Summary
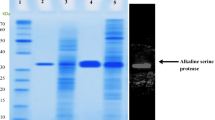
Alkaliphilic Bacillus sp. no. AH-101 produces an extremely thermostable alkaline serine protease that has a high optimum pH (pH 12–13) and shows keratinolytic activity. The gene encoding this protease was cloned in Escherichia coli and expressed in B. subtilis. The cloned protease was identical to the AH-101 protease in its optimum pH and thermostability at high alkaline pH. An open reading frame of 1083 bases, identified as the protease gene, was preceded by a putative Shine-Dalgarno sequence (AAAGGAGG) with a spacing of 11 bases. The deduced amino acid sequence revealed a pre-pro-peptide of 93 residues followed by the mature protease comprising 268 residues. AH-101 protease showed slightly higher homology to alkaline proteases from alkaliphilic bacilli (61.2% and 65.3%) than to those from neutrophilic bacilli (54.9–56.7%). Also AH-101 protease and other proteases from alkaliphilic bacilli shared common amino acid changes and a four amino acid deletion when compared to the proteases from neutrophilic bacilli. AH-101 protease, however, was distinct among the proteases from alkaliphilic bacilli in showing the lowest homology to the others.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523
Carter P, Wells JA (1988) Dissecting the catalytic triad of a serine protease. Nature 332:564–568
Chang SC, Cohen SN (1979) High frequency of transformation of Bacillus subtilis protoplast by plasmid DNA. Mol Gen Genet 168:111–115
Enquist L, Sternberg N (1979) In vitro packaging of λ Dam vectors and their use in cloning DNA fragments. Methds Enzymol 68:282–283
Estell DA, Graycar TP, Miller JV, Powers DB, Burnier JP, Ng PG, Wells JA (1986) Probing steric and hydrophobic effects on enzyme-substrate interactions by protein engineering. Science 233:659–663
Horikoshi K (1971) Production of alkaline enzymes by alkalophilic microorganisms. Part II. Alkaline amylase produced by Bacillus no. A-e-2. Agric Biol Chem 35:1783–1791
Horikoshi K (1991) Industrial applications. In: Miccoorganisms in Alkaline Environments. VCH, Weinheim, pp 187–194
Horinouchi S, Weisblum B (1982a) Nucleotide sequence and functional map of pE194, a plasmid that specifies inducible resistance to macrolide, lincosamide, and streptogramin type B antibiotics. J Bacteriol 150:804–814
Horinouchi S, Weisblum B (1982b) Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol. J Bacteriol 150:815–825
Jacobs M, Eliasson M, Uhlen M, Flock JI (1985) Cloning, sequencing and expression of subtilisin Carlsberg from Bacillus licheniformis. Nucleic Acids Res 13:8913–8926
Kaneko R, Koyama N, Tasi YC, Juang RY, Yoda K, Yamasaki M (1989) Molecular cloning of the structural gene for alkaline elastase, a new subtilisin produced by an alkalophilic Bacillus strain. J Bacteriol 171:5232–5236
Kawamura F, Doi RH (1984) Construction of a Bacillus subtilis double mutant deficient in extracellular alkaline and neutral proteases. J Bacteriol 160:442–444
Laan JC, Gerritse G, Mulleners LJSM, Hoek RAC (1991) Cloning, characterization, and multiple chromosomal integration of a Bacillus alkaline protease gene. Appl Environ Microbiol 57:901–999
Mandel M, Higa A (1970) Calcium dependent bacteriophage DNA infection. J Mol Biol 53:159–162
Oste C (1989) PCR Automation. In: Erlich HA (ed) PCR technology, principles and applications for DNA amplification. Stockton Press, New York, pp 23–30
Robertus JD, Alden RA, Birktoft AJ, Kraut J, Powers JC, Wilcox PE (1972) An X-ray crystallographic study of the binding of peptide chloromethyl ketone inhibitors to subtilisin BPN'. Biochemistry 11:2439–2449
Saito H, Miura K (1963) Preparation of transforming deoxyribonucleic acid by phenol treatment. Biochim Biophys Acta 72:619–629
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain terminating inhibitions. Proc Natl Acad Sci USA 74:5463–5467
Shine J, Dalgarno L (1974) The 3′-terminal sequence of E. coli 16S ribosomal RNA: complementary to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci USA 71:1342–1346
Southern EM (1975) Detection of specific sequences among DNA fragments by gel electrophoresis. J Mol Biol 98:503–517
Stahl ML, Ferrari E (1984) Replacement of Bacillus subtilis subtilisin structural gene with an in vitro derived deletion mutation. J Bacteriol 158:411–418
Takami H, Akiba T, Horikoshi K (1989) Production of extremely thermostable alkaline protease from Bacillus sp. no. AH-101. Appl Microbiol Biotechnol 30:120–124
Takami H, Akiba T, Horikoshi K (1990) Characterization of an alkaline protease from Bacillus sp. no. AH-101. Appl Microbiol Biotechnol 33:519–523
Takami H, Akiba T, Horikoshi K (1992) Substrate specificity of thermostable alkaline protease from Bacillus sp. no. AH-101. Biosci Biotechnol Biochem 56:333–334
Watson MEE (1984) Compilation of published signal sequences. Nucleic Acids Res 12:5145–5164
Wells JA, Ferrari E, Henner DJ, Estell DA, Chen EY (1983) Cloning, sequencing, and secretion of Bacillus amyloliquefaciens subtilisin in Bacillus subtilis. Nucleic Acids Res 11:7911–7925
Wright CS, Alden RA, Kraut J (1969) Structure of subtilisin BPN' at 2.5 Å resolution. Nature 221:235–242
Yoshimoto T, Oyama H, Honda T, Tone H, Takeshita T, Kamiyama T, Tsuru D (1988) Cloning and expression of subtilisin amylosacchariticus gene. J Biochem 103:1060–1065
Zuker M, Stiegler P (1981) Optical computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9:133–148
Author information
Authors and Affiliations
Additional information
Correspondence to: H. Takami
Rights and permissions
About this article
Cite this article
Takami, H., Kobayashi, T., Aono, R. et al. Molecular cloning, nucleotide sequence and expression of the structural gene for a thermostable alkaline protease from Bacillus sp. no. AH-101. Appl Microbiol Biotechnol 38, 101–108 (1992). https://doi.org/10.1007/BF00169427
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00169427