Abstract
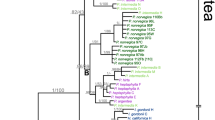
Bromus hordeaceus (sectionBromus, Poaceae), a predominantly self-fertilizing tetraploid (2n=28), is an annual weed native to the Mediterranean Basin, which now has a world-wide distribution. High morphological variation led to the recognition of four subspecies, three of which correlated with habitat-type. We examined genetic diversity at enzyme loci in 15 populations from the Mediterranean and the Atlantic region. Although sampled over a larger range of ecological and geographical conditions, the North-African populations appeared less genetically differentiated than populations from Brittany, suggesting higher levels of gene flow among the first ones (Nm=3.756 and 1.066 respectively). No genetic differentiation was encountered among the four subspecies. The populations were homozygous at homologous loci, suggesting high rates of selfing, but they frequently exhibited fixed intergenomic heterozygosity. The meiotic chromosome behaviour and disomic inheritance encountered are in accordance with the previously proposed allopolyploid origin of the species. The diploidsB. arvensis andB. scoparius have been previously implicated in the parentage ofB. hordeaceus on the basis of morphology and serology. We comparedB. hordeaceus with related diploid species belonging to the same section (sectionBromus) using different sources of data (flow cytometry, karyotypes, RAPD and DNA sequences). Molecular phylogeny based on internal transcribed spacer sequences of nuclear ribosomal genes provided the first clear scheme of relationships among monogenomic species of the section. A new hypothesis is proposed concerning the origin ofB. hordeaceus: We found that it diverged earlier than all other species of sectionBromus excluding the diploidB. caroli-henrici which is basal in this group. The 13 autapomorphies accumulated byB. hordeaceus, and the absence of intra-individual sequence heterogeneity are also consistent with the relatively ancient origin of the species within the section.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Adams R.P. &Rieseberg L.H. (1998): The effects of non-homology in RAPD bands on similarity and multivariate statistical ordination inBrassica andHelianthus.Theor. Appl. Genet. 97: 323–326.
Ainouche M. (1993):Les populations diploides et tétraploides du genre BromusL. section BromusSM. (Poaceae):Analyse de la diversité génétique. PhD. thesis, University of Rennes 1 Rennes.
Ainouche M., Misset M. T. &Huon A. (1995): Genetic diversity in Mediterranean diploid and tetraploidBromus L. (sectionBromus Sm.) populations.Genome 38: 879–888.
Ainouche M., Misset M. T. &Huon A. (1996): Patterns of genetic differentiation in two annual bromegrasses,Bromus lanceolatus andB. hordeaceus (Poaceae), Pl. Syst. Evol. 199: 65–68.
Ainouche M. &Bayer R. J. (1997): On the origins of the tetraploidBromus species (sectionBromus, Poaceae): insights from internal transcribed spacer sequences of nuclear ribosomal DNA.Genome 40: 730–743.
Armstrong K.C. (1991): Chromosome evolution ofBromus. In:Tsuchiya T. &Gupta P.K. (eds.),Chromosome engineering in plants, Part B, Elsevier Science Publishers, Amsterdam, pp. 363–377.
Bachmann K. (1997): Nuclear DNA markers in plant biosystematic research.Opera Bot. 132: 137–148.
Baker H.G. (1974): The evolution of weeds.Annual Rev. Ecol. Syst. 5: 1–24.
Barrett S.C.H. &Shore J.S. (1989): Isozyme variation in colonizing plants. In:Soltis D.E. &Soltis P.S. (eds.),Isozymes in plant biology, Chapman and Hall, London, pp. 106–126.
Baum D.A., Systsma K.J. &Koch P.C. (1994): A phylogenetic analysis ofEpilobium (Onagraceae) based on nuclear ribosomal DNA sequences.Syst. Bot. 19: 363–388.
Bremer K. (1988): The limits of amino acid sequence data in angiosperm phylogenetic reconstruction.Evolution 42: 795–803.
Brown A.H.D. &Marshall D.R. (1981): Evolutionary changes accompanying colonization in plants. In:Scudder G.C.E. &Reveal J.L. (eds.),Evolution today, Hunt Institute for Botanical Documentation, carnegie-Mellon University Press, Pittsburgh, pp. 351–363.
Da Silva J.A.G. &Sobral B.W.S. (1996): Genetics of polyploids. In:Sobral B.W.S. (ed.),The impact of plant molecular genetics, Birkhäuser, Boston, pp. 3–37.
Daehler C.C. (1998): The taxonomic distribution of invasive angiosperm plants: ecological insights and comparison to agriculture weeds.Biol. Conservation 84: 167–180.
Donoghue M., Olmstead R., Smith J. &Palmer J. (1992): Phylogenetic relationships ofDipsacales based onrbcL sequences.Ann. Missouri Bot. Gard. 79: 333–345.
Feldman M., Liu B., Segal G., Abbo S., Levy A.A. &Vega J.M. (1997): Rapid elimination of low-copy DNA sequences in polyploid wheat: A possible mechanism for differentiation of homoeologous chromosomes.Genetics 147: 1381–1387.
Felsenstein J. (1985): Confidence limits on phylogenies: an approach using the bootstrap.Evolution 39: 783–791.
Govindajaru R.D. (1989): Variation in gene flow levels among predominantly self-pollinated plants.J. Evol. 2: 173–181.
Gower J.C. (1966): Some distance properties of latent root and vector use in multivariate analysis.Biometrika 53: 326–338.
Jain S.K. (1978): Inheritance of phenotypic plasticity in soft chess,Bromus mollis L. (Gramineae).Experientia 34: 385–386.
Hsiao C., Chatterton N.J., Asay K.H. &Jensen K.B. (1994): Phylogenetic relationships of 10 grass species: an assessment of phylogenetic utility of the internal transcribed spacer region in nuclear ribosomal DNA in monocots.Genome 37: 117–120.
Hsiao C., Chatterton N.J., Asay K.H. &Jensen K.B. (1995): Molecular phylogeny of thePooideae (Poaceae) based on nuclear rDNA (ITS) sequences.Theor. Appl. Genet. 90: 389–398.
Lamboy W.F. (1994): Computing genetic similarity coefficients from RAPD data: the effects of PCR artefacts.PCR Meth. Appl. 4: 31–37.
Liu B., Vega J.M., Segal G., Abbo S. Rodova M. &Feldman M. (1998): Rapid genomic changes in newly synthesized amphiploids ofTriticum andAegilops. I. Changes in low-copy noncoding DNA sequences.Genome 41: 272–277.
Lönn M. (1993): Genetic structure and allozyme-microhabitat associations inBromus hordeaceus.Oikos 68: 99–106.
Meerts P. (1995): Phenotypic plasticity in the annual weedPolygonum aviculare.Bot. Acta 108: 414–424.
Meerts P., Baya T. &Lefèbvre C. (1998): Allozyme variation in the annual weed species complexPolygonum aviculare (Polygonaceae) in relation to ploidy level and colonizing ability.Pl. Syst. Evol. 211: 239–256.
Misset M.T. &Gourret J.P. (1996): Flow cytometric analysis of the different ploidy levels observed in the genusUlex L. (Faboideae-Genisteae) in Brittany, France.Bot. Acta 109: 72–79.
Nei M. (1973): Analysis of gene diversity in subdivided populations.Proc. Natl. Acad. Sci. U.S.A. 70: 3321–3323.
Nei M. (1978): Estimation of average heterozygosity and genetic distance from a small number of individuals.Genetics 89: 583–590.
Nei M. &Li W.H. (1979): Mathematical models for studying genetic variation in terms of restriction endonucleases.Proc. Natl. Acad. Sci. U.S.A. 76: 5269–5273.
Novack S.J. &Mack R.N. (1993): Genetic variation inBromus tectorum (Poaceae): comparison between native and introduced populations.Heredity 71: 167–176.
Pavlick L. (1995): BromusL. of North America. Royal British Columbia Museum, Victoria.
Ramsey J. &Schemske D.W. (1998): Pathways, mechanisms, and rates of polyploid formation in flowering plants.Annual Rev. Ecol. Syst. 29: 467–501.
Rieseberg L.H. (1996): Homology among RAPD fragments in interspecific comparisons.Molec. Ecol. 5: 99–105.
Rohlf F.J. (1990):NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 1.6. Applied Biostatistics, New York.
Romero-Zarco C. (1986): A new method for estimating karyotype asymmetry.Taxon 35: 526–530.
Roy J., Navas M.L. &Sonie L. (1991): Invasions by annual bromegrasses: a case study challenging the homoclime approach to invasions. In:Grove R.N. &di Castri F. (eds.),Biogeography of Mediterranean invasions, Cambridge University Press, Cambridge, pp. 205–221
Scholz H. (1998):Bromus molliformis. Soc. Ech. Pl. Vasc. Eur. Bass. Médit., Bull. 27 19–20.
Sang T., Crawford D.J. &Stuessy T.F. (1995): Documentation of reticulate evolution in peonies (Paeonia) using internal transcribed spacer sequences of nuclear ribosomal DNA: implications for biogeography and concerted evolution.Proc. Natl. Acad. Sci. U.S.A. 92: 6813–6817.
Song K., Lu P., Tang K. &Osborn T.C. (1995): Rapid genome change in synthetic polyploids ofBrassica and its implications for polyploid evolution.Proc. Natl. Acad. Sci. U.S.A. 92: 7719–7723.
Slatkin M. &Barton N.H. (1989): A comparison of three indirect methods for estimating average levels of gene flow.Evolution 43: 1349–1368.
Smith P.M. (1970): Taxonomy and nomenclature of the bromegrasses.Notes Roy. Bot. Gard. Edinburgh 30: 361–375.
Smith P.M. (1972): Serology and relationships in annual bromes (Bromus L. sect.Bromus).Ann. Bot. (London), 36: 1–30.
Smith P.M. (1980):Bromus. In:Tutin T.G., Heywood V.H., Burges N.A., Valentine D.H., Walters S.M. &Webb D.A. (eds.),Flora europaea 5, Cambridge University Press, Cambridge, pp. 319–323.
Smith P.M. (1981): Ecotypes and subspecies in annual brome-grasses (Bromus, Gramineae),Bot. Jahrb. Syst. 102: 497–509.
Smith P.M. (1983): Proteins, mimicry and microevolution in grasses. In:Jensen U. &Fairbrothers D.E. (eds.),Proteins and nucleic acids in plant systematics, Springer Verlag, Berlin, pp. 319–323.
Smith P.M. (1986): Native or introduced? Problems in the taxonomy and plant geography of some widely introduced annual bromegrasses.Proc. Roy. Soc. Edinb. B. 89: 273–281.
Soltis D.E. &Soltis P.S. (1993): Molecular data and the dynamic nature of polyploidy.C.R.C. Crit Rev. Pl. Sci. 12: 243–273.
Soltis P. &Doyle J. (1998):Molecular systematics of plants II. Kluwer Academic Publishers, Boston.
Stebbins G.L. (1981): Chromosomes and evolution in the genusBromus (Gramineae) Bot. Jahrb. Syst. 102: 359–379.
Swofford D.L. (1993):PAUP: phylogenetic analysis using—parsimony version 3.1.1. Illinois Natural History Survey, Champaign.
Warwick S.I. (1990): Allozyme and life history variation in five northwardly colonizing North American weed species.Pl. Syst. Evol. 169: 41–51.
Wendel J.F., Schnabel A. &Seelanan T. (1995): Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton (Gossypium).Proc. Natl. Acad. Sci. U.S.A. 92: 280–284.
Wendel J.F. &Doyle J.J. (1998): Phylogenetic incongruence: Window into genome history and molecular evolution. In:Soltis P. &Doyle J.J. (eds.),Molecular systematics of plants II, Kluwer Academic Publishers, Boston, pp. 265–296.
Whitkus R. (1988): Modified version of GENESTAT. A program for computing genetic statistics from allelic frequency data.Pl. Genet. Newslett. 4: 10.
Williams G.K., Kubelik A.R., Livak K.L., Rafalski J.A. &Tingey S.V. (1990): DNA polymorphisms amplified by arbitrary primers are useful as genetic markers.Nucl. Acid Res. 18: 6531–6535.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ainouche, M.L., Bayer, R.J., Gourret, JP. et al. The allotetraploid invasive weedBromus hordeaceus L. (Poaceae): Genetic diversity, origin and molecular evolution. Folia Geobot 34, 405–419 (1999). https://doi.org/10.1007/BF02914919
Issue Date:
DOI: https://doi.org/10.1007/BF02914919