Abstract
Computer-aided image analysis for better understanding of images has been time-honored approaches in the medical computing field. In the conventional machine learning approach, the domain experts in medical images are mandatory for image annotation that subsequently to be used for feature engineering. However, in deep learning, a big jump has been made to help the researchers do segmentation, feature extraction, classification, and detection from raw medical images obtained using digital breast tomosynthesis, digital mammography, magnetic resonance imaging, and ultrasound imaging modalities. As a result, deep learning (DL) has gained a state-of-the-art in many application areas, for example, breast cancer image analysis. In this survey paper, we reviewed the most common breast cancer imaging modalities, public, most cited and recently updated breast cancer databases, histopathological based breast cancer image analysis, and DL application types in medical image analysis. We finally conclude by pointing out the research gaps to be addressed in the future.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Breast cancer is the most prevalent type of cancer in women next to lung cancer and early detection has significantly increased the survival rate as proven by clinical reports (Zhang et al. 2018; Kim et al. 2016; Yousefi et al. 2018; Shin et al. 2017). There are a number of breast cancer imaging modalities from old (screen-film mammography) to recent (digital breast tomosynthesis) that have been used by radiologists to screen breast cancer. These imagining modalities have shown remarkable success in detecting breast cancer abnormalities that include masses, microcalcifications, architectural distortions, and bilateral asymmetry. However, they suffer from issues like breast tissue overlapping that hides breast information and which lets suspicious lesions out of sight (Yousefi et al. 2018).
Breast cancer abnormality can be categorized into in-situ and invasive ductal carcinoma (IDC). In-situ represents approximately 20–30\(\%\) of all new breast cancer diagnoses (Brennan et al. 2011; Zhu et al. 2018) whereas IDC is the most common type of breast cancer which almost accounts for 80\(\%\). For example, in-situ has started to be treated using active surveillance without undergoing surgical treatment (Grimm et al. 2017; Zhu et al. 2018) which is not true for IDC. Therefore, early differentiation of breast cancer type as in-situ and invasive is very important for patients so as to define treatment strategy (Grimm et al. 2017; Zhu et al. 2018). For a better understanding of the readers, we identified medical terms used in this survey paper and presented their definition in Table 1.
In Sect. 2 of this survey paper, we discuss the methodology adopted to search papers from selected search databases. In Sects. 3 and 4, we reviewed most common breast cancer imaging modalities and breast cancer databases that are most cited in articles, respectively. In Sect. 5, we reviewed the application areas of deep learning (DL) in medical image analysis in general and breast cancer image analysis in particular. Finally, we made the conclusion of the survey paper in Sect. 6 with highlighting the research gaps for further improvement.
2 Methods
We reviewed articles from 2004 to 2018 to (1) evaluate the use of imaging modalities, (2) compare breast cancer imaging modalities, (3) point out the most cited and publicly available breast cancer databases with different formats and modalities, (4) evaluate the use of DL application in medical image analysis specifically to breast cancer image analysis, (5) evaluate the application of DL using histopathological based breast cancer image analysis. Our general search criteria for this survey paper consisted keywords like ‘breast imaging technology’, ‘deep learning and medical image analysis’, ‘application of deep learning in medical image analysis’, and ‘application of deep learning to breast cancer’. However, we specifically used different search criteria for some of the search databases. The searches were carried out from eight databases: (1) Web of Science, (2) PubMed, (3) Science Direct, (4) IEEE Xplore Digital Library, (5) Google Scholar, (6) arxiv, (7) MICCAI, and (8) SPIE. PubMed was searched for papers containing “convolutional neural network” OR “deep learning” OR “medical imaging” OR “histology”. Arxiv was searched using search terminologies related to medical imaging with search string ’abs:((medical OR mri OR “magnetic resonance” OR(medical OR “histology” OR “ultrasound” OR sfm OR “screen-film mammography” OR “digital mammography” OR “breast cancer”) AND (“deep learning” OR “deep learning application” OR convolutional OR cnn OR “neural network”))’. IEEE Xplore Digital Library is searched for paper containing “convolutional neural network” OR “deep learning” OR “medical imaging”. Conference proceedings for MICCAI and SPIE were searched based on search terminologies that include: DL in breast cancer and MRI, DL in breast cancer and US, DL in breast cancer and DBT, DL in breast cancer and DM OR GM, DL in breast cancer and histology, DL and medical image analysis and application of deep learning in medical image analysis.
3 Breast cancer imaging modalities
In breast cancer image analysis, breast abnormality detection starts with imaging modalities for screening (Zhang et al. 2018). When an abnormality is found early, it is easy to treat the patients, but if evidence appears, cancer may start to spread and by then might be difficult to treat. Among several, some selected breast cancer screening methods are discussed here (Ethiopian Cancer Association 2016). There are different imaging technologies used for breast cancer screening. The performance of breast cancer imaging modalities can be evaluated mostly by sensitivity, specificity, recall rates, positive predicted value (PPV), AUC, F-score, and accuracy.
3.1 Screen-film mammography (SFM)
Screen-film mammography has been the standard imaging modality (still in use in some countries including Ethiopia) for detecting suspicious lesions at an early stage. In the past five decades, the SFM became a useful medium in breast screening. SFM has a high sensitivity (100\(\%\)) in detecting suspicious lesions in breasts composed primarily of fatty tissue (Duijm et al. 1997). However, decreases significantly for breasts with dense glandular tissue. Consequently, 10–20\(\%\) of breast cancers are not visualized (Burrell et al. 1996). Besides, the decrease in lesion conspicuousness may be due to the film itself since it serves as the medium of image acquisition, display, and storage. Once the film is produced, then further improvement is not possible and part of the image may be displayed with lesser contrast. If image improvements cannot be carried out for images with lesser contrast, then patients need to undergo another mammographic image and consequently be exposed to more radiation dose. Other drawbacks of the film are that different regions of the breast image are represented according to the characteristic response of the mammographic film. There is a trade-off between the dynamic range (latitude) and contrast resolution (gradient) (Helvie 2010). Another significant problem of SFM is that it is not digital.
3.2 Digital mammography (DM)
Early-stage breast cancer screening using digital mammography is an effective imaging modality (Gilbert et al. 2015; Liu et al. 2018). It has been the most effective and standard breast imaging modality in the detection and diagnosis of abnormalities of the female breast (Jalalian et al. 2013). However, it has some limitations which include low specificity. As a consequence there may be a higher number of unnecessary biopsies and this limitation increases costs and stress on the patients (Gilbert et al. 2015; Jalalian et al. 2013). Besides, low specificity and high cost the digital mammography exposes the patients to ionizing radiation which endangers the patient’s health (Jalalian et al. 2013). In cases where there is overlapping of breast tissue, then there is a high possibility to leave out some cancers in the retro-mammary space as a result of insufficient positioning of deep tissue (Gilbert et al. 2015; Kevin et al. 2010). Digital mammography offers several advantages over SFM (Patterson and Roubidoux 2014). Besides, computer-assisted detection (CAD) system has revealed favorable results in mammography and is used in the clinical routine to improve the radiologist’s sensitivity (Becker et al. 2018). However, it has also three limitations: high false positive results which imply higher recall rates, higher false negative results, and high radiation exposure (Liu et al. 2018).
3.3 Ultrasound (US)
Ultrasound is an imaging modality that has been used for breast lesion detection and differentiation even though it is operator dependent. Breast lesion detection and differentiation is only possible with the help of an operator who can properly locate the lesion using ultrasound scanner (Byra et al. 2018). However, in contrast to mammography ultrasound doesn’t require the use of ionizing radiation (Becker et al. 2018). According to a review made in Sudarshan et al. (2016) and Jalalian et al. (2013), ultrasound imaging modality is used for detection and diagnose of abnormalities in breast cancer as the second choice to DM. In Jalalian et al. (2013), it is indicated that ultrasound achieved high accuracy in detecting and discriminating benign and malignant masses and supported by Shin et al. (2017). This enabled US imaging modality to bring down unneeded biopsies. According to Byra et al. (2018) and Shin et al. (2017) US is found to be safe, accurate, low cost and highly universalize compared to magnetic resonance imaging, DM, and digital breast tomosynthesis imaging modalities. For every specific lesion types, deep knowledge of image features are required to interpret and this makes the Ultrasound image interpretation not to be straightforward. Ultrasound showed high sensitivity for identifying abnormalities in dense breasts and for women younger than 35 years of age (Sudarshan et al. 2016; Becker et al. 2018). Ultrasound is well recommended to be used as a supplement to DM because of its availability, inexpensiveness compared to other modalities and well-tolerated by patients (Kevin et al. 2010; Leach et al. 2005; Becker et al. 2018).
3.4 Magnetic resonance imaging (MRI)
MRI imaging is based on radio frequency absorption of nuclei in the existence of potent magnetic fields. It is used in case of presence of high patient risk and for clinical diagnosis and monitoring of breast cancer (Amit et al. 2017; Antropova and Giger 2018; Morrow et al. 2011; Kuhl et al. 2014; Saslow et al. 2007; Lin and Brown 2007). In previous studies MRI was used for breast segmentation (Gubern-Mèrida et al. 2015; Wu et al. 2013), breast abnormality detection (Chang et al. 2014; Renz et al. 2012), and breast abnormality classification (Gallego-Ortiz and Martel 2015; Agliozzo et al. 2012; Agner et al. 2011; Pang et al. 2015) using computer aided detection/diagnosis (CAD) system. The technologically enhanced form of MRI, DCE-MRI (dynamic contrast-enhanced MRI), has provided higher volumetric resolution for better lesion visualization and lesions temporal pattern enhancement to extract valuable information for better cancer management (Antropova and Giger 2018; Turkbey 2009). Studies have shown that DCE-MRI provides a useful tool for breast cancer diagnosis (Mahrooghy et al. 2015; Zhang et al. 2018), prognosis (Mazurowski et al. 2015a; Zhang et al. 2018), and correlation with genomics (Mazurowski 2015b; Zhang et al. 2018). In comparison with other imaging modalities like mammography and ultrasound, MRI has shown high sensitivity to breast cancer diagnosis (Antropova and Giger 2018; Zhang et al. 2018; Lin and Brown 2007). CE-MRI is an improved MRI technology and it has shown to have high sensitivity for cancer detection, even in dense breasts (Leach et al. 2005). Even though recommended for women with high-risk breast cancer, MRI might not be optimal imaging modality because of its higher cost and lower specificity (Griebsh et al. 2006; Kuhl et al. 2007).
3.5 Digital breast tomosynthesis (DBT)
Digital Breast Tomosynthesis is an imaging modality that produces a 3D image of the breast using low dose X-rays received at different angles (Regina et al. 2017; Helvie 2010). It is a new breast cancer imaging modality in which the breast is placed and compressed in the same way as a mammogram but the tube with the X-ray moves in a circular arc around the breast (Gur et al. 2009; Gennaro et al. 2010; Wallis et al. 2012; Andersson et al. 2008; Zhang et al. 2018; Poplack et al. 2007). It takes less time for the imaging (Fotin et al. 2016) and provides better detail of dense tissue in the breast compared to conventional mammography (Zhang et al. 2018; Poplack et al. 2007). 3D breast images are produced using computer based on information received from X-rays. The X-ray dose for a tomosynthesis image is similar to that of a regular mammogram (American College of Radiology Imaging Network 2017). After digital mammography, DBT has appeared to be a favorable breast cancer imaging modality to enhance the sensitivity and accuracy of screening (Gur et al. 2009; Gennaro et al. 2010; Wallis et al. 2012; Andersson et al. 2008; Poplack et al. 2007). DBT has emerged as a new breast cancer imaging modalities with a lot of benefits. However, DBT was not able to detect malignant micro-calcifications if those calcification were not on the DBT slice plane (Regina et al. 2017) and increases recall rates for architectural distortion type of breast cancer abnormality (Lourenco et al. 2015). It has also substantially increased the reading time compared to a digital mammogram (DM) in terms of mammogram reading (Samala et al. 2016b) (Table 2).
3.6 Combination of breast cancer imaging modalities
The radiologists and researchers started to use combined imaging modalities during screening to enhance the rate of early detection. In this survey paper we included a few papers and presented as follows:
Gilbert et al. (2015), evaluated the performance of three breast imaging modalities (DM, DBT and synthetic DM) and their combinations (DM + DBT and synthetic DM + DBT). The comparison was made using datasets containing 7060 cases collected randomly from 8869 women of age between 29 and 85. Then, independent radiologists, blind reviewers are considered to review images in DM + DBT, DM, and synthetic DM + DBT without access to the previous examination results. The blind review was made in terms of specificity and sensitivity. The sensitivity for DM, DM + DBT, and synthetic DM + DBT were 87\(\%\), 89\(\%\) and 88\(\%\), respectively. The blind review assured that for the age ranging from 50 to 59, the sensitivity of patients became significantly higher (p = 0.01) for DM + DBT than for DM. In the study the patients with dense breast were included and for those patients that had breast density of 50\(\%\) and higher, the sensitivity was 93\(\%\) for DM + DBT and 86\(\%\) for DM with a p-value of 0.03. The specificity for DM, DM + DBT, and synthetic DM + DBT were 57\(\%\), 70\(\%\) and 72\(\%\), respectively. Finally the study in Gilbert et al. (2015) has proved that adding DBT to DM increased the sensitivity value for patients with dense breasts and increased specificity for all age groups. More importantly, DBT has shown that it has potential benefits especially for dense breasts in younger women.
Mariscotti et al. (2014) compared the efficiency of four imaging modalities (DM, DBT, US, MRI) using 200 patients with age ranging from 26 to 79 and who undergo the screening. Their target was to compare the DM and MRI to DBT alone and a combination of imaging modalities. That means, comparing DM with DBT and MRI with DM + DBT + US. Parameters used for evaluation were sensitivity, specificity, and overall accuracy. DBT scored higher sensitivity than DM alone. The sensitivity of DBT and DM are 90.7\(\%\) and 85.2\(\%\), respectively. The three combined imaging modalities (DM + DBT + the US) achieved a sensitivity value of 97.7\(\%\). The sensitivity of MRI alone is 98.8\(\%\). However, combining it with the other three imaging modalities (DM + DBT + the US) didn’t show improvement to overall sensitivity. The overall accuracy of MRI and DM + DBT + the US were 93.3\(\%\) and 93.7\(\%\), respectively. Breast density affects the sensitivity of some imaging modalities, for example, it affects DM and DBT but not MRI.
Kuhl et al involved 529 participants for screening with 43 different cases where 34 invasive and 9 were ductal carcinoma in-situ type of cancers Kuhl et al. (2005). In their study three imaging modalities (DM, US, and MRI) were considered for comparison and they found out that the sensitivity of MRI, 91\(\%\), is significantly higher than that of DM (33\(\%\)), US (40\(\%\)) and DM + US (49\(\%\)). However, DM and MRI have scored almost the same specificity value, 97.2\(\%\) for MRI and 96.8\(\%\) mammography.
Leach et al. (2005) performed a comparative analysis between DM and contrast-enhanced magnetic resonance imaging (CE MRI) in terms of sensitivity and specificity. The study involved 649 women patients between 35 and 49 with breast cancer history from family (BRCA1 and BRCA2). The specificity and sensitivity were computed after annual screening for 2–7 years and CE-MRI, DM, and CE-MRI + DM scored a sensitivity of 77\(\%\), 40\(\%\) and 94\(\%\), respectively. The imaging modalities like CE-MRI, DM, and CE-MRI + DM scored the specificity of 81\(\%\), 93\(\%\) and 77\(\%\), respectively.
Warner et al. (2004) in their study targeted to compare three breast cancer imaging modalities (DM, US, and MRI) and breast examination in the clinic (CBE) in terms of sensitivity and specificity. The patients they considered were patients related to BRCA1 or BRCA2 mutation. The study, CBE, recommended every 6 months to carry out breast screening from age 25 onward for those with mutation (BRCA1 or BRCA2). In their study, they confirmed that the sensitivity of MRI is more for detecting breast cancers compared to DM, US, or CBE. The specificity and sensitivity scored were 77\(\%\) and 95.4\(\%\) for MRI, 36\(\%\) and 99.8\(\%\) for DM, 33\(\%\) and 96\(\%\) for the US, and 9.1\(\%\) and 99.3\(\%\) for CBE, respectively (Warner et al. 2004). Additionally, they did screening using MRI + DM + the US + CBE to compare with DM + CBE and achieved a sensitivity of 95\(\%\) and 45\(\%\), respectively.
Patient screening for breast cancer using MRI + DM scored higher sensitivity than DM alone in all age ranges (Phi et al. 2017). For example, the sensitivity of DM + MRI achieved 95\(\%\) and 51\(\%\) for DM and 50\(\%\) for MRI alone. For women aged between 40 and 49, the researchers found that the sensitivity of MRI + DM was 98\(\%\) and that of DM and MRI alone were 57\(\%\) and 47\(\%\), respectively. The sensitivity of DM enhanced to some level with increasing age but low in women less than the age of 40.
Phi et al. (2016), evaluated the performance of two breast imaging modalities (DM and MRI) and their combination (DM + MRI) for two mutation status indicators, BRCA1 and BRCA2. The study divided the patients into four age groups (all ages, \(\le\) 40, ages between 41 and 50 years, and above 50) to do age based performance analysis using specificity and sensitivity for the two imaging modalities. For all age groups and BRCA1 mutation status, the sensitivity and specificity were 35.7\(\%\) and 93.8\(\%\) for DM, 88.6\(\%\) and 84.4\(\%\) for MRI and 92.5\(\%\) and 80.4\(\%\) for DM + MRI, respectively. For all age groups and BRCA2 mutation status, the specificity and sensitivity were 44.6\(\%\) and 93.4\(\%\) for DM, 80.1\(\%\) and 85.3\(\%\) for MRI and 92.7\(\%\) and 80.5\(\%\) for DM + MRI, respectively. The sensitivity and specificity for other age groups and BRCA1 and BRCA2 were also presented in Phi et al. (2016) (Table 3).
4 Breast cancer image databases
Over the last few decades, a lot of databases/datasets was produced and published in a different repository where some of them were publicly available for use. Most datasets exist in two formats [CSV and image (jpg, pgm,png, DICOM and jpeg)]. Breast cancer image analysis has mainly used these databases. For example, Mammography Image Analysis Society(MIAS) database is the most popular and applicable by many researchers. It contains 322 image samples where 208 are normal and 114 are abnormal (63 benign cases and 51 malignant cases). The other popular database is Digital Database for Screening Mammography (DDSM) with 2500 images. The summary of most cited and recently updated breast cancer databases are presented in Table 4.
5 Deep learning and breast cancer image analysis
In this section we present breast cancer image analysis in two perspectives: in Sect. 5.1, we present breast cancer image analysis by deep convolutional neural network with datasets developed using various breast cancer imaging modalities; and in Sect. 5.2, we reviewed the histopathology based breast cancer analysis using deep convolutional neural network.
5.1 Imaging modalities and deep learning based breast cancer image analysis
Over the last decades, we have witnessed the importance of medical imaging, e.g., screen-film mammography, computed tomography (CT), magnetic resonance imaging (MRI), positron emission tomography (PET), digital mammography, ultrasound, and so on, for the early detection, diagnosis, and treatment of diseases (Antropova and Giger 2018). In the clinic, the medical image interpretation has mostly been performed by human experts such as radiologists and physicians. However, due to large variations in pathology and potential fatigue of human experts, researchers and doctors have recently begun to benefit from computer-assisted interventions. As compared to the advances in medical imaging technologies, it is belated for the advances in computational medical image analysis and recently it has been improving with the help of machine learning techniques. The most common application areas of DL in medical health care include: breast cancer image analysis (Rodriguez-Ruiz et al. 2018; Kooi et al. 2017a; Wang et al. 2017; Debelee et al. 2018), brain image analysis (Shen et al. 2017; Hosseini-Asl et al. 2016; Burgh et al. 2017; Ghafoorian et al. 2017), retinal image analysis (Wu et al. 2016; Zilly et al. 2017), chest X-ray image analysis (Rajkomar et al. 2017; Kim and Hwang 2016; Anavi et al. 2015, 2016; Bar et al. 2015, 2016; Hwang et al. 2016; Shin et al. 2016a; Wang et al. 2016a), abdominal image analysis (Shah et al. 2016; Zhu et al. 2017) and musculoskeletal image analysis (Forsberg et al. 2017; Spampinato et al. 2017).
Deep learning algorithms with deeper layers like Convolutional Neural Networks (DCNN) have recently shown success in different medical image analysis tasks like segmentation, detection, and classification (Kooi et al. 2016) for urinary bladder (Cha et al. 2016), thoracic-abdominal lymph nodes, interstitial lung disease (Gao et al. 2016), and pulmonary perifissural nodules (Ciompi et al. 2015; Shin et al. 2016b). Angelov and Sperduti (2016) have made an impressive and concise review of the challenges in DL. They started with how multiple layers in a DL approach help in letting efficient learning of hidden representations in datasets and exponential gain in depictive power of each feature in the datasets (Angelov and Gu 2017). Besides its computational cost, they added that fine-tuning the hyper-parameters of the models and structural features selection is not yet realized for DL techniques. But, the availability of pre-trained models has enabled the researchers to either extract features in different points of DL (Sargano et al. 2017) or use it for incremental training to adapt the models to other domain on which the models are trained on Angelov and Gu (2018). The summary of DL application types are given in Table 6. The acronyms of the databases used in papers that we considered in this survey paper are given in Table 5.
Samala et al. (2016a), evaluated their proposed DCNN layer built of 12 hidden layers by comparing it with a CNN with 8 hidden layers in terms of AUC. The DCNN with 5 kernel size and CNN with 3 kernel size were intended for the classification of true microcalcifications and false positives and achieved the AUC value of 0.93 for DCNN and 0.89 for CNN. The dataset used in this research work includes 64 DBT cases collected at the University of Michigan.
Samala et al. (2016b) proposed feature-based and DCNN based CAD system. In DCNN, transfer learning is applied to train the first four convolutional layers and the last three fully connected layers of DCNN using only mammographic images for lesion recognition and false-positive reduction. The transfer learning scored a training AUC of 0.99 and validated using DBT image dataset and achieved an AUC value of 0.81. However, after training using only DM, additional training was held using DBT images to improve the validation score of the model and scored an AUC of 0.90. Data used in their study was obtained using three imaging modalities (SFM, DM and DBT) where 2282 images using digitized SFM and DM, and 324 images was DBT. The source of the image dataset was the department of Radiology at the University of Michigan Health System and University of South Florida. Morphological and texture features were used as feature-based CAD system for detection of mass in the mammograms with the intention of false-positive reduction result. Finally, the feature-based and the DCNN-based CAD systems achieved a sensitivity of 83\(\%\) and 91\(\%\), respectively, at 1 FP/DBT volume.
Kim et al. (2016) proposed a latent bilateral feature representation learned from DCNN to classify masses and FPs through abstraction of data at multiple levels to get accurate representation of image dataset. This approach is applied for the latent bilateral feature representation of masses in DBT and compared with hand-crafted features. The AUC value for hand-crafted features was 0.826 and 0.847 for latent bilateral features.
Fotin et al. (2016) proposed a comparative analysis between the conventional approach and DCNN using 3D (DBT) images to detect (ROIs) and classify the two breast cancer abnormalities (mass and architectural distortions). In the conventional approach, hand-crafted features (contrast, histogram, gradient, texture, shape and topology descriptors) are extracted from the ROIs and given to the ensemble of boosted decision trees. However, in DCNN approach instead of hand-crafted features, a resized \(256\times 256\) ROIs are given to DCNN to detect and classify the abnormalities. The sensitivity of a conventional and DCNN approach was 83.2\(\%\) and 89.3\(\%\), respectively for suspicious ROIs and 85.2\(\%\) and 93.0\(\%\), respectively for malignant ROIs.
Junzhang et al. collected a weakly annotated image datasets for mass using expertise to use in the proposed approach, fully CNN based heatmap regression, for mass detection (Zhang et al. 2018). The weakly annotated mammograms were given as input to the fully CNN model and the model generated the heatmap for the breast mass. The trained model then used for two different purposes: first for estimating the probability of map of mass locations for the 439 mammograms and then 40 images of DBT were used to evaluate the performance of transfer learning by only fine-tuning the last two layers of the pre-trained U-Net model which was trained using mammographic images. The evaluation parameters used in this paper were precision and recall value. The precision and recall value of the approach using mammographic images were 0.85 and 0.92 and that of other approach using tomosynthesis images were 0.33 and 0.41, respectively.
Samala et al. (2018a) explained how too many parameters in the pre-trained models have become a major challenge in training deep learning. The pre-trained models like AlexNet, VGGNet16, GoogLeNet17 and ResNet18 use 60 million, 138 million, 4 million and 60 million parameters, respectively (Krizhevsky et al. 2012; Simonyan and Zisserman 2014; Szegedy et al. 2015; He et al. 2016). The limited amount of medical images is also another challenge to train these models and the applied practice to overcome this challenge is training the models with non-medical images. In this study (Samala et al. 2018a), imageNet pre-trained deep CNN model is selected for a transfer learning. The images used in their experiment was 2, 282 ROIs out of 2461 mass lesions from mammographic image dataset and 230 ROIs from 228 DBT mass lesions. Data augmentation is applied to these images resulting in a total of 19,688 mammographic images and 9120 DBT images. The authors added two additional FC layers to avoid divergence occurred by cross-domain transfer learning. The first added FC layer contained 100 nodes and the second with two nodes. According to (Samala et al. 2017), deactivating the first convolutional layer of the ImageNet pre-trained model with non-medical images became best for transfer learning to mammographic images. Training DCNN using mammographic images has performed well when validated with DBT image data (Samala et al. 2017). In the final stage of this approach transfer learning using DBT was made by deactivating layers from the first convolutional layer to third fully connected layer. These layers are used as a feature extractor to generate 1000 features, and then the recursive feature reduction method was applied to select 240 features. After feature reduction, genetic algorithm and layered pathway evolution were used to compress the frozen deep CNN. The AUC based classification performance of the method applied in this paper was 0.88 before compression and 0.90 after compression for deep CNN.
Samala et al. (2018b) proposed a two stages cross-domain transfer learning approach using DCNN (ImageNet) with five convolutional layers (C1-C5) trained with 1.2 million non-medical images. In their first stage, some convolutional layers of the DCNN were frozen and trained with 20K ROIs from mammographic images. In this stage, the convolutional layer was frozen in three ways: firstly, C1 was frozen, secondly C1 to C3 were frozen and lastly, C1 to C5 were frozen. In their second stage, mammographic trained DCNN was further trained using 9K ROIs from DBT in all the three cases considered in the first stage. Finally, the efficiency of the designed transfer learning approach was evaluated in terms of AUC and achieved an AUC value of 0.76 for C1, 0.73 for C1–C3 and 0.73 for all frozen convolutional layers. The result indicates that deactivating only C1 gives a higher performance during transfer learning.
Semi-automated breast mass segmentation was proposed by Zhang et al. (2018) using DBT images. In most recent years, mass detection and segmentation using machine learning approach has gained remarkable result (Zhang et al. 2016, 2017; Lian et al. 2015, 2017; Zhu et al. 2016, 2017; Liu et al. 2017). However, DCNN based methods have become even more robust and precise to detect and segment masses in the breast (Zhang et al. 2018). In their study, an auto encoder-decoder networks type of DL approach is used to do a mass segmentation in training and application stages. In the training stage, a breast mass mask was used to build an auto encoder-decoder model to realize the mass segmentation. In the application stage, mass region annotated by radiologists were extracted from each DBT image and feed to a pre-trained model, with U-Net network architecture, to do pixel-based mass segmentation. The network used in this study has two parts: an encoding path with two convolution operations, two rectified linear unit and one max-pooling operation for feature extraction and a decoding path (one up-pooling operation, one feature map, and two convolution operations) for image expansion. In the experiment, n-fold cross-validation was applied to measure the efficiency of the proposed mass segmentation in terms of Dice similarity coefficient (DSC) and achieved a value of 0.59.
Yousefi et al. (2018) introduced a three different CAD framework, hand-crafted, feature-based MIL framework, DCNN Multiple Instance-Random Forest (DCNN MI-RF) and deep cardinality-restricted Boltzmann machine Multiple Instance-Random Forest (DCaRBM MI-RF), for automatic detection of speculated mass. The 5040 2D slices collected from 87 DBT volumes were preprocessed that include data augmentation, noise removal, and the pectoral muscle removal of slices. For DCNN and deep CaRBM, data augmentation was carried out before noise and pectoral muscle removal. The efficiency of all the three frameworks was measured based on sensitivity, AUC, specificity, and accuracy. In hand-crafted, four features (morphological, statistical, gray-level, texture) were extracted from ROIs and given to MI-RF classifier to classify the DBT slices. The performance of this framework in terms of specificity, sensitivity, accuracy, and AUC were 75\(\%\), 66.6\(\%\), 69.2\(\%\), 0.75, respectively. In DCNN MI-RF based framework, DCNN was embedded into the framework to get the optimum high-level feature representation out of pre-processed \(256\times 256\) resized DBT slices. Then these features are given to MI-RF classifier as input to classify the DBT slices. The performance of DCNN MI-RF framework in terms of AUC, accuracy, sensitivity, and specificity were 0.87, 86.81\(\%\), 86.6\(\%\) and 87.5\(\%\), respectively. The CaRBM based CAD framework was similar to the DCNN except that the DCNN is replaced by deep CaRBM for feature representation. Then, these features are given to MI-RF classifier as input to classify the DBT slices. Its performance in terms of AUC, accuracy, specificity, and sensitivity were 0.70, 78.5\(\%\), 66.6\(\%\) and 81.8\(\%\), respectively.
Mendel et al. (2018) designed a method of feature extraction using CNN from the ROIs obtained from DM, synthesized 2D images and DBT slices. These images are collected from 76 patients using DBT and DM. Expert radiologists identified the 78 lesions (ROIs) with dimensions \(512\times 512\) pixels from these datasets where 48 of them were benign and the rest were malignant. Some of the lesions were visible in CC views and some in MLO views. These features were given to a pre-trained DCNN (VGGNet19) to extract the features (LeCun et al. 2015; Shin et al. 2016b). Feature extraction was followed by feature reduction through eliminating features with zero-values f or 50\(\%\) of the ROIs. Finally, the reduced features were given to linear SVM. The achievement of the SVM was measured in terms of AUC for three datasets. The AUC value of DM, synthesized mammographic images and DBT slice were 0.755, 0.814 and 0.743, respectively for CC view. The AUC value of DM synthesized 2D images and DBT slice were 0.757, 0.881 and 0.832, respectively for MLO view.
Rodriguez-Ruiz et al. (2018) adopted a DCNN architecture for three-class (pectoral, breast or open field) classification which was similar to the one used in Ronneberger et al. (2015). The U-Net model was evaluated based on the Dice similarity coefficient (DSC) in which the area overlap between segmentation and ground truth is compared. Data used in this paper was collected from 100 patients to gain 172 DBT slices where 121 slices for training, 15 slices for validation and 36 slices for testing. The experimental result showed that the DSC value of the method became 0.970 for test data and found to be promising for the other modalities like mammography and synthetic mammograms (Tables 7, 8).
Kooi et al. (2017a) carried out a feature extraction approach using a DCNN to classify benign solitary cysts from malignant masses. In their work, they adopted both data augmentation with different image resolution but end up with no significant improvement in the performance. Their experiment achieved 0.80 AUC value.
Jadoon et al. (2017) introduced CNN-DW and CNN-CT based multi-class classification techniques to classify the mammograms from IRMA datasets into normal, benign, and malignant. The fusion of the CNN features and most descriptive features with wavelets performed well and achieved an accuracy of 83.74\(\%\) for SVM classifier.
Gallego-Posado et al. (2016) applied DCNN for breast tumor detection and diagnosis. The authors preprocessed (cropping and resizing) the original mammograms from MIAS. Then, the data augmentation was applied by rotating the original images to enhance the size of image datasets. They extracted features using CNN pre-trained model and gave these features to SVM and scored an accuracy of 64.52\(\%\).
Amit et al. (2017) introduced two DCNN techniques that classify breast images into benign and malignant. The annotated images were cropped using a square bounding box around the annotated boundaries and obtained 891 malignant (BIRADS 5) and 365 benign (BI-RADS 2). These images were augmented using rotation (90\(^{\circ }\), 180\(^{\circ }\), 270\(^{\circ }\)) and flipping (right-left, down-up). In the first approach, the CNN with three convolutional layers was trained with the labeled datasets. And in the second approach, the same labeled datasets were used as input to pre-trained VGGNet to extract the features from fully connected layer to do classification using SVM. The first approach’s accuracy, sensitivity, specificity, and AUC were 83\(\%\), 84\(\%\), 82\(\%\) and 0.91, respectively. And the second approach’s accuracy, sensitivity, specificity, and AUC were 73\(\%\), 77\(\%\), 68\(\%\) and 0.81, respectively.
Antropova et al. (2017a) proposed two different ways to extract features with the aim of classifying the images into benign and malignant where the first is segmentation-based and the second is CNN based. The study was based on 640 images collected using DCE-MRIs imaging modality. Out of 640 images, 191 were benign and 449 were malignant. In a segmentation-based approach, 38 features of 6 different categories (enhancement texture, size, kinetics variance, morphology, shape, and kinetics) were extracted after segmentation for classification purpose. In a CNN-based approach, the extracted \(148\times 148\)-pixel sized ROIs were given as input to the AlexNet pre-trained model and 4096 feature vectors were extracted from FC layers. However, only 518 features were used for analysis after leaving 80\(\%\) of the feature vectors with zero value. The performance of this study was evaluated using LDA classifier with round-robin cross-validation for three cases (segmentation-based features (38), CNN-based features (518) and fused features (556)). The performance results in terms of AUC for segmentation-based, CNN features and combined features were 0.88, 0.76 and 0.91, respectively.
Antropova et al. (2017b) collected three datasets using three imaging modalities (mammography, ultrasound, and DCE-MRI). The number of patients considered in mammographic imaging modalities was 245, 1125 patients for ultrasound and 690 for DCE-MRI. However, the number of ROIs for mammographic imaging modality was 739 (328 benign, 411 malignant), 2393 (1978 benign, 415 malignant) for an ultrasound and 690 (212 benign, 478 malignant) for DCE-MRI. For all datasets, CNN-based features from FC and Max-pool layers of VGGNet (VGG19) and conventional features (handcrafted features) were collected. However, the performance of max-pool features outperformed the FC features according to the comparison made based on AUC and hence the performance of the conventional features was made only with max-pool features. The two features (Conventional features and CNN features) were fed to a non-linear SVM with Gaussian RBF kernel and achieved an AUC value of 0.79 and 0.81 for mammographic images, 0.84 and 0.87 for ultrasound images and 0.86 and 0.87 for DCE-MRI, respectively. The performance of SVM was also evaluated for the combined or fused features (Conventional features + CNN features) and achieved an AUC value of 0.86, 0.90 and 0.89 for mammography, ultrasound, and DCE-MRI datasets, respectively.
Antropova and Giger (2018) extracted CNN-based features from all five max-pooling layers using DCE-MRI images from 690 cases. Based on a report from pathologists and radiologists, out of 690 cases, 212 cases were benign and 478 cases were malignant. The extracted features were first normalized with Euclidean norm before concatenated to form fused CNN feature vectors. Then, these features were given to linear SVM to classify MRI images as malignant and benign. The discriminating power of the features from the three ROIs was evaluated using AUC with 80\(\%\) of the features for training and 20\(\%\) for testing. The AUC value of the central slice of the second postcontrast, a central slice of the second postcontrast subtracted and MIP were 0.80, 0.83 and 0.88, respectively.
Antropova et al. (2018) extracted features from all five max-pool layers using VGGNet with 19 layers to classify as benign and malignant. Out of the 703 images datasets collected using DCE-MRI imaging modality 221 were benign and 482 were malignant. They separately extracted the features from images before and after contrast enhancement and fed to LSTM and SVM with RBF kernel. The parameters of the LSTM and SVM classifiers were tuned on a grid search with cross-validation (5 fold). The efficiency of the classifiers and distinguishing power of the features were measured based on the AUC analysis. AUC for SVM classifier was 0.81 and 0.85 for LSTM classifier.
Zhu et al. (2018) used VGGNet with 16 layers to extract features (deep features, Conv11, Conv12, Conv13, FC1, and FC2 features) from MRI images. Images were collected from a total of 131 patients where 35 of them were invasive and the rest were DCIS diagnosed patients. After generating ROIs from the original images, data augmentation was applied using random translation and rotation. The SVM with kernel functions of different types (polynomial, linear and RBF) was trained and evaluated in terms of AUC and validated using cross-validation (10 fold). Compared to other features, the best AUC value (0.68) was achieved with deep features from convolutional layer 13.
Zhang et al. (2018) proposed a CNN based segmentation technique in two stages for images collected from 272 patients using DCE-MRI imaging modality. In the first stage, rough segmentation of breast tumor is obtained and followed by refining as the second stage of FCN. The efficiency of segmentation was evaluated using three measurements (Dice similarity coefficient, sensitivity, and PPV) and comparing with manually annotated ground-truth. The DSC, sensitivity and PPV values were 0.7176, 75.04\(\%\) and 77.33\(\%\), respectively.
Li et al. (2017) proposed a 2D CNN and 3D CNN classification of 143 breast images as benign and malignant. There were 77 malignant and 66 benign for classification and AUC, accuracy, sensitivity and specificity as evaluation parameters. The value of AUC, sensitivity, specificity, and accuracy on test data without augmentation were 0.841, 81.4\(\%\), 77.3\(\%\) and 80.4\(\%\), respectively for 3D CNN and 0.752, 76.1\(\%\), 67.4\(\%\) and 71.1\(\%\), respectively for 2D CNN.
Benjamin et al. (2017) applied cropping to extract 561 ROIs with \(111\times 111\) pixel size from 64 images. VGGNet was used to extract features from the five convolutional layers to enrich spatial information both in lower-level features and higher-level features. All the five convolutional layers from 1 to 5 presented 64, 128, 256, 512, and 512 features, respectively and fused features became 1472 features. Standardization technique was applied to all features to achieve a mean of zero value and a variance of 1. Following the removal of features with zero variance, the prediction power of LDA classifier for a retort to therapy was measured in terms of AUC and scored 0.85 as AUC value.
Becker et al. (2018) targeted 632 patients that undertaking breast ultrasound in 2014 for their study. Out of 632 patients, 550 patients were found to have malignant and the remaining 82 have benign lesions. The authors proposed a generic DL approach to compare with the performance of human readers (radiologists, residents, medical students) with different expertise (experienced and intermediate readers, inexperienced readers) to classify the ultrasound images into benign or malignant. Hold-out validation technique was used where 70\(\%\) of the dataset used for training and 30\(\%\) for testing. The performance analysis was made using AUC and DL method scored 0.84, experienced and intermediate readers scored 0.88 and inexperienced readers scored 0.79.
Han et al. (2017) have carried out an experiment by modifying the architecture of GoogleNet. The modifications targeted on removing two auxiliary classifiers and shifting the input layer to deal with grayscale images instead of color images. The modifications include the reduction of output classes of the target architecture from 100 to 2 classes. For their work, the authors collected 7408 biopsy-confirmed ultrasound breast images (ROIs) associated with masses. Semi-automatically segmentation technique was used to collect ROIs from 5151 patients lesions. Their dataset covered 4254 benign and 3154 malignant lesions. In their pre-processing, histogram equalization, image cropping and margin augmentation were considered. Image cropping was done using a margin with 180 pixels. Data augmentation was achieved using cropping with two different margins (120 pixels and 150 pixels) and translation to increase the number of the training dataset. Out of 7408 ROIs, 6579 ROIs [benign (3765) and malignant (2814)] were used for training and 829 were for testing.
Shin et al. (2017), proposed a CNN based framework to localize and classify masses in breast ultrasound (BUS) images. The CNN (VGGNet-16 and RESNET-101) was trained using large and weakly annotated (DX) dataset and small but strongly annotated (DX + Loc, 600 benign and 600 malignant) dataset. The evaluation is conducted on DX + Loc-Test using correct localization (CorLoc) measure; it is the percentage of images in which a method correctly localizes an object of the target class. Better results were obtained when both the weakly annotated and strongly annotated datasets were used to train the network. DX image dataset used image-level loss whereas DX + Loc image datasets used region-level losses. VGGNet-16 scored a CorLoc value of 0.8450 and RESNET-101 scored 0.8325.
Yap et al. (2018a) proposed three different DL approaches named Patch-based LeNet, U-Net and transfer learning using a fully connected network, AlexNet, for breast ultrasound lesion detection. The authors used two different datasets named dataset A [malignant (60), benign(246)) and B (malignant (53), benign(110)] and the overall best performance was achieved when the two datasets were combined using LeNet.
Yap et al. (2018) proposed an end-to-end breast ultrasound lesions detection using a fully connected network version of AlexNet (FCN-AlexNet). The dataset used in Yap et al. (2018) was identical to the one used in Yap et al. (2018a) and the proposed approach was found to be good for benign lesions detection compared to malignant lesions based on the performance assessment made using hold-out techniques (70\(\%\) for training, 10\(\%\) for validation and 20\(\%\) for testing).
5.2 Histopathology and deep learning based breast cancer image analysis
Histopathology is a technique applied for cancer diagnosis and prognostication for many decades where Pathologists analyze tissue cells under different microscopic standards (Ahmad and Khurshid 2019; Mobadersany et al. 2018). However, the pathologists’ chance to come to one final decision is rare since the assessment is subjective and hence frequent use of this method become tiresome and not repeatable (Ahmad and Khurshid 2019; Mobadersany et al. 2018). In addition, issues related to slide preparation, variations in scanning across sites and staining, and biological variance (Janowczyk and Madabhushi 2016) among patients made the histopathological based breast cancer analysis very challenging.
Ahmed and Khurshid have applied the histopathological method for breast cancer image analysis using deep convolutional neural networks as a supervised classification method cite Ahmad2019. They adopted three deep convolutional neural network architectures (AlexNet, GoogleNet, and ResNet) in their study to classify the 260 images into four classes (normal, benign, in-situ and invasive). The original image dataset distribution for the four classes were 51 for normal, 74 for benign, 68 for In-situ and 67 for Invasive. The classification was made patch-wise and image-wise, but the performance of image-wise classification better than patch-wise for all three CNN models.
Xie et al. (2019) have adopted two deep convolutional neural network models (Inception-V3 and Inception ResNet-V2) to classify the BreaKHis histology image dataset into binary classes (Benign and Malignant) and multi-classes. The multi-class is imposed as a result of malignant subtypes that include ductal carcinoma (DC), lobular carcinoma (LC), mucinous carcinoma (MC), and papillary carcinoma (PC). In their experimental analysis, they found that histopathological based image classification using the two selected DCNN models were superior compared to the existing methods. And they proved that Inception-ResNet-V2 is the most performing DCNN architecture for diagnosing breast cancer using histopathological images.
Sun and Binder (2017) has applied three deep convolutional neural network architectures (CaffeNet, GoogleNet, and ResNet-50). In their study, they used breast cancer biopsies from BreaKHis dataset with resolutions of \(40\,\times\), \(100\,\times\), \(200\,\times\), \(400\,\times\). The whole networks of CaffeNet, GoogleNet and ResNet-50 were fine-tuned with different crop sizes of histopathology images from the target dataset with specified resolutions and their performance was evaluated using accuracy. The best result was achieved at 200X resolution. The accuracy of CaffeNet, GoogleNet and ResNet-50 were 89.40\(\%\), 89.86\(\%\) and 89.60\(\%\), respectively.
Jiang et al. (2019) have ushered in a novel DCNN composed of a convolutional layer, small SE-ResNet module, and fully connected layer to classify the histopathology images from BreaKHis dataset into binary classes (benign and malignant) and multi-classes. The multi-classes include other malignant subtypes like ductal carcinoma (DC), lobular carcinoma (LC), mucinous carcinoma (MC), and papillary carcinoma (PC). In their architecture, they introduced a new module which is the combination of residual module and squeeze-and-excitation block. They top up a new learning rate scheduler to avoid the complicated fine-tuning process to achieve better performance (Table 9).
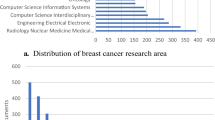
In our final stage of this survey paper, we selected papers with a publication year from 2016 to 2019 as indicated in Fig. 1 to show (a) the number of papers that use the particular database/dataset considered in this survey paper, (b) the distribution of papers that addresses the application types of DL in breast cancer image analysis, and (c) the frequency of breast cancer abnormality type that most diagnosed.
Distribution of papers for publication. a Shows the number of papers that used a particular database from the year 2016–2019 and b shows the number papers that considered a particular breast abnormality type from the year 2016–2018 c shows the number of papers that we considered for a particular DCNN application type from the year 2016–2018
6 Conclusion
Medical image analysis using DL has proven to be better for scientific researchers compared to conventional machine learning approach. A recently remarkable change has been made on deep learning for medical images analysis has enabled it to discover feature patterns in raw images except continued demanding of huge image datasets. Some of the application types that are commonly used in today’s DL based research work are feature extraction, classification, detection, and segmentation. In this survey paper, all these DL application types are considered for review. Since DL methods have succeeded in state-of-the-art achievement over different medical applications like breast image analysis, brain image analysis, retinal image analysis, abdominal image analysis, and musculoskeletal image analysis, using it for further improvement is the major step in analyzing medical images. However, there are some gaps that need to be addressed in medical image analysis using DL. First, building big datasets using medical images and making it available for researchers so that there will be different available pre-trained models trained on medical images, which in turn eases image requirement for transfer learning. Second, developing a new algorithm in which lesser image datasets are required to train deep models to specific domains in medical applications.
References
Agliozzo S et al (2012) Computer-aided diagnosis for dynamic contrast-enhanced breast MRI of mass-like lesions using a multiparametric model combining a selection of morphological, kinetic, and spatiotemporal features. Med Phys 39(4):1704–1715
Agner SC et al (2011) Textural kinetics: a novel dynamic contrast-enhanced (DCE)-MRI feature for breast lesion classification. J Digit Imaging 24(3):446–463
Ahmad, Khurshid (2019) Classification of breast cancer histology images using transfer learning. In: 16th IEEE international Bhurban conference on applied sciences and technology (IBCAST), Pakistan. https://doi.org/10.1109/IBCAST.2019.8667221
American Cancer Society (2015) Breast cancer facts and figures 2015–2016. http://www.cancer.org/acs/groups/content/@research/documents/document/acspc-046381.pdf. Accesed 14 Apr 2015
American College of Radiology Imaging Network (2017) ABOUT mammography and tomosynthesis—ACRIN. https://www.acrin.org. Accessed June 2017
Amit G et al (2017) Classification of breast MRI lesions using small-size training sets: comparison of deep learning approaches. In: Proceedings of SPIE 10134, medical imaging 2017: computer-aided diagnosis, 101341H. https://doi.org/10.1117/12.2249981
Anavi Y et al (2015) A comparative study for chest radiograph image retrieval using binary texture and deep learning classification. In: 2015 37th annual international conference of the IEEE engineering in medicine and biology society (EMBC), pp 2940–2943. https://doi.org/10.1109/EMBC.2015.7319008
Anavi Y et al (2016) Visualizing and enhancing a deep learning framework using patients age and gender for chest X-ray image retrieval. In: Medical imaging, vol 9785 of Proceedings of the SPIE, p 978510
Andersson I et al (2008) Breast tomosynthesis and digital mammography: a comparison of breast cancer visibility and BIRADS classification in a population of cancers with subtle mammographic findings. Eur Radiol 18(12):2817–25
Angelov P, Gu X (2017) MICE: multi-layer multi-model images classifier ensemble. In: 3rd IEEE international conference on cybernetics (CYBCONF), pp 1–8. https://doi.org/10.1109/CYBConf.2017.7985788
Angelov P, Gu X (2018) Deep rule-based classifier with human-level performance and characteristics. Inf Sci 463:196–213
Angelov P, Sperduti A (2016) Challenges in deep learning. In: ESANN 2016 proceedings, European symposium on artificial neural networks, Computational intelligence and machine learning. Bruges, Belgium, pp 27–29
Antropova N et al (2017b) A deep feature fusion methodology for breast cancer diagnosis demonstrated on three imaging modality datasets. Med Phys 44(10):5162–5171
Antropova HA, Giger ML (2018) Use of clinical MRI maximum intensity projections for improved breast lesion classification with deep convolutional neural networks. J Med Imaging 5(1):014503. https://doi.org/10.1117/1.JMI.5.1.014503
Antropova N, Huynh B, Giger M (2018) Recurrent neural networks for breast lesion classification based on DCE-MRIs. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, 105752M. https://doi.org/10.1117/12.2293265
Antropova N, Huynh B, Giger Maryellen (2017) Performance comparison of deep learning and segmentation-based radiomic methods in the task of distinguishing benign and malignant breast lesions on DCE-MRI. In: Proceedings of SPIE 10134, medical imaging 2017: computer-aided diagnosis, 101341G. https://doi.org/10.1117/12.2255582
Baker JA, Lo JY (2011) Breast tomosynthesis: state-of-theart and review of the literature. Acad Radiol 18(10):1298–310
Bar Y et al (2016) Chest pathology identification using deep feature selection with non-medical training. Comput Methods Biomech Biomed Eng Imaging Vis 6(3):259–263. https://doi.org/10.1080/21681163.2016.1138324
Bar Y, Diamant I, Wolf L, Greenspan H (2015) Deep learning with non-medical training used for chest pathology identification. In: Medical imaging, vol 9414 of Proceedings of the SPIE, p 94140V
Becker AS et al (2018) Classification of breast cancer in ultrasound imaging using a generic deep learning analysis software: a pilot study. Br J Radiol 91:20170576
Beroud C et al (2016) BRCA share: a collection of clinical BRCA gene variants. Hum Mutat 37(12):1318–1328
Brandt KR et al (2013) Can digital breast tomosynthesis replace conventional diagnostic mammography views for screening recalls without calcifications? A comparison study in a simulated clinical setting. Am J Roentgenol 200:291–298
Brennan ME, Turner RM, Ciatto S, Marinovich ML, French JR, Macaskill P, Houssami N (2011) Ductal carcinoma in situ at core-needle biopsy: meta-analysis of underestimation and predictors of invasive breast cancer. Radiology 260(1):119–128
Burgh V et al (2017) Deep learning predictions of survival based on MRI in amyotrophic lateral sclerosis. Neuro Image Clin 13:361–369
Burrell HC, Sibbering D, Wilson A et al (1996) Screening interval breast cancers: mammographic features and prognostic factors. Radiology 199(3):811–817
Byra M, Sznajder T, Korzinek D (2018) Impact of ultrasound image reconstruction method on breast lesion classification with neural transfer learning. arXiv:1804.02119v1
CBIS-DDSM (2019) Image dataset. https://wiki.cancerimagingarchive.net/display/Public/CBIS-DDSM. Accessed June 2019
Cha KH et al (2016) Urinary bladder segmentation in CT urography using deep-learning convolutional neural network and level sets. Med Phys 43:1882–1896
Chan H-P et al (2008) Computer-aided detection of masses in digital tomosynthesis mammography: comparison of three approaches. Med Phys 35(9):4087–4095
Chang YC et al (2014) Computerized breast lesions detection using kinetic and morphologic analysis for dynamic contrast-enhanced MRI. Magn Reson Imaging 32(5):514–522
Ciatto S et al (2013) Integration of 3D digital mammography with tomosynthesis for population breast-cancer screening (STORM): a prospective comparison study. Lancet Oncol 14:583–589
Ciompi F et al (2015) Automatic classification of pulmonary peri-fissural nodules in computed tomography using an ensemble of 2D views and a convolutional neural network out-of the-box. Med Image Anal 26:195–202
Conant EF et al (2016) Breast cancer screening using tomosynthesis in combination with digital mammography compared to digital mammography alone: a cohort study within the PROSPR consortium. Breast Cancer Res Treat 156:109–116
Dataset (2017) Breast histopathology images. https://toolbox.google.com/datasetsearch/search?query=Breast%20Histopathology&docid=ZhIlh%2BXjZZi2Abu5AAAAAA%3D%3D. Accessed June 2019
Dataset (2018) Breast Cancer Wisconsin (Diagnostic) data set. https://toolbox.google.com/datasetsearch/search?query=Breast%20Cancer%20Wiscosin%20(Prognostic)&docid=lqkM7t0bmGplzzTuAAAAAA%3D%3D. Accessed June 2019
Debelee TG et al (2018) Classification of mammograms using convolutional neural network based feature extraction. ICT4DA 2017 LNICST 244:89–98
Duijm LEM et al (1997) Sensitivity, specificity and predictive values of breast imagig in the detection of cancer. Br J Cancer 76(3):377–381
Durand MA et al (2015) Early clinical experience with digital breast tomosynthesis for screening mammography. Radiology 274:85–92
Ethiopian Cancer Association (2016) Learn about cancer. http://www.yeeca.org/Learnaboutcancer. Accessed Apr 2017
Faridah Y (2008) Digital versus screen film mammography: a clinical comparison. Biomed Imaging Interv J 4(4):e31
Forsberg D, Sjoblom E, Sunshine JL (2017) Detection and labeling of vertebrae in MR images using deep learning with clinical annotations as training data. J Digit Imaging 30(4):406–412
Fotin SV et al (2016b) Detection of soft tissue densities from digital breast tomosynthesis: comparison of conventional and deep learning approaches. In: Medical imaging, vol 9785 of Proceedings of the SPIE, p. 97850X
Freer PE, Wang JL, Rafferty EA (2014) Digital breast tomosynthesis in the analysis of fat-containing lesions. Radiographics 34:343–358
Gallego-Ortiz C, Martel AL (2015) Improving the accuracy of computer-aided diagnosis for breast MR imaging by differentiating between mass and nonmass lesions. Radiology 278(3):679–688. https://doi.org/10.1148/radiol.2015150241
Gallego-Posado JD et al (2016) Detection and diagnosis of breast tumors using deep Convolutional Neural Networks. In: Research Group on Mathematical Modeling School of Mathematical Sciences Universidad EAFIT Medell in, Colombia, pp 115-121
Gao M et al (2016) Holistic classification of CT attenuation patterns for interstitial lung diseases via deep convolutional neural networks. Comput Methods Biomech Biomed Eng Imaging Vis 6(1):1–6. https://doi.org/10.1080/21681163.2015.1124249
Gennaro G et al (2010) Digital breast tomosynthesis versus digital mammography: a clinical performance study. Eur Radiol 20(7):1545–53
Ghafoorian M et al (2017) Deep multi-scale location aware 3D convolutional neural networks for automated detection of lacunes of presumed vascular origin. NeuroImage Clin 14:391–399
Gilbert FJ et al (2015) Accuracy of digital breast tomosynthesis for depicting breast cancer subproups in a UK retrospective reading study. Radiology 277(3):697–706
Grabowski P (2016) Breast cancer proteomes. https://toolbox.google.com/datasetsearch/search?query=Breast%20Cancer%20Dataset&docid=472Uf%2BgVuRh3EsIoAAAAAA%3D%3D. Accessed May 2019
Griebsh I et al (2006) Cost-effectiveness of screening with contrast enhanced magnetic resonance imaging vs X-ray mammography of women at a high familial risk of breast cancer. Br J Cancer 95:801–810
Grimm LJ, Ryser MD, Partridge AH, Thompson AM, Thomas JS, Wesseling J, Hwang ES (2017) Surgical upstaging rates for vacuum assisted biopsy proven DCIS: implications for active surveillance trials. Ann Surg Oncol 24:3534–3540
Gubern-Mèrida A et al (2015) Breast segmentation and density estimation in breast MRI: a fully automatic framework. IEEE J Biomed Heal Inform 19(1):349–357
Gur D et al (2009) Digital breast tomosynthesis: observer performance study. Am J Roentgenol 193(2):586–591
Haas B et al (2013) Performance of digital breast tomosynthesis compared to conventional digital mammography for breast cancer screening. Radiology 269:694–700
Hagen AL et al (2007) Sensitivity of MRI versus conventional screening in the diagnosis of BRCA-associated breast cancer in a national prospective series. Breast 16(4):367–74
Han S et al (2017) A deep learning framework for supporting the classification of breast lesions in ultrasound images. Phys Med Biol 62:7714–7728
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition, In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE, Las Vegas, NV, USA, pp 770–778
Helvie MA (2010) Digital mammography imaging: breast tomosynthesis and advanced applications. Radiol Clin N Am 48(5):917–929. https://doi.org/10.1016/j.rcl.2010.06.009
Hosseini-Asl E, Gimel’farb G, El-Baz A (2016) Alzheimer’s disease diagnostics by a deeply supervised adaptable 3D convolutional network 1(23):584–596. arXiv: 1607.00556
Huynh BQ et al (2017) Comparison of breast DCE-MRI contrast time points for predicting response to neoadjuvant chemotherapy using deep convolutional neural network features with transfer learning. In: Proceedings of SPIE 10134, medical imaging 2017: computer-aided diagnosis, p 101340U. https://doi.org/10.1117/12.2255316
Hwang S, Kim H-E, Jeong J, Kim H-J (2016) A novel approach for tuberculosis screening based on deep convolutional neural networks. In: Medical imaging, vol 9785 of Proceedings of the SPIE, pp 97852W-1
Jadoon MM et al (2017) Three-class mammogram classification based on descriptive CNN features. Hindawi Biomed Res Int. https://doi.org/10.1155/2017/3640901
Jalalian A et al (2013) Computer-aided detection/diagnosis of breast cancer in mammography and ultrasound: a review. Clin Imaging 37:420–426
Janowczyk A, Madabhushi A (2016) Deep learning for digital pathology image analysis: a comprehensive tutorial with selected use cases. J Pathol Inf 7:29. https://doi.org/10.4103/2153-3539.186902
Jiang Y, Chen L, Zhang H, Xiao X (2019) Breast cancer histopathological image classification using convolutional neural networks with small SE-ResNet module. PLoS One 14(3):e0214587. https://doi.org/10.1371/journal.pone.0214587
Kallenberg et al (2016) Unsupervised deep learning applied to breast density segmentation and mammographic risk scoring. IEEE Trans Med Imaging 35:1322–1331
Kevin KM et al (2010) Breast cancer detection using automated whole breast ultrasound and mammography in radiographically dense breasts. Eur Radiol 20:734–742
Kim DH et al (2016) Latent feature representation with 3-D multi-view deep convolutional neural network for bilateral analysis in digital breast tomosynthesis. In: 2016 IEEE international conference on acoustics, speech and signal processing (ICASSP), Shanghai, 2016, pp 927–931. https://doi.org/10.1109/ICASSP.2016.7471811
Kim H, Hwang S (2016) Scale-invariant feature learning using deconvolutional neural networks for weakly-supervised semantic segmentation. ArXiv: 1602.04984
Kooi T et al (2016) A comparison between a deep convolutional neural network and radiologists for classifying regions of interest in mammography. In: Proceedings of the 13th international workshop on digital mammography. Springer International Publishing, Geneva, pp 51–56
Kooi et al (2017) Discriminating solitary cysts from soft tissue lesions in mammography using a pre-trained deep convolutional neural network. Med Phys 44(3):1017–1027
Kooi T et al (2017b) Large scale deep learning for computer aided detection of mammographic lesions. Med Image Anal 35:303–312
Kopans DB (2014) Digital breast tomosynthesis from concept to clinical care. Am J Roentgenol 202(2):299–308
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet classification with deep convolutional neural networks. In: Proceedings of the 25th international conference on neural information processing systems, vol 1, pp 1097–1105
Kuhl CK et al (2005) Mammography, breast ultrasound, and magnetic resonance imaging for surveillance of women at high familial risk for breast cancer. J Clin Oncol 23(33):8469–76
Kuhl CK et al (2007) MRI for diagnosis of pure ductal carcinoma in situ: a prospective observational study. Lancet 370:485–492
Kuhl CK et al (2014) Abbreviated breast magnetic resonance imaging (MRI): first postcontrast subtracted images and maximum-intensity projection-a novel approach to breast cancer screening with MRI. J Clin Oncol 32:2304–2310
Lang K et al (2016) Performance of oneview breast tomosynthesis as a stand-alone breast cancer screening modality: results from the Malmo Breast Tomosynthesis Screening Trial, a population based study. Eur Radiol 26:184–190
Leach MO, Boggis CR, Dixon AK, Easton DF, Eeles RA, Evans DG, Gilbert FJ, Griebsch I, Hoff RJ, Kessar P, Lakhani SR, Moss SM, Nerurkar A, Padhani AR, Pointon LJ, Thompson D, Warren RM, MARIBS study group (2005) Screening with magnetic resonance imaging and mammography of a UK population at high familial risk of breast cancer: a prospective multicentre cohort study (MARIBS). Lancet 365(9473):1769–1778. https://doi.org/10.1016/S0140-6736(05)66481-1
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521(7553):436–444
Lee RS et al (2017) A curated mammography data set for use in computer-aided detection and diagnosis research. Sci Data 4:170177
Lee et al (2016) Curated breast imaging subset of DDSM. Cancer Imaging Arch. https://doi.org/10.7937/K9/TCIA.2016.7O02S9CY
Lian C et al (2017) Spatial evidential clustering with adaptive distance metric for tumor segmentation in FDG-PET images. IEEE Trans Biomed Eng 65(1):21–30
Lian C, Ruan S, Denoeux T (2015) An evidential classifier based on feature selection and two-step classification strategy. Pattern Recogn 48(7):2318–2327
Li J, Fan M, Zhang J, Li L (2017) Discriminating between benign and malignant breast tumors using 3D convolutional neural network in dynamic contrast enhanced-MR images. In: Proceedings of SPIE 10138, medical imaging 2017: imaging informatics for healthcare, research, and applications, p 1013808. https://doi.org/10.1117/12.2254716
Lin SP, Brown JJ (2007) MR contrast agents: physical and pharmacologic basics. J Magn Reson Imaging 25:884–899
Liu J et al (2018) Radiation dose reduction in digital breast tomosynthesis (DBT) by means of deep-learning-based supervised image processing. In: Proceedings of SPIE 10574, medical imaging 2018: image processing, p 105740F. https://doi.org/10.1117/12.2293125
Liu M et al (2017) View-aligned hypergraph learning for Alzheimer’s disease diagnosis with incomplete multi-modality data. Med Image Anal 36:123–134
Lourenco AP et al (2015) Changes in recall type and patient treatment following implementation of screening digital breast tomosynthesis. Radiology 274:337–342
Mader K (2017) MIAS mammography. https://toolbox.google.com/datasetsearch/search?query=MIAS&docid=xPm6sBCQBOJ0yA5MAAAAAA%3D%3D. Accessed June 2019
Mahrooghy M et al (2015) Pharmacokinetic tumor heterogeneity as a prognostic biomarker for classifying breast cancer recurrence risk. IEEE Trans Biomed Eng 62(6):1585–1594
Mall S et al (2017) The role of digital breast tomosynthesis in the breast assessment clinic: a review. J Med Radiat Sci 64:203–211
Mariscotti G et al (2014) Accuracy of mammography, digital breast tomosynthesis, ultrasound and MR imaging in preoperative assessment of breast cancer. Anticancer Res 34:1219–1226
Mazurowski MA et al (2015) Recurrence-free survival in breast cancer is associated with MRI tumor enhancement dynamics quantified using computer algorithms. Eur J Radiol 84(11):2117–2122
Mazurowski MA (2015) Radiogenomics: what it is and why it is important. J Am Coll Radiol 12(8):862–866
McCarthy AM et al (2014) Screening outcomes following implementation of digital breast tomosynthesis in a general population screening program. J Natl Cancer Inst 2014:106
McDonald ES et al (2015) Baseline screening mammography: performance of full field digital mammography versus digital breast tomosynthesis. AJR 205:1143–1148
Mendel KR, Li H, Sheth D, Giger ML (2018) Transfer learning with convolutional neural networks for lesion classification on clinical breast tomosynthesis. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, p 105750T. https://doi.org/10.1117/12.2294973
Michell MJ et al (2012) A comparison of the accuracy of filmscreen mammography, full-field digital mammography, and digital breast tomosynthesis. Clin Radiol 67(10):976–981
Mobadersany P et al (2018) Predicting cancer outcomes from histology and genomics using convolutional networks. PNAS 115(13):E2970–E2979. https://doi.org/10.1073/pnas.1717139115
Moreira et al (2011) INbreast: toward a full-field digital mammographic database. Acad Radiol 19(236):48. https://doi.org/10.1016/j.acra.2011.09.014
Morrow M, Waters J, Morris E (2011) MRI for breast cancer screening, diagnosis, and treatment. Lancet 378:1804–1811
National Cancer Institute (2018) BRCA mutations: cancer risk and genetic testing. https://www.cancer.gov/about-cancer/causes-prevention/genetics/brca-fact-sheet. Accessed June 2019
NHS Digital (2010) Incidence of breast cancer(all). https://toolbox.google.com/datasetsearch/search?query=Breast%20Cancer%20Dataset&docid=3ilU5NrmvbmkKQkYAAAAAA%3D%3D. Accessed June 2019
Oliver MA (2007) Automatic mass segmentation in mammographic images. Ph.D. Thesis, Universitat De Girona
Palma G, Bloch I, Muller S (2014) Detection of masses and architectural distortions in digital breast tomosynthesis images using fuzzy and a contrario approaches. Pattern Recogn 47(7):2467–2480
Pang Z et al (2015) A computer-aided diagnosis system for dynamic contrast enhanced MR images based on level set segmentation and Relief feature selection. Comput Math Methods Med 2015:450531
Patterson SK, Roubidoux MA (2014) Update on new technologies in digital mammography. Int J Women’s Health 6:781–788
Phi XA et al (2016) Contribution of mammography to MRI screening in BRCA mutation carriers by BRCA status and age: individual patient data meta-analysis. Br J Cancer 114(6):631–637
Phi XA et al (2017) Accuracy of screening women at familial risk of breast cancer without a known gene mutation: Individual patient data meta-analysis. Eur J Cancer 85:31–38
Poplack SP, Tosteson TD, Kogel CA, Nagy HM (2007) Digital breast tomosynthesis: initial experience in 98 women with abnormal digital screening mammography. Am J Roentgenol 189(3):616–623
Rafferty EA (2007) Digital mammography: novel applications. Radiol Clin N Am 45:831–843
Rafferty EA et al (2013) Assessing radiologist performance using combined digital mammography and breast tomosynthesis compared with digital mammography alone: results of a multicenter. Multireader Trial Radiol 266(1):104–113
Rafferty EA et al (2016) Breast cancer screening using tomosynthesis and digital mammography in dense and non-dense breasts. JAMA 315:1784–1786
Rajkomar A, Lingam S, Taylor AG, Blum M, Mongan J (2017) High-throughput classification of radiographs using deep convolutional neural networks. J Digit Imaging 30:95–101
Rakhlin A et al (2018) Deep convolutional neural networks for breast cancer histology image analysis. 1–9, ArXiv:1802.00752v2
Ramanan D (2018) NKI breast cancer data. https://toolbox.google.com/datasetsearch/search?query=Breast%20Cancer%20Dataset&docid=Fj%2BIDVyi5Wdm3sS7AAAAAA%3D%3D. Accessed June 2019
Regina RJ et al (2017) Advances in digital breast tomosynthesis. AJR 208:256–266
Reiser I et al (2006) Computerized mass detection for digital breast tomosynthesis directly from the projection images. Med Phys 33(2):482–491
Renz DM et al (2012) Detection and classification of contrast-enhancing masses by a fully automatic computer assisted diagnosis system for breast MRI. J Magn Reson Imaging 35(5):1077–1088
Rodrigues PS (2017) Breast ultrasound image, Mendeley data, vol 1. https://doi.org/10.17632/wmy84gzngw.1
Rodriguez-Ruiz A et al (2018) Pectoral muscle segmentation in breast tomosynthesis with deep learning. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, p 105752J. https://doi.org/10.1117/12.2292920
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. arXiv: 1505.04597v1
Samala RK et al (2016a) Deep-learning convolution neural network for computer aided detection of micro-calciications in digital breast tomosynthesis. In: Medical imaging, vol 9785 of Proceedings of the SPIE, p 97850Y
Samala RK et al (2016b) Mass detection in digital breast tomosynthesis: Deep convolutional neural network with transfer learning from mammography. Med Phys 43(12):6654–6666
Samala RK et al (2017) Multi-task transfer learning deep convolutional neural network: application to computer-aided diagnosis of breast cancer on mammograms. Phys Med Biol 62:8894–8908
Samala R, Chan H-P, Hadjiiski LM, Helvie MA, Richter C, Cha K (2018a) Compression of deep convolutional neural network for computer-aided diagnosis of masses in digital breast tomosynthesis. Proceedings of SPIE, medical imaging: computer-aided diagnosis, 72. https://doi.org/10.1117/12.2293400
Samala R, Chan H-P, Hadjiiski LM, Helvie MA, Richter C, Cha K (2018b) Cross-domain and multi-task transfer learning of deep convolutional neural network for breast cancer diagnosis in digital breast tomosynthesis. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis. https://doi.org/10.1117/12.2293412
Sampat M, Markey M, Bovik A (2005) Computer-aided detection and diagnosis in mammography. In: Handbook of image and video processing. Elsevier, Academic Press, pp 1195-1217. https://doi.org/10.1016/B978-012119792-6/50130-3
Sarah VCH (2018) Breast cancer wisconsin (prognostic) dataset. https://toolbox.google.com/datasetsearch/search?query=Breast%20Cancer%20Wisconsin%20(Prognostic)%20Data%20Set&docid=B7RP5OldrjAWVn1HAAAAAA%3D%3D. Accessed 8 May 2019
Sargano AB et al (2017b) Human action recognition using transfer learning with deep representations. In: International joint conference on neural networks (IJCNN), pp 463–469
Saslow D et al (2007) American Cancer Society guidelines for breast screening with MRI as an adjunct to mammography. CA Cancer J Clin 57:75–89
Scuccimarra EA (2018) DDSM mammography. https://toolbox.google.com/datasetsearch/search?query=DDSM%20Mammography&docid=%2BIlkfJsgufHU7GpiAAAAAA%3D%3D. Accessed June 2019
Shah A, Conjeti S, Navab N, Katouzian A (2016) Deeply learnt hashing forests for content based image retrieval in prostate MR images. Med Imaging 9784:1–8
Shen D, Wu G, Suk H-I (2017) Deep learning in medical image analysis. Annu Rev Biomed Eng 36(5):1172–1181
Shin HC et al (2016a) Learning to read chest X-rays: recurrent neural cascade model for automated image annotation. ArXiv:1603.08486
Shin HC et al (2016b) Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Trans Med Imaging 35(5):1285–1298
Shin SY et al (2017) Joint weakly and semi-supervised deep learning for localization and classification of masses in breast ultrasound images. arXiv: 1710.03778v1
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. arXiv:1409.1556
Skaane P (2009) Studies comparing screen-film mammography and full-field digital mammography in breast cancer screening: updated review. Acta Radiologica 501:3–14
Skaane P et al (2013) Comparison of digital mammography alone and digital mammography plus tomosynthesis in a population-based screening program. Radiology 267(1):47–56
Spampinato C et al (2017) Deep learning for automated skeletal bone age assessment in X-ray images. Med Image Anal 36:41–51
Sudarshan VK et al (2016) Application of wavelet techniques for cancer diagnosis using ultrasound images: a review. Comput Biol Med 69:97–111
Sumkin JH et al (2015) Recall rate reduction with tomosynthesis during baseline screening examinations. Acad Radiol 22:1477–1482
Sun J, Binder A (2017) Comparison of deep learning architectures for H\&E histopathology images. In: 2017 IEEE Conference on Big Data and Analytics (ICBDA). IEEE, Kuching, Malaysia, pp 43–48. https://doi.org/10.1109/ICBDAA.2017.8284105
Szegedy C et al. (2015) Proceedings of the IEEE conference on computer vision and pattern recognition, pp 1–9
Turkbey B et al (2009) The role of dynamic contrast enhanced MR imaging in cancer diagnosis and treatment. Diagn Interv Radiol 13:45–53
van Schie G et al (2013) Mass detection in reconstructed digital breast tomosynthesis volumes with a computer aided detection system trained on 2D mammograms. Med Phys 40(4):041902
Wallis MG, Moa E, Zanca F, Leifland K, Danielsson M (2012) Two-view and single-view tomosynthesis versus full-field digital mammography: high-resolution X-ray imaging observer study. Radiology 262(3):788–96
Wang J et al (2017) Detecting cardiovascular disease from mammograms with deep learning. IEEE Trans Med Imaging 36(5):1172–1181
Wang C, Elazab A, Wu J, Hu Q (2016a) Lung nodule classification using deep feature fusion in chest radiography. Comput Med Imaging Gr 57:10–18
Warner E et al (2004) Surveillance of BRCA1 and BRCA2 mutation carriers with magnetic resonance imaging, ultrasound, mammography, and clinical breast examination. JAMA 292(11):1317–25
Warner E et al (2008) Systematic review: using magnetic resonance imaging to screen women at high risk for breast cancer. Ann Intern Med 148(9):671–679
Wu S, Weinstein SP, Conant EF, Schnall MD, Kontos D (2013) Automated chest wall line detection for whole-breast segmentation in sagittal breast MR images. Med Phys 40(4):042301
Wu A, Xu Z, Gao M, Buty M, Mollura DJ (2016) Deep vessel tracking: a generalized probabilistic approach via deep learning. IEEE Int Symp Biomed Imaging 5(6):1363–1367
Xie et al (2019) Deep learning based analysis of histopathological images of breast cancer. Front Genet 10:80. https://doi.org/10.3389/fgene.2019.00080
Yap MH et al (2018b) End-to-end breast ultrasound lesions recognition with a deep learning approach. In: Proceedings of SPIE 10578, medical imaging 2018: biomedical applications in molecular. structural, and functional imaging, p 1057819. https://doi.org/10.1117/12.2293498
Yap MH et al (2018a) Automated breast ultrasound lesions detection using convolutional neural networks. IEEE J Biomed Health Inform 22(4):1218–1226
Yousefi M, Krzyzak Adam, Suen Ching Y (2018) Mass detection in digital breast tomosynthesis data using convolutional neural networks and multiple instance learning. Comput Biol Med 96:283–293
Zhang J et al (2018) Automatic deep learning-based normalization of breast dynamic contrast-enhanced magnetic resonance images. 1–11. arXiv:1807.02152v1
Zhang J et al (2018) Breast mass detection in mammography and tomosynthesis via fully convolutional network-based heatmap regression. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, p 1057525. https://doi.org/10.1117/12.2295443
Zhang J et al (2018) Breast tumor segmentation in DCE-MRI using fully convolutional networks with an application in radiogenomics. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, 105750U. https://doi.org/10.1117/12.2295436
Zhang J et al (2018) Convolutional encoder-decoder for breast mass segmentation in digital breast tomosynthesis. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, p 105752V. https://doi.org/10.1117/12.2295437
Zhang J et al (2016) Automatic craniomaxillofacial land mark digitization via segmentation-guided partially-joint regression forest model and multiscale statistical features. IEEE Trans Biomed Eng 63(9):1820–1829
Zhang J et al (2017) Alzheimer’s disease diagnosis using landmark-based features from longitudinal structural MR images. IEEE J Biomed Health Inform 21(3):1607–1616
Zhu Z et al (2018) Deep learning-based features of breast MRI for prediction of occult invasive disease following a diagnosis of ductal carcinoma in situ: preliminary data. In: Proceedings of SPIE 10575, medical imaging 2018: computer-aided diagnosis, 105752W. https://doi.org/10.1117/12.2295470
Zhu Z et al (2016) Faithful completion of images of scenic landmarks using internet images. IEEE Trans Vis Comput Gr 22(8):1945–1958
Zhu Y et al (2017) MRI based prostate cancer detection with high-level representation and hierarchical classification. Med Phys 44(3):1028–1039
Zhu Z et al (2017) An optimization approaches for localization refinement of candidate traffic signs. IEEE Trans Vis Comput Gr 23(5):1561–1573
Zilly J et al (2017) Glaucoma detection using entropy sampling and ensemble learning for automatic optic cup and disc segmentation. Comput Med Imaging Gr 55:28–41
Acknowledgements
The corresponding author would like to thank the Ethiopian Ministry of Education (MoE) and the Deutscher Akademischer Auslandsdienst (DAAD) for funding this research work (Funding number 57162925).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Debelee, T.G., Schwenker, F., Ibenthal, A. et al. Survey of deep learning in breast cancer image analysis. Evolving Systems 11, 143–163 (2020). https://doi.org/10.1007/s12530-019-09297-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12530-019-09297-2