Abstract
The G-protein coupled estrogen receptor (GPER), a proposed tumor suppressor, relays short-term non-genomic responses in target cells and tissues. It frequently undergoes down-modulation in primary tumors of the breast, ovary, and endometrium. Liu and co-workers recently reported loss of GPER expression in colorectal cancer and attributed it to DNA methylation-dependent silencing. We hypothesized that GPER expression is inversely correlated with methylation in the upstream CpG island (upCpGi) in the GPER locus. Methylation in the upCpGi was analysed by bisulfite sequencing and correlated with GPER expression in a panel of colon cancer cell lines. Eight downstream CpGs of the upCpGi was differentially methylated across the cell lines. Methylation in this differentially methylated region (DMR) correlated inversely with GPER expression. Two cell lines, namely SW620 and COLO-320DM, were compared in terms of their viability in response to varying concentrations of G1, a GPER specific agonist. SW-620 cells, which had the least methylated DMR and the highest level of GPER expression, showed significant loss of viability with 1 µM G1. COLO-320DM, which had the most methylated DMR and the lowest level of GPER expression, did not show a significant response to 1 µM G1. At 5 µM G1, SW620 cells showed a greater reduction in viability than COLO-320DM cells. DNA methylation in the DMR is inversely correlated with GPER expression. DNA methylation-dependent silencing of GPER may be, at least in part, the underlying reason behind the loss of estrogen’s oncoprotective effect via GPER in the colon.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer is the second most diagnosed cancer in both men and women. It is also the second most leading cause of cancer-related deaths [1]. The incidence of colorectal cancer in women is significantly lower compared to men [2]. Women receiving hormone replacement therapy have a significantly lower risk of colorectal cancer [3, 4]. Premenopausal women diagnosed with colorectal cancer have better survival than age-matched men [5]. These findings posit estrogen as an important etiological factor in the colorectal tissue and indicate its oncoprotective action.
Estrogen exerts its effects on target cells and tissues via genomic and non-genomic pathways. The genomic effects of estrogen are mediated by the canonical estrogen receptors, namely ERα and ERβ. These are ligand-dependent transcription factors encoded by ESR1 and ESR2 genes, respectively [6]. The non-genomic effects of estrogen are mediated by membrane-tethered canonical estrogen receptors [7,8,9,10], ERα36 [11, 12], a splice variant of ERα, or non-canonical estrogen receptors such as the G-protein coupled estrogen receptor (GPER) [13]. Low ERα expression in the colon makes it an unlikely mediator of estrogen actions in this tissue [14]. ERβ is abundantly expressed in the normal colon [15]. Its expression is reduced in colorectal cancer, and its loss is associated with higher Dukes’ stages and poor prognosis [16,17,18,19]. ERβ knockout mice develop colitis-associated neoplasia [20]. Thus, ERβ may be a crucial mediator of estrogen’s oncoprotective effects in the colon, and a target for colorectal cancer prevention [21].
GPER is the most recent entry to the list of membrane-associated ERs (mER). Upon estrogen binding, it produces short-term non-genomic effects, such as increased cAMP, increased cytoplasmic calcium, and activation of PI3K and MAPK. GPER has been studied in almost all physiological systems, such as immune, reproductive, cardiovascular, neuroendocrine, urinary and musculoskeletal [22, 23]. It is an emerging prognostic marker and a potential therapeutic target in endocrine cancers [22, 24,25,26,27,28].
Recent studies provided insights into the role and relevance of GPER in the physiology and pathology of the colonic environment. They projected it as a significant mediator of estrogen actions in the colon [29]. The importance of GPER expression, or lack of it, in the etiology of colorectal cancer and its progression is ambiguous. Liu and co-workers reported reduced GPER expression in colorectal cancer samples compared to normal counterparts. Reduced GPER expression was associated with poor prognosis, suggesting that GPER is a tumor suppressor [30]. On the other hand, hypoxia induces GPER expression. Bustos and co-workers found evidences that support the pro-tumorigenic role of GPER, especially in the face of reduced ERβ expression in the hypoxic tumor microenvironment [31]. Despite the contradiction, the association between altered GPER expression and colorectal cancer is apparent.
Recent studies, including those reported from our laboratory, show that epigenetic silencing is, at least in part, responsible for the loss of GPER expression in breast and colorectal cancer [30, 32,33,34]. In a study on breast cancer cell lines as models, we found that GPER expression had an inverse relationship with methylation in the upstream CpG island (upCpGi) in the GPER locus. We also described a differentially methylated region (DMR) comprising of terminal eight CpG dinucleotides in the upCpGi, which was differentially methylated in two breast cancer cell lines with contrasting GPER expression levels [34]. The methylation status in the DMR has not been examined in colorectal cancer. In the present study, we have examined GPER expression and upCpGi methylation in a panel of colon cancer cell lines.
Materials and methods
Plasticwares, chemicals, and reagents
Cell culture plasticwares were purchased from Eppendorf (Hamburg, Germany). Dulbecco’s modified eagle’s medium (DMEM) and Roswell Park Memorial Institute (RPMI) 1640 with phenol red were purchased from HiMedia (Mumbai, India). Fetal bovine serum (FBS) was from Invitrogen Corporation (Grand Island, NY, USA). Trypsin-EDTA, antibiotics, and Dulbecco’s phosphate-buffered saline (DPBS) were purchased from HiMedia (Mumbai, India). EmeraldAmp RR320B MAX PCR Master Mix was from DSS TAKARA BIO India Pvt. Ltd. (New Delhi, India). The GPER-specific agonist, G1, was purchased from Cayman Chemical (Cat. No. CAS 881639-98-1) (MI, USA). All other chemicals, salts, and buffers were from Merck and SRL (Mumbai, India).
Cell culture
Colon cancer cell lines HT-29, HCT-15, HCT-116, SW-480, SW-620, COLO-205, COLO-320DM, and breast cancer cell line MCF-7 were obtained from the National Centre for Cell Science (NCCS, Pune, India). They were routinely cultured and expanded under standard conditions of 37 °C and 5% CO2 in phenol red-containing DMEM for HT-29 and MCF-7, and RPMI 1640 for the remaining cell lines. Media were supplemented with 10% heat-inactivated FBS, 100 units/mL penicillin, and 100 µg/mL streptomycin.
Total RNA isolation and cDNA synthesis
Total RNA and protein were isolated from cell lines as described previously [34]. 2 µg of total RNA was reverse transcribed using High Capacity cDNA Reverse Transcription kit (Invitrogen, USA) as per manufacturer’s instructions.
RT-PCR
cDNA equivalent to 20 ng of total RNA was used as the template for PCR reactions with gene-specific primers (Table 1). RPL35a served as an internal control. PCR products were resolved in 2% agarose gels, and the images of ethidium bromide-stained bands were captured using Gel Doc EZ Imager (Bio-Rad Laboratories Inc., CA, USA).
Western blotting
Total protein was isolated from cell lines as described previously [35] and quantified by Lowry’s protein estimation method [36]. Proteins were fractionated on 10% denaturing SDS page and transferred to nitrocellulose membranes. The membranes were blocked with 1% gelatin in TBST for 2 h at room temperature followed by overnight incubation with primary antibody at 4 °C in 0.1% gelatin in TBST. The blots were then washed (3 × 10 min) with Tris buffered saline containing 0.05% Tween 20 (TBST) followed by incubation with goat anti-rabbit IgG HRP (secondary antibody) for 1 h. The blots were then washed (3 × 10 min) with TBST and developed using Clarity Western ECL Substrate (Bio-Rad Laboratories Inc., CA, USA). Colorimetric images at suitable exposures were captured using the Bio-Rad Gel Doc EZ Imager. For GPER1 western blots, peptide affinity-purified polyclonal antibody was used [35]. ERα (Cat. No. 8644S) and PR (Cat. No. 8757S) antibodies were purchased from Cell Signaling Technology (MA, USA). Histone H3, which was probed by anti-H3 antibody (Cat. No. BB-AB0055, BioBharati LifeScience, India) served as the internal control. Anti-rabbit secondary antibody (Cat. No. 7074S) from Cell Signaling Technology (MA, USA) was used.
Bisulfite sequencing
Genomic DNA (gDNA) from the cell lines was extracted using PureLink Quick Gel Extraction Kit (Invitrogen, CA, USA) and subjected to bisulfite conversion using the EpiJET Bisulfite Conversion kit (Thermo Fisher Scientific, USA) as per manufacturer’s instructions. The bisulfite-converted gDNA was used as a template to generate upCpGi containing amplicons using specific primers reported earlier [34]. The amplified products were eluted from agarose gels and ligated into the pMD20 vector (Takara Bio, India) as per the manufacturer’s instructions. The ligated products were then transformed into E.coli Top10 competent cells, and recombinant colonies were screened using ampicillin resistance as the selectable marker. Plasmids were isolated using GSure Plasmid Kit (GCC Biotech, India), and the inserts of 10 independent recombinant plasmids per cell line were sequenced. Methylated and unmethylated CpG sites were identified and represented as lollipop plots. The proportion of CpGs methylated in the upCpGi was determined for each cell line. The sequencing was done by Eurofins Scientific (Bangalore, India).
G1 treatment and determination of cell viability
G1 was dissolved in ethanol to make a stock concentration of 2 mM. The stock solution was diluted appropriately with the medium to achieve the desired concentrations. Colon cancer cell lines were seeded (4000–5000 cells/well) in 96 well plates. After 48 h, the monolayer was washed with DPBS. The cells were then processed directly for MTT assay to determine baseline viability or treated with indicated concentrations of G1 or ethanol (vehicle control) for 120 h before MTT assay. In case of the latter, media containing G1 or ethanol were replenished after every 48 h. MTT assays were performed as described previously [37]. The spent medium was removed and monolayer of cells was washed with DPBS and incubated with 100 µL MTT reagent (0.5 mg/mL) for 3 hours in 37 °C. After incubation, MTT reagent was removed, and the formazan crystals were dissolved in DMSO. Absorbance was measured at 570 nm in the Infinite Pro M200 microplate reader (Tecan Life Sciences, Switzerland), with 690 nm as the reference wavelength. The difference in the absorbance (A570-A690) was considered as a measure of cell viability. For each cell line, the change in viability due to treatment by ethanol or G1 was determined by subtracting the baseline viability. The change in viability of ethanol treated cells were assigned the value of 100, and those of G1 treated cells were expressed relative to control. COLO-205 cells, which are semi-adherent cells, were not amenable to 120 h MTT assays, because there would be loss in cell number with each replenishment after every 48 h.
Statistical analysis
Data generated from MTT assays were analyzed using one-way ANOVA followed by Tukey’s HSD using the R statistical package. Methylation in the upCpGi of GPER locus and comparison of percent methylation across all cell lines were performed as described earlier [38]. In all the tests, p < 0.05 was considered statistically significant.
Results
GPER expression in colon cancer cell lines
There are three variants of GPER mRNA, namely NM_001505.2 (GPERv2), NM_001039966.1 (GPERv3) and NM_001098201.1 (GPERv4). They have exactly the same open reading frame and 3′UTR, but have different exon-intron organization and 5′UTR. Using variant-specific primer combinations described earlier [34], we studied the expression of GPER mRNA in a panel of colon cancer cell lines. MCF-7 cell line which expresses the three variants [34], was used as a reference. The colon cancer cell lines expressed varying degrees of GPER mRNA variants (Fig. 1a). In SW-480 and SW-620 cells, the GPERv2 and GPERv3 were the dominant variants. GPERv3 was the major variant in HT-29 cells with very low and undetectable levels of GPERv4 and GPERv2, respectively. HCT-15 showed almost no expression of GPER mRNA. Other cell lines had a moderate expression of GPER mRNA variants (Fig. 1a). We examined GPER protein expression in these cell lines. The protein levels, by and large, were consistent with the levels of GPER mRNA (Fig. 1b). HT-29 was an exception. Despite appreciable level of GPERv3, negligible protein could be detected (Fig. 1a, b). ERα expression was undetectable by RT-PCR and western blotting in the cell lines. RT-PCR analysis of ERβ expression showed very faint amplicons (Supplementary Fig. S2).
GPER expression in colon cancer cell lines. Total RNA and protein were isolated from the indicated cell lines. GPER expression was determined by RT-PCR (a) using primers and PCR conditions described previously [34], and western blotting (b). The raw western blotting data in b is provided as Supplementary Fig. S1
CpG methylation in the upCpGi
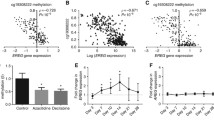
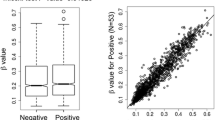
Using bisulfite sequencing approach as described earlier [34], we analysed DNA methylation in the upCpGi, which harbors 32 CpG sites. Figure 2a shows the methylation pattern deduced from bisulfite sequencing data in the form of lollipop models. The upCpGi in COLO-320DM was extensively methylated, which was in sharp contrast to the observation in SW-620 cells. The remaining cell lines showed varying levels of methylation, but not as extensive as that in COLO-320DM. Interestingly, much of the methylation was observed in the downstream terminal eight CpGs in the upCpGi (referred to as the DMR). This was similar to our previous observation in MCF-7 and MDA-MB-231 breast cancer cell lines [34]. The proportion of methylated CpGs in the DMR for each of the cell lines is indicated in parenthesis in Fig. 2a. For each pair of cell lines, the difference in the proportion of methylated CpGs was tested for statistical significance. The adjusted p-values are indicated in Fig. 2b. The level of GPER protein significantly correlated inversely with the proportion of methylated CpGs in the DMR (Fig. 2c, Spearman’s rho = − 0.78, p = 0.048).
upCpGi methylation in the GPER locus. Genomic DNA isolated from the indicated colon cancer cell lines were bisulfite converted and used for PCR reaction with primers described earlier to amplify the upCpGi. The PCR amplified products were cloned in pMD20. Inserts from 10 independent TA clones per cell line were sequenced and analyzed for methylated and unmethylated CpG sites. There are 32 CpG sites in the upCpGi. a Lollipop models depicting the status of each CpG site in the indicated cell lines. Filled circles represent methylated CpGs, and open circles represent unmethylated CpGs. Grey circles represent the absence of a CpG site. Since there are 8 CpGs in the DMR, a total of 80 CpGs were sampled for each cell line (n = 10 clones for each sample). The numbers in parentheses denote the proportion of methylated CpGs sampled in the DMR. b Adjusted p-values for the test of significant difference in proportion of CpGs methylated in the DMR. *p < 0.05, **p < 0.01, ***p < 0.001, ns – not significant. c Relationship between the proportion of CpGs methylated in the DMR and GPER expression in colon cancer cell lines. The GPER protein expression data was obtained from a western blot experiment shown in Fig. 1b, wherein the densitometric readings of GPER bands normalized with histone H3 were considered as a measure of GPER protein expression. The data were analysed by Spearman’s correlation test
Response to GPER stimulation
Liu et al., showed that G1, a specific GPER agonist, induces cell cycle arrest and apoptosis in colon cancer cell lines [30]. We tested whether the response of the cell lines under study to G1 treatment was commensurate with the levels of GPER protein expression. We designed an MTT assay to examine the change in viability of cells following 120 h of treatment with varying concentrations of G1 after due correction of the zero-hour baseline viability. Up to 500 nM G1, none of the cell lines had reduced viability (Fig. 3), except HCT15. HCT15 had significantly reduced viability with 500 nM G1 (Fig. 3c). All the cell lines under study showed significantly reduced viability when treated with 5 µM G1. Notably, SW-620, which had the highest expression of GPER protein, showed significantly reduced viability in response to 1 µM G1 (Figs. 1b, 3b). COLO-320DM, with no or negligible expression of GPER, did not show significant reduction of viability at 1 µM G1 (Figs. 1b, 3e). The two cell lines significantly differed in their response to 5 µM G1. At this concentration the viability of SW620 was reduced to 8.44%, whereas the viability of COLO-320DM was reduced to 29.36% (Fig. 3b, e).
Effect of G1 treatment on viability of colon cancer cell lines. The indicated cell lines were treated with varying concentrations of G1 for a period of 120 h. The change in viability was determined as described in materials and methods. Bars represent mean relatively viability ± SD [n = 04 (SW620) and n = 03 (for other cell lines); each biological replicate comprising of 6 technical replicate wells for each of the treatment groups]
Discussion
The recognition of GPER as a mediator of estrogen effects not only brought complexity to estrogen signaling, but also ushered a new dimension to the etiology, progression, and therapeutic resistance of endocrine cancers. Recent studies have expanded the scope and relevance of GPER expression and regulation beyond the traditionally conceived endocrine (estrogen) responsive tissues [26]. Several reports implicated GPER in the pathophysiology of colon cancer, although there is ambiguity about the role of GPER. There are evidences for tumor suppressor [30] and pro-tumorigenic [31] role for GPER in the colon. Results from Santolla et al., suggested that GPER1 activation may lead to tumor progression via induction of FASN, a putative metabolic oncogene [39]. GPER1 may also play a tumor augmenting role in the face of dysregulated estrogen metabolism that favours estradiol; a caveat in the use of hormone replacement therapy in individuals with high risk of colon cancer [40, 41]. The present work is set against the backdrop of growing literature on epigenetic silencing of GPER in diverse pathological contexts [30, 32,33,34, 42, 43], and the proposed tumor suppressor role of GPER [30].
Liu and co-workers found that GPER down-modulation in colorectal cancer was associated with promoter methylation and histone deacetylation [30]. They analyzed a CpG island (between − 461 and − 781) in the GPER promoter by bisulfite sequencing and found significantly greater proportion of methylated CpGs in colorectal cancer compared to normal tissues. Liu and co-workers also analysed the CpG island in colorectal cancer cell lines. However, their conclusion was based on few sequences, with no statistical analysis. The region analyzed by Liu and co-workers overlapped with the upCpGi. However, it did not include the DMR, which we had described earlier in breast cancer cell lines [34]. The technology used to generate TCGA methylation data does not have probes to interrogate the DMR. Hence, TCGA colorectal cancer data was not useful in the analysis of methylation in the DMR. The present work was motivated by the need to address the following missing links- (a) the status of methylation in the DMR in colorectal cancer cells, and (b) the correlation between GPER expression and methylation in the upCpGi.
To our knowledge, this is the first study to analyze GPER expression in colon cancer cell lines and correlate it with the extent of methylation in the upCpGi, with special reference to the DMR. One of the striking observations from our bisulfite sequencing data is the concentration of methylated CpGs in the DMR of the upCpGi. This is reminiscent of the methylation pattern reported earlier in MCF-7 and MDA-MB-231 breast cancer cells. In the COLO-320DM cell line the methylated CpGs were found throughout the upCpGi, and not just in the DMR. COLO-320DM and SW-620 cells represented two extreme states of methylation in the DMR. The remaining cell lines showed varying degrees of CpG methylation in the DMR. Interestingly, inserts in 6 clones from HCT-15 cells, which were sequenced showed the absence of the twenty-fifth CpG dinucleotide. Similarly, one clone from HT-29 had an absence of the first CpG dinucleotide (Fig. 2a). We believe that this is due to the heterogeneity brought about by mutations that accumulate during cell passages. The inverse correlation between methylation in the DMR and GPER expression suggests that DNA methylation dependent silencing of GPER could be the underlying cause of GPER downmodulation in colon cancer.
Estrogen’s oncoprotective effect on the colorectal tissue has a sound epidemiological basis. The receptors that transduce the protective effects of estrogen merit special attention on the following counts. First, they hold the key to our understanding of the effects of estrogen in a tissue traditionally considered non-endocrine. Second, they may inspire novel therapeutic strategies against colorectal cancer. Given the absence or undetectable expression of ERα in the cell lines used in this study (Supplementary Fig. S2), the study also attempted to correlate the status of methylation in the DMR, or GPER expression, with response to G1. We were not successful in finding a significant correlation. However, the pattern of methylation, GPER expression, and response to G1 in COLO-320DM and SW-620 cells, which have contrasting levels of methylation and GPER expression, suggest that reduced GPER expression brought about by DNA methylation results in reduced response to G1. Given the low or undetectable levels of ERα and ERβ (Supplementary Fig. S2), GPER serves as a mediator for estrogen signaling to influence the physiology of colon cancer cell lines. Low ERα expression in the colonic epithelium [14] makes GPER a crucial mediator of estrogen’s effects in the colon; more so in the face of ERβ loss under hypoxia [31]. GPER activation induces cell cycle arrest and apoptosis in colon cancer cells [30]. Hence, we propose that DNA methylation dependent loss of GPER expression negatively impacts estrogen’s oncoprotective effect in the colonic epithelium. This in turn contributes to colon cancer.
Tumor suppressors are frequently inactivated through methylation of DNA in neoplastic conditions. The biomarker and prognostic potential of methylated CpG islands in specific tumor suppressors are appreciated generally. Despite the ambiguity in the perceived role of GPER in the colorectal tissue, its expression level in primary tumors appears to be an important clinical marker. The present study confirms the inverse relationship between upCpGi methylation and GPER expression in cell line models. Furthermore, it underscores the importance of the DMR that encompasses the terminal eight CpGs in methylation-dependent silencing of GPER. More investigations are warranted to explore DMR hypermethylation as an indicator of GPER expression in colon cancer and its utility in diagnosis and prognosis.
References
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Simon MS, Chlebowski RT, Wactawski-Wende J et al (2012) Estrogen plus progestin and colorectal cancer incidence and mortality. J Clin Oncol 30:3983–3990. https://doi.org/10.1200/JCO.2012.42.7732
Lin KJ, Cheung WY, Lai JY-C, Giovannucci EL (2012) The effect of estrogen vs. combined estrogen-progestogen therapy on the risk of colorectal cancer. Int J cancer 130:419–430. https://doi.org/10.1002/ijc.26026
Rossouw JE, Anderson GL, Prentice RL et al (2002) Risks and benefits of estrogen plus progestin in healthy postmenopausal women: principal results From the Women’s Health Initiative randomized controlled trial. JAMA 288:321–333. https://doi.org/10.1001/jama.288.3.321
Quaresma M, Coleman MP, Rachet B (2015) 40-Year trends in an index of survival for all cancers combined and survival adjusted for age and sex for each cancer in England and Wales, 1971–2011: a population-based study. Lancet 385:1206–1218. https://doi.org/10.1016/S0140-6736(14)61396-9
Nilsson S, Mäkelä S, Treuter E et al (2001) Mechanisms of estrogen action. Physiol Rev 81:1535–1565. https://doi.org/10.1152/physrev.2001.81.4.1535
Hammes SR, Levin ER (2007) Extranuclear steroid receptors: nature and actions. Endocrinol Rev 28:726–741. https://doi.org/10.1210/er.2007-0022
Hammes SR, Levin ER (2011) Minireview: recent advances in extranuclear steroid receptor actions. Endocrinology 152:4489–4495. https://doi.org/10.1210/en.2011-1470
Lange CA, Gioeli D, Hammes SR, Marker PC (2007) Integration of rapid signaling events with steroid hormone receptor action in breast and prostate cancer. Annu Rev Physiol 69:171–199. https://doi.org/10.1146/annurev.physiol.69.031905.160319
Losel RM, Falkenstein E, Feuring M et al (2003) Nongenomic steroid action: controversies, questions, and answers. Physiol Rev 83:965–1016. https://doi.org/10.1152/physrev.00003.2003
Chaudhri RA, Schwartz N, Elbaradie K et al (2014) Role of ERα36 in membrane-associated signaling by estrogen. Steroids 81:74–80. https://doi.org/10.1016/j.steroids.2013.10.020
Sołtysik K, Czekaj P (2015) ERα36–another piece of the estrogen puzzle. Eur J Cell Biol 94:611–625. https://doi.org/10.1016/j.ejcb.2015.10.001
Xu S, Yu S, Dong D, Lee LTO (2019) G protein-coupled estrogen receptor: a potential therapeutic target in cancer. Front Endocrinol (Lausanne) 10:725. https://doi.org/10.3389/fendo.2019.00725
Kennelly R, Kavanagh DO, Hogan AM, Winter DC (2008) Oestrogen and the colon: potential mechanisms for cancer prevention. Lancet Oncol 9:385–391. https://doi.org/10.1016/S1470-2045(08)70100-1
Papaxoinis K, Triantafyllou K, Sasco AJ et al (2010) Subsite-specific differences of estrogen receptor beta expression in the normal colonic epithelium: implications for carcinogenesis and colorectal cancer epidemiology. Eur J Gastroenterol Hepatol 22:614–619. https://doi.org/10.1097/MEG.0b013e328335ef50
Konstantinopoulos PA, Kominea A, Vandoros G et al (2003) Oestrogen receptor beta (ERbeta) is abundantly expressed in normal colonic mucosa, but declines in colon adenocarcinoma paralleling the tumour’s dedifferentiation. Eur J Cancer 39:1251–1258. https://doi.org/10.1016/s0959-8049(03)00239-9
Jassam N, Bell SM, Speirs V, Quirke P (2005) Loss of expression of oestrogen receptor beta in colon cancer and its association with Dukes’ staging. Oncol Rep 14:17–21
Elbanna HG, Ebrahim MA, Abbas AM et al (2012) Potential value of estrogen receptor beta expression in colorectal carcinoma: interaction with apoptotic index. J Gastrointest Cancer 43:56–62. https://doi.org/10.1007/s12029-010-9214-4
Rudolph A, Toth C, Hoffmeister M et al (2012) Expression of oestrogen receptor β and prognosis of colorectal cancer. Br J Cancer 107:831–839. https://doi.org/10.1038/bjc.2012.323
Saleiro D, Murillo G, Benya RV et al (2012) Estrogen receptor-β protects against colitis-associated neoplasia in mice. Int J cancer 131:2553–2561. https://doi.org/10.1002/ijc.27578
Williams C, DiLeo A, Niv Y, Gustafsson J-Å (2016) Estrogen receptor beta as target for colorectal cancer prevention. Cancer Lett 372:48–56. https://doi.org/10.1016/j.canlet.2015.12.009
Prossnitz ER, Barton M (2009) Signaling, physiological functions and clinical relevance of the G protein-coupled estrogen receptor GPER. Prostaglandins Other Lipid Mediat 89:89–97. https://doi.org/10.1016/j.prostaglandins.2009.05.001
Maggiolini M, Picard D (2010) The unfolding stories of GPR30, a new membrane-bound estrogen receptor. J Endocrinol 204:105–114. https://doi.org/10.1677/JOE-09-0242
Smith HO, Leslie KK, Singh M et al (2007) GPR30: a novel indicator of poor survival for endometrial carcinoma. Am J Obstet Gynecol 196:386.e1-386.e11. https://doi.org/10.1016/j.ajog.2007.01.004
Smith HO, Arias-Pulido H, Kuo DY et al (2009) GPR30 predicts poor survival for ovarian cancer. Gynecol Oncol 114:465–471. https://doi.org/10.1016/j.ygyno.2009.05.015
Prossnitz ER, Barton M (2014) Estrogen biology: new insights into GPER function and clinical opportunities. Mol Cell Endocrinol 389:71–83. https://doi.org/10.1016/j.mce.2014.02.002
Gaudet HM, Cheng SB, Christensen EM, Filardo EJ (2015) The G-protein coupled estrogen receptor, GPER: the inside and inside-out story. Mol Cell Endocrinol 418 Pt 3:207–219. https://doi.org/10.1016/j.mce.2015.07.016
Barton M (2016) Not lost in translation: Emerging clinical importance of the G protein-coupled estrogen receptor GPER. Steroids 111:37–45. https://doi.org/10.1016/j.steroids.2016.02.016
Jacenik D, Beswick EJ, Krajewska WM, Prossnitz ER (2019) G protein-coupled estrogen receptor in colon function, immune regulation and carcinogenesis. World J Gastroenterol 25:4092–4104. https://doi.org/10.3748/wjg.v25.i30.4092
Liu Q, Chen Z, Jiang G et al (2017) Epigenetic down regulation of G protein-coupled estrogen receptor (GPER) functions as a tumor suppressor in colorectal cancer. Mol Cancer 16:87. https://doi.org/10.1186/s12943-017-0654-3
Bustos V, Nolan ÁM, Nijhuis A et al (2017) GPER mediates differential effects of estrogen on colon cancer cell proliferation and migration under normoxic and hypoxic conditions. Oncotarget 8:84258–84275. https://doi.org/10.18632/oncotarget.20653
Weißenborn C, Ignatov T, Poehlmann A et al (2014) GPER functions as a tumor suppressor in MCF-7 and SK-BR-3 breast cancer cells. J Cancer Res Clin Oncol 140:663–671. https://doi.org/10.1007/s00432-014-1598-2
Weissenborn C, Ignatov T, Nass N et al (2017) GPER promoter methylation controls GPER expression in breast cancer patients. Cancer Invest 35:100–107. https://doi.org/10.1080/07357907.2016.1271886
Manjegowda MC, Gupta PS, Limaye AM (2017) Hyper-methylation of the upstream CpG island shore is a likely mechanism of GPER1 silencing in breast cancer cells. Gene 614:65–73. https://doi.org/10.1016/j.gene.2017.03.006
Manjegowda MC, Gupta PS, Limaye AM (2016) Validation data of a rabbit antiserum and affinity purified polyclonal antibody against the N-terminus of human GPR30. Data Br 7:1015–1020. https://doi.org/10.1016/j.dib.2016.03.054
LOWRY OH, ROSEBROUGH NJ, RANDALL FARRAL RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Manjegowda MC, Deb G, Limaye AM (2014) Epigallocatechin gallate induces the steady state mRNA levels of pS2 and PR genes in MCF-7 breast cancer cell. Indian J Exp Biol 52:312–316
John Mary DJS, Sikarwar G, Kumar A, Limaye AM (2020) Interplay of ERα binding and DNA methylation in the intron-2 determines the expression and estrogen regulation of cystatin A in breast cancer cells. Mol Cell Endocrinol 504:110701. https://doi.org/10.1016/j.mce.2020.110701
Santolla MF, Lappano R, De Marco P et al (2012) G protein-coupled estrogen receptor mediates the up-regulation of fatty acid synthase induced by 17β-estradiol in cancer cells and cancer-associated fibroblasts. J Biol Chem 287:43234–43245. https://doi.org/10.1074/jbc.M112.417303
Gilligan LC, Gondal A, Tang V et al (2017) Estrone sulfate transport and steroid sulfatase activity in colorectal cancer: implications for hormone replacement therapy. Front Pharmacol 8:103. https://doi.org/10.3389/fphar.2017.00103
Gilligan LC, Rahman HP, Hewitt A-M et al (2017) Estrogen activation by steroid sulfatase increases colorectal cancer proliferation via GPER. J Clin Endocrinol Metab 102:4435–4447. https://doi.org/10.1210/jc.2016-3716
Weißenborn C, Ignatov T, Ochel H-J et al (2014) GPER functions as a tumor suppressor in triple-negative breast cancer cells. J Cancer Res Clin Oncol 140:713–723. https://doi.org/10.1007/s00432-014-1620-8
Lv Q-Y, Xie B-Y, Yang B-Y et al (2017) Increased TET1 expression in inflammatory microenvironment of hyperinsulinemia enhances the response of endometrial cancer to estrogen by epigenetic modulation of GPER. J Cancer 8:894–902. https://doi.org/10.7150/jca.17064
Acknowledgements
The work was supported by financial assistance from the Indian Council of Medical Research, Govt. of India (Sanction letter No. 2019 − 0687/CMB/ADHOC-BMS, dated 13.11.2019). Infrastructural support from the Department of Biosciences and Bioengineering at IIT Guwahati is acknowledged. SG was a student of the M.Tech program supported by the Department of Biotechnology, Govt. of India, which also provided her fellowship.
Author information
Authors and Affiliations
Contributions
UP and AML designed the experiments. UP and SG performed the experiments. UP and AML analysed the results. UP, SG, and AML wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Research involving human and animal rights
The work involves cell line models. No human participants and animals are involved in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pal, U., Ghosh, S. & Limaye, A.M. DNA methylation in the upstream CpG island of the GPER locus and its relationship with GPER expression in colon cancer cell lines. Mol Biol Rep 47, 7547–7555 (2020). https://doi.org/10.1007/s11033-020-05817-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05817-5