Abstract
Human induced Pluripotent Stem Cells (iPSCs) have enormous potential in understanding developmental biology, disease modeling, drug discovery, and regenerative medicine. The initial human iPSC studies used fibroblasts as a starting cell source to reprogram them; however, it has been identified to be a less appealing somatic cell source by numerous studies due to various reasons. One of the important criteria to achieve efficient reprogramming is determining an appropriate starting somatic cell type to induce pluripotency since the cellular source has a major influence on the reprogramming efficiency, kinetics, and quality of iPSCs. Therefore, numerous groups have explored various somatic cell sources to identify the promising sources for reprogramming into iPSCs with different reprogramming factor combinations. This review provides an overview of promising easily accessible somatic cell sources isolated in non-invasive or minimally invasive manner such as keratinocytes, urine cells, and peripheral blood mononuclear cells used for the generation of human iPSCs derived from healthy and diseased subjects. Notably, iPSCs generated from one of these cell types derived from the patient will offer ethical and clinical advantages. In addition, these promising somatic cell sources have the potential to efficiently generate bona fide iPSCs with improved reprogramming efficiency and faster kinetics. This knowledge will help in establishing strategies for safe and efficient reprogramming and the generation of patient-specific iPSCs from these cell types.
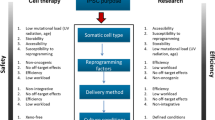
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Embryonic Stem Cells (ESCs) are unspecialized/undifferentiated cells derived from the inner cell mass of the blastocyst during early embryonic development [1,2,3]. ESCs have the ability to self-renew indefinitely in vitro, maintaining a stable karyotype and pluripotency. Because of the same, one can manipulate the culture conditions to obtain the desired specialized cells in sufficient numbers for downstream biomedical applications. However, the potential usage of these cells in cell therapy is restricted to pre-clinical animal studies only, more so due to associated ethical issues and their unsuitability for autologous cell therapy. To obviate this, a pioneering study was carried out by Yamanaka’s group to utilize fibroblasts as the starting cellular source, and they could succeed in deriving pluripotent cells with a molecular and functional resemblance to ESCs, termed as induced pluripotent stem cells (iPSCs) [4]. The initial studies involved derivation of human iPSCs from fibroblasts by overexpression of a quartet of reprogramming factors, namely OCT4, SOX2, KLF4, and c-MYC (OSKM; also known as Yamanaka factors) or OCT4, SOX2, NANOG, and LIN28 (OSNL; also known as Thomson factors) [5, 6]. Subsequent studies by various investigators worldwide has brought about enormous improvements in the technology in terms of efficiency and time regimen required in generating iPSCs [7,8,9,10,11]. Moreover, human iPSCs have also been differentiated successfully into a wide variety of desired cell types, namely cardiomyocytes, neurons, β-cells, hepatocytes, blood lineages, and so forth for research and biomedical applications [7, 12, 13]. Notably, iPSCs allow derivation of autologous pluripotent stem cells circumventing the embryonic cell source [14]. Thus, the generation of patient-specific and disease-specific iPSCs has revolutionized stem cell research to influence regenerative medicine, drug discovery and improve our fundamental understanding of early embryonic development and specific disease mechanisms.
iPSCs have been generated from a variety of somatic cell sources, namely fibroblasts, myoblasts, keratinocytes, melanocytes, hepatocytes, β-cells, dental pulp cells, blood cells, urine-derived renal epithelial cells, amniotic fluid stem cells, muse cells, adult stem cells, and so forth [12, 13, 15,16,17,18,19,20,21,22,23] with varied reprogramming efficiencies and kinetics, demonstrating that the origin of starting cell source is a crucial determining factor. Among these, fibroblasts are the most widely used somatic cell source employed for the generation of iPSCs due to commercial availability, easy handling, economical culture media, and well-established cell culture and reprogramming protocols [24]. However, this cell type is not an optimal cell source due to several disadvantages. It requires skin biopsy for its isolation, which is undesirable in children and patients with abnormal wound healing or coagulation or skin disorders. It generally takes at least four weeks for its expansion to have sufficient cells for reprogramming. The patient age and the passage number used for reprogramming also play a deterministic role in successful reprogramming to obtain iPSCs [25,26,27]. Hence, reprogramming fibroblasts from aged patients is generally highly inefficient. The epigenetic state of fibroblasts also acts as a major barrier to efficient reprogramming and requires remodeling of the epigenome. Furthermore, fibroblasts are generally considered a heterogeneous cell population comprising mesenchymal and non-mesenchymal cell types [28, 29]. In addition, the presence of several reprogramming barriers and the requirement to undergo mesenchymal-to-epithelial (MET) transition also add to the lower reprogramming efficiency in fibroblasts [10, 30, 31]. Also, fibroblasts have a potential risk of accumulating mutations due to constant exposure to stressors, such as UV rays. Due to this, high frequencies of pre-existing coding mutations have been observed in the original somatic cell source and the iPSCs derived from them [32,33,34].
For future biomedical applications, it is ideal if genetically stable iPSCs could be generated from a somatic cell source with these essential criteria: (i) should be abundantly present in a tissue, (ii) should be accessible with ease, so that it can be isolated using minimally invasive procedure, (iii) should be easy to culture and expand to get a sufficient number of cells for reprogramming in a shortest possible time, (iv) primarily free from critical somatic mutations and chromosomal aberrations, (v) should be easy to reprogram with high efficiency and faster kinetics, (vi) should reprogram cells from subjects of different ages and diseased states. This review provides an overview of the most promising somatic cell sources [keratinocytes, urine cells, and peripheral blood mononuclear cells (PBMCs)] that fulfill these criteria and aid in yielding bona fide iPSCs with higher efficiency.
Keratinocytes
Keratinocytes are one of the most promising cell sources utilized for the generation of iPSCs [35]. These cells are keratin protein-enriched epithelial cells that generate the outer protective epidermal layer of the skin and appendages such as nails and hairs. The process of keratinocyte generation continues throughout life due to the presence of self-renewing keratinocyte stem cells [36]. Keratinocytes are one of the most easily accessible cell sources from the human body (from skin epidermis and hair follicles) [35], and hair is the most readily available source for keratinocytes. Keratinocytes can be obtained in a non-invasive manner by merely plucking a hair (Fig. 1). This results in the isolation of transit-amplifying cells having short-term culture potential [37]. They have three growth phases, namely anagen (growth phase), catagen (regression phase), and telogen (resting phase) [38]. Hair root in the anagen phase is active and is preferred for keratinocyte culture, whereas it is inactive in the catagen or telogen phase [39]. Only the outer root sheath needs to be cultured in the medium, whereas the rest of the hair shaft can be removed (Fig. 1). Once the hair is plucked, it can be kept in the media at room temperature for about two days to grow and expand keratinocytes [40]. Several media have been developed for keratinocyte culture, which share a common feature of low calcium concentration to prevent early senescence [40, 41]. Notably, the number of passages for reprogramming experiments should not be high (not more than five passages) to prevent complete differentiation and chromosomal aberration(s) [40]. Even though several research groups investigated keratinocytes as a starting cell source for the generation of iPSCs due to their non-invasive derivation from healthy and diseased subjects and easy handling [42,43,44], it is still not commonly used to date due to their incompatibility with available reprogramming media and certain reprogramming methods like RNA-based approaches [40].
Keratinocytes were first reprogrammed to pluripotency in 2008 using retroviral transduction of Yamanaka factors or three (OSK) factors [42]. Reprogramming keratinocytes with four factors was reported to be 100-fold more efficient and two times faster than reprogramming human fibroblasts [42]. iPSCs derived from keratinocytes had fewer retroviral integrations than fibroblast-derived iPSCs [42]. Similarly, other studies have also reported that keratinocytes are more amenable to efficient reprogramming with faster kinetics when compared to fibroblasts [44,45,46,47] and cord blood-derived cells [46]. Notably, keratinocytes are found to have the endogenous expression of pluripotency-associated genes, namely KLF4 and c-MYC and stem cell marker CD24 [42]. Therefore, attempts have been made to reprogram them using fewer factors. Although keratinocytes endogenously express KLF4, reprogramming without it was unsuccessful [42]. However, retroviral transduction of three-factor combination (OSK) generated iPSCs in approximately 20 days [42, 46]. Inclusion of Tranylcypromine (an inhibitor of lysine-specific demethylase 1 and H3K4 demethylation) and CHIR99021 (Glycogen synthase kinase inhibitor) resulted in reprogramming of human keratinocytes with just two factors, OCT4 and KLF4 [48].
In addition, iPSCs have been derived using a single polycistronic excisable lentiviral vector harboring Yamanaka factors with molecular and functional characteristics similar to human ESCs [49]. However, integrating viral-based approaches involve permanent undesirable integrations of transgenes into the target cell genome [11, 50,51,52]. Therefore, various integration-free reprogramming approaches have been employed to convert keratinocytes into iPSCs efficiently. Non-integrating techniques like Sendai viral vectors [43, 53], adenoviral vectors [54], and episomal vectors [27, 39, 44, 55] have been successfully used to derive iPSCs. But using non-integrative approaches for reprogramming affects the kinetics as it takes around 18 to 34 days [27, 39, 43, 44, 53, 56, 57] when compared to integrative approaches, which takes around 10-14 days [42, 46, 58]. Various studies reporting reprogrammed human keratinocytes are summarized in Table 1.
iPSCs are either cultured on growth-arrested feeder cells [59] or feeder-free conditions, the latter using matrix proteins like collagen or protein mixtures like Matrigel [60]. Therefore, iPSCs either require a matrix or a feeder layer to adhere and grow, maintaining their stem cell identity and function. The most commonly used cells for the feeder layer are irradiated MEFs [42, 61]. Interestingly, a study showed that the type of cell used as a feeder layer could also influence reprogramming efficiency. Using rat embryonic fibroblasts as a feeder layer, a study has reported enhanced reprogramming efficiency of keratinocytes due to the upregulation of secreted growth factors, namely Tgfb1, Inhba, and Grem1, and downregulation of Bmp4 [58].
Keratinocytes have multiple advantages over fibroblasts for reprogramming. For instance, fibroblasts have to undergo a mesenchymal-to-epithelial transition to be successfully reprogrammed to iPSCs [30], whereas keratinocytes are of epithelial origin, and therefore this step is not required [62]. The exogenous expression of the pluripotency-associated genes, namely OSKM, plays a major role in the mesenchymal-to-epithelial transition process in fibroblasts [30]. Also, keratinocytes retain their epithelial gene signature, which might help improve reprogramming [62]. Importantly, genomic characterization of cells from different stages of reprogramming suggests that in most somatic cells, pluripotency-associated genes are hypermethylated, thereby inhibiting their expression. Therefore, reactivation of these hypermethylated pluripotency-associated genes is vital for successful reprogramming into iPSCs [63]. Evidently, the hypermethylation pattern at CpGs and tissue-specific genes in keratinocytes are more similar to ESCs than fibroblasts [64]. This might be the reason why keratinocytes are efficiently reprogrammed to iPSCs than fibroblasts. Moreover, iPSCs derived from keratinocytes have a methylation profile very much similar to ESCs than that of iPSCs from fibroblasts [64]. Furthermore, analyzing the cellular metabolism of iPSCs has revealed that they are highly glycolytic compared to somatic cells [65]. Interestingly, comparing cellular metabolism of fibroblasts and keratinocytes has indicated that keratinocytes have a close resemblance to iPSCs bioenergetically than fibroblasts, and this might also be the reason for the higher reprogramming efficiency of keratinocytes [65, 66]. Furthermore, the tumor suppressor genes decrease cell proliferation rate and compromise reprogramming efficiency [67]. Therefore, inhibition of these tumor suppressor genes and their pathways will pave the way for enhanced reprogramming [67]. A gene named Ink4a/Arf activates the p53 pathway by activation of effector genes, which eventually affects the efficiency of reprogramming. Hence, silencing of Ink4a/Arf locus in keratinocytes helped to enhance reprogramming efficiency by almost 100-fold [68]. Besides, it was found that the inherent expression of p53 and p21 in keratinocytes is low, which might account for its high reprogramming efficiency [67]. Nevertheless, further suppression of the p53 pathway in keratinocytes lends an additive effect to help improve the reprogramming efficiency [67]. Notably, downregulation of pRb in keratinocytes resulted in a 3-fold increase in reprogramming efficiency [69]. In addition, the potent senescence roadblock in keratinocytes can be overcome by telomerase overexpression for efficient derivation of mouse and human iPSCs [70]. Thus, keratinocytes have multiple intrinsic advantages that enable their efficient reprogramming.
Certain caveats still remain to be addressed pertaining to the generation of iPSCs from keratinocytes. First, keratinocytes have the propensity to undergo senescence after a few passages [37]. Second, they have a longer doubling time, leading to a need to carefully cultivate these cells [37]. Also, non-integrating and safer approaches like the Sendai viral method have yielded a lower reprogramming efficiency of 0.01% [53]. This might be because the protocols for Sendai viral transduction are sub-optimal for keratinocytes. In addition, reprogramming using an RNA-based approach, which has been successfully carried out for other cell types [40, 71, 72], is problematic in keratinocytes as it requires multiple passages and hence may trigger their senescence.
Despite these disadvantages, keratinocytes are an appropriate cell source for reprogramming because of their collection in a non-invasive manner, easy isolation and capability to generate iPSCs. Therefore, keratinocyte-derived iPSCs are already finding applications to develop disease models and form organoids [73]. Specifically, they are preferred to derive cells of neural origin as they have a higher tendency than other iPSCs to differentiate towards neural precursor cells, possibly due to common ectodermal origin [74]. Therefore, they are being explored as a means to generate disease models for conditions like Attention Deficit Hyperactivity Disorder [43] as well as a source of autologous stem cells for Spinal Cord Injuries [24]. Disease-specific iPSCs have also been generated from keratinocytes by various studies for disease modeling and developing novel cell therapy applications (Table 2). Thus, with more research on keratinocytes, it can serve as a non-invasive, easily accessible, and viable cell source for the generation of iPSCs [35]. These keratinocyte-derived iPSCs can be widely employed for ‘disease-in-a-dish’ modeling and cell-based therapeutics.
Urine Cells
The search for better cell sources for iPSCs generation perhaps leads to an unlikely source of cells for reprogramming, i.e., human urine (Fig. 2). Human urine satisfies the mentioned criteria for an ideal cell source; i.e., it should be readily available, universal (regardless of age, sex, and disease condition), and involve non-invasive collection [14, 81]. Moreover, they can be collected without any medical assistance [81]. In mammals, the urinary tract consists of the kidney, ureter, urinary bladder, and urethra [82]. Approximately 2000 to 7000 cells of various types detach from the urinary tract and are excreted via urine daily [83]. Urine cells are a heterogeneous population of diverse cells, including renal tubular epithelial cells, fibroblast-like cells, urothelial cells, and urine-derived adult stem cells [82], which endogenously express KLF4 and c-MYC with high telomerase activity [84] and cell surface markers such as TRA-1-60 and TRA-1-81 [85]. These urine-derived stem cells are capable of myogenic and uroepithelial differentiation [86, 87]. The idea of culturing urine cells began as early as 1972. The fact that during the gestation period, fetal urine contributes to amniotic fluid lead to a reasonable possibility that cells in urine also can be cultured [88].
Collection of urine is routinely done for medical diagnosis. Thus, people find it to be an acceptable practice to donate urine samples. Around 50 to 200 ml of urine collected from midstream during micturition was shown to have a 90% success rate in generating epithelial cell cultures [89]. These cells can then be used for reprogramming experiments. Contrastingly, another study reported a lower success rate for human urine cell culture [57]. The study found that osmolality is an important factor for determining successful human urine cell isolation. They deduced that the optimal osmolality range for efficient isolation of human urine cells was between 241 to 598 mmol/l [57]. Moreover, urine epithelial cells can be successfully isolated from urine samples stored for 48 hours at 4 °C [90]. This is advantageous for transporting samples using simple storage conditions [90]. Generally, a minimum of 50 ml of urine is required to successfully isolate urine cells and subsequent reprogramming [91]. Moreover, the collection of urine should be carried out under aseptic conditions. Interestingly, Mulder and his colleagues successfully isolated urine cells obtained from urine as minimum as 10 ml, collected in urine bags [91]. This study showed that the collection of urine samples is one of the easiest and universal methods. All these characteristics make urine one of the ideal cell sources for reprogramming.
In 2011, the researchers reported for the first time that exfoliated renal tubular cells extracted from urine can be infected with retroviral vectors encoding human OCT4, SOX2, KLF4, and c-MYC to generate iPSCs with high efficiency (0.1-4%) [81]. In addition to integrating, several non-integrating approaches have also been applied to reprogram urine cells to develop safe iPSC lines that can be used for therapeutic purposes (Table 3). Sendai virus [92], episomal plasmids [89], RNA-based approaches [93,94,95], and a combination of episomal plasmids and small molecules [57] have all been able to reprogram human urine cells successfully. The urine-derived cells were reprogrammed successfully using an mRNA approach with faster kinetics than that of fibroblasts and dental pulp cells [93]. One caveat of using mRNA-based reprogramming is that the expression of reprogramming factors is transient, and therefore, multiple transfections are needed to reprogram utilizing this approach [93, 94]. But this can be overcome by using the self-replicative RNA approach, which can generate iPSCs in a single transfection [94, 95]. In addition, several small molecule cocktails have also been reported to promote the reprogramming of human urine cells to iPSCs [57]. Moreover, iPSCs can also be generated under feeder-free conditions from human urine cells, which circumvents the problem of cross-contamination of feeder cells [89, 93, 96]. Various studies that have reprogrammed human urine cells are summarized in Table 3.
Besides being a readily available and non-invasive cell source, human urine cells have other intrinsic advantages over other cell sources. Being epithelial, they do not have to undergo MET, which significantly enhances their reprogramming efficiency and kinetics than fibroblasts [84, 85]. As mentioned earlier, these cells endogenously express stem cell-specific genes such as c-MYC and KLF4 with high telomerase activity [84] and stem cell surface markers such as TRA-1-60 and TRA-1-81 [85]. Apart from introducing pluripotency-associated genes, additional factors like miR302-367 and simian virus 40 large T antigen (SV40LT) have proven to be effective in the generation of iPSCs [85, 89]. In addition, miR302-367 can also be used as a substitute for the oncogene c-MYC in the reprogramming process [89]. This substitution has proven to be effective only in the case of urine cells, not in fibroblasts, which demonstrates that urine cells are easier to reprogram [89].
Stem cells, in general, largely depend on their extracellular environment or otherwise called “niche,” for their function and maintenance [97]. The extracellular environment plays a vital role in cell-matrix adhesion and influences cell-cell adhesion. Therefore, modifying the extracellular microenvironment of iPSCs might favor enhanced reprogramming efficiency. Recently, reprogramming human urine cells in a three-dimensional self-assembling peptide hydrogel Puramatrix, rather than a regular two-dimensional Matrigel, has shown efficient generation of iPSC colonies [98]. Although reprogramming efficiencies were comparable between the two stated approaches, the iPSC colonies in the former case were found to be more homogeneous [98]. Studies have also shown that human iPSCs have remnants of epigenetic memory of their starting cell source [74], which makes them biased towards differentiating into specific lineages only. Interestingly, iPSCs derived from urine cells have shown no such propensity to differentiate into particular lineages only [99]. However, the major issue concerning the derivation of iPSCs from human urine cells is that the efficiency differs significantly from donor to donor [81]. The reprogramming efficiency also decreases with increasing passages of human urine cells [81]. Moreover, urine cannot be collected from people with rare renal defects like renal insufficiency and cystectomy [82].
Notably, the applicability of urine-derived iPSCs in the field of therapeutics has been widely studied and analyzed. Urine-derived iPSCs have been of immense help for developing disease models for diseases like dystrophic cardiomyopathy [84, 92, 100], ventricular septal defects [101], hemophilia A [102], diabetes mellitus [103], hepatocyte polarity [104], systemic lupus erythematosus [105], and neurodevelopment diseases like autism spectrum disorder [106, 107], and multiple sclerosis [108]. Furthermore, the application of urine-derived iPSCs have been investigated by developing patient-specific cells or organoids like microvascular grafts for hemophilia A [109], kidney organoids for congenital anomalies [91], tooth-like structures [110], retinal organoids [111], neural progenitor cells for spinal cord injuries [112], and cerebral organoids [113]. These applications undoubtedly exemplify the enormous potential and utility of urine-derived iPSCs. Patient- and disease-specific iPSCs have also been generated from urine cells by several studies for disease modeling and developing novel cell therapy applications (Table 4). Human urine cells are therefore considered as a viable source for the generation of iPSCs, which then have broad applications in the field of disease modeling and therapeutics.
Peripheral Blood Mononuclear Cells (PBMCs)
PBMCs can be isolated from various sources, namely cord blood, bone marrow, and peripheral blood. PBMCs isolated from blood serve as one of the most popular somatic cell sources for iPSCs generation (Fig. 3). Culturing and expansion of PBMCs are simpler and easier. While the collection of peripheral blood and cord blood for isolation of mononuclear cells is less invasive, the same for bone marrow is highly invasive. PBMCs are usually extracted from the buffy coat layer by Ficoll-Paque density gradient centrifugation from whole blood (Fig. 3) [126]. Isolated PBMCs consist of monocytes, natural killer cells, lymphocytes, dendritic cells, and hematopoietic stem cells/hematopoietic progenitor cells. Among these, lymphocytes contribute to a large fraction of the PBMCs population.
The initial studies reprogrammed terminally differentiated mature T and B lymphocytes to give rise to iPSCs using viral-based techniques [127,128,129,130]. Also, integration-free methods have been successfully employed to efficiently reprogram B and T lymphocytes and Epstein Barr Virus immortalized lymphoblastoid cells [19, 127, 131,132,133,134,135,136]. Although B and T lymphocytes are ample in PBMCs and easier to culture, they are not preferred for reprogramming as they are subjected to intrinsic DNA rearrangements at the V(D)J as well as T cell receptor loci to give rise to a vast and highly diverse repertoire of antigen-specific surface immunoglobulins from a relatively limited number of genes [137]. These irreversible DNA rearrangements are then perpetuated in iPSCs derived from these cells, and their potential effect on the proper differentiation of iPSCs is not yet investigated. Additionally, reprogrammed T cells have been shown to induce spontaneous T cell lymphomas in mice [138], limiting their broad applications in regenerative medicine. Therefore, a protocol that enriches erythroblast-like cells and eliminates lymphocytes is preferred [139]. Interestingly, B cells isolated from the human fetal liver are reprogrammed much more efficiently than B cells isolated from peripheral blood or cord blood [136]. This result shows that cell ontogeny might influence reprogramming efficiency. On the other hand, the CD34+ cells, present in PBMCs, are highly proliferative and efficiently reprogrammable [129, 137, 140,141,142,143,144]. But this cell population is rare (<0.01%) in PB [137], which is why patients are subcutaneously injected with stem cell mobilizers such as granulocyte colony-stimulating factor to obtain these progenitor cells in large numbers from circulating peripheral blood. This is only possible if the human subject is in good medical condition. Also, stem cell mobilization is associated with major side effects in patients and requires a multi-day dosing regimen [145, 146]. A study has reported functional differences between cells from mobilized and non-mobilized blood samples, indicating that the intrinsic characteristics of the cells have been altered [147]. In addition, it has been reported that chromosomal aneuploidy can occur in cells mobilized with granulocyte colony-stimulating factor [148]. In agreement with this, genetic and epigenetic abnormalities have been observed in cells isolated from patients mobilized with granulocyte colony-stimulating factor [148, 149]. These observations highlight the possibility of introducing similar genomic and epigenomic modifications that would perpetuate into iPSCs derived from mobilized cells and potentially affect their function. Therefore, it is more desirable to acquire blood that does not necessitate donors to receive these mobilizers to circumvent the adverse side effects and chromosomal abnormalities. In addition, the iPSCs derived from non-lymphocytes having intact genomes will be more appropriate for therapeutic applications [139].
The first study to reprogram PBMCs (from mobilized human CD34+ cells from PB) to iPSCs was carried out by Loh and colleagues [140]. In this study, the researchers delivered Yamanaka factors retrovirally to generate iPSCs, which were indistinguishable from human ESCs with respect to morphology, expression of cell surface and pluripotency-associated markers, DNA methylation at key pluripotency genes, and the ability to differentiate in vitro and in vivo into all three germ layers. Subsequently, the same lab derived iPSCs from both CD34+ cells and mononuclear cells from the PB (nonmobilized) obtained from healthy donors via lentiviral transduction of Yamanaka factors [129]. Another study generated iPSCs from nonmobilized human PBMCs by retroviral transduction of Yamanaka factors [150]. The authors in this study observed that 5x105 PBMCs, which corresponds to less than 1 mL of peripheral blood, were sufficient for the derivation of numerous iPSC colonies. Although the usage of viral vectors gives higher reprogramming efficiencies, their integration into the host genome causes genetic instability in iPSCs due to high frequencies of insertional mutagenesis [151], which, very likely, will affect its biological function. Therefore, non-integrative approaches like the Sendai virus and episomal vectors are preferred for reprogramming for potential clinical applications [8, 9, 11]. Therefore, studies have demonstrated that integration-free approaches such as Sendai virus and episomal vectors can also be efficiently employed for reprogramming PBMCs isolated from healthy subjects [131, 142,143,144, 152,153,154,155,156,157,158,159,160,161,162,163,164,165] and patients [153]. The iPSCs derived in these studies were free from any genomic integration, expressed pluripotency-associated genes, karyotypically normal, and efficiently differentiated into various cell types in vitro and in vivo. Notably, studies have also demonstrated that iPSCs can be generated from PBMCs isolated from less than 10 ml of blood [142, 143, 150, 153, 156,157,158,159,160,161, 163, 165,166,167,168,169,170,171,172,173,174]. Importantly, the epigenetic signatures and gene expression patterns of PBMCs are closer to ESCs and iPSCs than those of the age-matched fibroblasts, which results in faster reprogramming (14 days) of PBMCs compared to age-matched fibroblasts (28 days) [141, 175]. Incorporating a histone deacetylase inhibitor (sodium butyrate or valproic acid) has also been reported to enhance the reprogramming efficiency [175]. Reprogramming kinetics and efficiency can also be further improved by including some additional factors apart from OSKM. Specifically, the inclusion of an anti-apoptotic factor, BCL-XL, increases the reprogramming efficiency by 10-fold [161, 176]. Recently, the episomal vector combination was fine-tuned to demonstrate ~100-fold improvement in the reprogramming of human adult PBMCs [177]. To accomplish this, OCT4 and SOX2 were cloned in a single episomal vector and connected by a 2A peptide linker for their equimolar expression. Additionally, two separate episomal vectors having c-MYC and KLF4 were used to achieve a higher and steadier increase in c-Myc over KLF4 expression during the course of cell reprogramming. These alterations, along with the expression of BCL-XL, contributed to improved efficiency [177]. However, addition of other factors episomally along with OSKM like LIN28 in PBMCs, and LIN28 and shRNA against p53 in lymphoblastoid cells gave comparable efficiencies to that of iPSCs generated from PBMCs by transfection of OSKM factors episomally [134, 178]. It is reported that PBMCs are 50-fold less efficient than cord blood mononuclear cells for iPSCs generation. The efficiency can also be further improved by including additional factors like SV40 Large T antigen and EBNA1 [137, 160, 175]. Numerous studies that have reprogrammed human PBMCs are summarized in Table 5. Patient- and disease-specific iPSCs have also been generated from PBMCs by numerous studies for disease modeling and developing novel cell therapy applications (Table 6).
Most methods require 10 ml of whole blood, requiring minimal invasiveness for PBMC isolation. Interestingly, iPSCs can also be generated successfully by reprogramming cells isolated from whole blood as little as 10 µl collected by the finger-prick method [157]. The finger-prick method of collecting blood is considered one of the least invasive and less complicated procedures [157]. Besides, blood cells can also be cryopreserved and reprogrammed in the future without any compromise in reprogramming kinetics and efficiency [165]. Also, PBMCs are stable at room temperature for up to 24-48 hours and can resist multiple freeze-thaw cycles while maintaining their genomic integrity and reprogramming ability [153, 163]. This ease in access, isolation, and maintenance makes them an attractive candidate as a somatic cell source for reprogramming. However, the major limitation of using PBMCs as a source for iPSC generation is that whole blood samples from patients with clotting disorders may not give quality iPSCs. Further, the epigenetic marks from PBMCs like V(D)J rearrangements in B and T lymphocytes might transfer to iPSCs, hindering its differentiation potential [137]. Therefore, a thorough screening of iPSC clones reprogrammed only from purified hematopoietic progenitors or myeloid-erythroid cells is required before its further use. Also, the reprogramming efficiency relies on the donor, i.e., different reprogramming efficiency is reported for different donors even when the same reprogramming technique and factors were used [175].
In general, PBMCs serves as the most abundant and convenient source of somatic cells because sample collection is less cumbersome, cheap, easily accessible, less exposed to environmental mutagens, and devoid of ethical complications than other somatic cell sources such as hepatocytes, β-cells, and so forth. Also, it allows access to several frozen samples stored at blood banks worldwide, and iPSCs can also be derived from these frozen human peripheral blood samples [160, 179]. iPSCs derived from such samples can provide abundant cells to screen for genetic factors and to elucidate the molecular mechanisms fundamental to lymphoid and myeloid blood disorders. Moreover, the “epigenetic memory” of blood-derived iPSCs will drive their differentiation, particularly towards blood cell type [66, 74], making them suitable for the development of disease models, pre-clinical drug screening, and identifying a cure for various hematopoietic diseases [180]. Despite this, PBMC-derived iPSCs can be coaxed to differentiate towards other cell types such as mesenchymal stem cells, neural stem cells, cardiomyocytes, hepatocytes, and so forth [144, 164, 165, 180, 181]. Importantly, the iPSCs generated using PBMCs are of good quality with better efficiencies and kinetics. Notably, all the colonies picked from reprogrammed PBMCs established stable iPSC clones, which could be expanded and cryopreserved compared to those picked from reprogrammed fibroblasts [166]. In fact, the clinical significance and advantage of using PBMCs as a somatic cell source for iPSCs generation can be seen in Japan’s iPSC stock project, which started in 2013 [182]. Their main objective is to generate a human leukocyte antigen homozygous haplobank and support research related to it. The prospect of iPSC banking seems exciting and opens up new opportunities for personalized medicine, although there is still a long way to go, and PBMCs seem to be one of the most practical sources with promising results.
Conclusion and Future Perspectives
Numerous studies have reported the derivation of iPSCs from a variety of cell sources using integration-based and integration-free approaches [7,8,9, 11, 211, 212]. The potential applications of iPSCs are limited due to several barriers that act as roadblocks to prevent efficient reprogramming [7, 10, 211, 213]. To realize the full potential of this technology, attempts have been made to reprogram a diverse array of cell types derived from different cell sources, some of which are discussed in this review, namely keratinocytes which are easier to obtain, isolate and reprogram, urine cells which are easier to collect and universally applicable to all irrespective of age and gender, and PBMCs which is the most commonly collected sample for diagnostics (also other promising cell sources such as adult stem cells; not discussed in this review due to their isolation in an invasive manner) to generate mouse and human iPSCs efficiently. The origin of cell type dictates the path to pluripotency and involves crucial events such as loss of somatic cell identity, MET (in cell types which are of non-epithelial origin), the degree to which they undergo developmental reversion, and reactivation of the pluripotency network [62]. It is very likely that other promising cell sources are still unexplored and may have a high reprogramming potential, which is still unknown due to limited knowledge on good biomarkers. For example, late-outgrowth endothelial progenitor cells (CD34+ cells) derived from blood showed a 10-fold increase in reprogramming efficiency (0.22%; 10-fold more efficiently than human fibroblasts) and faster kinetics (10 days vs. 15 days for human fibroblasts) when used as a starting cell source [214].
It is also crucial that iPSCs formed have minimal or are ideally devoid of any crucial genetic mutations and chromosomal aberrations in them. Also, the epigenetic memory, which is a characteristic of the somatic cell source of origin, is required to be erased in iPSCs to avoid any impact on their differentiation potential. One of the inherent properties of iPSCs is their indefinite self-renewal and their capability to form teratoma. The presence of undifferentiated residual iPSCs during differentiation may lead to tumor formation and hence pose a serious challenge in its clinical applications. However, this can be circumvented by sorting pure populations of differentiated cells or removal of pluripotent stem cells using different strategies before transplantation [53, 215,216,217,218,219,220,221]. Another aspect of improving the safety of iPSCs is using integration-free reprogramming strategies instead of the commonly used integration-based approaches [7, 8, 11, 212, 213, 222]. In addition, one of the major requirements is to establish a GMP-compliant system for the derivation of clinical-grade iPSCs. To accomplish this, a fast, robust, and simple iPSC generation strategy from an ideal somatic cell source under feeder-free, serum-free, and xeno-free (ideally chemically defined culture conditions) conditions using an integration-free approach is highly desirable. Further extensive research to characterize and identify ideal somatic cell sources, which will yield genetically stable iPSCs with improved efficiency and kinetics, is crucial for biobanking and various biomedical applications. This will eventually translate this promising technology to generate patient-specific iPSCs for clinical applications.
References
Evans, M. J., & Kaufman, M. H. (1981). Establishment in culture of pluripotential cells from mouse embryos. Nature, 292(5819), 154–156. https://doi.org/10.1038/292154a0
Martin, G. R. (1981). Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proceedings of the National Academy of Sciences, 78(12), 7634–7638. https://doi.org/10.1073/pnas.78.12.7634
Thomson, J. A. (1998). Embryonic Stem Cell Lines Derived from Human Blastocysts. Science, 282(5391), 1145–1147. https://doi.org/10.1126/science.282.5391.1145
Takahashi, K., & Yamanaka, S. (2006). Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell, 126(4), 663–676. https://doi.org/10.1016/j.cell.2006.07.024
Takahashi, K., Tanabe, K., Ohnuki, M., Narita, M., Ichisaka, T., Tomoda, K., & Yamanaka, S. (2007). Induction of Pluripotent Stem Cells from Adult Human Fibroblasts by Defined Factors. Cell, 131(5), 861–872. https://doi.org/10.1016/j.cell.2007.11.019
Yu, J., Vodyanik, M. A., Smuga-Otto, K., Antosiewicz-Bourget, J., Frane, J. L., Tian, S., & Thomson, J. A. (2007). Induced Pluripotent Stem Cell Lines Derived from Human Somatic Cells. Science, 318(5858), 1917–1920. https://doi.org/10.1126/science.1151526
Omole, A. E., & Fakoya, A. O. J. (2018). Ten years of progress and promise of induced pluripotent stem cells: Historical origins, characteristics, mechanisms, limitations, and potential applications. PeerJ, 2018 (MAY), 1–47. https://doi.org/10.7717/peerj.4370
Haridhasapavalan, K. K., Borgohain, M. P., Dey, C., Saha, B., Narayan, G., Kumar, S., & Thummer, R. P. (2019). An insight into non-integrative gene delivery approaches to generate transgene-free induced pluripotent stem cells. Gene, 686. https://doi.org/10.1016/j.gene.2018.11.069
Borgohain, M. P., Haridhasapavalan, K. K., Dey, C., Adhikari, P., & Thummer, R. P. (2019). An Insight into DNA-free Reprogramming Approaches to Generate Integration-free Induced Pluripotent Stem Cells for Prospective Biomedical Applications. Stem Cell Reviews and Reports, 15(2), 286–313. https://doi.org/10.1007/s12015-018-9861-6
Haridhasapavalan, K. K., Raina, K., Dey, C., Adhikari, P., & Thummer, R. P. (2020). An Insight into Reprogramming Barriers to iPSC Generation. Stem Cell Reviews and Reports, 16(1), 56–81. https://doi.org/10.1007/s12015-019-09931-1
Dey, C., Raina, K., Haridhasapavalan, K. K., Thool, M., Sundaravadivelu, P. K., Adhikari, P., Thummer, R. P. (2021). An overview of reprogramming approaches to derive integration-free induced pluripotent stem cells for prospective biomedical applications. In Recent Advances in iPSC Technology (pp. 231–287). Elsevier. https://doi.org/10.1016/B978-0-12-822231-7.00011-4
Singh, V. K., Kalsan, M., Kumar, N., Saini, A., & Chandra, R. (2015). Induced pluripotent stem cells: applications in regenerative medicine, disease modeling, and drug discovery. Frontiers in Cell and Developmental Biology, 3. https://doi.org/10.3389/fcell.2015.00002
Menon, S., Shailendra, S., Renda, A., Longaker, M., & Quarto, N. (2016). An Overview of Direct Somatic Reprogramming: The Ins and Outs of iPSCs. International Journal of Molecular Sciences, 17(1), 141. https://doi.org/10.3390/ijms17010141
Liu, G., David, B. T., Trawczynski, M., & Fessler, R. G. (2020). Advances in Pluripotent Stem Cells: History, Mechanisms, Technologies, and Applications. Stem Cell Reviews and Reports, 16(1), 3–32. https://doi.org/10.1007/s12015-019-09935-x
Winder, M. L., & Trokovic, R. (2021). Induced pluripotent stem cell derivation from myoblasts. In Cell Sources for iPSCs (pp. 37–55). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00009-4
Raab, S., Klingenstein, M., Liebau, S., & Linta, L. (2014). A Comparative View on Human Somatic Cell Sources for iPSC Generation. Stem Cells International, 2014. https://doi.org/10.1155/2014/768391
Saha, B., Krishna Kumar, H., Borgohain, M. P., & Thummer, R. P. (2018). Prospective applications of induced pluripotent stem cells in military medicine. Medical Journal Armed Forces India, 74(4), 313–320. https://doi.org/10.1016/j.mjafi.2018.03.005
Petzendorfer, E., & Guillot, P. V. (2021). Induced pluripotent stem cells derived from amniotic fluid stem cells. In Cell Sources for iPSCs (pp. 1–13). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00010-0
Chahine, M. (2021). Lymphoblastoid-derived human-induced pluripotent stem cells. In Cell Sources for iPSCs (pp. 57–70). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00005-7
Jamal, M., Bashir, A., Al-Sayegh, M., & Huang, G. T.-J. (2021). Oral tissues as sources for induced pluripotent stem cell derivation and their applications for neural, craniofacial, and dental tissue regeneration. In Cell Sources for iPSCs (pp. 71–106). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00007-0
Disler, E. R., Ng, N. W., Nguyen, T. G., Anchan, C. J., Waldman, I. N., & Anchan, R. M. (2021). Induced pluripotent stem cell derived from ovarian tissue. In Cell Sources for iPSCs (pp. 107–135). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00011-2
Li, G., Wakao, S., Kuroda, Y., Kushida, Y., & Dezawa, M. (2021). Muse cells as a robust source of induced pluripotent stem cells. In Cell Sources for iPSCs (Vol. 13, pp. 137–161). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00006-9
Pellicano, R., Caviglia, G. P., Ribaldone, D. G., Altruda, F., & Fagoonee, S. (2021). Induced pluripotent stem cells from spermatogonial stem cells. In Cell Sources for iPSCs (pp. 15–35). Elsevier. https://doi.org/10.1016/B978-0-12-822135-8.00001-X
Khazaei, M., Ahuja, C. S., & Fehlings, M. G. (2017). Induced pluripotent stem cells for traumatic spinal cord injury. Frontiers in Cell and Developmental Biology, 4(JAN), 1–9. https://doi.org/10.3389/fcell.2016.00152
Streckfuss-Bömeke, K., Wolf, F., Azizian, A., Stauske, M., Tiburcy, M., Wagner, S., & Guan, K. (2013). Comparative study of human-induced pluripotent stem cells derived from bone marrow cells, hair keratinocytes, and skin fibroblasts. European Heart Journal, 34(33), 2618–2629. https://doi.org/10.1093/eurheartj/ehs203
Rohani, L., Johnson, A. A., Arnold, A., & Stolzing, A. (2014). The aging signature: a hallmark of induced pluripotent stem cells? Aging Cell, 13(1), 2–7. https://doi.org/10.1111/acel.12182
Diecke, S., Lu, J., Lee, J., Termglinchan, V., Kooreman, N. G., Burridge, P. W., & Wu, J. C. (2015). Novel codon-optimized mini-intronic plasmid for efficient, inexpensive and xeno-free induction of pluripotency. Scientific Reports, 5(1), 8081. https://doi.org/10.1038/srep08081
Driskell, R. R., & Watt, F. M. (2015). Understanding fibroblast heterogeneity in the skin. Trends in Cell Biology, 25(2), 92–99. https://doi.org/10.1016/j.tcb.2014.10.001
Eminli, S., Utikal, J., Arnold, K., Jaenisch, R., & Hochedlinger, K. (2008). Reprogramming of Neural Progenitor Cells into Induced Pluripotent Stem Cells in the Absence of Exogenous Sox2 Expression. Stem Cells, 26(10), 2467–2474. https://doi.org/10.1634/stemcells.2008-0317
Li, R., Liang, J., Ni, S., Zhou, T., Qing, X., Li, H., & Pei, D. (2010). A mesenchymal-to-Epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell, 7(1), 51–63. https://doi.org/10.1016/j.stem.2010.04.014
Samavarchi-Tehrani, P., Golipour, A., David, L., Sung, H., Beyer, T. A., Datti, A., & Wrana, J. L. (2010). Functional Genomics Reveals a BMP-Driven Mesenchymal-to-Epithelial Transition in the Initiation of Somatic Cell Reprogramming. Cell Stem Cell, 7(1), 64–77. https://doi.org/10.1016/j.stem.2010.04.015
Gore, A., Li, Z., Fung, H.-L., Young, J. E., Agarwal, S., Antosiewicz-Bourget, J., & Zhang, K. (2011). Somatic coding mutations in human induced pluripotent stem cells. Nature, 471(7336), 63–67. https://doi.org/10.1038/nature09805
Abyzov, A., Mariani, J., Palejev, D., Zhang, Y., Haney, M. S., Tomasini, L., & Vaccarino, F. M. (2012). Somatic copy number mosaicism in human skin revealed by induced pluripotent stem cells. Nature, 492(7429), 438–442. https://doi.org/10.1038/nature11629
Young, M. A., Larson, D. E., Sun, C.-W., George, D. R., Ding, L., Miller, C. A., & Ley, T. J. (2012). Background Mutations in Parental Cells Account for Most of the Genetic Heterogeneity of Induced Pluripotent Stem Cells. Cell Stem Cell, 10(5), 570–582. https://doi.org/10.1016/j.stem.2012.03.002
Zhang, Y., Hu, W., Ma, K., Zhang, C., & Fu, X. (2019). Reprogramming of Keratinocytes as Donor or Target Cells Holds Great Promise for Cell Therapy and Regenerative Medicine. Stem Cell Reviews and Reports, 15(5), 680–689. https://doi.org/10.1007/s12015-019-09900-8
Fuchs, E. (2007). Scratching the surface of skin development. Nature, 445(7130), 834–842. https://doi.org/10.1038/nature05659
Aasen, T., & Belmonte, J. C. I. (2010). Isolation and cultivation of human keratinocytes from skin or plucked hair for the generation of induced pluripotent stem cells. Nature Protocols, 5(2), 371–382. https://doi.org/10.1038/nprot.2009.241
Alonso, L., & Fuchs, E. (2006). The hair cycle. Journal of Cell Science, 119(3), 391–393. https://doi.org/10.1242/jcs02793
Hung, S. S. C., Pébay, A., & Wong, R. C. B. (2015). Generation of integration-free human induced pluripotent stem cells using hair-derived keratinocytes. Journal of Visualized Experiments, 2015(102), 1–6. https://doi.org/10.3791/53174
Klingenstein, S., Klingenstein, M., Kleger, A., & Liebau, S. (2020). From Hair to iPSCs—A Guide on How to Reprogram Keratinocytes and Why. Current Protocols in Stem Cell Biology, 55(1). https://doi.org/10.1002/cpsc.121
Elsholz, F., Harteneck, C., Muller, W., & Friedland, K. (2014). Calcium - A central regulator of keratinocyte differentiation in health and disease. European Journal of Dermatology, 24(6), 650–661. https://doi.org/10.1684/ejd.2014.2452
Aasen, T., Raya, A., Barrero, M. J., Garreta, E., Consiglio, A., Gonzalez, F., & Belmonte, J. C. I. (2008). Efficient and rapid generation of induced pluripotent stem cells from human keratinocytes. Nature Biotechnology, 26(11), 1276–1284. https://doi.org/10.1038/nbt.1503
Re, S., Dogan, A. A., Ben-Shachar, D., Berger, G., Werling, A. M., Walitza, S., & Grünblatt, E. (2018). Improved generation of induced pluripotent stem cells from hair derived keratinocytes – A tool to study neurodevelopmental disorders as ADHD. Frontiers in Cellular Neuroscience, 12(September). https://doi.org/10.3389/fncel.2018.00321
Piao, Y., Hung, S.S.-C., Lim, S. Y., Wong, R.C.-B., & Ko, M. S. H. (2014). Efficient Generation of Integration-Free Human Induced Pluripotent Stem Cells From Keratinocytes by Simple Transfection of Episomal Vectors. STEM CELLS Translational Medicine, 3(7), 787–791. https://doi.org/10.5966/sctm.2013-0036
Lim, S. J., Ho, S. C., Mok, P. L., Tan, K. L., Ong, A. H. K., & Gan, S. C. (2016). Induced pluripotent stem cells from human hair follicle keratinocytes as a potential source for in vitro hair follicle cloning. PeerJ, 2016(11), 1–17. https://doi.org/10.7717/peerj.2695
Giorgetti, A., Montserrat, N., Aasen, T., Gonzalez, F., Rodríguez-Pizà, I., Vassena, R., & Belmonte, J. C. I. (2009). Generation of Induced Pluripotent Stem Cells from Human Cord Blood Using OCT4 and SOX2. Cell Stem Cell, 5(4), 353–357. https://doi.org/10.1016/j.stem.2009.09.008
Maherali, N., Ahfeldt, T., Rigamonti, A., Utikal, J., Cowan, C., & Hochedlinger, K. (2008). A High-Efficiency System for the Generation and Study of Human Induced Pluripotent Stem Cells. Cell Stem Cell, 3(3), 340–345. https://doi.org/10.1016/j.stem.2008.08.003
Li, W., Zhou, H., Abujarour, R., Zhu, S., Joo, J. Y., Lin, T., & Ding, S. (2009). Generation of Human Induced Pluripotent Stem Cells in the Absence of Exogenous Sox2. Stem Cells, N/A-N/A. https://doi.org/10.1002/stem.240
Novak, A., Shtrichman, R., Germanguz, I., Segev, H., Zeevi-Levin, N., Fishman, B., & Itskovitz-Eldor, J. (2010). Enhanced Reprogramming and Cardiac Differentiation of Human Keratinocytes Derived from Plucked Hair Follicles, Using a Single Excisable Lentivirus. Cellular Reprogramming, 12(6), 665–678. https://doi.org/10.1089/cell.2010.0027
Sommer, C. A., & Mostoslavsky, G. (2013). The evolving field of induced pluripotency: Recent progress and future challenges. Journal of Cellular Physiology, 228(2), 267–275. https://doi.org/10.1002/jcp.24155
Brouwer, M., Zhou, H., & Nadif Kasri, N. (2016). Choices for Induction of Pluripotency: Recent Developments in Human Induced Pluripotent Stem Cell Reprogramming Strategies. Stem Cell Reviews and Reports, 12(1), 54–72. https://doi.org/10.1007/s12015-015-9622-8
Kadari, A., Lu, M., Li, M., Sekaran, T., Thummer, R. P., Guyette, N., & Edenhofer, F. (2014). Excision of viral reprogramming cassettes by Cre protein transduction enables rapid, robust and efficient derivation of transgene-free human induced pluripotent stem cells. Stem Cell Research & Therapy, 5(2), 47. https://doi.org/10.1186/scrt435
Sułkowski, M., Konieczny, P., Chlebanowska, P., & Majka, M. (2018). Introduction of exogenous HSV-TK suicide gene increases safety of keratinocyte-derived induced pluripotent stem cells by providing genetic “emergency exit” switch. International Journal of Molecular Sciences, 19(1), 1–16. https://doi.org/10.3390/ijms19010197
Li, C., Ding, L., Sun, C. W., Wu, L. C., Zhou, D., Pawlik, K. M., & Townes, T. M. (2016). Novel HDAd/EBV Reprogramming Vector and Highly Efficient Ad/CRISPR-Cas Sickle Cell Disease Gene Correction. Scientific Reports, 6(July), 1–10. https://doi.org/10.1038/srep30422
Peters, A., & Zambidis, E. T. (2011). Generation of Nonviral Integration-Free Induced Pluripotent Stem Cells from Plucked Human Hair Follicles, (3), 203–227. https://doi.org/10.1007/978-1-61779-267-0_16
Park, T. S., Huo, J. S., Peters, A., Talbot, C. C., Verma, K., Zimmerlin, L., Zambidis, E. T. (2012). Growth factor-activated stem cell circuits and stromal signals cooperatively accelerate non-integrated iPSC reprogramming of human myeloid progenitors. PLoS One, 7(8). https://doi.org/10.1371/journal.pone.0042838
Li, D., Wang, L., Hou, J., Shen, Q., Chen, Q., Wang, X., & Pan, G. (2016). Optimized Approaches for Generation of Integration-free iPSCs from Human Urine-Derived Cells with Small Molecules and Autologous Feeder. Stem Cell Reports, 6(5), 717–728. https://doi.org/10.1016/j.stemcr.2016.04.001
Linta, L., Stockmann, M., Kleinhans, K. N., Böckers, A., Storch, A., Zaehres, H., & Liebau, S. (2012). Rat embryonic fibroblasts improve reprogramming of human keratinocytes into induced pluripotent stem cells. Stem Cells and Development, 21(6), 965–976. https://doi.org/10.1089/scd.2011.0026
Llames, S., García-Pérez, E., Meana, Á., Larcher, F., & Del Río, M. (2015). Feeder Layer Cell Actions and Applications. Tissue Engineering - Part B: Reviews, 21(4), 345–353. https://doi.org/10.1089/ten.teb.2014.0547
Ohmine, S., Dietz, A. B., Deeds, M. C., Hartjes, K. A., Miller, D. R., Thatava, T., & Ikeda, Y. (2011). Induced pluripotent stem cells from GMP-grade hematopoietic progenitor cells and mononuclear myeloid cells. Stem Cell Research and Therapy, 2(6), 46. https://doi.org/10.1186/scrt87
Lowry, W. E., Richter, L., Yachechko, R., Pyle, A. D., Tchieu, J., Sridharan, R., & Plath, K. (2008). Generation of human induced pluripotent stem cells from dermal fibroblasts. Proceedings of the National Academy of Sciences, 105(8), 2883–2888. https://doi.org/10.1073/pnas.0711983105
Nefzger, C. M., Rossello, F. J., Chen, J., Liu, X., Knaupp, A. S., Firas, J., & Polo, J. M. (2017). Cell Type of Origin Dictates the Route to Pluripotency. Cell Reports, 21(10), 2649–2660. https://doi.org/10.1016/j.celrep.2017.11.029
Mikkelsen, T. S., Hanna, J., Zhang, X., Ku, M., Wernig, M., Schorderet, P., & Meissner, A. (2008). Dissecting direct reprogramming through integrative genomic analysis. Nature, 454(7200), 49–55. https://doi.org/10.1038/nature07056
Barrero, M. J., Berdasco, M., Paramonov, I., Bilic, J., Vitaloni, M., Esteller, M., & Belmonte, J. C. I. (2012). DNA hypermethylation in somatic cells correlates with higher reprogramming efficiency. Stem Cells, 30(8), 1696–1702. https://doi.org/10.1002/stem.1138
Panopoulos, A. D., Yanes, O., Ruiz, S., Kida, Y. S., Diep, D., Tautenhahn, R., & Belmonte, J. C. I. (2012). The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Research, 22(1), 168–177. https://doi.org/10.1038/cr.2011.177
Khoo, T. S., Jamal, R., Abdul Ghani, N. A., Alauddin, H., Hussin, N. H., & Abdul Murad, N. A. (2020). Retention of Somatic Memory Associated with Cell Identity, Age and Metabolism in Induced Pluripotent Stem (iPS) Cells Reprogramming. Stem Cell Reviews and Reports, 16(2), 251–261. https://doi.org/10.1007/s12015-020-09956-x
Kawamura, T., Suzuki, J., Wang, Y. V., Menendez, S., Morera, L. B., Raya, A., & Belmonte, J. C. I. (2009). Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature, 460(7259), 1140–1144. https://doi.org/10.1038/nature08311
Li, H., Collado, M., Villasante, A., Strati, K., Ortega, S., Cãamero, M., & Serrano, M. (2009). The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature, 460(7259), 1136–1139. https://doi.org/10.1038/nature08290
Ruiz, S., Panopoulos, A. D., Herrerías, A., Bissig, K. D., Lutz, M., Berggren, W. T., & Izpisua Belmonte, J. C. (2011). A high proliferation rate is required for cell reprogramming and maintenance of human embryonic stem cell identity. Current Biology, 21(1), 45–52. https://doi.org/10.1016/j.cub.2010.11.049
Utikal, J., Polo, J. M., Stadtfeld, M., Maherali, N., Kulalert, W., Walsh, R. M., & Hochedlinger, K. (2009). Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature, 460(7259), 1145–1148. https://doi.org/10.1038/nature08285
Weber, J., Weber, M., Steinle, H., Schlensak, C., Wendel, H. P., & Avci-Adali, M. (2021). An alternative in vivo model to evaluate pluripotency of patient-specific iPSCs. ALTEX. https://doi.org/10.14573/altex.2005221
Warren, L., Manos, P. D., Ahfeldt, T., Loh, Y.-H., Li, H., Lau, F., & Rossi, D. J. (2010). Highly Efficient Reprogramming to Pluripotency and Directed Differentiation of Human Cells with Synthetic Modified mRNA. Cell Stem Cell, 7(5), 618–630. https://doi.org/10.1016/j.stem.2010.08.012
Hohwieler, M., Renz, S., Liebau, S., Lin, Q., Lechel, A., Klaus, J., & Kleger, A. (2016). “Miniguts” from plucked human hair meet Crohn’s disease. Zeitschrift für Gastroenterologie, 54(08), 748–759. https://doi.org/10.1055/s-0042-105520
Kim, K., Zhao, R., Doi, A., Ng, K., Unternaehrer, J., Cahan, P., & Daley, G. Q. (2011). Donor cell type can influence the epigenome and differentiation potential of human induced pluripotent stem cells. Nature Biotechnology, 29(12), 1117–1119. https://doi.org/10.1038/nbt.2052
Boonkaew, B., Tapeng, L., Netsrithong, R., Vatanashevanopakorn, C., Pattanapanyasat, K., & Wattanapanitch, M. (2018). Induced pluripotent stem cell line MUSIi006-A derived from hair follicle keratinocytes as a non-invasive somatic cell source. Stem Cell Research, 31(July), 79–82. https://doi.org/10.1016/j.scr.2018.07.007
Nakayama, C., Fujita, Y., Matsumura, W., Ujiie, I., Takashima, S., Shinkuma, S., & Shimizu, H. (2018). The development of induced pluripotent stem cell-derived mesenchymal stem/stromal cells from normal human and RDEB epidermal keratinocytes. Journal of Dermatological Science, 91(3), 301–310. https://doi.org/10.1016/j.jdermsci.2018.06.004
Shrestha, R., Wen, Y.-T., Ding, D.-C., & Tsai, R.-K. (2019). Aberrant hiPSCs-Derived from Human Keratinocytes Differentiates into 3D Retinal Organoids that Acquire Mature Photoreceptors. Cells, 8(1), 36. https://doi.org/10.3390/cells8010036
Chlebanowska, P., Sułkowski, M., Skrzypek, K., Tejchman, A., Muszyńska, A., Noroozi, R., & Majka, M. (2020). Origin of the induced pluripotent stem cells affects their differentiation into dopaminergic neurons. International Journal of Molecular Sciences, 21(16), 1–23. https://doi.org/10.3390/ijms21165705
Fleischer, A., Lorenzo, I. M., Palomino, E., Aasen, T., Gómez, F., Servera, M., & Bachiller, D. (2018). Generation of two induced pluripotent stem cell (iPSC) lines from p. F508del Cystic Fibrosis patients. Stem Cell Research, 29, 1–5. https://doi.org/10.1016/j.scr.2018.03.004
Kolundzic, N., Khurana, P., Devito, L., Donne, M., Hobbs, C., Jeriha, J., Ilic, D. (2019). Induced pluripotent stem cell line heterozygous for p.R2447X mutation in filaggrin: KCLi002-A. Stem Cell Research, 38 (March), 101462. https://doi.org/10.1016/j.scr.2019.101462
Zhou, T., Benda, C., Duzinger, S., Huang, Y., Li, X., Li, Y., & Esteban, M. A. (2011). Generation of Induced Pluripotent Stem Cells from Urine. Journal of the American Society of Nephrology, 22(7), 1221–1228. https://doi.org/10.1681/ASN.2011010106
Benda, C., Zhou, T., Wang, X., Tian, W., Grillari, J., Tse, H. F., Esteban, M. A. (2013). Urine as a source of stem cells. In Advances in Biochemical Engineering/Biotechnology, 129, 19–32. https://doi.org/10.1007/10_2012_157
Rahmoune, H., Thompson, P. W., Ward, J. M., Smith, C. D., Hong, G., & Brown, J. (2005). Glucose transporters in human renal proximal tubular cells isolated from the urine of patients with non-insulin-dependent diabetes. Diabetes, 54(12), 3427–3434. https://doi.org/10.2337/diabetes.54.12.3427
Guan, X., Mack, D. L., Moreno, C. M., Strande, J. L., Mathieu, J., Shi, Y., & Childers, M. K. (2014). Dystrophin-deficient cardiomyocytes derived from human urine: New biologic reagents for drug discovery. Stem Cell Research, 12(2), 467–480. https://doi.org/10.1016/j.scr.2013.12.004
Drozd, A. M., Walczak, M. P., Piaskowski, S., Stoczynska-Fidelus, E., Rieske, P., & Grzela, D. P. (2015). Generation of human iPSCs from cells of fibroblastic and epithelial origin by means of the oriP/EBNA-1 episomal reprogramming system. Stem Cell Research and Therapy, 6(1), 1–17. https://doi.org/10.1186/s13287-015-0112-3
Bharadwaj, S., Liu, G., Shi, Y., Markert, C., Andersson, K. E., Atala, A., & Zhang, Y. (2011). Characterization of urine-derived stem cells obtained from upper urinary tract for use in cell-based urological tissue engineering. Tissue Engineering - Part A, 17(15–16), 2123–2132. https://doi.org/10.1089/ten.tea.2010.0637
Lang, R., Liu, G., Shi, Y., Bharadwaj, S., Leng, X., Zhou, X., & Zhang, Y. (2013). Self-Renewal and Differentiation Capacity of Urine-Derived Stem Cells after Urine Preservation for 24 Hours. PLoS ONE, 8(1), e53980. https://doi.org/10.1371/journal.pone.0053980
Sutherland, G. R., & Bain, A. D. (1972). Culture of Cells from the Urine of Newborn Children. Nature, 239(5369), 231–231. https://doi.org/10.1038/239231a0
Xue, Y., Cai, X., Wang, L., Liao, B., Zhang, H., Shan, Y., Pan, G. (2013). Generating a Non-Integrating Human Induced Pluripotent Stem Cell Bank from Urine-Derived Cells. PLoS One, 8(8). https://doi.org/10.1371/journal.pone.0070573
Sauer, V., Tchaikovskaya, T., Wang, X., Li, Y., Zhang, W., Tar, K., & Roy-Chowdhury, J. (2016). Human urinary epithelial cells as a source of engraftable hepatocyte-like cells using stem cell technology. Cell Transplantation, 25(12), 2221–2243. https://doi.org/10.3727/096368916X692014
Mulder, J., Sharmin, S., Chow, T., Rodrigues, D. C., Hildebrandt, M. R., D’Cruz, R., & Rosenblum, N. D. (2020). Generation of infant- and pediatric-derived urinary induced pluripotent stem cells competent to form kidney organoids. Pediatric Research, 87(4), 647–655. https://doi.org/10.1038/s41390-019-0618-y
Afzal, M. Z., & Strande, J. L. (2015). Generation of induced pluripotent stem cells from muscular dystrophy patients: Efficient integration-free reprogramming of urine derived cells. Journal of Visualized Experiments, 95, 1–8. https://doi.org/10.3791/52032
Gaignerie, A., Lefort, N., Rousselle, M., Forest-Choquet, V., Flippe, L., Francois-Campion, V., & David, L. (2018). Urine-derived cells provide a readily accessible cell type for feeder-free mRNA reprogramming. Scientific Reports, 8(1), 2–11. https://doi.org/10.1038/s41598-018-32645-2
Steinle, H., Weber, M., Behring, A., Mau-Holzmann, U., von Ohle, C., Popov, A. F., & Avci-Adali, M. (2019). Reprogramming of Urine-Derived Renal Epithelial Cells into iPSCs Using srRNA and Consecutive Differentiation into Beating Cardiomyocytes. Molecular Therapy - Nucleic Acids, 17 (September), 907–921. https://doi.org/10.1016/j.omtn.2019.07.016
Bouma, M. J., Arendzen, C. H., Mummery, C. L., Mikkers, H., & Freund, C. (2020). Reprogramming Urine-Derived Cells using Commercially Available Self-Replicative RNA and a Single Electroporation. Current Protocols in Stem Cell Biology, 55(1), 1–17. https://doi.org/10.1002/cpsc.124
Lee, K.-I., Kim, H.-T., & Hwang, D.-Y. (2014). Footprint- and xeno-free human iPSCs derived from urine cells using extracellular matrix-based culture conditions. Biomaterials, 35(29), 8330–8338. https://doi.org/10.1016/j.biomaterials.2014.05.059
Gattazzo, F., Urciuolo, A., & Bonaldo, P. (2014). Extracellular matrix: A dynamic microenvironment for stem cell niche. Biochimica et Biophysica Acta (BBA) - General Subjects, 1840(8), 2506–2519. https://doi.org/10.1016/j.bbagen.2014.01.010
Sun, W., Zhang, S., Zhou, T., Shan, Y., Gao, F., Zhang, Y., & Zhang, X. (2020). Human Urinal Cell Reprogramming: Synthetic 3D Peptide Hydrogels Enhance Induced Pluripotent Stem Cell Population Homogeneity. ACS Biomaterials Science and Engineering, 6(11), 6263–6275. https://doi.org/10.1021/acsbiomaterials.0c00667
Shi, L., Cui, Y., Luan, J., Zhou, X., & Han, J. (2016). Urine-derived induced pluripotent stem cells as a modeling tool to study rare human diseases. Intractable and Rare Diseases Research, 5(3), 192–201. https://doi.org/10.5582/irdr.2016.01062
Pioner, J. M., Guan, X., Klaiman, J. M., Racca, A. W., Pabon, L., Muskheli, V., & Regnier, M. (2020). Absence of full-length dystrophin impairs normal maturation and contraction of cardiomyocytes derived from human-induced pluripotent stem cells. Cardiovascular Research, 116(2), 368–382. https://doi.org/10.1093/cvr/cvz109
Cao, Y., Xu, J., Wen, J., Ma, X., Liu, F., Li, Y., & Huang, G. (2018). Generation of a Urine-Derived Ips Cell Line from a Patient with a Ventricular Septal Defect and Heart Failure and the Robust Differentiation of These Cells to Cardiomyocytes via Small Molecules. Cellular Physiology and Biochemistry, 50(2), 473–488. https://doi.org/10.1159/000494167
Jia, B., Chen, S., Zhao, Z., Liu, P., Cai, J., Qin, D., & Pan, G. (2014). Modeling of hemophilia A using patient-specific induced pluripotent stem cells derived from urine cells. Life Sciences, 108(1), 22–29. https://doi.org/10.1016/j.lfs.2014.05.004
Tang, L., Wang, H., Dai, B., Wang, X., Zhou, D., Shen, J., & Liang, P. (2020). Human induced pluripotent stem cell-derived cardiomyocytes reveal abnormal TGFβ signaling in type 2 diabetes mellitus. Journal of Molecular and Cellular Cardiology, 142 (April), 53–64. https://doi.org/10.1016/j.yjmcc.2020.03.016
Overeem, A. W., Klappe, K., Parisi, S., Klöters-Planchy, P., Mataković, L., du Teil Espina, M., & van IJzendoorn, S. C. D. . (2019). Pluripotent stem cell-derived bile canaliculi-forming hepatocytes to study genetic liver diseases involving hepatocyte polarity. Journal of Hepatology, 71(2), 344–356. https://doi.org/10.1016/j.jhep.2019.03.031
Chen, Y., Luo, R., Xu, Y., Cai, X., Li, W., Tan, K., & Dai, Y. (2013). Generation of systemic lupus erythematosus-specific induced pluripotent stem cells from urine. Rheumatology International, 33(8), 2127–2134. https://doi.org/10.1007/s00296-013-2704-5
Song, J., Yang, X., Zhou, Y., Chen, L., Zhang, X., Liu, Z., & Li, W. (2019). Dysregulation of neuron differentiation in an autistic savant with exceptional memory. Molecular Brain, 12(1), 1–12. https://doi.org/10.1186/s13041-019-0507-7
Yang, X., Liu, Y., Zhou, T., Zhang, H., Dong, R., Li, Y., Gai, Z. (2019). An induced pluripotent stem cells line (SDQLCHi014-A) derived from urine cells of a patient with ASD and hyperactivity carrying a 303 kb de novo deletion at chr3p26.1 implicating GRM7 gene. Stem Cell Research, 41 (September), 2–5. https://doi.org/10.1016/j.scr.2019.101635
Massa, M. G., Gisevius, B., Hirschberg, S., Hinz, L., Schmidt, M., Gold, R., & Haghikia, A. (2016). Multiple sclerosis patient-specific primary neurons differentiated from urinary renal epithelial cells via induced pluripotent stem cells. PLoS ONE, 11(5), 1–14. https://doi.org/10.1371/journal.pone.0155274
Neumeyer, J., Lin, R. Z., Wang, K., Hong, X., Hua, T., Croteau, S. E., & Melero-Martin, J. M. (2019). Bioengineering hemophilia a-specific microvascular grafts for delivery of full-length factor VIII into the bloodstream. Blood Advances, 3(24), 4166–4176. https://doi.org/10.1182/bloodadvances.2019000848
Cai, J., Zhang, Y., Liu, P., Chen, S., Wu, X., Sun, Y., Pei, D. (2013). Generation of tooth-like structures from integration-free human urine induced pluripotent stem cells. Cell Regeneration, 2(1), 2:6. https://doi.org/10.1186/2045-9769-2-6
Li, G., Xie, B., He, L., Zhou, T., Gao, G., Liu, S., Zhong, X. (2018). Generation of retinal organoids with mature rods and cones from urine-derived human induced pluripotent stem cells. Stem Cells International, 2018. https://doi.org/10.1155/2018/4968658
Liu, Y., Zheng, Y., Li, S., Xue, H., Schmitt, K., Hergenroeder, G. W., & Cao, Q. (2017). Human neural progenitors derived from integration-free iPSCs for SCI therapy. Stem Cell Research, 19, 55–64. https://doi.org/10.1016/j.scr.2017.01.004
Lin, V. J. T., Hu, J., Zolekar, A., Yan, L. J., & Wang, Y. C. (2020). Urine Sample-Derived Cerebral Organoids Suitable for Studying Neurodevelopment and Pharmacological Responses. Frontiers in Cell and Developmental Biology, 8 (May), 1–18. https://doi.org/10.3389/fcell.2020.00304
Wang, L., Wang, L., Huang, W., Su, H., Xue, Y., Su, Z., & Pei, D. (2013). Generation of integration-free neural progenitor cells from cells in human urine. Nature Methods, 10(1), 84–89. https://doi.org/10.1038/nmeth.2283
Rossbach, B., Hildebrand, L., El-Ahmad, L., Stachelscheid, H., Reinke, P., & Kurtz, A. (2016). Generation of a human induced pluripotent stem cell line from urinary cells of a healthy donor using an integration free vector. Stem Cell Research, 16(2), 314–317. https://doi.org/10.1016/j.scr.2015.12.018
Steichen, C., Si-Tayeb, K., Wulkan, F., Crestani, T., Rosas, G., Dariolli, R., Krieger, J. E. (2017). Human Induced Pluripotent Stem (hiPS) Cells from Urine Samples: A Non-Integrative and Feeder-Free Reprogramming Strategy. Current Protocols in Human Genetics, 2017 (January), 21.7.1-21.7.22. https://doi.org/10.1002/cphg.26
Uhm, K. O., Jo, E. H., Go, G. Y., Kim, S. J., Choi, H. Y., Im, Y. S., & Koo, S. K. (2017). Generation of human induced pluripotent stem cells from urinary cells of a healthy donor using a non-integration system. Stem Cell Research, 21, 44–46. https://doi.org/10.1016/j.scr.2017.03.019
Wang, L., Chen, Y., Guan, C., Zhao, Z., Li, Q., Yang, J., & Li, J. (2017). Using low-risk factors to generate non-integrated human induced pluripotent stem cells from urine-derived cells. Stem Cell Research and Therapy, 8(1), 1–13. https://doi.org/10.1186/s13287-017-0698-8
Shi, L., Cui, Y., Zhang, G., Zhou, X., Luan, J., & Han, J. (2020). Establishment of a control induced pluripotent stem cell line SMBCi006-A from a healthy male donor. Stem Cell Research, 49 (September), 102025. https://doi.org/10.1016/j.scr.2020.102025
Hildebrand, L., Rossbach, B., Kühnen, P., Gossen, M., Kurtz, A., Reinke, P., & Stachelscheid, H. (2016). Generation of integration free induced pluripotent stem cells from fibrodysplasia ossificans progressiva (FOP) patients from urine samples. Stem Cell Research, 16(1), 54–58. https://doi.org/10.1016/j.scr.2015.11.017
Sochacki, J., Devalle, S., Reis, M., Mattos, P., & Rehen, S. (2016). Generation of urine iPS cell lines from patients with Attention Deficit Hyperactivity Disorder (ADHD) using a non-integrative method. Stem Cell Research, 17(1), 102–106. https://doi.org/10.1016/j.scr.2016.05.015
Li, S., Zhao, H., Huang, R., He, L., Tian, C., Huang, H., & Li, Z. (2019). Generation of iPSC line (GIBHi001-A) from a patient with autism spectrum disorder. Stem Cell Research, 40 (September), 101571. https://doi.org/10.1016/j.scr.2019.101571
Dong, Y., Peng, T., Wu, W., Tan, D., Liu, X., & Xie, D. (2019). Efficient introduction of an isogenic homozygous mutation to induced pluripotent stem cells from a hereditary hearing loss family using CRISPR/Cas9 and single-stranded donor oligonucleotides. Journal of International Medical Research, 47(4), 1717–1730. https://doi.org/10.1177/0300060519829990
Jouni, M., Si-Tayeb, K., Es-Salah-Lamoureux, Z., Latypova, X., Champon, B., Caillaud, A., & Gaborit, N. (2015). Toward personalized medicine: Using cardiomyocytes differentiated from urine-derived pluripotent stem cells to recapitulate electrophysiological characteristics of type 2 long QT syndrome. Journal of the American Heart Association, 4(9), 1–13. https://doi.org/10.1161/JAHA.115.002159
He, L., Zhao, H., Li, S., Han, X., Chen, Z., Wang, C., & Jiang, H. (2020). Generation of induced pluripotent stem cell line (CSUXHi002-A) from a patient with spinocerebellar ataxia type 1. Stem Cell Research, 45 (April), 101816. https://doi.org/10.1016/j.scr.2020.101816
Higdon, L. E., Lee, K., Tang, Q., & Maltzman, J. S. (2016). Virtual Global Transplant Laboratory Standard Operating Procedures for Blood Collection, PBMC Isolation, and Storage. Transplantation Direct, 2(9), e101. https://doi.org/10.1097/txd.0000000000000613
Hanna, J., Markoulaki, S., Schorderet, P., Carey, B. W., Beard, C., Wernig, M., & Jaenisch, R. (2008). Direct Reprogramming of Terminally Differentiated Mature B Lymphocytes to Pluripotency. Cell, 133(2), 250–264. https://doi.org/10.1016/j.cell.2008.03.028
Brown, M. E., Rondon, E., Rajesh, D., Mack, A., Lewis, R., Feng, X., Nuwaysir, E. F. (2010). Derivation of induced pluripotent stem cells from human peripheral blood T lymphocytes. PLoS One, 5(6). https://doi.org/10.1371/journal.pone.0011373
Loh, Y. H., Hartung, O., Li, H., Guo, C., Sahalie, J. M., Manos, P. D., & Daley, G. Q. (2010). Reprogramming of T cells from human peripheral blood. Cell Stem Cell, 7(1), 15–19. https://doi.org/10.1016/j.stem.2010.06.004
Hong, H., Takahashi, K., Ichisaka, T., Aoi, T., Kanagawa, O., Nakagawa, M., & Yamanaka, S. (2009). Suppression of induced pluripotent stem cell generation by the p53–p21 pathway. Nature, 460(7259), 1132–1135. https://doi.org/10.1038/nature08235
Seki, T., Yuasa, S., Oda, M., Egashira, T., Yae, K., Kusumoto, D., & Fukuda, K. (2010). Generation of induced pluripotent stem cells from human terminally differentiated circulating t cells. Cell Stem Cell, 7(1), 11–14. https://doi.org/10.1016/j.stem.2010.06.003
Choi, S. M., Liu, H., Chaudhari, P., Kim, Y., Cheng, L., Feng, J., & Jang, Y.-Y. (2011). Reprogramming of EBV-immortalized B-lymphocyte cell lines into induced pluripotent stem cells. Blood, 118(7), 1801–1805. https://doi.org/10.1182/blood-2011-03-340620
Rajesh, D., Dickerson, S. J., Yu, J., Brown, M. E., Thomson, J. A., & Seay, N. J. (2011). Human lymphoblastoid B-cell lines reprogrammed to EBV-free induced pluripotent stem cells. Blood, 118(7), 1797–1800. https://doi.org/10.1182/blood-2011-01-332064
Barrett, R., Ornelas, L., Yeager, N., Mandefro, B., Sahabian, A., Lenaeus, L., & Sareen, D. (2014). Reliable Generation of Induced Pluripotent Stem Cells From Human Lymphoblastoid Cell Lines. STEM CELLS Translational Medicine, 3(12), 1429–1434. https://doi.org/10.5966/sctm.2014-0121
Kishino, Y., Seki, T., Fujita, J., Yuasa, S., Tohyama, S., Kunitomi, A., & Fukuda, K. (2014). Derivation of transgene-free human induced pluripotent stem cells from human peripheral T cells in defined culture conditions. PLoS ONE, 9(5), 1–8. https://doi.org/10.1371/journal.pone.0097397
Muñoz-López, Á., Van Roon, E. H. J., Romero-Moya, D., López-Millan, B., Stam, R. W., Colomer, D., & Menendez, P. (2016). Cellular ontogeny and hierarchy influence the reprogramming efficiency of human B cells into induced pluripotent stem cells. Stem Cells, 34(3), 581–587. https://doi.org/10.1002/stem.2303
Chou, B.-K., Gu, H., Gao, Y., Dowey, S. N., Wang, Y., Shi, J., & Cheng, L. (2015). A Facile Method to Establish Human Induced Pluripotent Stem Cells From Adult Blood Cells Under Feeder-Free and Xeno-Free Culture Conditions: A Clinically Compliant Approach. STEM CELLS Translational Medicine, 4(4), 320–332. https://doi.org/10.5966/sctm.2014-0214
Serwold, T., Hochedlinger, K., Swindle, J., Hedgpeth, J., Jaenisch, R., & Weissman, I. L. (2010). T-cell receptor-driven lymphomagenesis in mice derived from a reprogrammed T cell. Proceedings of the National Academy of Sciences, 107(44), 18939–18943. https://doi.org/10.1073/pnas.1013230107
Dowey, S. N., Huang, X., Chou, B. K., Ye, Z., & Cheng, L. (2012). Generation of integration-free human induced pluripotent stem cells from postnatal blood mononuclear cells by plasmid vector expression. Nature Protocols, 7(11), 2013–2021. https://doi.org/10.1038/nprot.2012.121
Loh, Y. H., Agarwal, S., Park, I. H., Urbach, A., Huo, H., Heffner, G. C., & Daley, G. Q. (2009). Generation of induced pluripotent stem cells from human blood. Blood, 113(22), 5476–5479. https://doi.org/10.1182/blood-2009-02-204800
Ye, Z., Zhan, H., Mali, P., Dowey, S., Williams, D. M., Jang, Y. Y., & Cheng, L. (2009). Human-induced pluripotent stem cells from blood cells of healthy donors and patients with acquired blood disorders. Blood, 114(27), 5473–5480. https://doi.org/10.1182/blood-2009-04-217406
Merling, R. K., Sweeney, C. L., Choi, U., De Ravin, S. S., Myers, T. G., Otaizo-Carrasquero, F., & Malech, H. L. (2013). Transgene-free iPSCs generated from small volume peripheral blood nonmobilized CD34+ cells. Blood, 121(14), e98–e107. https://doi.org/10.1182/blood-2012-03-420273
Mack, A. A., Kroboth, S., Rajesh, D., & Wang, W. B. (2011). Generation of Induced Pluripotent Stem Cells from CD34+ Cells across Blood Drawn from Multiple Donors with Non-Integrating Episomal Vectors. PLoS ONE, 6(11), e27956. https://doi.org/10.1371/journal.pone.0027956
de Leeuw, V. C., van Oostrom, C. T. M., Imholz, S., Piersma, A. H., Hessel, E. V. S., & Dollé, M. E. T. (2020). Going Back and Forth: Episomal Vector Reprogramming of Peripheral Blood Mononuclear Cells to Induced Pluripotent Stem Cells and Subsequent Differentiation into Cardiomyocytes and Neuron-Astrocyte Co-cultures. Cellular Reprogramming, 22(6), 300–310. https://doi.org/10.1089/cell.2020.0040
Sohn, S. K., Kim, J. G., Seo, K. W., Chae, Y. S., Jung, J. T., Suh, J. S., & Lee, K. B. (2002). GM-CSF-based mobilization effect in normal healthy donors for allogeneic peripheral blood stem cell transplantation. Bone Marrow Transplantation, 30(2), 81–86. https://doi.org/10.1038/sj.bmt.1703598
Cashen, A. F., Lazarus, H. M., & Devine, S. M. (2007). Mobilizing stem cells from normal donors: is it possible to improve upon G-CSF? Bone Marrow Transplantation, 39(10), 577–588. https://doi.org/10.1038/sj.bmt.1705616
Szilvassy, S. J., Meyerrose, T. E., Ragland, P. L., & Grimes, B. (2001). Differential homing and engraftment properties of hematopoietic progenitor cells from murine bone marrow, mobilized peripheral blood, and fetal liver. Blood, 98(7), 2108–2115. https://doi.org/10.1182/blood.V98.7.2108
Nagler, A., Korenstein-Ilan, A., Amiel, A., & Avivi, L. (2004). Granulocyte colony-stimulating factor generates epigenetic and genetic alterations in lymphocytes of normal volunteer donors of stem cells. Experimental Hematology, 32(1), 122–130. https://doi.org/10.1016/j.exphem.2003.09.007
Hernández, J. M., Castilla, C., Gutiérrez, N. C., Isidro, I. M., Delgado, M., & de las Rivas, J., San Miguel, J. F. . (2005). Mobilisation with G-CSF in healthy donors promotes a high but temporal deregulation of genes. Leukemia, 19(6), 1088–1091. https://doi.org/10.1038/sj.leu.2403753
Kunisato, A., Wakatsuki, M., Shinba, H., Ota, T., Ishida, I., & Nagao, K. (2011). Direct generation of induced pluripotent stem cells from human nonmobilized blood. Stem Cells and Development, 20(1), 159–168. https://doi.org/10.1089/scd.2010.0063
Zheng, W., Wang, Y., Chang, T., Huang, H., & Yee, J.-K. (2013). Significant differences in genotoxicity induced by retrovirus integration in human T cells and induced pluripotent stem cells. Gene, 519(1), 142–149. https://doi.org/10.1016/j.gene.2013.01.009
Potirat, P., Wattanapanitch, M., Kheolamai, P., & Issaragrisil, S. (2018). Establishment of a human iPSC line (MUSIi007-A) from peripheral blood of normal individual using Sendai viral vectors. Stem Cell Research, 32 (July), 43–46. https://doi.org/10.1016/j.scr.2018.08.014
Okumura, T., Horie, Y., Lai, C. Y., Lin, H. T., Shoda, H., Natsumoto, B., & Otsu, M. (2019). Robust and highly efficient hiPSC generation from patient non-mobilized peripheral blood-derived CD34+ cells using the auto-erasable Sendai virus vector. Stem Cell Research and Therapy, 10(1), 4–6. https://doi.org/10.1186/s13287-019-1273-2
Vlahos, K., Sourris, K., Mayberry, R., McDonald, P., Bruveris, F. F., Schiesser, J. V., Elefanty, A. G. (2019). Generation of iPSC lines from peripheral blood mononuclear cells from 5 healthy adults. Stem Cell Research, 34 (November 2018), 101380. https://doi.org/10.1016/j.scr.2018.101380
Hamada, A., Akagi, E., Obayashi, F., Yamasaki, S., Koizumi, K., Ohtaka, M., & Okamoto, T. (2020). Induction of Noonan syndrome-specific human-induced pluripotent stem cells under serum-, feeder-, and integration-free conditions. Vitro Cellular and Developmental Biology - Animal, 56(10), 888–895. https://doi.org/10.1007/s11626-020-00515-9
Ye, L., Muench, M. O., Fusaki, N., Beyer, A. I., Wang, J., Qi, Z., & Kan, Y. W. (2013). Blood Cell-Derived Induced Pluripotent Stem Cells Free of Reprogramming Factors Generated by Sendai Viral Vectors. STEM CELLS Translational Medicine, 2(8), 558–566. https://doi.org/10.5966/sctm.2013-0006
Tan, H.-K., Toh, C.-X.D., Ma, D., Yang, B., Liu, T. M., Lu, J., & Loh, Y.-H. (2014). Human Finger-Prick Induced Pluripotent Stem Cells Facilitate the Development of Stem Cell Banking. STEM CELLS Translational Medicine, 3(5), 586–598. https://doi.org/10.5966/sctm.2013-0195
Su, R. J., Neises, A., & Zhang, X.-B. (2016). Generation of iPS Cells from Human Peripheral Blood Mononuclear Cells Using Episomal Vectors. In Methods in Molecular Biology (pp. 57–69). https://doi.org/10.1007/7651_2014_139
Sharma, A., Mücke, M., & Seidman, C. E. (2018). Human Induced Pluripotent Stem Cell Production and Expansion from Blood using a Non‐Integrating Viral Reprogramming Vector. Current Protocols in Molecular Biology, 122(1). https://doi.org/10.1002/cpmb.58
Okita, K., Yamakawa, T., Matsumura, Y., Sato, Y., Amano, N., Watanabe, A., & Yamanaka, S. (2013). An efficient nonviral method to generate integration-free human-induced pluripotent stem cells from cord blood and peripheral blood cells. Stem Cells, 31(3), 458–466. https://doi.org/10.1002/stem.1293
Su, R. J., Baylink, D. J., Neises, A., Kiroyan, J. B., Meng, X., Payne, K. J., & Zhang, X. B. (2013). Efficient Generation of Integration-Free iPS Cells from Human Adult Peripheral Blood Using BCL-XL Together with Yamanaka Factors. PLoS ONE, 8(5), 1–12. https://doi.org/10.1371/journal.pone.0064496
Nakagawa, M., Taniguchi, Y., Senda, S., Takizawa, N., Ichisaka, T., Asano, K., & Yamanaka, S. (2014). A novel efficient feeder-Free culture system for the derivation of human induced pluripotent stem cells. Scientific Reports, 4, 1–7. https://doi.org/10.1038/srep03594
Agu, C. A., Soares, F. A. C., Alderton, A., Patel, M., Ansari, R., Patel, S., & Kirton, C. M. (2015). Successful generation of human induced pluripotent stem cell lines from blood samples held at room temperature for up to 48 hr. Stem Cell Reports, 5(4), 660–671. https://doi.org/10.1016/j.stemcr.2015.08.012
Liu, J., Brzeszczynska, J., Samuel, K., Black, J., Palakkan, A., Anderson, R. A., & Ross, J. A. (2015). Efficient episomal reprogramming of blood mononuclear cells and differentiation to hepatocytes with functional drug metabolism. Experimental Cell Research, 338(2), 203–213. https://doi.org/10.1016/j.yexcr.2015.08.004
Zhou, H., Martinez, H., Sun, B., Li, A., Zimmer, M., Katsanis, N., & Chang, S. (2015). Rapid and efficient generation of transgene-free iPSC from a small volume of cryopreserved blood. Stem Cell Reviews and Reports, 11(4), 652–665. https://doi.org/10.1007/s12015-015-9586-8
Sommer, A. G., Rozelle, S. S., Sullivan, S., Mills, J. A., Park, S. M., Smith, B. W., & Mostoslavsky, G. (2012). Generation of human induced pluripotent stem cells from peripheral blood using the STEMCCA lentiviral vector. Journal of visualized experiments : JoVE, 68, 1–5. https://doi.org/10.3791/4327
Chen, I. P., Fukuda, K., Fusaki, N., Iida, A., Hasegawa, M., Lichtler, A., & Reichenberger, E. J. (2013). Induced pluripotent stem cell reprogramming by integration-free sendai virus vectors from peripheral blood of patients with craniometaphyseal dysplasia. Cellular Reprogramming, 15(6), 503–513. https://doi.org/10.1089/cell.2013.0037
Quintana-Bustamante, O., & Segovia, J. C. (2016). Generation of patient-specific induced pluripotent stem cell from peripheral blood mononuclear cells by sendai reprogramming vectors. Methods in Molecular Biology, 1353(17), 1–11. https://doi.org/10.1007/7651_2014_170
Kim, Y., Rim, Y. A., Yi, H., Park, N., Park, S. H., & Ju, J. H. (2016). The Generation of Human Induced Pluripotent Stem Cells from Blood Cells: An Efficient Protocol Using Serial Plating of Reprogrammed Cells by Centrifugation. Stem Cells International, 2016. https://doi.org/10.1155/2016/1329459
Ali, M., Kabir, F., Thomson, J. J., Ma, Y., Qiu, C., Delannoy, M., & Riazuddin, S. A. (2019). Comparative transcriptome analysis of hESC- and iPSC-derived lentoid bodies. Scientific Reports, 9(1), 1–9. https://doi.org/10.1038/s41598-019-54258-z
Isogai, S., Yamamoto, N., Hiramatsu, N., Goto, Y., Hayashi, M., Kondo, M., & Imaizumi, K. (2018). Preparation of induced pluripotent stem cells using human peripheral blood monocytes. Cellular Reprogramming, 20(6), 347–355. https://doi.org/10.1089/cell.2018.0024
Ye, H., & Wang, Q. (2018). Efficient Generation of Non-Integration and Feeder-Free Induced Pluripotent Stem Cells from Human Peripheral Blood Cells by Sendai Virus. Cellular Physiology and Biochemistry, 50(4), 1318–1331. https://doi.org/10.1159/000494589
Guan, J., Liu, X., Zhang, H., Yang, X., Ma, Y., Li, Y., Liu, Y. (2020). Reprogramming of human Peripheral Blood Mononuclear Cell (PBMC) from a Chinese patient suffering Duchenne muscular dystrophy to iPSC line (SDQLCHi007-A) carrying deletion of 49–50 exons in the DMD gene. Stem Cell Research, 42 (September 2019). https://doi.org/10.1016/j.scr.2019.101666
Trokovic, R., Weltner, J., Nishimura, K., Ohtaka, M., Nakanishi, M., Salomaa, V., & Kyttälä, A. (2014). Advanced Feeder-Free Generation of Induced Pluripotent Stem Cells Directly From Blood Cells. STEM CELLS Translational Medicine, 3(12), 1402–1409. https://doi.org/10.5966/sctm.2014-0113
Chou, B. K., Mali, P., Huang, X., Ye, Z., Dowey, S. N., Resar, L. M. S., & Cheng, L. (2011). Efficient human iPS cell derivation by a non-integrating plasmid from blood cells with unique epigenetic and gene expression signatures. Cell Research, 21(3), 518–529. https://doi.org/10.1038/cr.2011.12
Morales Pantoja, I. E., Smith, M. D., Rajbhandari, L., Cheng, L., Gao, Y., Mahairaki, V., & Whartenby, K. A. (2020). IPSCs from people with MS can differentiate into oligodendrocytes in a homeostatic but not an inflammatory milieu. PLoS ONE, 15(6), 1–19. https://doi.org/10.1371/journal.pone.0233980
Wen, W., Zhang, J. P., Xu, J., Su, R. J., Neises, A., Ji, G. Z., & Zhang, X. B. (2016). Enhanced generation of integration-free iPSCs from human adult peripheral blood mononuclear cells with an optimal combination of episomal vectors. Stem Cell Reports, 6(6), 873–884. https://doi.org/10.1016/j.stemcr.2016.04.005
Ustyantseva, E. I., Medvedev, S. P., Vetchinova, A. S., Illarioshkin, S. N., Leonov, S. V., & Zakian, S. M. (2020). Generation of an induced pluripotent stem cell line, ICGi014-A, by reprogramming peripheral blood mononuclear cells from a patient with homozygous D90A mutation in SOD1 causing Amyotrophic lateral sclerosis. Stem Cell Research, 42 (September 2019), 101675. https://doi.org/10.1016/j.scr.2019.101675
Staerk, J., Dawlaty, M. M., Gao, Q., Maetzel, D., Hanna, J., Sommer, C. A., & Jaenisch, R. (2010). Reprogramming of Human Peripheral Blood Cells to Induced Pluripotent Stem Cells. Cell Stem Cell, 7(1), 20–24. https://doi.org/10.1016/j.stem.2010.06.002
Riedel, M., Jou, C. J., Lai, S., Lux, R. L., Moreno, A. P., Spitzer, K. W., & Benjamin, I. J. (2014). Functional and pharmacological analysis of cardiomyocytes differentiated from human peripheral blood mononuclear-derived pluripotent stem cells. Stem Cell Reports, 3(1), 131–141. https://doi.org/10.1016/j.stemcr.2014.04.017
Fuerstenau-Sharp, M., Zimmermann, M. E., Stark, K., Jentsch, N., Klingenstein, M., Drzymalski, M., & Hengstenberg, C. (2015). Generation of highly purified human cardiomyocytes from peripheral blood mononuclear cell-derived induced pluripotent stem cells. PLoS ONE, 10(5), 1–21. https://doi.org/10.1371/journal.pone.0126596
Hanatani, T., & Takasu, N. (2021). CiRA iPSC seed stocks (CiRA’s iPSC Stock Project). Stem Cell Research, 50 (April 2019), 102033. https://doi.org/10.1016/j.scr.2020.102033
Churko, J. M., Burridge, P. W., & Wu, J. C. (2013). Generation of human iPSCs from human peripheral blood mononuclear cells using non-integrative sendai virus in chemically defined conditions. In Methods in Molecular Biology (Vol. 1036, pp. 81–88). https://doi.org/10.1007/978-1-62703-511-8_7
Varga, E., Hansen, M., Wüst, T., von Lindern, M., & van den Akker, E. (2017). Generation of human erythroblast-derived iPSC line using episomal reprogramming system. Stem Cell Research, 25, 30–33. https://doi.org/10.1016/j.scr.2017.10.001
Varga, E., Nemes, C., Bock, I., Varga, N., Fehér, A., Dinnyés, A., & Kobolák, J. (2016). Generation of Mucopolysaccharidosis type II (MPS II) human induced pluripotent stem cell (iPSC) line from a 1-year-old male with pathogenic IDS mutation. Stem Cell Research, 17(3), 482–484. https://doi.org/10.1016/j.scr.2016.09.033
Lopez-Onieva, L., Montes, R., Lamolda, M., Romero, T., Ayllon, V., Lozano, M. L., & Real, P. J. (2016). Generation of induced pluripotent stem cells (iPSCs) from a Bernard-Soulier syndrome patient carrying a W71R mutation in the GPIX gene. Stem Cell Research, 16(3), 692–695. https://doi.org/10.1016/j.scr.2016.04.013
Marsoner, F., Marcatili, M., Karnavas, T., Bottai, D., D’Agostino, A., Scarone, S., & Conti, L. (2016). Generation and characterization of an induced pluripotent stem cell (iPSC) line from a patient with clozapine-resistant Schizophrenia. Stem Cell Research, 17(3), 661–664. https://doi.org/10.1016/j.scr.2016.11.005
Lee, H.-K., Morin, P., & Xia, W. (2016). Peripheral blood mononuclear cell-converted induced pluripotent stem cells (iPSCs) from an early onset Alzheimer’s patient. Stem Cell Research, 16(2), 213–215. https://doi.org/10.1016/j.scr.2015.12.050
Táncos, Z., Varga, E., Kovács, E., Dinnyés, A., & Kobolák, J. (2016). Establishment of induced pluripotent stem cell (iPSC) line from a 75-year old patient with late onset Alzheimer’s disease (LOAD). Stem Cell Research, 17(1), 81–83. https://doi.org/10.1016/j.scr.2016.05.013
Griscelli, F., Oudrirhi, N., Feraud, O., Divers, D., Portier, L., Turhan, A. G., & Bennaceur Griscelli, A. (2017). Generation of induced pluripotent stem cell (iPSC) line from a patient with triple negative breast cancer with hereditary exon 17 deletion of BRCA1 gene. Stem Cell Research, 24, 135–138. https://doi.org/10.1016/j.scr.2017.09.003
Zhang, S., Liu, L., Hu, Y., Lv, Z., Li, Q., Gong, W., & Wu, H. (2017). Derivation of human induced pluripotent stem cell (iPSC) line from a 79 year old sporadic male Parkinson’s disease patient. Stem Cell Research, 19, 43–45. https://doi.org/10.1016/j.scr.2016.12.025
Bhargava, N., Jaitly, S., Goswami, S. G., Jain, S., Chakraborty, D., & Ramalingam, S. (2019). Generation and characterization of induced pluripotent stem cell line (IGIBi001-A) from a sickle cell anemia patient with homozygous β-globin mutation. Stem Cell Research, 39 (June), 101484. https://doi.org/10.1016/j.scr.2019.101484
Bozaoglu, K., Gao, Y., Stanley, E., Fanjul-Fernández, M., Brown, N. J., Pope, K., & Lockhart, P. J. (2019). Generation of seven iPSC lines from peripheral blood mononuclear cells suitable to investigate Autism Spectrum Disorder. Stem Cell Research, 39 (June), 101516. https://doi.org/10.1016/j.scr.2019.101516
Gatinois, V., Desprat, R., Becker, F., Pichard, L., Bernex, F., Corsini, C., & Lemaitre, J. M. (2019). Reprogramming of Human Peripheral Blood Mononuclear Cell (PBMC) from a patient suffering of a Werner syndrome resulting in iPSC line (REGUi003-A) maintaining a short telomere length. Stem Cell Research, 39 (July), 101515. https://doi.org/10.1016/j.scr.2019.101515
Gubert, F., Vasques, J. F., Cozendey, T. D., Domizi, P., Toledo, M. F., Kasai-Brunswick, T. H., & Mendez-Otero, R. (2019). Generation of four patient-specific pluripotent induced stem cell lines from two Brazilian patients with amyotrophic lateral sclerosis and two healthy subjects. Stem Cell Research, 37 (February), 101448. https://doi.org/10.1016/j.scr.2019.101448
Lamolda, M., Montes, R., Simón, I., Perales, S., Martínez-Navajas, G., Lopez-Onieva, L., Real, P. J. (2019). GENYOi005-A: An induced pluripotent stem cells (iPSCs) line generated from a patient with Familial Platelet Disorder with associated Myeloid Malignancy (FPDMM) carrying a p.Thr196Ala variant. Stem Cell Research, 41 (September), 101603. https://doi.org/10.1016/j.scr.2019.101603
Montes, R., Mollinedo, P., Perales, S., Gonzalez-Lamuño, D., Ramos-Mejía, V., Fernandez-Luna, J. L., & Real, P. J. (2019). GENYOi004-A: An induced pluripotent stem cells (iPSCs) line generated from a patient with autism-related ADNP syndrome carrying a pTyr719* mutation. Stem Cell Research, 37, 101446. https://doi.org/10.1016/j.scr.2019.101446
Piovani, G., Lanzi, G., Ferraro, R. M., Masneri, S., Barisani, C., Savio, G., & Giliani, S. C. (2019). Generation of induced pluripotent stem cells (iPSCs) from patient with Cri du Chat Syndrome. Stem Cell Research, 35 (December 2018), 101393. https://doi.org/10.1016/j.scr.2019.101393
Tong, J., Lee, K. M., Liu, X., Nefzger, C. M., Vijayakumar, P., Hawi, Z., Bellgrove, M. A. (2019). Generation of four iPSC lines from peripheral blood mononuclear cells (PBMCs) of an attention deficit hyperactivity disorder (ADHD) individual and a healthy sibling in an Australia-Caucasian family. Stem Cell Research, 34 (November 2018), 101353. https://doi.org/10.1016/j.scr.2018.11.014
Zhang, Y., Li, A., Huang, C. L. H., Wang, G., & Wang, D. (2019). Generation of induced pluripotent stem cells (iPSCs) from an infant with catecholaminergic polymorphic ventricular tachycardia carrying the double heterozygous mutations A1855D in RyR2 and Q1362H in SCN10A. Stem Cell Research, 39 (December 2018), 101509. https://doi.org/10.1016/j.scr.2019.101509
Zhu, Y.-J., Zhang, S.-J., Wu, X.-H., Lian, T.-Y., He, Y.-Z., Zhang, Z.-J., Jing, Z.-C. (2020). Generation of an induced pluripotent stem cell line (PUMCHi006-A) derived from a patient with pulmonary arterial hypertension carrying heterozygous c.1339 G > A mutation in PTGIS gene. Stem Cell Research, 49, 102088. https://doi.org/10.1016/j.scr.2020.102088
Thakur, P., Bhargava, N., Jaitly, S., Gupta, P., Kumar Bhattacharya, S., Padma, G., & Ramalingam, S. (2021). Establishment and characterization of induced pluripotent stem cell line (IGIBi002-A) from a β-thalassemia patient with IVS1-5 mutation by non-integrating reprogramming approach. Stem Cell Research, 50, 102124. https://doi.org/10.1016/j.scr.2020.102124
Wang, Y., Yu, H., Chen, Y., Li, G., Lei, Y., & Zhao, J. (2018). Derivation of induced pluripotent stem cells TUSMi006 from an 87-year old Chinese Han Alzheimer’s disease patient carrying GRINB and SORL1 mutations. Stem Cell Research, 31 (July), 127–130. https://doi.org/10.1016/j.scr.2018.07.018
Zhang, L., Xu, M., Liu, G., Wu, R., Meng, S., Xiahou, K., & Zhou, W. (2019). Generation of induced pluripotent stem cell line (IPTi001-A) from a 62-year old sporadic Alzheimer’s disease patient with APOE3 (ε3/ε3) genotype. Stem Cell Research, 41 (September), 101589. https://doi.org/10.1016/j.scr.2019.101589
Zhao, Z., Ji, S., Shi, Z., & Liu, H. (2018). Generation of CSi001-A, a transgene-free, induced pluripotent stem cell line derived from a Parkinson Disease (PD) patient. Stem Cell Research, 33 (September), 1–5. https://doi.org/10.1016/j.scr.2018.09.020
Kamath, A., Ternes, S., McGowan, S., & Moy, A. B. (2018). Virus-free and oncogene-free induced pluripotent stem cell reprogramming in cord blood and peripheral blood in patients with lung disease. Regenerative Medicine, 13(8), 899–915. https://doi.org/10.2217/rme-2018-0041
Paredes, B. D., Martins, G. L. S., Azevedo, C. M., de Sampaio, G. L., & A., Nonaka, C. K. V., Silva, K. N. da, Souza, B. S. de F. . (2019). Generation of three control iPS cell lines for sickle cell disease studies by reprogramming erythroblasts from individuals without hemoglobinopathies. Stem Cell Research, 38 (March), 101454. https://doi.org/10.1016/j.scr.2019.101454
Laperle, A. H., Sances, S., Yucer, N., Dardov, V. J., Garcia, V. J., Ho, R., & Svendsen, C. N. (2020). iPSC modeling of young-onset Parkinson’s disease reveals a molecular signature of disease and novel therapeutic candidates. Nature Medicine, 26(2), 289–299. https://doi.org/10.1038/s41591-019-0739-1
Gao, X., Liao, X., Zhang, J., Lin, J., & Tan, M. (2021). Derivation of an induced pluripotent stem cell line (GWCMCi001-A) from PBMCs of a four-year-old male patient with Immunoglobulin A nephropathy. Stem Cell Research, 50, 102123. https://doi.org/10.1016/j.scr.2020.102123
Wang, B., Liu, C., Zhang, H., Gai, Z., & Liu, Y. (2021). An induced pluripotent stem cell line (SDQLCHi033-A) derived from a patient with maple syrup urine disease type Ib carrying a homozygous mutation in BCKDHB gene. Stem Cell Research, 50, 102146. https://doi.org/10.1016/j.scr.2020.102146
Saha, B., Borgohain, M., Dey, C., & Thummer, R. (2018). Annals of Stem Cell Research & Therapy iPS Cell Generation : Current and Future Challenges. Annals of Stem Cell Research & Therapy, 1(2), 2–5.
Dey, C., Narayan, G., Krishna Kumar, H., Borgohain, M., & Lenka, N. (2017). Cell-Penetrating Peptides as a Tool to Deliver Biologically Active Recombinant Proteins to Generate Transgene-Free Induced Pluripotent Stem Cells. Studies on Stem Cells Research and Therapy, 3(1), 006–015. https://doi.org/10.17352/sscrt.000011
Ebrahimi, B. (2015). Reprogramming of adult stem/progenitor cells into iPSCs without reprogramming factors. Journal of Medical Hypotheses and Ideas, 9(2), 99–103. https://doi.org/10.1016/j.jmhi.2015.09.003
Geti, I., Ormiston, M. L., Rouhani, F., Toshner, M., Movassagh, M., Nichols, J., & Morrell, N. W. (2012). A Practical and Efficient Cellular Substrate for the Generation of Induced Pluripotent Stem Cells from Adults: Blood-Derived Endothelial Progenitor Cells. STEM CELLS Translational Medicine, 1(12), 855–865. https://doi.org/10.5966/sctm.2012-0093
Naujok, O., Kaldrack, J., Taivankhuu, T., Jörns, A., & Lenzen, S. (2010). Selective Removal of Undifferentiated Embryonic Stem Cells from Differentiation Cultures Through HSV1 Thymidine Kinase and Ganciclovir Treatment. Stem Cell Reviews and Reports, 6(3), 450–461. https://doi.org/10.1007/s12015-010-9148-z
Ben-David, U., Mayshar, Y., & Benvenisty, N. (2011). Large-Scale Analysis Reveals Acquisition of Lineage-Specific Chromosomal Aberrations in Human Adult Stem Cells. Cell Stem Cell, 9(2), 97–102. https://doi.org/10.1016/j.stem.2011.06.013
Lee, M.-O., Moon, S. H., Jeong, H.-C., Yi, J.-Y., Lee, T.-H., Shim, S. H., & Cha, H.-J. (2013). Inhibition of pluripotent stem cell-derived teratoma formation by small molecules. Proceedings of the National Academy of Sciences, 110(35), E3281–E3290. https://doi.org/10.1073/pnas.1303669110
Mohseni, R. (2014). Safe Transplantation Of Pluripotent Stem Cell By Preventing Teratoma Formation. Journal of Stem Cell Research & Therapy, 04(06). https://doi.org/10.4172/2157-7633.1000212
Parr, C. J. C., Katayama, S., Miki, K., Kuang, Y., Yoshida, Y., Morizane, A., & Saito, H. (2016). MicroRNA-302 switch to identify and eliminate undifferentiated human pluripotent stem cells. Scientific Reports, 6(1), 32532. https://doi.org/10.1038/srep32532
Kuang, Y., Miki, K., Parr, C. J. C., Hayashi, K., Takei, I., Li, J., & Saito, H. (2017). Efficient, Selective Removal of Human Pluripotent Stem Cells via Ecto-Alkaline Phosphatase-Mediated Aggregation of Synthetic Peptides. Cell Chemical Biology, 24(6), 685-694.e4. https://doi.org/10.1016/j.chembiol.2017.04.010
Sperling., E. L. (2013). Embryonic Stem Cell Therapy – From Bench to Bed. In Pluripotent Stem Cells. InTech. https://doi.org/10.5772/54368
Borgohain, M. P., Narayan, G., Krishna Kumar, H., Dey, C., & Thummer, R. P. (2018). Maximizing Expression and Yield of Human Recombinant Proteins from Bacterial Cell Factories for Biomedical Applications. In P. Kumar, J. K. Patra, & P. Chandra (Eds.), Advances in Microbial Biotechnology (1st Edition, pp. 447–486). New York: Apple Academic Press. https://doi.org/10.1201/9781351248914
Acknowledgements
We thank all the members of the Laboratory for Stem Cell Engineering and Regenerative Medicine (SCERM) for their excellent support. This work was supported by North Eastern Region – Biotechnology Programme Management Cell (NERBPMC), Department of Biotechnology, Government of India (BT/PR16655/NER/95/132/2015), and also by IIT Guwahati Institutional Top-Up on Start-Up Grant.
Author information
Authors and Affiliations
Contributions
All authors contributed to the conception and design of this manuscript. Data collection and interpretation were performed by all the authors. The first draft of the manuscript was written by Arnab Ray (wrote the section on urine cells) and Jahnavy Madhukar Joshi (wrote the section on PBMCs) and Pradeep Kumar Sundaravadivelu (wrote the section on keratinocytes) and thoroughly cross-checked by Khyati Raina. All the authors commented on the previous versions of the manuscript. All authors read and approved the final draft of the manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ray, A., Joshi, J.M., Sundaravadivelu, P.K. et al. An Overview on Promising Somatic Cell Sources Utilized for the Efficient Generation of Induced Pluripotent Stem Cells. Stem Cell Rev and Rep 17, 1954–1974 (2021). https://doi.org/10.1007/s12015-021-10200-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-021-10200-3