescence program in human diploid fibroblasts. Cell Death Differ 19(4):713–721. doi:10.1038/cdd.2011.143
Hackl M, Brunner S, Fortschegger K, Schreiner C, Micutkova L, Mück C, Laschober GT, Lepperdinger G, Sampson N, Berger P, Herndler-Brandstetter D, Wieser M, Kühnel H, Strasser A, Rinnerthaler M, Breitenbach M, Mildner M, Eckhart L, Tschachler E, Trost A, Bauer JW, Papak C, Trajanoski Z, Scheideler M, Grillari-Voglauer R, Grubeck-Loebenstein B, Jansen-Dürr P, Grillari J (2010) miR-17, miR-19b, miR-20a, and miR-106a are down-regulated in human aging. Aging Cell 9(2):291–296
Benhamed M, Herbig U, Ye T, Dejean A, Bischof O (2012) Senescence is an endogenous trigger for microRNA-directed transcriptional gene silencing in human cells. Nat Cell Biol 14(3):266–275. doi:10.1038/ncb2443
Mudhasani R, Zhu Z, Hutvagner G, Eischen CM, Lyle S, Hall LL, Lawrence JB, Imbalzano AN, Jones SN (2008) Loss of microRNA biogenesis induces p19Arf-p53 signaling an senescence in primary cells. J Cell Biol 181(7):1055–1063
Kosar M, Bartkova J, Hubackova S, Hodny Z, Lukas J, Bartek J (2011) Senescence-associated heterochromatin foci are dispensable for cellular senescence, occur in a cell type- and insult-dependent manner and follow expression of p16(ink4a). Cell Cycle 10(3):457–468
Williams GC (1957) Pleiotropy, natural selection, and the evolution of senescence. Evolution 11(4):398–411
Smith-Vikos T, Slack FJ (2012) MicroRNAs and their roles in aging. J Cell Sci 125(Pt 1):7–17
Abdelmohsen K, Srikantan S, Kang MJ, Gorospe M (2012) Regulation of senescence by microRNA biogenesis factors. Ageing Res Rev [Epub ahead of print]
Pincus Z, Smith-Vikos T, Slack FJ (2011) MicroRNA predictors of longevity in Caenorhabditis elegans. PLoS Genet 7(9):e1002306
Soares AR, Pereira PM, Santos B, Egas C, Gomes AC, Arrais J, Oliveira JL, Moura GR, Santos MA (2009) Parallel DNA pyrosequencing unveils new zebrafish microRNAs. BMC Genomics 10:195
Heimberg AM, Cowper-Sal-lari R, Sémon M, Donoghue PC, Peterson KJ (2010) microRNAs reveal the interrelationships of hagfish, lampreys, and gnathostomes and the nature of the ancestral vertebrate. Proc Natl Acad Sci USA 107(45):19379–19383
Acknowledgments
Thomas Andl received support from a Dermatology Foundation Career Development Award. Support of this work by the National Institute of Arthritis and Musculoskeletal and Skin Diseases of the National Institutes of Health (grant no.: 1R01AR061474-01) is also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ning, M.S., Andl, T. Control by a hair’s breadth: the role of microRNAs in the skin. Cell. Mol. Life Sci. 70, 1149–1169 (2013). https://doi.org/10.1007/s00018-012-1117-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-012-1117-z