Abstract
In natural environments, the average abundance of heavy metals is generally low and much of that sequestered in sediments, soil and mineral deposits may be biologically unavailable. Microorganisms have ability to adapt and live in all ecological condition. In natural habitat, the cause for microbes on heavy metal depends on the physico-chemical properties of the environmental condition. Microbes can metabolize the metal ion and yield energy through oxidation and reduction process by dissolving them. Many trace metals are necessary for growth and metabolism at low concentrations, (e.g. Co, Cu, Ni, Mo, Fe, Zn), and microorganism acquires mechanisms of varying specificity for the intracellular increase from the external environment. The molecular mechanism of microorganism and plants in the removal of toxic heavy metals into nontoxic form using plants and microorganisms is well studied, and this has many biotechnology implications in the bioremediation of heavy metal contaminated sites.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
There are nearly sixty-five elements, which may be termed as heavy metal as they exhibit metallic properties. In the ecosystem, organic toxic contaminants cause harmful effect because of releasing hazardous waste such as lead, cadmium and chromium by the increasing technological advancement. A wide variety of activities by industries have generated the mobilization of large heave metal in the elevated point to the natural geochemical system (Gaur 2014; Dixit 2015; Tak 2013).
The non-degradable nature of toxic heavy metal pollution is the important environmental trouble caused by metal ions that continue to present in the environment that is controlled biological system. Based on the dose of adsorption and course of exposure, toxic heavy metal causes a serious lethal effect at very high concentration and the toxic effect depends on the amount present to organisms (Sand et al. 1992). Bioremediation techniques are used to reduce or completely remove the toxic heavy metals from the environment (Dixit 2015; Akcil et al. 2015).
Microbes and plants are used to remediate the polluted environment through sustained technology and bring back natural environment condition (Doelman 1994). The change in the microbial make up is due to the degradation of function group in the heavy metal or alteration in the conformation of the molecules (Wood 1983; Li 1994).
The reaction of microbes to heavy metals is because of the concentration, and accessibility of toxic metal is a completed process which is managed by many factors like metal chemical property, medium type and the microbes (Goblenz et al. 1994). A perceptive of microbial responses towards heavy metals is also significant for the preservation and protection of natural and synthetic materials, since inorganic and organometallic compounds are still widely used as microbicidal agent in domestic and agricultural pest and pathogen control operations.
2 Toxic Heavy Metals in the Environment
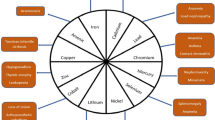
Industrial actions have increased the mobilization of many heavy metals above rates of natural geochemical cycling, and there is improved deposition in aquatic and terrestrial ecosystem, as well as release into the atmosphere. The heavy metal contamination of the soil and water is mainly due to manmade activity and by natural. Table 1 shows the sources of certain heavy metals in the environment and its impact.
3 Bioremediation
Bioremediation is process of using microorganisms to remove polluted environment or to prevent pollution. The basis of bioremediation is the inherent natural capacity of microorganism to degrade organic compound present in the environment.
4 Bioremediation Methods
The toxic metal may remain in the form of solid or liquid phase; bioremediation technology will vary accordingly the waste material remains in its natural form or is taken to the fermentor. Bioremediation of toxic waste material treatment has two methods: in situ bioremediation and ex situ bioremediation.
5 Microbes in Metal-Containing Habitats
A wide variety of organism from all the major groups may be found in the metal-polluted habitats such as bacteria, fungi, yeast and algae (White et al. 1997; Vieira and Volesky 2000). Microorganisms can be isolated from almost any environmental conditions. In contaminated environments, there are some dominant microbes, which can tolerate the metal stress and grow more or less normally or even better, due to easy availability of resources.
Microorganisms can be subdivided into the following groups:
-
a.
Aerobic
Pseudomonas, Alcaligenes, Sphingomonas, Rhodococcus and Mycobacterium. These microbes have ability to degrade pesticides and hydrocarbons, both alkanes and polyaromatic compounds. Microorganisms use the contaminant as the prime source of carbon and energy.
-
b.
Anaerobic
Anaerobic bacteria are used for bioremediation of dechlorination of the solvent trichloroethylene (TCE) and chloroform, polychlorinated biphenyls (PCBs) in river sediments.
-
c.
Lignilolytic fungi
Phanerochaete chrysosporium has the capability to remove an extremely diverse range of persistent or toxic pollutants from environment.
-
d.
Methylotrophs
Aerobic bacteria take methane for carbon and energy. The primary enzyme in the metabolic pathway for aerobic degradation is the methane monooxygenase, has a wide range of substrate and is active against a large amount of compounds, together with the chlorinated aliphatic trichloroethylene and 1,2-dichloroethane. Table 2 shows some microbes utilizing heavy metals.
6 Metal and Microbe Interaction
The capability of microbes to survive and grow in a contaminated metal habitat may depend on genetical and physiological adaptation. The mechanisms of metal ions binding to the cell surface include electrostatic interactions, van der Waals forces, covalent bonding, redox interactions and extracellular precipitation or combination of these processes (Blanco 2000). The negatively charged groups (carboxyl, hydroxyl and phosphoryl) of bacterial cell wall adsorb metal cations, which are then retained by mineral nucleation. Biosorption of metal such as U, Zn, Pb, Cd, Ni, Cu, Hg, Th, Cs, Au, Ag, Sn and Mn shows the extent of sorption which varies clearly with the metal and microorganisms.
7 Microbes with Metal Tolerance
Some microorganisms are supposed to have developed metal resistance because of their contact with toxic metals in an already metal-polluted world (Gupta et al. 1993). In response to metal resistance in the environment, microorganisms have evolved ingenious mechanisms of metal resistance and detoxification. Some resistance mechanisms are plasmid encoded and tend to be specific for a particular metal. Others are general resistance to a variety of toxic heavy metals. There are two mechanisms of metal resistance:
-
General mechanism of metal resistance,
-
Metal-dependent mechanism of metal resistance.
7.1 General Mechanism of Metal Resistance
Metal binding on to extracellular materials immobilizes the metals and prevents the entry into the cell. Metal binding to anionic cell surfaces occurs with a large number of cationic metals, including cadmium, lead, zinc and iron. For example, algal cell surfaces contain carboxylic, amino, thio, hydroxo and hydroxyl carboxylic groups that strongly bind metals. Phosphoryl groups and phospholipids in bacterial lipopolysaccharides in the outer membrane also strongly interact with cationic metals.
7.1.1 Exopolymer Binding
The major molecules which constitute the exopolymeric substances include nucleic acids, carbohydrates, fatty acids and polysaccharides. These extracellular polymeric substances (EPSs) are very general in natural environments, and they provide protection against desiccation, phagocytosis and parasitism in addition to their role in heavy metal binding. They bind metal such as lead, cadmium and uranium.
7.1.2 Siderophore Complexation
It is an iron-chelating organic molecule. Their biological function is to concentrate iron in the environment where their concentration is very low and to transport iron into the cell. The interactions of siderophore with other metals are chemically similar to iron, viz., aluminium, gallium and chromium; for example, siderophores in cyanobacteria reduce copper toxicity.
7.1.3 Biosurfactants Complexation
They have ability to complex metals such as lead, cadmium and zinc. This apparently enhances the solubility of metals.
7.1.4 Precipitation by Metal Reduction
In this, soluble metals are reduced to less soluble metal salts, as well as sulfidic and phosphatidic metal salts. For example, Citrobacter spp. under aerobic condition can enzymatically produce phosphates which results in the precipitation of lead and copper. In anaerobic condition, Desulfovibrio spp., can have ability to precipitate sulphate metal.
7.2 Metal-Dependent Mechanism of Metal Resistance
Metallothioneins are proteins that play pivotal role in heavy metal metabolism and in the management of various forms of stress. Metallothioneins are similar to phytochelatins of plants and have high number of cysteine residues in the protein and also in the fact that both are responsible for the detoxification of heavy metals. Metallothioneins have three characteristics, low molecular weight proteins, and have high cysteine content and high metal with organization of metal ions in clusters.
The metallothioneins have two categories. The class-I MTs are found in all the animal metallothioneins, and the class-II is present in microorganisms and plants.
In mammalian archetype, the class-I MTs have been very well characterized and represent. The invariant arrangement of Cys within the protein provides the thiol for mercaptide bonds in arrangements typical of metal-thiolate clusters. These proteins are isolated from the cyanobacterium Synechococcus spp., Escherichia coli and Pseudomonas putida (Gupta et al. 1993). Table 3 shows microbial metal resistance mechanisms.
8 Microbial Immobilization and Transformation of Metals
Bioremediation strategy involves mechanisms like precipitation and solubilization which shows the microbial transformations of metal as a result of metal resistance mechanisms.
8.1 Extracellular Complexation
Extracellular polysaccharides have anionic properties which show the purpose of efficient biosorbents for metal cations. Metal ions interaction is generally considered a direct effect of the presence of negatively charged functional groups on these exopolymers; they are pyruvate, phosphate, hydroxyl succinyl and uronic acids. A pH reliant binding of positively charged cations can quickly occur, with stability constants in excess of those generally measured for humic substances and other naturally occurring ligands.
Many organic metabolites can be significant on detoxification of metals because of the chelating properties or complexation properties. Organic or inorganic acids produced by microorganism, including Thiobacillus, Serratia, Pseudomonas, Bacillus, Penicillium and Aspergillus, are competent to extract metals from solid substrates. The oxalic acid can precipitate metals as insoluble oxalates around cell wall efficiently done by metal chelator citric acid.
8.2 Extracellular Precipitation and Crystallization
Microbes like sulphate-reducing bacteria, e.g. Desulfovibrio, are concerned in the formation of sulphide deposits which contain large amounts of metals. Metal-resistant strains of Klebsiella aerogenes precipitate Pb, HG or Cd as insoluble sulphide granules on outer surfaces of cells, and particles of AgS are deposited on thiobacilli when growth in silver-containing sulphide leaching systems. Strains of the green algae Cyanidium caldarium can grow in acidic waters at 45 °C containing high concentrations of metal ions. Iron, copper, nickel, aluminium and chromium can be removed from solution by precipitation, at cell surfaces, as metal sulphides. Cells can contain up to 20% metal on a dry weight basis. Yeasts can also precipitate metals as sulphides in and around cells walls, and colonies may appear dark brown in the presence of copper (Gadd 1990).
Algae and fungi can promote Mn2+ oxidation in a variety of habitats and can become coated with manganic oxides. Other bacteria can become coated with oxidized iron compounds by metabolism-dependent and -independent processes. In Thiobacillus ferrooxidans and certain algae, Au3+ can be adsorbed to cell walls and the plasma membrane and reduced to particles of elemental Au. Reducing compounds produced by silver-resistant bacteria can reduce Agy2+ to metallic AgO, which results in silver deposition on glass surfaces of culture vessels or in growing colonies.
8.3 Transport-Related Metal Resistance
Metabolism-dependent intracellular transport may be a slower process than binding and is inhibited or halted by metabolic inhibitors, low temperatures, the absence of an energy source (e.g. glucose in heterotrophs, light in phototrophs) and uncouplers. In several algae, bacteria and yeasts, amounts accumulated by transport may greatly exceed amounts taken up by general biosorption. Rates of transport are also affected by the metabolic state of the cell and nature and composition of the growth medium. Integral to heavy metal transport systems are ionic gradients across the cell membrane, e.g. H+ and/or K+ together, generating the electrochemical gradient across the cell membrane, and the membrane potential.
A relationship between heavy metal transport into microbial cells and toxicity is often observed, with sensitive strains taking up more metal ions than resistant strains. When transport rates do not vary, the sensitive organisms have more intracellular toxic metal species than the resistant ones. It is therefore not surprising that decreased transport, impermeability or the occurrence of metal efflux systems constitute resistance mechanisms in many organisms. In several bacteria, including Staphylococcus aureus and Bacillus subtilis, active transport of Cd2+ depends on the cross-membrane electrical potential and this uptake system is highly specific for Mn2+, as well as Cd2+. Resistance may arise from reduced Cd2+ transport in gram-negative bacteria. The efflux of cadmium from the cells of actively growing S. aureus has been found to be an energy-consuming process, involving cell membrane ATPase. This Cd2+/2H+ antiport is presented in resistant strains which results in a net reduction of Cd2+ uptake. Cadmium-resistant P. putida also exhibits reduced uptake, and this may also be a result of defective transport and/or the presence of a Cd2+ efflux system. In an Alcaligenes sp., adaptation to Cd2+ resistance was associated with the synthesis of a new membrane protein, which may be involved in the prevention of Cd2+ influx or the cause of Cd2+ efflux. Membrane proteins that reduce bacterial uptake of Hg2+ have also been described. Some Cu2+ resistance strains of E. coli exhibit reduced uptake and/or exclude Cu2+, due to the disappearance of outer membrane proteins which may be involved in Cu2+ transport.
8.4 Intracellular Compartmentation and Detoxification
Certain bacteria, yeasts, fungi and algae can actively accumulate intracellular metal ions against a gradient. Once inside cells, metal ions may be compartmentalized and/or converted to more innocuous forms. Such processed can be effective detoxification mechanism, and microbes expressing them may be able to accumulate metals to high intracellular concentrations. It has been suggested that such mechanisms may be temporary and precede other means of expulsion of accumulated metals from the cells, possibly by means of vacuoles in eukaryotic microbes. A variety of electron-dense deposits, many of unknown compositions, have been recorded in cyanobacteria, bacteria, algae, protozoa and fungi after exposure to heavy metals. Polyphosphate has been implicated as a metal sequestering agent in P. putida, Anabaena cylindrica and Plectonema boryanum, a variety of eukaryotic algae and certain fungi and yeasts. In eukaryotic microbes, e.g. algae, yeasts and fungi, there may be compartmentation of accumulated Co2+, Mn2+, Zn2+ and K+ that is located in the vacuoles, where it may be in ionic form or bound to low molecular weight polyphosphates.
A common metal-induced response in many microorganisms is the synthesis of intracellular metal-binding proteins which function in detoxification and also the storage and regulation of intracellular metal ion concentrations. Metallothioneins are the most widely studied, and these are low molecular weight, cysteine-rich proteins that can bind metal like Cd, Zn and/or Cu. A Cd-binding prokaryotic metallothionein (molecular weight approx. 8100) was first described in the cyanobacterium Synechococcus sp. and others have since been reported in several bacteria including P. putida inducible Cd-binding proteins also occur in E. coli, but these are larger than metallothioneins and may be more related to recovery from cadmium toxicity.
8.5 Enzymatic Transformation of Metals
Microbes transform heavy metals, such as oxidation, reduction, methylation and demethylation, and these are significant not only to biogeochemical cycling, but may also constitute mechanisms of resistance. Metal transformation has concentrated on bacterial involvement in the mercury cycle, which can be simply represented as:
Mercury resistance to gram-positive and gram-negative bacteria is a common property, the determinants usually being plasmid encoded, particularly in gram-negative bacteria. The most ubiquitous mechanism of mercury resistance in bacteria is the enzymatic reduction of Hg2+, by cytoplasmic mercuric reductase, to metallic HgO, which is less toxic than Hg2+, volatile and rapidly lost from the environment. This has also been recorded in certain fungi and yeasts. Organomercurial compounds are enzymatically detoxified by organomercurial lyase which cleaves the Hg–C bond of, e.g. methyl-, ethyl- and phenyl-mercury, to form Hg2+ and methane, ethane and benzene, respectively. The Hg2+ can then be volatile by mercuric reductase. More than one kind of organomercurial lyase may be present in a given bacterium. In organomercurial resistance E. coli, two enzymes are active against methyl- and ethylmercury chloride. In a Pseudomonas strain, also there are two enzymes of similar specifies, as well as being able to cleave ρ-hydroxy-mercury benzoate.
In FAD-containing mercuric reductase is a flavoprotein has been studies for metal detoxification enzyme this, and organomercurial lyase, require an excess of thiols for activity. NADPH is the preferred reducing cofactor for mercuric reductase as well as organomercurial lyase, but in some bacterial strains, NADH is effective. The thiols prevent the formation of NADPH-Hg2+ complex and ensure that the Hg2+ is present as a dimercaptide. The reaction catalysed by mercuric reductase in vivo can be represented as:
Structurally and mechanically mercuric reductase is related to glutathione reductase and lipoamide dehydrogenase. The mechanism of mercuric reductase probably involves electrons being transferred from NADPH via FAD to reduce the active site cystine, converting it to two cysteine residues with titratable SH groups. One Cys residue forms a charge transfer complex with FAD. The active site cysteines then reduce Hg2+, bound to the C-terminal cysteines, forming HgO. Mercuric reductase has been reported to exist as a monomer, dimer and trimer with a subunit molecular weight between 54,000 and 69,000, depending on the source, though it seems probable that the dimeric structure is active in vivo.
9 Genetic Aspects of Heavy Metal Resistance
The metal microbe genetic relations have been inadequate for mutant strains, primarily because understanding of the physiology and biochemistry of resistance is fragmentary in many cases. The areas, for which significant genetic advances have been made to date, are in heavy metal resistance in bacteria, cyanobacteria and the metallothionein of Saccharomyces cerevisiae.
9.1 Genetic Aspects of Heavy Metal Resistance in Bacteria
Many bacteria isolated from metal-contaminated environment shown mercury resistance. The mercury resistance determinants in bacteria (gram-positive and gram-negative) are usually plasmid encoded, predominantly in gram-negative bacteria. A common feature of the Hg2+ resistance determinants of bacteria from the environment is the absence of association to drug resistance markers in contrast to clinical bacterial isolates, where Hg2+ resistance is usually linked to genes for antibiotic resistance. Several mercury resistance transposons could be responsible for the widespread occurrence of Hg2+ resistance in gram-negative bacteria. To date, the most recent advances have stemmed from DNA sequencing studies of the mer genes located on transposons Tn501 and Tn21 from gram-negative bacteria, and such studies have been carried out with broad spectrum determinants, in both gram-negative and gram-positive bacteria (Gadd 1990).
9.2 Genetic Aspects of Heavy Metal Resistance in Fungi and Other Eukaryotes
In fungi, the copper-induced metallothionein of S. cerevisiae (molecular weight approx, 6573) has received most attention although a copper inducible metallothionein has also been described from Neurospora crassa. The yeast protein is inducible only by copper not cadmium or zinc and is therefore often referred to as Cu-MT or yeast MT.
Inducible Cd-binding proteins that are structurally different to Cu-MT have been isolated from Schizosaccharomyces pombe. A common name for these sulphur-rich metal-binding polypeptides is phytochelatin. The metal-binding peptides are composed of only three amino acids, l-cysteine, l-Glutamic and glycine and are analogous to similar peptides found in plant cell exposed to have metals like Cd, Cu, Hg, Pb and Zn. The general structure of two peptides found in Cd-exposed S. pombe is (g-Glu-Cys) Gly (n = 2 and 3). The unit peptides are called cadystin, and the phytochelatins n = 2 and n = 3 are called cadystin A and B, respectively. In addition to these, five homologous peptides with chain lengths from n = 4 and n = 8 are also observed in S. pombe, indicating that these are synthesized by elongation of the peptide with one (g-Glu-Cys) unit at the expense of and possibly, starting from glutathione. The cysteine-rich peptides of S. pombe can confiscate several transition metals and therefore share some basic features with metallothioneins. Multiple enzymes may be involved in phytochelatin synthesis, and therefore, mechanisms must exist for metal-dependent induction or activation of such enzymes.
10 Conclusion
The main strategy is to reduce the bioavailability, mobility and toxicity of the metal. Toxic heavy metals have adverse effects on aquatic and terrestrial ecosystems. All though various microorganisms are commonly found in polluted habitats and possess a range of morphological and physiological attributes that enable survival. Certain kinds of resistances may be genetically determined and transferrable, whereas, in other cases, tolerance may depend on intrinsic properties of the organism and/or detoxification due to environmental components. Many organisms can express a variety of survival strategies, and besides providing excellent model systems for scientific endeavour in fields ranging from ecology to molecular biology and genetics, they are also of biotechnological relevance to the detoxification of metals and for containing environmental metal recovery and reclamation for preservation of natural, synthetic materials and treatment of infection.
In relation to industrial application of microbial metal removal/recovery, most attention has so far been focused on high value elements of commercial importance. With non-precious metals and radionuclides, the economics are frequently unattractive and there is the problem of contaminated biomass. However, the containment of low-volume waste or elute is preferable to its entry into the environment. At the moment, environmental protection receives only limited political and industrial attention. However, with increasing pollution and public awareness of this, and the dangers of heavy metals and radionuclides to all living components of the biosphere, action may be taken and the fuller involvement of microbe-based technologies may be part of this.
References
Ahmad A, Mukherjee P, Mandal D, Senapati S, Khan MI, Kumar R, Sastry M (2002) Enzyme mediated extracellular synthesis of CdS nanoparticles by the fungus, Fusarium oxysporum. J Am Chem Soc 124:12108–12109
Akcil A, Erust C, Ozdemiroglu S, Fonti V, Beolchini F (2015) A review of approaches and techniques used in aquatic contaminated sediments: metal removal and stabilization by chemical and biotechnological processes. J Clean Prod 86:24–36
Amoroso MJ, Castro GR, Durain A, Peroud O, Oliver G, Hill RT (2001) Chromium accumulation by two Streptomyces spp. isolated from riverine sediments. J Ind Microbiol Biotechnol 26:p210–p215
Blanco A (2000) Immobilization of nonviable cyanobacteria and their use for heavy metal adsorption from water. In Oluguin EJ, Sanchez, Hernandez E (eds) Environmental biotechnology and cleaner bioprocesses. Taylor and Amp Francis, Philadelphia, p 135
Culotta VC, Howard WR, Liu XF (1994) CRS5 encodes a metallothionein-like protein in Saccharomyces cerevisiae. J Biol Chem 269:25295–25302
Dixit R, Malaviya D, Pandiyan K, Singh UB, Sahu A, Shukla R, Singh BP, Rai JP, Sharma PK, Lade H (2015) Bioremediation of heavy metals from soil and aquatic environment: an overview of principles and criteria of fundamental processes. Sustainability 7:2189–2212
Doelman P, Jansen E, Michels M, van Til M (1994) Effects of heavy metals in soil on microbial diversity and activity as shown by the sensitivity-resistance index, an ecologically relevant parameter. Biol Fertil Soil 17:177–1784
Favero N, Costa P, Massimino ML (1991) In vitro uptake of cadmium by basidiomycete Pleurotusostreatus. Biotechnol Lett 10:701–704
Gabriel J, Mokrejs M, Bily J, Rychlovsky P (1994) Accumulation of heavy metal by some Woodrooting fungi. Folia Microbiol 39:115–118
Gabriel J, Kofronova O, Rychlovsky P, Krenzelok M (1996) Accumulation and effect of cadmium in the wood rotting basidiomycete, Daedaleaquercina. Bull Environ Contam Toxicol 57:383–390
Gadd GM (1990) Heavy metal accumulation by bacteria and other microorganisms, vol 46, no 8. Springer, Berlin, pp 834–840
Gaur N, Flora G, Yadav M, Tiwari A (2014) A review with recent advancements on bioremediation-based abolition of heavy metals. Environ Sci Process Impacts 16:180–193
Goblenz A, Wolf K, Bauda P (1994) The role of glutathione biosynthesis in heavy metal resistance in the fission yeast Schizosaccharomyces pombe. FEMS Microbiol Rev 14:303–308
Gunasekaran P, Muthukrishnan J, Rajendran P (2003) Microbes in heavy metal remediation. Indian J Exp Biol 41(1):935–944
Gupta A, Morby AP, Turner JS, Whitton BA, Robinson NJ (1993) Deletion within the metallothionein locus of cadmium-tolerant Synechococcus PCC-6301 involving a highly iterated palindrome (hip1). Mol Microbiol 7:189–195
Li F, Tan TC (1994) Monitoring BOD in the presence of heavy metal ions using a poly (4-vinylpyridine) coated microbial sensor. Biosens Bioelectron 9:445–455
Lioyd JR (2003) Microbial reduction of metals and radionuclides. FEMS Microbiol Rev 27:411–425
Meyer J, Schmidt A, Michalke K, Hensel R (2007) Volatilization of metals and metalloids by the microbial population of an alluvial soil. Syst Appl Microbiol 31:81–87
Mohamed ZA (2001) Removal of cadmium and manganese by a non-toxic strain of the fresh water Cyanobacterium Gloeothece magna. Water Res 35(18):p4405–p4409
Nair B, Pradeep T (2002) Coalescence of nanoclusters and formation of submicron crystallites assisted by Lacto strains. Cryst Growth Des 2:293–298
Nair A, Juwarkar AA, Singh SK (2007) Production and characterization of siderophores and its application in arsenic removal from contaminated soil. Water Air Soil Pollut 180:p199–p212
Nies DH (2003) Efflux mediated heavy metal resistance in prokaryotes. FEMS Microbiol Rev 27:p313–p339
Nies DH, Silver S (1989) Plasmid determined inducible efflux is responsible for resistance to cadmium, zinc and cobalt in Alcaligenes eutrophus. J Bacteriol 171:896–900
Philip L, Iyengar L, Venkobacher L (2000) Site of interaction of copper on. Water Air Soil Pollut 119:11–21
Roane TM, Josephson KL, Pepper IL (2001) Dual-bioaugmentation strategy to enhance remediation of cocontaminated soil. Appl Environ Microbiol 67:p3208–p3215
Sand W, Rohde K, Sabotke B, Zenneck C (1992) Evaluation of Leptospirillum ferrooxidans for leaching. Appl Environ Microbiol 58:85–92
Sar P, D’Souza SF (2001) Biosorptive uranium uptake by Pseudomonas strain: characterization and equilibrium studies. J Chem Technol Biotechnol 76:1286–1294
Smedley PL, Kinniburgh DG (2002) A review of the source behaviour and distribution of arsenic in natural waters. Appl Geochem 17:517–568
Spark DL, Chemistry Environmental soil (2003) Academic Press. San Diego, California
Tak HI, Ahmad F, Babalola OO (2013) Advances in the application of plant growth-promoting rhizobacteria in phytoremediation of heavy metals. In: Reviews of environmental contamination and toxicology. Springer, New York, pp 33–52
Vieira R, Volesky B (2000) Biosorption: a solution to pollution? Int Microbiol 3:17–24
White C, Sayer JA, Gadd GM (1997) Microbial solubilization and immobilization of toxic metals: key biochemical processes for treatment of contamination. FEMS Microbiol Rev 20:503–516
Wood JM, Wang HK (1983) Microbiol resistance to heavy metals. Environ Sci Technol 17:582–590
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Raja Sathendra, E., Praveen Kumar, R., Baskar, G. (2018). Microbial Transformation of Heavy Metals. In: Varjani, S., Gnansounou, E., Gurunathan, B., Pant, D., Zakaria, Z. (eds) Waste Bioremediation. Energy, Environment, and Sustainability. Springer, Singapore. https://doi.org/10.1007/978-981-10-7413-4_13
Download citation
DOI: https://doi.org/10.1007/978-981-10-7413-4_13
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-7412-7
Online ISBN: 978-981-10-7413-4
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)