Summary
Hydrogenases are efficient biological catalysts of H2 oxidation and production. Most of them are inhibited by oxygen, and a prerequisite for their use in biotechnological applications under air is to improve their oxygen tolerance. A few bacteria, however, contain hydrogenases that activate H2 even in the presence of O2. Intriguingly, molecular, kinetic and spectroscopic studies lead to assume that different mechanisms might be responsible for the resistance, depending on the enzyme type. In order to better understand the molecular bases of resistance to O2 inhibition, this chapter focuses on the hydrogenases and their reaction with O2 and examines the different strategies to lead to engineer kinetically efficient hydrogenases operating under aerobic conditions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Electron Paramagnetic Resonance
- Hydrogen Production
- Heterologous Expression
- Biofuel Cell
- Allochromatium Vinosum
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
I. Introduction
Climate change, along with the rapid depletion of oil and gas reserves, prompt the world to turn to a search for a clean energy sources to provide the energy necessary for present and projected human activities. A variety of possible fuel sources are being examined at present. Among these, dihydrogen (H2) has been identified as a clean and renewable energy carrier and is found to be one potential alternative to fossil fuel energy and has drawn a worldwide attention as a future energy source (Mason 2007). Interestingly, the pioneering notion of a “Dihydrogen Energy System” drew inspiration from the French science-fiction novel “The mysterious Island” by Jules Vernes (1874), where the idea of using H2 as an energy carrier first appeared. However, over 90 % of the production of H2 remains based upon steam reforming of hydrocarbons and coal gasification, which starts from fossil fuels and requires high temperature and pressure conditions. So, a sustainable, renewable supply of H2 to power this economy is required. Alternative methods of H2 generation include electrolysis of water and biological means. The development of new biotechnological processes, designed to meet the future energy demand, may take advantage of microbes that have been using H2 from very early in the evolution of life (Perez-Arellano et al. 1998; Andersson and Kurland 1999). Many organisms including some Bacteria, Archaea and unicellular eukaryotes have an active H2 metabolism, utilizing the cleavage of H2 to gain energy, or H2 production to release reducing power (Casalot and Rousset 2001). It has been estimated that these microorganisms produce or consume more than 200 million tons of H2 per year (Richardson and Stewart 1990). These processes, carried out by hydrogenase occur via the inter-conversions between the molecular hydrogen and two protons plus two electrons (H2 ↔ 2H+ + 2e−).
In this sense, an outlook in the production of hydrogen from water and light energy would be to use photosynthetic microorganisms, such as cyanobacteria and green algae (Antal et al. 2011; Carrieri et al. 2011). Green algae have many areas of potential improvements that often overlap with those of cyanobacteria and are currently recognized as better photobiological hydrogen producers from a demonstrated solar efficiency standpoint (Ghirardi et al. 2009). In principle, there is absorption of light in the form of photons by the photosystem which are going generate a strong oxidant that can oxidize water into protons, electrons/reducing equivalents and O2. Thereafter, the electrons reduce protons to form H2, carried out by hydrogenases (Fig. 3.1). In this sense, photosynthetic production of H2 using water as a source of electrons and sunlight as the source of energy, driving proton reduction, is the most desirable process. Both water and light are available to an almost unlimited extent and in addition to H2 only O2 is formed, whereas greenhouse gases are avoided. Moreover, photosynthetic production of H2 is potentially very efficient in terms of energy conservation, since 10 % of the incident light energy can theoretically be recovered into H2 (Prince and Kheshgi 2005). For example, an average sun-light flux of 46 Mwh/ha/day can be converted, with a 10 % yield, in 1,650 m3 of H2 per hectare per day, which represents 145 TOE (ton of oil equivalent) per hectare per year.
H2 production by cyanobacteria and algae. Water (bold) is oxidized to O2 by photosystem II (PSII) and electrons are transferred to photosystem I (PSI) via the plastoquinone pool (PQ). Photosystem I transfers electrons to ferredoxin (Fd) which can donate electrons to [FeFe]-hydrogenase (H2ase) in algae or to nitrogenase (N2ase) in some cyanobacteria. Fd electrons can also be transferred to NAD(P)+ by a ferredoxins NAD reductase (FNR). NAD(P)H can donate electrons to [NiFe]-hydrogenase in cyanobacteria.
In addition to being used to the in vivo H2 production, hydrogenases are known to be used in a variety of biotechnological applications including biofuel cells, biosensors, prevention against microbial-induced corrosion, and the generation and regeneration of NAD(P) cofactors. In the case of biofuel cell, the nature of the hydrogenases has inspired researchers worldwide to use them as biocatalysts predominantly to replace platinum electrode in hydrogen fuel cells (Fig. 3.2). Indeed, platinum is limited in availability and very expensive and therefore the use of hydrogenases would be good candidates to replace this precious metal in fuel cells. Moreover the hydrogenase-coated electrode confers greater fuel specificity and turnover rates than the platinum and it could be used to as an alternative to allow operation of biofuel cells at neutral pH and ambient temperatures, which are the conditions much more favorable for the handling of fuel cells (Ikeda and Kano 2001; Morozov et al. 2002).
Most applications of H2-oxidizing and H2-producing catalysts require them to function in air. For example, an O2-stable H2-production catalyst is an essential component for the photobiological production of H2, a device that uses solar energy to split water into H2 and O2. Similarly, use of hydrogenases in biofuel cells requires them to remain active in the presence of O2 since biofuel cells must work necessarily with O2 (Figs. 3.1 and 3.2). However, the major barrier of developing an economically viable systems, is the oxygen sensitivity of the vast majority of hydrogenases because their active site react strongly with O2 (Cournac et al. 2004; Léger et al. 2004; Oh et al. 2011).
At present, structure function relationship studies in hydrogenases have mainly remained in the basic research realm, aimed at understanding the enzyme catalytic mechanism (De Lacey et al. 2005). In order to obtain a sufficient level of enzyme efficiency and robustness for technological purposes, hydrogenases must be functionally optimized by improving efficiency H2-oxidation or production while improving O2 tolerance. These challenges can be approached through genetic engineering by two different strategies: at the cellular level, by metabolic engineering, it is possible to create favorable conditions to improve the H2-production by deletion of hydrogen-uptake system (Liang et al. 2009), in inserting of non native hydrogenases more efficient and tolerant to O2 (Wells et al. 2011) or to simply avoid O2 exposure (Kruse et al. 2005; Rupprecht et al. 2006; Henstra et al. 2007). At the enzyme level, by protein engineering, the goal will be to improve hydrogenases so that they can outcompete other enzymes for substrate utilization, use of a thermodynamically more favorable substrate or become more O2-tolerant. In this chapter, we will review hydrogenase structure-function relationship studies in which new properties of modified enzymes might serve as an inspiration source for rational optimization of hydrogenases for biotechnological processes.
In the first part, we will describe the different hydrogenases, and then we focus on these which are naturally O2-tolerants. Thereafter, we will process the engineering approaches in three parts: (1) Improving H2 production by heterologous expression of non-native hydrogenases or their over-expression. (2) Improving the electron transfer by increasing the substrate specificity or redirect redox intermediates. (3) Engineering protein to improve the tolerance of the hydrogenases to O2.
II. Classification and Physiological Properties of Hydrogenases
Hydrogenases are metalloproteins which are involved in the metabolic machinery of a wide variety of microorganisms by catalyzing the reversible heterolytic splitting of dihydrogen according to the elementary reaction: H2 < == > H− + H+ < == > 2H+ + 2e−.
Since 1931, when hydrogenases were described by Stephenson and Stickland, extensive research has been conducted in this area. Biochemical, spectroscopic, phylogenetic studies made possible to separate three groups of hydrogenases on the basis of the metal content of their active site: [Fe]-, [FeFe]- and [NiFe]-hydrogenase (Vignais et al. 2001; Cournac et al. 2004).
A. Generalities
The three classes of hydrogenases are evolutionarily unrelated but share similar nonprotein ligand assemblies at their active site that are not observed elsewhere in biology. They all contain a complex active-site cofactor that consists of at least one Fe atom coordinated by varying numbers of cysteine-S ligands and biologically unique carbon monoxide (CO) and in most cases additional cyanides (CN) ligands (Fig. 3.3). Therefore, this structure Fe(CO)2 or Fe(CO-CN) likely represents the minimal cofactor making hydrogenase activity possible. These metal cofactors are synthesized in a coordinated post-translational process that involves up to nine hydrogenase-specific auxiliary proteins (Böck et al. 2006; Lenz et al. 2010; Mulder et al. 2011). In the bimetallic hydrogenases the active-site cofactor is electronically coupled to FeS clusters that direct the electrons from the active site to the protein surface or from an external electron donor to the active site. Moreover, analysis of crystals structure from all three classes of hydrogenases revealed a network of hydrophobic cavities and channels (Fig. 3.3), or packing defects that form pathways connecting the active site to the surface of the enzyme (Volbeda et al. 1995; Montet et al. 1997; Nicolet et al. 1999, 2002; Fontecilla-Camps et al. 2007; Hiromoto et al. 2009; Mulder et al. 2011; Hong and Pachter 2012; Nicolet and Fontecilla-Camps 2012). Molecular dynamics simulations and xenon mapping of [NiFe]-hydrogenases show that these channels facilitate the diffusion of H2 between the bulk of solvent and the active site (Montet et al. 1997; Cohen et al. 2005; Fontecilla-Camps et al. 2007; Leroux et al. 2008; Liebgott et al. 2010; Topin et al. 2012). In addition, the pathways can facilitate diffusion of small gas molecules such as CO and O2 to access the active site, which in the case of CO leads to a reversible inhibition and in the case of O2, to complete but non destructive inhibition.
Structure of [NiFe]-hydrogenase, [FeFe]-hydrogenase and [Fe]-hydrogenase. Panel (a) shows a structural model of the periplamsic [NiFe]-hydrogenase from D. fructosovorans (1YQW). The active site is buried in the large subunit (purple) whereas the small subunit contains three consecutive iron-sulfur clusters (light blue). Panel (b) shows the structure of [FeFe]-hydrogenase of C. pasteurianum (CpI: 3C8Y). All cofactors (active site and iron-sulfur) are localized in the same subunit (blue). The PyMOL Molecular Graphics System was used for visualization. The lower part of the figure shows the catalytic centers of three hydrogenases. The extremity of hydrophobic channels for gas diffusion were computed using Caver2.0 and appear as gray meshes for the panels a and b. Panel (a) residues, component the entrance gate of gases, are indicated in red and the multiple sequence alignment (ClustalW) of some [NiFe]-hydrogenase large subunits, is presented.
The redox chemistry of hydrogenases is rich and involves many intermediate states, as beside catalyzing H2 oxidation/production, they can also interact with gaseous molecules (i.e. CO, O2) and become inhibited. It is thus important to gain an understanding into how hydrogenases catalyze H2 production and oxidation, to determine the mechanisms by which they are inactivated under oxidizing conditions and how they may become re-activated. A combination of spectroscopic and electrochemical methods has provided structural knowledge on the oxidised and reduced forms of the enzyme, namely electron paramagnetic resonance (EPR) spectroscopy, Fourier transform infrared (FTIR) spectroscopy and protein film voltammetry (PFV).
EPR is a spectroscopic technique widely used to study the hydrogenase. This technique allows the detection of chemical species that have unpaired electrons designated as paramagnetic. A large number of molecules contain such paramagnetic atoms such as nickel at the active site, or the iron atoms of the iron-sulfur centers in [NiFe]-hydrogenases. Upon reduction or oxidation, the metal atoms of the prosthetic groups go through several redox states and in some of these states, they become paramagnetic and can unambiguously be identified by their corresponding EPR spectrum. Magnetic coupling between these atoms also provide information of the active or inactive states.
Another powerful method for monitoring reactions at the active sites of hydrogenases is infrared spectroscopy. This is an unusual technique to use for studying enzymes, but for hydrogenases it exploits the fact that CN− and (particularly) CO are strong infrared-active vibrational oscillators and their stretching frequencies appear in a spectral window where the rest of the protein and water do not absorb.
Beside spectroscopic studies that provide thermodynamic information at equilibrium states, kinetic parameters have equally been studied using protein film voltametry (PFV), where the hydrogenase is adsorbed to an electrode and its activity directly measured by electron transfer through the electrode under oxidizing or reducing potentials during gas exposure (Inhibitors: CO, O2 or substrate: H2) (Vincent et al. 2007; Armstrong et al. 2009).
B. Classification
[Fe]-Hydrogenase (EC 1.12.98.2)
This type of enzyme was found only in a small group of methanogenic Archaea and has been described for the first time in Methanothermobacter thermoautotrophicum (Zirngibl et al. 1992). This enzyme catalyzes CO2 reduction to methane using H2 (Vignais and Billoud 2007). Based on the metal content of their active site, although in the past they were considered as metal-free hydrogenases, they have been recently designated as “iron-sulfur-cluster free hydrogenase” or simply [Fe]-dihydrogenases (Armstrong and Albracht 2005). This enzyme is also known as H2-forming Methylenetetrahydromethanopterin Dehydrogenase (Hmd) (Corr and Murphy 2011). The structure of the active site and functional models have been reported in 2008 (Shima 2008; Hiromoto et al. 2009). This hydrogenase differs from the others hydrogenases not only by the primary and tertiary structures but also by the fact that the iron, required for enzyme activity is not redox active. Furthermore, the hydrogenase activity is rapidly lost under aerobic conditions and in presence of light (Lyon et al. 2004) which would make the isolation and the characterization very difficult.
[FeFe]-Hydrogenase (EC 1.12.7.2)
[FeFe]-hydrogenase have been found mainly in Gram positives and in eukaryotes, as well as in few anaerobic Gram negative (Atta and Meyer 2000; Horner et al. 2000, 2002). It should be emphasized that these are the only type of hydrogenase that is found in eukaryotes but that is absent in the Archaea domain (Cournac et al. 2004). In this sense, the hydrogenases present in eukaryotic microorganisms (Green algae) are [FeFe]-hydrogenases only. Generally, [FeFe]-hydrogenases are usually involved in H2 production but they were also reported to function as an uptake hydrogenase. Indeed, the location of hydrogenases in the bacterial cell reflects the enzyme’s function (Nicolet et al. 2000). For instance, the periplasmic Desulfovibrio desulfuricans [FeFe]-hydrogenase (DdH) is involved in dihydrogen uptake. Protons resulting from this dihydrogen oxidation create a gradient across the membrane that is thought to be coupled to ATP synthesis in the cytoplasm. Clostridium pasteurianum [FeFe]-hydrogenase I (CpI) is a cytoplasmic enzyme that accepts electrons from ferredoxin and generates dihydrogen with protons as electron acceptors.
The only [FeFe]-hydrogenase structures from anaerobic soil bacterium Cl. pasteurianum (CpI) (Peters et al. 1998) (pdb code 1FEH and 3C8Y) and sulfate reducing bacterium D. desulfuricans (DdH) (Nicolet et al. 1999) (pdb code 1HFE) revealed an unique active site metal cluster, termed as the H-cluster, where catalysis takes place. The H-cluster is composed of a binuclear [2Fe]H center bound to a [4Fe-4S]H subcluster by a bridging cystein, the [4Fe-4S] center is attached to the protein by four cystein ligands. The [2Fe]H center is coordinated by five diatomic CN− and CO ligands, as well as a non protein dithiomethylamine ligand (Fig. 3.3b). Molecular masses of [FeFe]-hydrogenases can vary from 45 to 130 kDa according to the number of subunits. [FeFe]-hydrogenases are mainly monomeric and contains only one catalytic subunit, but they often comprise additional domains, which accommodate FeS clusters. For example in the [FeFe]-hydrogenase I from Cl. pasteurianum (CpI), three accessory [4Fe4S] clusters and one [2Fe2S] cluster are believed to transfer electrons between the electron donor or acceptor at the protein surface and the active site at the center of the protein (Nicolet et al. 2002; Nicolet and Fontecilla-Camps 2012). Contrarily, the simplest characterized [FeFe]-hydrogenases are observed in the green algae, including Chlamydomonas reinhardtii, Chlorella fusca, and Scenedesmus obliquus, which express enzymes consisting of only the H-cluster without FeS-cluster domains (Florin et al. 2001; Horner et al. 2002; Forestier et al. 2003). These proteins have been exploited more recently for biochemical and spectroscopic characterization because they lack the additional FeS clusters observed in most native [FeFe]-hydrogenases that may complicate the direct examination of the H-cluster (Kamp et al. 2008; Silakov et al. 2009; Stripp et al. 2009; Mulder et al. 2011).
Even though [FeFe]-hydrogenases may appear as the best suited for hydrogen production purposes, enzyme engineering studies for these enzymes are still poorly developed because of their great sensitivity to oxidative damage, which makes any biochemical characterization very uncertain.
a. Reaction with O2 and CO
In most [FeFe]-hydrogenases, oxygen inhibits the enzyme, possibly by binding to the open coordination site on the distal Fe of the [2Fe]H center, then form a reactive oxygen species that destroys the [4Fe4S]H subcluster. This distal Fe would be equally the hydrogen binding site and also the site of reversible CO binding and inhibition (Stripp et al. 2009). The chemical nature of the oxygen species bound to the H-cluster after the exposure to O2 is not known, but density function theory (DFT) calculations on inactivated states of the H-cluster have proposed a FeII-FeII oxidation state for the [2Fe]H center, with a possible OH group terminally bound to the distal Fe (Liu and Hu 2002).
Oxygen inactivation has been studied using protein film electrochemistry and it was shown that the rate of inhibition of [FeFe]-hydrogenases would be limited by two steps: (1) the diffusion of oxygen through the protein to the active site pocket, and (2) the binding of oxygen to the [2Fe]H subcluster (Armstrong et al. 2009).
Interestingly, the level of O2 inhibition varies among [FeFe]-hydrogenases, with I50 values ranging from less than a few seconds for Chlamydomonas reinhardtii enzymes, to several minutes for the clostridial enzymes (Böck et al. 2006; Baffert et al. 2008). Algal hydrogenases, which lack the additional accessory cluster domain found in bacterial enzymes, are typically more sensitive to O2 inhibition than are the enzymes isolated from bacteria. However, it is clear that this elevated sensitivity is not solely due to the lack of the N-terminal accessory cluster (Böck et al. 2006; Stripp et al. 2009).
[NiFe]-Hydrogenase (EC 1.12.2.1)
The most numerous and best studied class of hydrogenases have been the [NiFe]-hydrogenases. This type of enzyme was found in Bacteria and Archaea domains. The core enzyme consists of two subunits; the large subunit is approximately 60 kDa and houses the Ni-Fe-active site, whereas the small subunit, of approximately 35 kDa, which can be of variable size and harbors typically three iron-sulfur clusters (Fig. 3.3a): a distal [4Fe-4S] cluster at the surface of the protein and furthest from the active site; a medial [3Fe-4S] cluster; and a proximal Fe-S cluster, with variable properties, the closest to the active site. The large and small subunit exhibit sequence homologies to subunits of NADH:ubiquinone oxidoreductase (Complex I) (Volbeda et al. 2012). In certain enzymes, additional subunits enable the interaction of these clusters with physiological electron carriers such as quinones, pyridine nucleotides (NAD(P)H), ferredoxins, and cytochromes (Cournac et al. 2004). Crystal structure analysis of heterodimeric [NiFe]-hydrogenases from Desulfovibrio species (Volbeda et al. 1995, 2002; Higuchi et al. 1997; Matias et al. 2001) (Fig. 3.3a), and photosynthetic bacterium Allochromatium vinosum (Ogata et al. 2010), revealed that the Ni-Fe cofactor is deeply buried in the large subunit. The Ni is coordinated to the protein via four thiol groups from conserved cysteine residues; two of these are bridging ligands that coordinate both Fe and Ni (Volbeda et al. 1995, 2002; Higuchi et al. 1997; Matias et al. 2001). Fourier Transform Infrared spectroscopy revealed that the Fe coordination sphere also possesses three diatomic ligands: one CO and two CN molecules (Volbeda et al. 1996; Pierik et al. 1999). The sixth iron coordination position is assumed to be occupied by a bridging hydride between iron and nickel (Pardo et al. 2006; De Lacey et al. 2007). Hydrophobic cavities which channel the gas substrate between the protein surface and the active site (Fig. 3.3a) (Montet et al. 1997; Volbeda et al. 2002; Teixeira et al. 2006) as well as a proton-conducting channel (Léger et al. 2004) were identified inside the hydrogenase.
a. Classification
Based on their primary protein sequences the [NiFe]-hydrogenases have been categorized into four different groups (Table 3.1): Group I, H2-uptake enzymes, localized in the bacterial or archaeal cell membrane, are primarily involved in H2 oxidation; Group II, (a) cyanobacterial uptake [NiFe]-hydrogenases, whose location is cytoplasmic and which are involved in N2 fixation and (b) H2 regulatory hydrogenases or hydrogenase sensors, which detect the presence of H2 in the environment and trigger a cascade of regulation controlling the synthesis of hydrogenases; Group III, cytoplasmic bidirectional enzymes, these water soluble multi-protein complexes are dependent on NAD(P)H or NAD(P)+ as cofactors; Group IV, energy conserving H2-evolving hydrogenases, these membrane-associated hydrogenases generate H2 from reduced ferredoxin with the concomitant generation/utilization of an ion gradient (Vignais and Billoud 2007).
b. Reaction with O2 and CO
It can be stated, as a general rule, that hydrogenases of either type are inhibited by O2, but individual sensitivities can vary in a wide extent. Thus, [NiFe]-hydrogenases are considered to be more robust than [FeFe]-hydrogenases because they can be totally reactivated after inhibition by O2. In the case of [NiFe]-hydrogenases, O2 has also been shown to oxidize directly the bimetallic active site (van der Zwaan et al. 1990) but the difference with [FeFe]-hydrogenases is that [NiFe]-hydrogenases are not damaged by O2 as they can be reactivated by reduction. X-ray diffraction studies showed that the main structural difference between oxidized and reduced states of the active site is that oxygen species bridge the metals in the oxidized state (Garcin et al. 1999; Carepo et al. 2002; Volbeda et al. 2005). Therefore, reductions of the oxidized states are triggered by the removal of the bridging oxygen species, which allows H2 to bind to the active site and catalytic turnover. Studies of EPR revealed that oxidized enzyme may exist under two different states which have been called Ni-A and Ni-B (Fig. 3.4). Another very important feature, that differentiates Ni-A and Ni-B is their reactivation kinetics (Cammack et al. 1986). Ni-B is called the ‘ready’ state because it quickly becomes active upon reduction, while Ni-A is called the ‘unready’ state because it needs a long period of incubation under reducing conditions before becoming active. This is illustrated by PFV experiments conducted on the Allochromatium vinosum [NiFe]-hydrogenase. After inactivation by O2, the enzyme is reactivated by reduction at low potential under H2 (Lamle et al. 2004). For example at -208 mV (SHE) and pH 6, the reactivation process occurs in two phases: a fast (instantaneous) phase corresponding to the reactivation of the Ni-B state and a slower phase (several hundreds of seconds) assigned to Ni-A. FTIR-spectroelectrochemical studies of different hydrogenases indicated that one-electron reduction of Ni-A and Ni-B leads to two different states: Ni-A leads to the Ni-SU state, and Ni-B leads to Ni-SI (De Lacey et al. 2007). The enzyme in the Ni-SI state is active, whereas the Ni-SU state is still inactive (De Lacey et al. 2007). The rate-limiting step of the reactivation process is the a gradual and spontaneous conversion of the Ni-SU to the active Ni-SI state (Lamle et al. 2005) (Fig. 3.4).
Proposed reaction scheme for the standard O2 sensitive [NiFe]-hydrogenase from D. fructosovorans based on electrochemical titrations, including both active and inactive intermediates. The nomenclature is based on that initially adopted to designate the EPR paramagnetic states (Ni-A, NiB, NiC) and the “Ni-S” designates an EPR silent state. In Ni-A and Ni-SU states, the binding oxygenic species presented by (Ox) has not been conclusively determined. There are multiple forms of Ni-C and Ni-R and these have been suggested to differ in terms of the redox state of the nearest (proximal) iron–sulfur cluster and/or protonation of the cysteinyl sulfur atoms.
As regards the structures in either oxidized state, this stir up still a matter of debate. Indeed, X-ray diffraction, (Volbeda et al. 1995, 2002) EPR (van der Zwaan et al. 1990), ENDOR (Carepo et al. 2002; van Gastel et al. 2006) and EXAFS (Davidson et al. 2000) data indicate that in both states an oxygen species is bridging nickel and iron atoms (De Lacey et al. 2007; Pandelia et al. 2010b). Most of the studies agree with the presence of a hydroxide in the Ni-B state (Black et al. 1994; Davidson et al. 2000; Stein et al. 2001; Stadler et al. 2002; Volbeda et al. 2005) whereas the nature of the oxygen species in Ni-A is more controversial. Indeed, either oxo (Carepo et al. 2002), hydroxo (Davidson et al. 2000; Stein et al. 2001; Stadler et al. 2002; Pandelia et al. 2010b) or peroxo (Lamle et al. 2005; Volbeda et al. 2005) species have been proposed. However, the “peroxo” hypothesis was not ruled out in recent DFT and spectroscopic studies (van Gastel et al. 2008; Pandelia et al. 2010b) but rather a hydroxo species. Moreover, an ENDOR study of a sample of [NiFe]-hydrogenase aerobically oxidized in H2 17O, demonstrated that the bridging [NiFe]- ligand in Ni-A originates from the solvent water (Carepo et al. 2002). Thus, this will mean that the debate continues on the question of the Ni-A structure with the aim to understand differences between Ni-A and Ni-B states (Fig. 3.4).
Most [NiFe]-hydrogenases are inhibited by CO in a competitive manner (Teixeira et al. 1987; Léger et al. 2004). It reaches the active site using the same gas channel as H2 and O2 (Fig. 3.3a) (Liebgott et al. 2010). It binds weakly to the Ni ion at the active site (Albracht 1994; Stadler et al. 2002; De Lacey et al. 2007; Lubitz et al. 2007), only after reductive activation of the enzyme to the Ni-SI forms, presumably when the bridging oxygen species had been removed from the active site. No binding of CO occurs when the enzyme is in the inactive states Ni-A, Ni-B, and Ni-SU. In addition, CO-inhibition blocks electron and proton transfer at the active site, although reduction at the proximal [4Fe-4S] cluster is detected (Stadler et al. 2002). The kinetics of CO inhibition has been studied by PFV experiments and the kinetics of CO binding was fast, about 108 s−1/M. Thus, it has been shown that diffusion was the rate-limiting step of CO-inhibition. As a result, this inhibitor has been used to probe gas diffusion in hydrogenases (Leroux et al. 2008; Liebgott et al. 2010).
c. Specific Characteristics of the Naturally Occurring O2-Tolerant [NiFe]-Hydrogenases
[NiFe] hydrogenases are considered to be more robust than [FeFe] hydrogenases because they can be totally reactivated after inhibition by O2. Moreover, there are even a few examples in nature of relatively O2-tolerant [NiFe] hydrogenases. Thus, the [NiFe] hydrogenase can be classified as either ‘standard O2-sensitive’ or ‘O2-tolerant’ based on their ability to function in the presence of O2. O2-tolerance defines one hydrogenase that retains some activity in the presence of O2. The level of residual activity can vary depending on the enzyme, but it should be remembered that trace amounts (a few μM) of O2 readily inhibit standard O2-sensitive [NiFe] hydrogenases.
The fact that a large number of microbes are able to use H2 as the sole energy source in the course of aerobic respiration indicates that some [NiFe] hydrogenases afford a specific protection against detrimental effects of O2 (Tremblay and Lovley 2012). This aerobic H2 oxidation occurs in phylogenetically diverse groups of prokaryotes such as the nitrogen-fixing bacterium Bradyrhizobium japonicum (Kaneko et al. 2002), the photosynthetic proteobacterium Rhodobacter capsulatus (Strnad et al. 2010), the hyperthermophilic bacterium Aquifex aeolicus (Deckert et al. 1998) and the well-studied proteobacterium Ralstonia eutropha H16 that contains even three indigenous O2-tolerant hydrogenases (Schwartz et al. 2003; Pohlmann et al. 2006; Lenz et al. 2010; Bürstel et al. 2011).
R. eutropha H16 is a bacterium that lives in soil and water and it is one of organisms able to grow chemolithoautotrophically using hydrogen as the sole energy source and dioxygen as terminal electron acceptor. This is called the Knallgas (detonating gas) reaction. The three distinct O2-tolerant [NiFe] hydrogenases, that each serve unique physiological roles, are: a bidirectional cytoplasmic Soluble Hydrogenase (ReSH) able to generate reducing equivalents by reducing NAD+ at the expense of hydrogen (Group III), a Regulatory Hydrogenase (ReRH) which acts in a signal transduction cascade to control transcription of hydrogenase genes (Group II) and a Membrane-Bound Hydrogenase (ReMBH) coupled to the respiratory chain (Group I) (Burgdorf et al. 2005). Interestingly, these O2-tolerant hydrogenases are usually less active than standard O2-sensitive enzymes. Their H2 oxidation activities are reduced by a factor of about 5 for the ReSH (Ludwig et al. 2009a), about 50 for the ReMBH and about 500 for the ReRH (Vignais and Billoud 2007). The H2 production activities are usually considerably weak, especially because of the strong inhibitory effect of H2 (Goldet et al. 2008). However, these O2-tolerant enzymes represent precious inspiration sources for the study of the molecular bases of O2 inhibition.
Three strategies seem to have been developed by R. eutropha H16 to allow its [NiFe] hydrogenases to be catalytically active in the presence of dioxygen.
In the case of the SH (Group III), the O2 resistance was assumed to be due to the presence of extra CN ligands at the active site (Happe et al. 2000) that might be incorporated by a specific maturation protein HypX (Bleijlevens et al. 2004). Deletion of hypX led to a lower O2 resistance of the ReSH enzyme (Bleijlevens et al. 2004) while the activities of ReMBH (Buhrke and Friedrich 1998) and of the regulatory hydrogenase (ReRH) remained unaffected (Buhrke et al. 2001). However, the presence of a Ni-Bound cyanide under native conditions has been recently ruled out (Horch et al. 2010). In this context, the supply of low-potential electrons from the oxidation of NAD(P)H appears to play a major role in preserving catalytic activity under aerobic conditions in vivo. However, despite considerable efforts and promising insights (Horch et al. 2012), the structural and mechanistic basis for this property has still to be resolved. Bidirectional cytoplasmic soluble hydrogenases are of particular interest for biotechnological applications as they are suited for light driven hydrogen production in vivo (Prince and Kheshgi 2005) and the regeneration of NAD(P)H in biocatalytic processes (Okura et al. 1990; Ratzka et al. 2011). Such applications are particularly promising as some of these enzymes are oxygen tolerant in contrast to most other hydrogenases (Happe et al. 2000; Horch et al. 2012).
The ReRH and related enzymes adopted apparently another strategy consisting in reducing the gas channel size at the level of the interface with the active site cavity. At the end of the hydrophobic channel, near the active site, two hydrophobic residues, usually valine and leucine that are conserved in O2-sensitive hydrogenases, are replaced by larger residues, respectively isoleucine and phenylalanine, in the O2-tolerant hydrogen-sensors (Volbeda et al. 2002) (Fig. 3.3a). It has therefore been suggested that increasing the bulk of residues occupying these two positions may act like a molecular sieve, reducing the channel diameter at that point, thereby preventing efficient dioxygen access to the active site. This hypothesis was supported by two experiments in which the bulky amino-acids from ReRH were substituted by valine and leucine. In both cases, determination of the inactivation kinetics in the presence of dioxygen revealed that the mutated enzymes were inactivated after prolonged incubation and required a reductive activation to reach the maximum activity (Burgdorf et al. 2005; Duche et al. 2005). Even though the mutated enzymes became more sensitive to dioxygen than the wild-type, it should be noted that they retain a significant level of activity after prolonged dioxygen exposure, and therefore still belong to the O2-tolerant group of hydrogenases.
Membrane bound hydrogenases of the group I represent the best studied group of O2-tolerant hydrogenases and has recently gained extensive attention, due to their potential biotechnological importance. Thus, the O2 tolerance of MBH (group I) has been recently discovered through the hydrogenase crystal structures of R. eutropha H16 (ReMBH, pdb accession number 3RGW) (Fritsch et al. 2011), E. coli (EcHyd-1, pdb 3UQY, 3USC and 3USE) (Volbeda et al. 2012), and Hydrogenovibrio marinus (HmMBH, pdb 3AYX, 3AYY, 3AYZ) (Shomura et al. 2011) and spectroscopic data obtained from hydrogenases of A. aeolicus (AaHyd-1) (Pandelia et al. 2011) and ReMBH (Fritsch et al. 2011; Goris et al. 2011; Lukey et al. 2011). Therefore, the recent crystallographic structures from three members of the Group I revealed that there are no significant differences between the [NiFe] catalytic centres of standard or O2-tolerance hydrogenases. The presence of the usual nickel signatures detected by EPR indicates that the chemistry at the active site is identical to that catalyzed by standard hydrogenases.
Remarkably, there are two additional cysteine residues in the close vicinity of the proximal FeS cluster that are absent in O2-sensitive standard [NiFe] hydrogenases which lead to the construction of a unique proximal [4Fe-3S] cluster with in-total six coordinating cysteine residues (Fig. 3.5). These two additional cysteines are fully conserved in the O2-tolerant hydrogenase (Pandelia et al. 2010a). The sulfur atom of an extra cysteine replaces one of the inorganic sulfides and thus becomes an intrinsic cluster ligand, whereas the second cysteine terminally coordinates one of the Fe atoms (Fig. 3.5). This particular structure studied by EPR in its reduced or oxidized form in HmMBH, AaHyd-1 and ReMBH, indicate that the [4Fe-3S] cluster is stable in three oxidation states. These redox-dependent structural changes promoted by the surplus of cysteine coordination, give the potential to the proximal cluster to theoretically deliver two electrons for the O2-reduction instead of one (Goris et al. 2011; Lukey et al. 2011). Thus when O2-tolerant [NiFe]-hydrogenases are attacked by O2, they might fully reduce O2 to water, thereby avoiding the production of reactive oxygen species that would damage or block the active site. As a result, it is currently assumed that the electron deficiency during O2 attack might be responsible for the formation of the Ni-A inactive form (‘unready’ state) (Ogata et al. 2009, 2010). O2-tolerant hydrogenases would escape the ‘unready’ state by forming only the ‘ready’ conformation (Ni-B) (in which O2 has been fully reduced), which re-activates very easily to re-join the catalytic cycle (Armstrong et al. 2009; Lenz et al. 2010). However, in the case of AaHyd-1, a weak Ni-A signal has been reported to appear after O2 exposure (Guiral et al. 2006), while only Ni-B was detected in a recent study (Pandelia et al. 2010b). The EcHyd-1 exhibits a Ni-A signal upon aerobic isolation but this signal is then barely detectable when the enzyme is exposed to O2 after activation (Lukey et al. 2010).
Multiple sequence alignment and structural comparison of the proximal clusters between O2-sensitive and O2-tolerant hydrogenases. Panel (a) shows the multiple sequence alignment comparing the cysteines which ligate the proximal cluster in O2-sensitive [NiFe]-hydrogenases and O2-tolerant enzymes. Panels (b) and (c) shows the structural comparison between the proximal FeS clusters of D. fructosovorans (O2-sensitive hydrogenase, 1YQW) and H. marinus (O2-tolerant hydrogenase, 3AYY and 3AYX). Panel (c) Details of the structural changes associated with ‘super-oxidation’ of the proximal cluster in membrane bound O2-tolerant [NiFe]-hydrogenases (Adapted from Parkin and Sargent 2012).
In PFV experiments, the activity of the ReMBH and AaHyd-1 recover extremely fast after O2 exposure (Armstrong et al. 2009; Pandelia et al. 2010b). The O2-tolerance properties of these enzymes are therefore likely to be due to a fast reactivation rate, as shown in recent electrochemical studies (Armstrong et al. 2009; Liebgott et al. 2010; Pandelia et al. 2010b) and by the lack of the Ni-A signal after aerobic inactivation.
The exploitation of these hydrogenases and their molecular determinants is a major challenge for a broad range of biotechnological applications. Indeed, microorganisms harboring optimized hydrogenases may play a major role in H2 generation for fuels. Bio-fuel cells and biosensors also represent an important potential application of these hydrogenases as immobilized enzymes. At last, these hydrogenases might also allow searches in the metabolic engineering in the aim to improve the H2 production or in protein engineering to mimic molecular determinants responsible to the tolerance towards of O2.
III. Maturation of Hydrogenases: Specific and Complex Process
In metal-containing enzymes, complex active sites generally require specific machineries for their synthesis and assembly. Indeed, assembly of the active hydrogenase involves sophisticated biological processes, such as careful co-ordination of cofactor biosynthesis and insertion, subunit recruitment, and protein target processes (Vignais et al. 2001; Paschos et al. 2002; Böck et al. 2006).
A. [FeFe]-Hydrogenase Maturation: Protein Machinery
In order to be catalytically active after its synthesis, the [FeFe]-hydrogenase polypeptide encoded by the hydA gene has to incorporate the H-cluster and, when required, accessory [Fe-S] clusters. This post-translational process is extraordinarily complex as it involves a number of difficult reactions including: (i) the synthesis of CO, CN and the dithiolate bridging ligand; (ii) the assembly of the di-iron active site subcluster; (iii) its incorporation into the enzyme already containing the [4Fe-4S] component of the H-cluster and (iv) the assembly and transfer of the accessory FeS clusters.
The [FeFe]-hydrogenase maturation protein machinery was initially discovered in the eukaryotic green alga Chlamydomonas reinhardtii incapable of H2 production (Posewitz et al. 2004). The disruption of either the hydEF or hydG (Hyd machinery) gene resulted in a mutant that proved to be unable to produce hydrogen, even though full-length hydrogenase accumulated. Genes encoding for HydE, HydF, and HydG are present in all organisms capable of synthesizing an active [FeFe]-hydrogenase (HydA) (Meyer 2007). Thus, it can be concluded that HydEF and HydG provide the minimal protein machinery necessary for the synthesis and assembly of the H-cluster. Whether other proteins are required for an optimal maturation process has not been demonstrated so far. Moreover, several reports of heterologous expression of active [FeFe]-hydrogenases have demonstrated that the Hyd machinery from one organism can be successfully used for the maturation of an enzyme from another. For example, expression of an active HydA1 enzyme from C. reinhardtii or Scenedesmus obliquus (green algae) has been shown to be possible using Clostridium acetobutycicum, another [FeFe]-hydrogenase synthesizing organism, as the expression host (Girbal et al. 2005). Further evidence for the lack of selectivity of the Hyd machinery came from the observation that co-expression of HydE, HydF and HydG from the bacterium Cl. acetobutylicum with various algal and bacterial [FeFe]-hydrogenases in E. coli resulted in purified enzymes with specific activities that were not very different from those of their counterparts from native sources (Böck et al. 2006). Finally, the bacterium Shewanella oneidensis proved to be an efficient system for the expression and maturation of HydA1 from C. reinhardtii (Sybirna et al. 2008).
B. [NiFe]-Hydrogenase Maturation: Protein Machinery
At present, cloning [NiFe]-hydrogenases is still very difficult and the progresses realised recently remain very limited (Burgdorf et al. 2005; Ludwig et al. 2009b). Even though [NiFe]-hydrogenase operons are highly conserved and exhibit a high degree of similarity, each maturation system is specific to the corresponding structural subunits, probably because of tight protein-protein interactions occurring during processing (Leach et al. 2007). Indeed, the complex architecture of the active site of [NiFe]-hydrogenases with their diatomic ligand (CN and CO) requires a specific and complex maturation system. There are two main groups of genes responsible for maturation, which are differentiated by their resultant phenotypes. The first group of genes is mainly located on the same transcription unit as the structural genes. Disruption of this group of genes specifically impairs the processing or activity of the hydrogenase encoded in cis in the operon, without affecting the maturation of other hydrogenases. The maturation processes mediated by the products of this family of accessory genes cannot be complemented in trans by homologous genes from the other hydrogenase operons, regardless of the degree of similarity (Sauter et al. 1992; Menon et al. 1994; Bernhard et al. 1996). This specific barrier is one of the key reasons for the failure of the active hydrogenase production in heterologous hosts. The second group is another set of the hyp (‘p’ for pleiotropic) genes which encode proteins, involved in the insertion of Ni, Fe, CO and CN into the active site (Jacobi et al. 1992; Maier et al. 1996; Wolf et al. 1998; Böck et al. 2006; Mulder et al. 2011; Petkun et al. 2011). Mutations of these genes affect the synthesis and activity of all the hydrogenase isoenzymes. However, the functions of this set of genes can be complemented in trans by heterologous genes (Chaudhuri and Krasna 1990).
C. O2-Tolerant [NiFe]-Hydrogenase Maturation: Protein Machinery
It would be particularly fruitful to take advantage of the properties of the O2-tolerant hydrogenases by cloning their corresponding genes into organisms of biotechnological interests. Although [NiFe]-hydrogenase exhibit reversible inhibition by oxygen, the sensitivity of hydrogen production in presence of O2 is a multifaceted problem, since hydrogenase transcription, and likely maturation and assembly, might be also inhibited by exposure to atmospheric oxygen (Soboh et al. 2012). However, among O2-tolerant hydrogenases, some are synthesized solely under aerobic conditions (Lukey et al. 2011; Tremblay and Lovley 2012). E. coli could be one of the most informative model systems for understanding the biosynthesis of O2-tolerant enzymes because the bacterium produces both O2-tolerant MBH (Hyd-1) and standard O2-sensitive (Hyd-2) hydrogenases. Among the three types of O2-tolerant hydrogenases, only the MBH family appears to require specific maturation proteins, necessary for the synthesis of the unique [4Fe-3S] proximal cluster (Figs. 3.5 and 3.6). Hyd-1 is produced from an operon of six genes, hyaABCDEF, where HyaA is the small subunit, HyaB is the large subunit, HyaC is a cytochrome that anchors the [NiFe]-hydrogenase to the membrane and the HyaD is the specific protease required for large subunit maturation-terminal processing. There are therefore two extra genes hyaE and hyaF that are not required for the assembly of standard O2-sensitive hydrogenases (Hyd-2 and others) and are apparently only involved in the assembly of O2-tolerant respiratory enzymes (Schubert et al. 2007).
Schematic maturation process of the [NiFe] hydrogenase. The R. Eutropha MBH gene cluster is presented at the top. hoxK encodes the small subunit and hoxG encodes the large subunit. The genes in red and green encode the accessory proteins, the role of which in the assembly of the active is presented (Adapted from Fritsch et al. 2011).
In R. eutropha H16 the HyaE homolog is HoxO, which has been shown to interact with the small subunit during biosynthesis and is essential for MBH activity in that organism (Schubert et al. 2007) (Fig. 3.6). In Rhizobium leguminosarum the HyaE homolog, HupG, was shown to be only required for small subunit maturation under aerobic conditions (Manyani et al. 2005), and in E. coli, which only expresses Hyd-1 under anaerobic conditions, the hyaE gene was dispensable for Hyd-1 biosynthesis (Dubini and Sargent 2003). Similarly, the HyaF (HoxQ in R. eutropha H16 and HupH in R. leguminosarum) is also absolutely required for MBH activity. HyaF interacts with HyaE to form a complex together with the small subunit during assembly (Schubert et al. 2007). Genetic two-hybrid studies suggested that HyaE interacted strongly with HyaA, the small subunit, (Dubini and Sargent 2003). It is possible, therefore, that HyaE-like proteins have a role to play in assembly of the [4Fe-3S] cluster. Shomura et al. (2011) suggested that additional negative charges around the proximal [4Fe-3S] cluster in the final structure may be important for its stabilization. Interestingly, HyaE has a perfect thioredoxin-like fold (Parish et al. 2008), except that in the HyaE protein, acid residues (two aspartates and two glutamates) are found at the same position as the redox-active cysteines, originally present in true thioredoxins (Parish et al. 2008). Thus, it is possible that this negatively charged region mediates protein-protein interactions with the small subunit, though it can also be considered that it might be involved in protecting the proximal cluster until the large subunit has docked correctly with its small subunit partner.
IV. Enzyme and Metabolic Engineering to Improve H2 Production
Genetic modifications of hydrogen metabolism or hydrogenases, can be very promising strategies to achieve an efficient H2 production system or to improve hydrogenases as biocatalysts. During the last 5 years, many reviews dealt with this topic and summarized the scientific and technological hurdle encountered (Ghirardi et al. 2007; Germer et al. 2009; Brentner et al. 2010; McKinlay and Harwood 2010; Abo-Hashesh et al. 2011; Hallenbeck et al. 2012). In this section, we will discuss the different processes of engineering used to generate significant improvement in the production of hydrogen or hydrogenases, it will be divided in three points: (A) the heterologous expression and overexpression of interesting hydrogenases; (B) enhancing the efficiency of H2 production by redirecting the flow of reducing equivalents toward hydrogenases, and (C) increasing the O2 tolerance of hydrogenase by enzyme engineering.
A. Heterologous-Expression and Overexpression of Hydrogenases
In order to characterize their structure-function properties in greater detail, and to use hydrogenases for biotechnological applications, reliable methods for rapid, high-yield expression and purification are required. Owing to genetic manipulations, the purification of recombinant hydrogenases is greatly facilitated by the use of affinity tags, such as His or StrepII-tag, inserted at the N- or C-terminus (Kim et al. 2012). One approach that has been adopted in order to enhance the amount of enzymes or promote the H2 production, is the engineering of a stable hydrogenase either through its production in a heterologous host or through overexpression (English et al. 2009; Abo-Hashesh et al. 2011). The interest of heterologous expression systems is not to be demonstrated but is still limited to [FeFe] hydrogenases (Böck et al. 2006; Nicolet and Fontecilla-Camps 2012).
[FeFe]-Hydrogenase
In recent years, several groups have developed different strategies for the expression of recombinant [FeFe]-hydrogenases (English et al. 2009) (Table 3.2). These include using as hosts organisms bacteria expressing naturally native [FeFe]-hydrogenase (Girbal et al. 2005; Sybirna et al. 2008), or using E. coli for heterologous expression in a more common way. At present, in order to obtain large amounts of [FeFe]-hydrogenases for biochemical and biophysical studies, the studies are focused on the high yield heterologous production mainly in E. coli (Kuchenreuther et al. 2010; Yacoby et al. 2012). On the other hand, the overexpression of [FeFe]-hydrogenase is also an important factor for the improvement of H2 production. Indeed, the most efficient hydrogen-producing enzymes are [FeFe]-hydrogenases, which can have an activity about 10–100 times higher than that of [NiFe]-hydrogenases. As a result, overexpressing (HydA) in Cl. paraputrificum M-21 (Morimoto et al. 2005) improved the H2 yield from 1.4 to 2.4 mol H2 per mol of glucose. Jo et al. (2009) also reported that Cl. tyrobutyricum JM1 showed an improved H2 yield (1.8 mol H2/mol glucose) compared to the parental strain (1.2 mol H2/mol glucose), when HydA was overexpressed. Regarding metabolic engineering, the inactivation of ack, which encodes acetate kinase of Cl. tyrobutyricum (Liu et al. 2006) increased the H2 production yield by 1.5-fold compared to the wild-type strain.
[NiFe]-Hydrogenase
The main interests in producing [NiFe]-hydrogenases in heterologous hosts are to improve hydrogen production by focusing on (a) the heterologous expression of bidirectional [NiFe]-hydrogenases of the group III and (b) to take advantage of the O2 tolerance properties of some [NiFe]-hydrogenases (Carrieri et al. 2011) (Table 3.2). However, as discussed above, the development of heterologous expression systems for the biosynthesis and molecular engineering of [NiFe]-hydrogenases is challenging due to the complexity and the high specificity of the maturation process. There are several documented examples of non-functional heterologous expression, which have only recently been reported for a limited number of organisms. For example, heterologous expression of [NiFe]-hydrogenases from Rhodococcus opacus, Desulfovibrio vulgaris, and Synechocystis sp. PCC6803 all resulted in the production of non-functional hydrogenases (Voordouw et al. 1987; Grzeszik et al. 1997; Maeda et al. 2007). For this reason, expression and purification of [NiFe]-hydrogenase for structural and in vitro studies are most often accomplished through the development of plasmid -based expression in homologous strains or closely related species as expression hosts (Rousset et al. 1998; Burgdorf et al. 2005; Ludwig et al. 2009b).
E. coli is an ideal microorganism commonly used in genetic engineering due to its well-characterized genome, well known metabolism, and its ability to utilize a wide range of carbon sources including hexoses and pentoses. In addition, E. coli has the potential advantages over at least some other microorganisms to exhibit a rapid growth, has simple nutritional requirements and also harbors four [NiFe]-hydrogenases. Two hydrogenases, one O2-tolerant hydrogenase (Hyd-1) and one standard O2-sensitive hydrogenase (Hyd-2) encoded by the hya and hyb operons respectively, are involved in periplasmic hydrogen uptake. Two others, hydrogenase 3 and 4, are part of cytoplasmically oriented hydrogenase complexes. Hydrogenase 3, encoded by the hyc operon, produces hydrogen from formate as a part of the formate hydrogen lyase complex (FHL-1), which is active in hydrogen production during mixed-acid fermentation at acidic pHs (Vignais et al. 2001). Hydrogenase 4, encoded by the hyf operon, appears to be cryptic under normal circumstances (Self et al. 2004). Thus, E. coli has a set of specific maturation proteins, which might be used for the maturation of heterologous enzymes (Porthun et al. 2002; Maroti et al. 2003, 2009). Although E. coli is perhaps the most useful organism as a target for metabolic engineering, the lack of any NADH-dependent hydrogenases (Group III) is one major hurdle for the engineering of hydrogen metabolism in this organism. Indeed, these bidirectional cytoplasmic hydrogenases function reversibly in their physiological setting, coupling hydrogen uptake or emission, to oxidation and reduction of cellular coenzymes, such as NAD(P)H (Cournac et al. 2004). So, in E. coli, NAD(P)H generated through carbon metabolism cannot be used directly for H2 production (Schmitz et al. 2002). Therefore, expressing a heterologous NAD(P)H-dependent hydrogenase is one of the main goals for increasing the maximum H2 yield in E. coli.
a. Heterologous Expression of Bidirectional [NiFe]-Hydrogenases (Group III) in E. coli
The [NiFe]-hydrogenase of the cyanobacterium Synechocystis sp. PCC6803 is a well-studied representative of the bidirectional [NiFe]-hydrogenases from Group III. Therefore, the maturation process has been reconstituted to allow functional expression of this hydrogenase in E. coli (Maeda et al. 2007; Wells et al. 2011; Zheng et al. 2012). As a result, the introduction of the bidirectional [NiFe]-hydrogenase of Synechocystis sp. PCC6803 has altered the whole metabolism for hydrogen production in E. coli. Firstly, the hydrogen productivity was enhanced up to 41-fold in comparison with Synechocystis sp. PCC6803 (Maeda et al. 2007). Secondly the expression of this hydrogenase also showed a distinct H2 production pathway than the one initially presents in E. coli (Wells et al. 2011). Finally, this heterologous expression has suppressed the transcription of native uptake [NiFe]-hydrogenases (Hya and Hyb) (Maeda et al. 2007; Zheng et al. 2012), increasing significantly the production of H2. Similarly, the successful expression in E. coli of a recombinant cytoplasmic, NADP-dependent hydrogenase from Pyrococcus furiosus, an anaerobic hyperthermophile increased significantly the production of H2. Remarkably, the native E. coli maturation machinery was able to generate a functional hydrogenase when transformed with only the genes encoding the hydrogenase structural subunits and the C-terminal protease (Sun et al. 2010).
b. Heterologous Expression of O2-Tolerant [NiFe]-Hydrogenase (Group I)
The biotechnological goal of algal and cyanobacterial hydrogen production is to divert the reducing equivalents away from normal growth functions and to redirect them toward hydrogenases. All the enzymatic components for hydrogen production from water splitting and sunlight are present in cyanobacteria. As discussed above, hydrogenases are inactivated by molecular oxygen, which represent a major technological hurdle for hydrogen production from cyanobacteria at high solar efficiencies. Thus, strategies for overcoming this barrier include heterologous expression of a more oxygen-tolerant hydrogenase in cyanobacteria. In this connection, a system of heterologous expression has been developed to express O2-tolerant [NiFe]-H2ase belonging to group I from the bacteria Alteromonas macleodii and Thiocapsa roseopersicina (Vargas et al. 2011) in the cyanobacterium Synechococcus elongatus (Maroti et al. 2009); (Vargas et al. 2011). The cloned enzymes were active, indicating that it is possible to express hydrogenases in the cyanobacteria.
Recently, Kim et al. (2011) have succeeded in performing a heterologous expression of oxygen-tolerant Hydrogenovibrio marinus [NiFe]-hydrogenase in E. coli. Interestingly, recombinant H. marinus [NiFe]-hydrogenase produced of sevenfold to ninefold more hydrogen than did E. coli [NiFe]-hydrogenase (Hyd-1) in a gaseous environment containing 5–10 % (v/v) oxygen. Likewise, the same team (Kim et al. 2012) has improved the H2 production (1.3 fold more) of this recombinant H. marinus [NiFe]-hydrogenase in co-expressing it with the proteorhodopsin under light conditions.
B. Substrate Selectivity, Competition and Linking
When expressed in vivo, hydrogenases interact with electron carriers which are generally at the junction of numerous redox reactions (respiration, CO2 fixation, assimilation, etc.). A key point is that these reactions that appear as competitors for biotechnological purposes are often essential for cell survival or development. This explains in part the difficulty and the slow progress in biohydrogen research. One proposed research direction, to overcome this kind of limitation, would be to increase the amount of specific redox partners of hydrogenases. In this sense, a recent study has showed that the overexpression of ferredoxin-NADPH-reductase (FNR) coupled to that of ferredoxin (Fd) and a [FeFe]-hydrogenase, increased the H2 production when NADPH was added in the medium (Weyman et al. 2011).
Another strategy to favor H2 production would be to carry out an expression of synthetic or chimerical enzymes based on native hydrogenase fused to the electron transfer subunit (Kontur et al. 2012). In the case of algal [FeFe]-hydrogenases for instance, the redox partner is Fd (Fig. 3.1), which is also involved in photosynthetic carbon fixation via NADPH production by FNR. Indeed, under the anaerobic conditions that support hydrogen production, there is a significant loss of photosynthetic electrons toward NADPH production supplied by ferredoxin:NADP+ oxidoreductase (FNR). No algal [FeFe]-hydrogenase structure is available yet, but Horner et al. (2002) modeled algal hydrogenase structure and charge distribution, and identified a set of amino-acids likely to participate in electrostatic interaction with algal Fd. Mutagenesis experiments conducted at the (putative) Fd binding sites in hydrogenase (and also at the Fd binding site in FNR) could be a way to modify relative affinities of these enzymes for their substrate and ultimately tune these affinities for an optimal ratio between photosynthetic and hydrogen-producing capabilities. Some results illustrated the feasibility of utilizing directly attached redox partners for H2 production in vivo (Agapakis et al. 2010). Indeed, it has been reported that H2 production via Fd-dependent hydrogenase can be improved by manipulating the interaction between hydrogenase and Fd via protein surface engineering in E. coli. Some chimerical enzymatic complexes have been expressed in E. coli, in which a ferredoxin and heterologous [FeFe] hydrogenase were either immobilized in a modular protein scaffold or directly attached to each other via an amino acid linker. H2 production from cells containing these complexes showed 3-fold and 4.4-fold increases in H2 production, respectively, over cells containing separate versions of the same proteins (Agapakis et al. 2010). However, yields from these complexes were relatively low (<0.1 mol H2/mol glucose). More recently, a complex consisting of the Fd fused to the [FeFe]-hydrogenase HydA from C. reinhardtii (green algae), was shown to improve the light-dependent H2 production in vitro when incubated with purified PSI (Yacoby et al. 2011). This algal Fd-HydA fusion prevents the competition between FNR and HydA that both exhibit affinity with Fd. Moreover, Fd-HydA fusion improved HydA function in several respects. First, the specific activities were up to sixfold higher than for the native HydA. Second, the fusion successfully insulates its internal Fd electrons, because only 10 % of the electrons are lost with external competitors such as FNR. Third, the fusion was able to overcome the limitation caused by FNR, as more than 60 % of photosynthetic electrons were diverted to hydrogen production, compared to less than 10 % for nonfused HydA (Yacoby et al. 2011).
Another tempting approach to favor H2 production would be to tightly connect a specific electron carrier or a photosystem with hydrogenase, making the electrons flow directly from the photosystem to the hydrogenase or from the hydrogenase to the electron acceptor, avoiding competition with the bulk of electron carriers. In a study, Ihara et al. (2006) engineered a ‘hard-wired’ protein complex consisting of a hydrogenase and a photosystem. They designed an artificial fusion protein composed of the membrane-bound [NiFe]-hydrogenase from R. eutropha H16 and the peripheral photosystem I (PSI) subunit PsaE (involved in the docking of Fd to the PSI) of the cyanobacterium Thermosynechococcus elongatus. The resulting hydrogenase-PsaE fusion protein when associated with PsaE-free PSI spontaneously formed a complex which showed light-driven hydrogen production at a rate of 0.58 μmol H2/mg chlorophyll/h. The complex retained accessibility to the native electron acceptor Fd, which is necessary for autotrophic growth of these cells. But unfortunately, the activity was totally suppressed in the presence of the physiological PSI partners, Fd and FNR. In an attempt to establish a H2 photoproduction system in which the activity is not interrupted by Fd and FNR, the same group introduced a chimeric protein of PsaE and cytochrome c3 (cytc3) from D. vulgaris into the cyanobacterium Synechocystis sp. PCC6803 (Ihara et al. 2006). The covalent adduct of cytc3 and PsaE assembled with PsaE-free PSI and formed a complex which was still able to reduce Fd for photosynthesis (approximately 20 % of the original activity). Interestingly, this complex was able to drive hydrogen production when coupled with hydrogenase from D. vulgaris even in the presence of Fd and FNR, although the rate was limited (around 0.30 μmol H2/mg chlorophyll/h). These results suggest, however, that this type of complex may eventually be modified to produce H2 in vivo. More recently, Cl. acetobutylicum [FeFe]-hydrogenase was fused with Synechococcus sp. PCC 7002 PSI via a 1,6-hexanedithiol molecular wire (Lubner et al. 2010). However, while this complex also catalyzed light-dependent H2 production in vitro, the abiotic nature of the wire make the system difficult to implement in vivo. Another system in which a fused PSI-[NiFe]-hydrogenase complex was immobilized on a gold electrode (Krassen et al. 2009) produced H2 at the equivalent of 3 mmol H2/mg chlorophyll/h (Bürstel et al. 2011), it then was suggested that this system is analogous to the in vivo situation where membrane-bound PSI receives electrons from the photosynthetic electron transport chain (Bürstel et al. 2011).
C. Protein Engineering to Improve the O2-Tolerance of Hydrogenases
To develop a viable H2 technology, hydrogenases should work in presence of O2. Indeed, the direct biological photoproduction of hydrogen at the expense of water oxidation will unavoidably lead to a certain exposure to O2. Similarly, the bio-fuel cells operate necessarily with O2 to realize the oxidation of H2 by hydrogenases. Thus, improving hydrogenase oxygen resistance is then a major challenge for a broad range of biotechnological applications such as hydrogen photoproduction, bio-fuel cells, and biosensors. As said above, there are currently several projects in progress with the single objective of identifying and expressing natural O2-tolerant [NiFe]-hydrogenases in model host organisms in order to over-express the hydrogenases of interest or to improve H2 production.
Another strategy is to carry out molecular engineering studies directly on the hydrogenases by directed mutagenesis for instance. Significant interest surrounds molecular engineering studies aimed at achieving hydrogenases with low levels of sensitivity to O2 (Bürstel et al. 2011), and standard O2-sensitive [NiFe]-hydrogenases are often the target of these studies because they are reversibly inhibited by oxygen unlike [FeFe]-hydrogenases. One major difficulty, however, lies in the complexity of the maturation process associated with the low production of native [NiFe]-hydrogenases. One of the most studied is the periplasmic O2-sensitive [NiFe]-hydrogenase from Desulfovibrio fructosovorans (Rousset et al. 1998). This enzyme is soluble, highly produced, genetically accessible and it can be crystallized without much difficulties. All the mutants of interests were fully characterized, at the structural, spectroscopic and kinetic levels. Studies of this enzyme, at the molecular level, enabled to determine the mechanisms of electron transfer (Dementin et al. 2006, 2011), of proton transfer (Léger et al. 2004) and to increase the tolerance towards O2 (Dementin et al. 2009; Liebgott et al. 2010; Dementin et al. 2011). Research strategies to increase the tolerance towards oxygen have been primarily inspired by some key features observed in the O2-tolerant hydrogenase from R. eutropha H16 (Bleijlevens et al. 2004; Burgdorf et al. 2005; Fritsch et al. 2011). Thus, two general strategies have been followed: (1) slowing down the oxygen diffusion along the gas channel and (2) changing the reactivity of oxygen with the active site.
Slowing Down of the Oxygen Diffusion Along the Gas Channel
Firstly, it is important to understand how the structure of the tunnel in hydrogenases determines the diffusion rate and possibly the selectivity of the enzymes with respect to substrates and inhibitors of similar sizes. From a multiscale simulation approach, associated with the comparison between the sequences and biochemical properties of homologous [NiFe] hydrogenases, it has been proposed that diffusion in these enzymes is controlled by two gates, which guard the entrance of the active site (Fig. 3.3a), and may determine the accessibility of the active site and therefore the resistance to O2. One is located between residues 74 and 476 and the other between residues 74 and 122 (Fig. 3.3a) (Volbeda and Fontecilla-Camps 2004; Kim et al. 2012; Topin et al. 2012) (We use D. fructosovorans amino acids numbering throughout). This hypothesis has been erratically supported by the results of mutagenesis studies. Both R. eutropha H16 and R. capsulatus RH (regulatory hydrogenase) become oxygen sensitive when the two conserved bulky amino-acids, Phe122 and Ile74, are replaced with smaller residues, Leu and Val respectively, which are conserved in “standard,” O2-sensitive [NiFe] hydrogenases, such as those from D. fructosovorans and A. vinosum (Burgdorf et al. 2005; Duche et al. 2005).
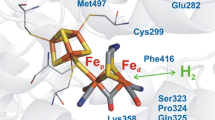
Conversely, inspired by the RH, substitutions of the Val74 and Leu122 with isoleucine and phenylalanine, respectively, were carried out in the [NiFe]-hydrogenase from D. fructosovorans (Fig. 3.7). Surprisingly, these substitutions did not improve O2 tolerance and did not significantly modify the catalytic properties of the enzyme under anoxic conditions (Dementin et al. 2009). Consequently, the residue bulkiness at these positions was not the only parameter affecting O2 tolerance. Indeed, the existence of two control points in different locations might explain why the reduction in the experimental diffusion rate does not simply correlate with the width of the main gas channel. Moreover, the orientation or chemical nature of the side chain are also crucial to slow down the diffusion rate and the inhibition rate by O2 (Leroux et al. 2008).
Rate of inhibition by O2 (\( {k}_{in}^{O_2} \)) plotted against the rate of binding of CO (k CO in ). The dashed line depicts y = x for which O2-inhibition rate would be a linear function of diffusion rate; the plain line is the best fit to equation: \( 1/{k}_{in}^{O_2}=1/{k}_{in}^{CO}+1/{k}_{in}^{O_2} \max \), with \( {k}_{in}^{O_2} \max \) = 32 s−1 mM(O2)−1. Error bars represent either the deviation from the average of three to five independent determinations or the estimated error introduced from the extrapolation to 40 °C (Adapted from Liebgott et al. 2010).
a. Slowing Down of the Diffusion Rate
In order to explore the respective roles of the bulk, hydrophobicity, charge and polarity, a number of single and double mutants at positions 122 and/or 74 have been constructed and purified. Val74 was replaced with aspartate (D), asparagine (N), tryptophan (W), glutamate (E), glutamine (Q), isoleucine (I), and phenylalanine (F) while Leu122 was replaced with phenylalanine, alanine (A) and methionine (M) (Liebgott et al. 2010). The mutants were screened using PFV to estimate different parameters such as the rate of inhibition by CO (kinCO; the limiting step is the diffusion rate) and the rate of O2-inhibition (kinO2; the limiting step is the reaction rate) (Leroux et al. 2008; Liebgott et al. 2010) (Fig. 3.7).
The effect of the mutations that keeps the side chain hydrophobic indicates a simple correlation between bulk and diffusion rate. Compared to the wild-type (WT), the diffusion rate is reduced by about two orders of magnitude for the V74F and about three orders of magnitude for the V74W. The increase of the molecular volume (calculated according to Ref. Häckel et al. 1999) of the amino-acid side chains lining the channel has a strong influence on the diffusion rate. In this set of experiments, the effect observed is only due the steric hindrance that shrivels the tunnel (Fig. 3.7).
In the case of polar amino-acids, two levels of impact were observed on diffusion, involving both bulk and charge. The influence of changing the size and polarity of the residue at position 74 was analyzed by comparing the V74E, V74D, V74Q and V74N mutants two at a time. Increasing the length of the side chain by one carbon (V74D to V74E, or V74N to V74Q) slows the diffusion rate about 30-fold. The magnitude of this effect lies within the same range as that observed with hydrophobic residues. Nevertheless, charge also matters as the substitution V74 to D74 slows the diffusion by a factor 40, while the molecular volume of aspartate is 40 % smaller than that of valine (Häckel et al. 1999). The polarity has an even stronger impact as the substitution V74 to Q74 reduces the diffusion by about four orders of magnitude, while the molecular volumes of these two amino acids are quite similar (Häckel et al. 1999). Within the polar amino acids, replacing a carboxylic acid with an amide, keeping the Van der Waals volume constant (V74E to V74Q, or V74D to V74N), slows diffusion by a factor of about 12. The two contributions, size and polarity, are independent of each other, and therefore the combination of the two (V74D to V74Q) decreases the rate of diffusion by more than two orders of magnitude. The electrostatic interaction of the amino acid side chains lining the tunnel is therefore very unfavourable to CO diffusion (Fig. 3.7). This could be due to a direct interaction of the polar group with the gas molecule or the Arg476 residue (residue strictly conserved within [NiFe]-hydrogenases, which lead to complications in actual experimental mutations), or to the stabilization of a water molecule that would be part of the barrier to ligand entry, as observed in certain myoglobin mutants (Nienhaus et al. 2003).
b. Slowing Down of the Inhibition Rate by O2
In [NiFe] hydrogenase, the rate of inhibition by CO is about four orders of magnitude faster than the rate of aerobic inhibition (3.104 s−1/M) (Léger et al. 2004; Liebgott et al. 2010). Considering that CO and O2 diffuse within the protein at about the same rate, this observation implies that the rate of inhibition by O2 is limited by the reaction at the active site. Mutations such as V74N or V74W decrease the rate of intramolecular diffusion by blocking the tunnel, but this has no effect on the overall reaction with O2 because the diffusion process does not limit the inhibition rate (Fig. 3.7). However, other mutations decrease the rate of diffusion in such a large extent (three orders of magnitude for V74E or four orders of magnitude for V74Q) that this step becomes limiting because is slower than the reaction of O2 at the active site, thus decreasing the overall rate of inhibition by O2.
Changing of the Reactivity of Oxygen with the Active Site
Contrary to the ReRH, the O2 tolerance of both MBH and SH is performed by a peculiar reaction of the active site with O2. In the case of MBH the presence of a unique proximal [4Fe-3S] cluster is thought to be responsible for the tolerance of these hydrogenases towards O2. The complexity and specificity of the maturation process of this unique [4Fe-3S] cluster will make the realization of variant enzymes, mimicking the capacities of O2-tolerant [NiFe] Hydrogenases (MBH), more difficult. However, drawing inspiration from ReSH various mutations have been carried out. Amino acids like cysteine and methionine have a high affinity for oxygen, which results in a strong reactivity with reactive oxygen species and they are also known to participate in oxidative stress responses and protection in several proteins (Kim et al. 2001; Stadtman et al. 2002; Stadtman 2004, 2006; Reddie and Carroll 2008). With this knowledge in mind, several mutants at position 74 and 122 have been performed to replace valine by methionine or cysteine (Dementin et al. 2009).
a. Introduction of Methionines in the Gas Channel Near the Active Site
In order to decrease the sensitivity of the [NiFe] hydrogenase from D. fructosovorans, the effects of replacing Val74 and Leu122 with methionines have been tested. Indeed, methionines placed at the entrance of the active site cavity at positions V74M and L122M may protect the Ni-Fe site from oxidation, either by reacting or at least by interacting with the oxygen species present at the active site under oxidizing conditions (Volbeda et al. 2005). The goal of that substitution was twofold: slowing diffusion and modifying the reactivity with O2.
At diffusion rate level, the molecular volume of methionine is 30 % larger than that of valine and about the same as leucine (Häckel et al. 1999). The diffusion rate in the case of the V74M-L122M is decreased by more than two orders of magnitude and even three orders of magnitude for the V74M (Fig. 3.7), which goes far beyond the expected effect of the volume increase on the diffusion. The interaction of gases with methionine is therefore stronger than a simple steric obstruction. Indeed, another interesting property of the V74M-L122M mutant was detected by PFV: unlike the WT enzyme, it partly reactivates in the presence of H2 even under very oxidizing conditions (Dementin et al. 2009) (Fig. 3.8). The V74M mutant also reactivates under H2 at high potential but to a lesser extent (Dementin et al. 2009). This process is slow and has a small amplitude, but it is significant because under the very oxidizing conditions used in the experiments, one would expect nothing but the inactivation of the enzyme (Vincent et al. 2007; Leroux et al. 2008) (Fig. 3.8).
The inhibition by O2 of D. fructosovorans [Ni-Fe]-hydrogenase selected mutants. (a) The change in dioxygen concentration plotted against time, reconstructed from the amount of O2 injected and the time constant of the exponential decay; the latter is calculated from fitting the change in current. (b) The plain lines show the change in [Ni-Fe]-hydrogenase activity (current i normalized by its value i(0) just before the inhibitor is added); (c, d) Enlarged views of the data in b, showing the decreases in current after the first exposure to O2 (c) and the partial reactivation of the L122M-V74M mutant (d). E = +200 mV versus SHE, T = 40 °C, pH 7, electrode rotation rate w = 2 kr.p.m (Adapted from Liebgott et al. 2010).
Crystallographic and spectroscopic studies showed that methionines are not modified in the oxidized enzymes but that they interact with the active site by modifying its environment. FTIR studies determined that the mutants were inactivated more slowly and reactivated more rapidly than the native enzyme. This slower inactivation is attributed to a reduced active site accessibility that is due to partial tunnel obstruction by the mutations (Leroux et al. 2008). However the faster activation necessarily involves a quicker removal of the bound oxygen species. This was assumed to involve methionine that would stabilize the rearrangement of the oxygen species that is necessary to allow its protonation, facilitating its escape from the oxidized enzyme. As a result, the phenotype of the V74M and V74M-L122M mutants is not a consequence of a modification of the structure of the active site, but rather reveals subtle changes in the kinetics of the reaction with O2.
These different studies clearly showed that the V74M-L122M and V74M mutated hydrogenases became O2 tolerant, since these mutant hydrogenases continued to operate in the presence of 150 μM of O2, which is close to the O2 concentration of 200 μM in air-equilibrated solutions (Dementin et al. 2009).
b. Introduction of a Cysteine in the Gas Channel Near the Active Site
As described above, the O2-tolerance of membrane-bound [NiFe]-hydrogenases, results from the fact that they only convert into the Ni-B state under O2. Moreover, the reactivation rate of this inactive state is greater than that of the same species in O2-sensitive enzymes. In order to test the reaction of a thiol function with O2, the valine 74 in the wild type enzyme, has been exchanged with a cysteine (Liebgott et al. 2011). The obtained mutant has showed an activity during several minutes under oxygenated atmosphere. Further FTIR and PFV experiments showed that the inhibition of V74C by oxygen led to the formation of Ni-B state only, unlike the wild-type enzyme (Fig. 3.9). Moreover, the Ni-B state of V74C reactivated 17–25 times faster than the same species in the wild type (Fig. 3.9). Two other point mutations, at the level of valine 74, have been constructed to further address the role of cysteine 74 in modulating oxygen reactivity. The V74S and V74N mutations have showed also a significant increase of the rate of reactivation after inactivation under aerobic conditions (~ 4 and 11 times faster than wild type), although these rates are smaller than that of V74C (Fig. 3.10) Thus, the V74C mutant exhibits similar features typical of the naturally-occurring oxygen-tolerant hydrogenases (MBH). However, in the resistant EcMBH, ReMBH and AaMBH enzymes, it has been hypothesized that an additional source of electrons supplied to the active site would be responsible for the conversion into the Ni-B species only. V74C cannot supply electrons but from a combination of biochemical, spectroscopic (EPR, FTIR), X-ray and PFV studies, the authors proposed that V74C accelerates the electron transfer from the FeS clusters to the active site. This may promote (i) complete reduction of O2 into H2O and OH−, which favors the formation of the Ni-B state and (ii) fast reactivation of this state (Dementin et al. 2011). Thus, with the V74M and V74M-L122M mutants (Dementin et al. 2009), the V74C mutant provides another evidence that it is possible to improve the oxygen tolerance of hydrogenases, which is a prerequisite for their biotechnological use in air. Interestingly, the V74C mutant was not designed to copy the sequence of the oxygen resistant enzymes, these results are totally unexpected but they suggest new strategies for conceiving oxygen-tolerant enzymes.
Reactivation of the WT (red) and V74C (purple) enzymes, at-90 mV, pH 5.5, 1 atm of H2, after aerobic inactivation at + 215 mV, under 1 atm of Argon. The WT reactivates in two phases (NiB and NiA or ‘ready’ and ‘unready’ states respectively). In contrast, the reactivation of the V74C mutant is essentially monophasic and has a time constant of reactivation 20 times faster than for WT (Adapted from Liebgott et al. 2011).
Aerobic inhibition of D. fructosovorans WT (red), V74C, (purple), V74S (cyan), and V74N (green) and A. aeolicus MBH (dark blue) hydrogenases adsorbed at a rotating electrode poised at +140 mV, 40 °C, pH 5.5, 1 atm H2. (a) Oxygen concentration in the cell during experiment. The three peaks correspond to injections of 100, 200, and 500 μL of an air-saturated solution (b) Effect of O2 on enzyme’s activity. The traces were recorded after a 200 s (WT) or 500 s (V74C) poise at +140 mV (Adapted from Liebgott et al. 2011).
The main outcome of these studies is the demonstration that it is possible to induce O2-tolerance in a [NiFe] hydrogenase, in such a way that modified enzymes are active in the presence of O2 at concentrations close to that in air-equilibrated solutions. The fact that the substitutions tested are located in conserved regions makes it possible to engineer [NiFe]-hydrogenases in a wide range of organisms, as heterologous expression of [NiFe]-hydrogenase is still difficult. This achievement opens the way for future development in the field of biological hydrogen production or utilization.
V. Conclusion
Hydrogenase engineering is a very difficult issue not only because of the complexity of these enzymes but also because of the vital processes in which they are involved. However, in spite of the difficulties, research is very active in several countries because of the potential spin offs that might participate in the development of a new hydrogen energy economy. The enzyme bias, substrate specificity and oxygen resistance are the main domains in which some progress have already been made, opening the way towards future applications. But other issues, like heterologous expression of [NiFe]-hydrogenases that would facilitate molecular research and organism engineering or deciphering the catalytic mechanism that would allow the development of biomimetic catalysts, are also the subjects of intense research and will contribute to biohydrogen implementation.
Abbreviations
- DFT:
-
Density function theory;
- ENDOR:
-
Electron nuclear double resonance;
- EPR:
-
Electron paramagnetic resonance;
- EXAFS:
-
Extended X-Ray absorption fine structure;
- Fd:
-
Ferredoxin;
- FHL:
-
Formate hydrogen lyase;
- FNR:
-
Ferredoxin NADPH reductase;
- FTIR:
-
Fourier Transform InfraRed spectroscopy;
- Hmd:
-
H2-forming methylenetetrahydromethanopterin dehydrogenase;
- MBH:
-
Membrane-bound hydrogenase;
- PFV:
-
Protein film voltammetry;
- RH:
-
Regulatory hydrogenase;
- SH:
-
Soluble hydrogenase;
- SHE:
-
Standard hydrogen electrode;
- WT:
-
Wild type;
References
Abo-Hashesh M, Wang R, Hallenbeck PC (2011) Metabolic engineering in dark fermentative hydrogen production; theory and practice. Bioresour Technol 102:8414–8422
Agapakis C, Ducat D, Boyle P, Wintermute E, Way J, Silver P (2010) Insulation of a synthetic hydrogen metabolism circuit in bacteria. J Biol Eng 4:3
Albracht SP (1994) Nickel hydrogenases: in search of the active site. Biochim Biophys Acta 1188:167–204
Andersson SGE, Kurland CG (1999) Origins of mitochondria and hydrogenosomes. Curr Opin Microbiol 2:535–541
Antal T, Krendeleva T, Rubin A (2011) Acclimation of green algae to sulfur deficiency: underlying mechanisms and application for hydrogen production. Appl Microbiol Biotechnol 89:3–15
Armstrong FA, Albracht SP (2005) [NiFe]-hydrogenases: spectroscopic and electrochemical definition of reactions and intermediates. Philos Transact A Math Phys Eng Sci 363:937–954
Armstrong FA, Belsey NA, Cracknell JA, Goldet G, Parkin A, Reisner E, Vincent KA, Wait AF (2009) Dynamic electrochemical investigations of hydrogen oxidation and production by enzymes and implications for future technology. Chem Soc Rev 38:36–51
Atta M, Meyer J (2000) Characterization of the gene encoding the [Fe]-hydrogenase from Megasphaera elsdenii. Biochim Biophys Acta 1476:368–371
Baffert C, Demuez M, Cournac L, Burlat B, Guigliarelli B, Bertrand P, Girbal L, Leger C (2008) Hydrogen-activating enzymes: activity does not correlate with oxygen sensitivity. Angew Chem Int Ed Engl 47:2052–2054
Bai LP, Wu XB, Jing LJ, Liu J, Long MN (2012) Hydrogen production by overexpression of hydrogenase subunit in oxygen-tolerant Klebsiella oxytoca. Int J Hydrogen Energy 37:13227–13233
Bernhard M, Schwartz E, Rietdorf J, Friedrich B (1996) The Alcaligenes eutrophus membrane-bound hydrogenase gene locus encodes functions involved in maturation and electron transport coupling. J Bacteriol 178:4522–4529
Black LK, Fu C, Maier RJ (1994) Sequences and characterization of hupU and hupV genes of Bradyrhizobium japonicum encoding a possible nickel-sensing complex involved in hydrogenase expression. J Bacteriol 176:7102–7106
Bleijlevens B, Buhrke T, van der Linden E, Friedrich B, Albracht SP (2004) The auxiliary protein HypX provides oxygen tolerance to the soluble [NiFe]-hydrogenase of Ralstonia eutropha H16 by way of a cyanide ligand to nickel. J Biol Chem 279:46686–46691
Böck A, King PW, Blokesch M, Posewitz MC (2006) Maturation of hydrogenases. Adv Microb Physiol 51:1–71
Brentner LB, Peccia J, Zimmerman JB (2010) Challenges in developing biohydrogen as a sustainable energy source: implications for a research agenda. Environ Sci Technol 44:2243–2254
Buhrke T, Friedrich B (1998) hoxX (hypX) is a functional member of the Alcaligenes eutrophus hyp gene cluster. Arch Microbiol 170:460–463
Buhrke T, Bleijlevens B, Albracht SP, Friedrich B (2001) Involvement of hyp gene products in maturation of the H(2)-sensing [NiFe] hydrogenase of Ralstonia eutropha. J Bacteriol 183:7087–7093
Buhrke T, Lenz O, Krauss N, Friedrich B (2005) Oxygen tolerance of the H2-sensing [NiFe] hydrogenase from Ralstonia eutropha H16 is based on limited access of oxygen to the active site. J Biol Chem 280:23791–23796
Burgdorf T, Lenz O, Buhrke T, van der Linden E, Jones AK, Albracht SP, Friedrich B (2005) [NiFe]-hydrogenases of Ralstonia eutropha H16: modular enzymes for oxygen-tolerant biological hydrogen oxidation. J Mol Microbiol Biotechnol 10:181–196
Bürstel I, Hummel P, Siebert E, Wisitruangsakul N, Zebger I, Friedrich B, Lenz O (2011) Probing the origin of the metabolic precursor of the CO ligand in the catalytic center of [NiFe] hydrogenase. J Biol Chem 286:44937–44944
Cammack R, Fernandez VM, Schneider K (1986) Activation and active sites of nickel-containing hydrogenases. Biochimie 68:85–91
Carepo M, Tierney DL, Brondino CD, Yang TC, Pamplona A, Telser J, Moura I, Moura JJ, Hoffman BM (2002) 17O ENDOR detection of a solvent-derived Ni-(OH(x))-Fe bridge that is lost upon activation of the hydrogenase from Desulfovibrio gigas. J Am Chem Soc 124:281–286
Carrieri D, Wawrousek K, Eckert C, Yu J, Maness P-C (2011) The role of the bidirectional hydrogenase in cyanobacteria. Bioresour Technol 102:8368–8377
Casalot L, Rousset M (2001) Maturation of the [NiFe] hydrogenases. Trends Microbiol 9:228–237
Chaudhuri A, Krasna AI (1990) Restoration of hydrogenase activity in hydrogenase-negative strains of Escherichia coli by cloned DNA fragments from Chromatium vinosum and Proteus vulgaris. J Gen Microbiol 136:1153–1160
Cohen J, Kim K, King P, Seibert M, Schulten K (2005) Finding gas diffusion pathways in proteins: application to O2 and H2 transport in CpI [FeFe]-hydrogenase and the role of packing defects. Structure 13:1321–1329
Corr MJ, Murphy JA (2011) Evolution in the understanding of [Fe]-hydrogenase. Chem Soc Rev 40:2279–2292
Cournac L, Guedeney G, Peltier G, Vignais PM (2004) Sustained photoevolution of molecular hydrogen in a mutant of Synechocystis sp. strain PCC 6803 deficient in the type I NADPH-dehydrogenase complex. J Bacteriol 186:1737–1746
Davidson G, Choudhury SB, Gu Z, Bose K, Roseboom W, Albracht SP, Maroney MJ (2000) Structural examination of the nickel site in chromatium vinosum hydrogenase: redox state oscillations and structural changes accompanying reductive activation and CO binding. Biochemistry 39:7468–7479
De Lacey AL, Fernandez VM, Rousset M (2005) Native and mutant nickel-iron hydrogenases: unravelling structure and function. Coord Chem Rev 249:1596–1608
De Lacey AL, Fernandez VM, Rousset M, Cammack R (2007) Activation and inactivation of hydrogenase function and the catalytic cycle: spectroelectrochemical studies. Chem Rev 107:4304–4330
Deckert G, Warren PV, Gaasterland T, Gaasterland T, Young WG, Lenox AL, Graham DE, Overbeek R, Snead MA, Keller M, Aujay M, Huber R, Feldman RA, Short JM, Olsen GJ, Swanson RV (1998) The complete genome of the hyperthermophilic bacterium Aquifex aeolicus. Nature 392:353–358
Dementin S, Belle V, Bertrand P, Guigliarelli B, Adryanczyk-Perrier G, De Lacey AL, Fernandez VM, Rousset M, Leger C (2006) Changing the ligation of the distal [4Fe4S] cluster in NiFe hydrogenase impairs inter- and intramolecular electron transfers. J Am Chem Soc 128:5209–5218
Dementin S, Leroux F, Cournac L, de Lacey AL, Volbeda A, Léger C, Burlat B, Martinez N, Champ S, Martin L, Sanganas O, Haumann M, Fernández VM, Guigliarelli B, Fontecilla-Camps JC, Rousset M (2009) Introduction of methionines in the gas channel makes [NiFe] hydrogenase aero-tolerant. J Am Chem Soc 131:10156–10164
Dementin S, Burlat B, Fourmond V, Leroux F, Liebgott PP, Abou Hamdan A, Léger C, Rousset M, Guigliarelli B, Bertrand P (2011) Rates of intra- and intermolecular electron transfers in hydrogenase deduced from steady-state activity measurements. J Am Chem Soc 133:10211–10221
Dubini A, Sargent F (2003) Assembly of Tat-dependent [NiFe] hydrogenases: identification of precursor-binding accessory proteins. FEBS Lett 549:141–146
Duche O, Elsen S, Cournac L, Colbeau A (2005) Enlarging the gas access channel to the active site renders the regulatory hydrogenase HupUV of Rhodobacter capsulatus O2 sensitive without affecting its transductory activity. FEBS J 272:3899–3908
English CM, Eckert C, Brown K, Seibert M, King PW (2009) Recombinant and in vitro expression systems for hydrogenases: new frontiers in basic and applied studies for biological and synthetic H2 production. Dalton Trans 45:9970–9978
Florin L, Tsokoglou A, Happe T (2001) A novel type of iron hydrogenase in the green alga Scenedesmus obliquus is linked to the photosynthetic electron transport chain. J Biol Chem 276:6125–6132
Fontecilla-Camps JC, Volbeda A, Cavazza C, Nicolet Y (2007) Structure/function relationships of [NiFe]- and [FeFe]-hydrogenases. Chem Rev 107:4273–4303
Forestier M, King P, Zhang L, Posewitz M, Schwarzer S, Happe T, Ghirardi ML, Seibert M (2003) Expression of two [Fe]-hydrogenases in Chlamydomonas reinhardtii under anaerobic conditions. Eur J Biochem 270:2750–2758
Fritsch J, Scheerer P, Frielingsdorf S, Kroschinsky S, Friedrich B, Lenz O, Spahn CMT (2011) The crystal structure of an oxygen-tolerant hydrogenase uncovers a novel iron-sulphur centre. Nature 479:249–252
Garcin E, Vernede X, Hatchikian EC, Volbeda A, Frey M, Fontecilla-Camps JC (1999) The crystal structure of a reduced [NiFeSe] hydrogenase provides an image of the activated catalytic center. Structure 7:557–566
Germer F, Zebger I, Saggu M, Lendzian F, Schulz R, Appel J (2009) Overexpression, isolation, and spectroscopic characterization of the bidirectional [NiFe] hydrogenase from Synechocystis sp. PCC 6803. J Biol Chem 284:36462–36472
Ghirardi ML, Posewitz MC, Maness PC, Dubini A, Yu J, Seibert M (2007) Hydrogenases and hydrogen photoproduction in oxygenic photosynthetic organisms. Annu Rev Plant Biol 58:71–91
Ghirardi ML, Dubini A, Yu J, Maness PC (2009) Photobiological hydrogen-producing systems. Chem Soc Rev 38:52–61
Girbal L, von Abendroth G, Winkler M, Benton PM, Meynial-Salles I, Croux C, Peters JW, Happe T, Soucaille P (2005) Homologous and heterologous overexpression in Clostridium acetobutylicum and characterization of purified clostridial and algal Fe-only hydrogenases with high specific activities. Appl Environ Microbiol 71:2777–2781
Goldet G, Wait AF, Cracknell JA, Vincent KA, Ludwig M, Lenz O, Friedrich B, Armstrong FA (2008) Hydrogen production under aerobic conditions by membrane-bound hydrogenases from Ralstonia species. J Am Chem Soc 130:11106–11113
Goris T, Wait AF, Saggu M, Fritsch J, Heidary N, Stein M, Zebger I, Lendzian F, Armstrong FA, Friedrich B, Lenz O (2011) A unique iron-sulfur cluster is crucial for oxygen tolerance of a [NiFe]-hydrogenase. Nat Chem Biol 7:310–318
Grzeszik C, Lubbers M, Reh M, Schlegel HG (1997) Genes encoding the NAD-reducing hydrogenase of Rhodococcus opacus MR11. Microbiology 143:1271–1286
Guiral M, Tron P, Belle V, Aubert C, Léger C, Guigliarelli B, Giudici-Orticoni M-T (2006) Hyperthermostable and oxygen resistant hydrogenases from a hyperthermophilic bacterium Aquifex aeolicus: physicochemical properties. Int J Hydrog Energy 31:1424–1431
Häckel M, Hinz H-J, Hedwig GR (1999) Partial molar volumes of proteins: amino acid side-chain contributions derived from the partial molar volumes of some tripeptides over the temperature range 10–90 °C. Biophys Chem 82:35–50
Hallenbeck PC, Abo-Hashesh M, Ghosh D (2012) Strategies for improving biological hydrogen production. Bioresour Technol 110:1–9
Happe RP, Roseboom W, Egert G, Friedrich CG, Massanz C, Friedrich B, Albracht SP (2000) Unusual FTIR and EPR properties of the H2-activating site of the cytoplasmic NAD-reducing hydrogenase from Ralstonia eutropha. FEBS Lett 466:259–263
Henstra AM, Sipma J, Rinzema A, Stams AJM (2007) Microbiology of synthesis gas fermentation for biofuel production. Curr Opin Biotechnol 18:200–206
Higuchi Y, Yagi T, Yasuoka N (1997) Unusual ligand structure in Ni-Fe active center and an additional Mg site in hydrogenase revealed by high resolution X-ray structure analysis. Structure 5:1671–1680
Hiromoto T, Ataka K, Pilak O, Vogt S, Stagni MS, Meyer-Klaucke W, Warkentin E, Thauer RK, Shima S, Ermler U (2009) The crystal structure of C176A mutated [Fe]-hydrogenase suggests an acyl-iron ligation in the active site iron complex. FEBS Lett 583:585–590
Hong G, Pachter R (2012) Inhibition of biocatalysis in [FeFe] hydrogenase by oxygen: molecular dynamics and density functional theory calculations. ACS Chem Biol 7:1268–1275
Horch M, Lauterbach L, Saggu M, Hildebrandt P, Lendzian F, Bittl R, Lenz O, Zebger I (2010) Probing the active site of an O2-tolerant NAD+−reducing [NiFe]-hydrogenase from Ralstonia eutropha H16 by in situ EPR and FTIR spectroscopy. Angew Chem Int Ed Engl 49:8026–8029
Horch M, Lauterbach L, Lenz O, Hildebrandt P, Zebger I (2012) NAD(H)-coupled hydrogen cycling-structure-function relationships of bidirectional [NiFe] hydrogenases. FEBS Lett 586:545–556
Horner DS, Foster PG, Embley TM (2000) Iron hydrogenases and the evolution of anaerobic eukaryotes. Mol Biol Evol 17:1695–1709
Horner DS, Heil B, Happe T, Embley TM (2002) Iron hydrogenases–ancient enzymes in modern eukaryotes. Trends Biochem Sci 27:148–153
Ihara M, Nishihara H, Yoon KS, Lenz O, Friedrich B, Nakamoto H, Kojima K, Honma D, Kamachi T, Okura I (2006) Light-driven hydrogen production by a hybrid complex of a [NiFe]-hydrogenase and the cyanobacterial photosystem I. Photochem Photobiol 82:676–682
Ikeda T, Kano K (2001) An electrochemical approach to the studies of biological redox reactions and their applications to biosensors, bioreactors, and biofuel cells. J Biosci Bioeng 92:9–18
Jacobi A, Rossmann R, Bock A (1992) The hyp operon gene products are required for the maturation of catalytically active hydrogenase isoenzymes in Escherichia coli. Arch Microbiol 158:444–451
Jo JH, Lee DS, Kim J, Park JM (2009) Effect of initial glucose concentrations on carbon material and energy balances in hydrogen-producing Clostridium tyrobutyricum JM1. J Microbiol Biotechnol 19:291–298
Kamp C, Silakov A, Winkler M, Reijerse EJ, Lubitz W, Happe T (2008) Isolation and first EPR characterization of the [FeFe]-hydrogenases from green algae. Biochim Biophys Acta 1777:410–416
Kaneko T, Nakamura Y, Sato S, Minamisawa K, Uchiumi T, Sasamoto S, Watanabe A, Idesawa K, Iriguchi M, Kawashima K, Kohara M, Matsumoto M, Shimpo S, Tsuruoka H, Wada T, Yamada M, Tabata S (2002) Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res 9:189–197
Kim YH, Berry AH, Spencer DS, Stites WE (2001) Comparing the effect on protein stability of methionine oxidation versus mutagenesis: steps toward engineering oxidative resistance in proteins. Protein Eng 14:343–347
Kim JY, Jo BH, Cha HJ (2011) Production of biohydrogen by heterologous expression of oxygen-tolerant Hydrogenovibrio marinus [NiFe]-hydrogenase in Escherichia coli. J Biotechnol 155(3):312–319
Kim S, Lu D, Park S, Wang G (2012) Production of hydrogenases as biocatalysts. Int J Hydrogen Energy. http://dx.doi.org/10.1016/j.ijhydene.2012.03.033
King PW, Posewitz MC, Ghirardi ML, Seibert M (2006) Functional studies of [FeFe] hydrogenase maturation in an Escherichia coli biosynthetic system. J Bacteriol 188:2163–2172
Kontur WS, Noguera DR, Donohue TJ (2012) Maximizing reductant flow into microbial H2 production. Curr Opin Biotechnol 23:382–389
Krassen H, Schwarze A, Friedrich B, Ataka K, Lenz O, Heberle J (2009) Photosynthetic hydrogen production by a hybrid complex of photosystem I and [NiFe]-hydrogenase. ACS Nano 3:4055–4061
Kruse O, Rupprecht J, Bader KP, Thomas-Hall S, Schenk PM, Finazzi G, Hankamer B (2005) Improved photobiological H2 production in engineered green algal cells. J Biol Chem 280:34170–34177
Kuchenreuther JM, Grady-Smith CS, Bingham AS, George SJ, Cramer SP, Swartz JR (2010) High-yield expression of heterologous [FeFe] hydrogenases in Escherichia coli. PLoS One 5:e15491
Lamle SE, Albracht SP, Armstrong FA (2004) Electrochemical potential-step investigations of the aerobic interconversions of [NiFe]-hydrogenase from Allochromatium vinosum: insights into the puzzling difference between unready and ready oxidized inactive states. J Am Chem Soc 126:14899–14909
Lamle SE, Albracht SP, Armstrong FA (2005) The mechanism of activation of a [NiFe]-hydrogenase by electrons, hydrogen, and carbon monoxide. J Am Chem Soc 127:6595–6604
Leach MR, Zhang JW, Zamble DB (2007) The role of complex formation between the Escherichia coli hydrogenase accessory factors HypB and SlyD. J Biol Chem 282:16177–16186
Léger C, Dementin S, Bertrand P, Rousset M, Guigliarelli B (2004) Inhibition and aerobic inactivation kinetics of Desulfovibrio fructosovorans [NiFe] hydrogenase studied by protein film voltammetry. J Am Chem Soc 126:12162–12172
Lenz O, Gleiche A, Strack A, Friedrich B (2005) Requirements for heterologous production of a complex metalloenzyme: the membrane-bound [NiFe] hydrogenase. J Bacteriol 187:6590–6595
Lenz O, Ludwig M, Schubert T, Bürstel I, Ganskow S, Goris T, Schwarze A, Friedrich B (2010) H2 conversion in the presence of O2 as performed by the membrane-bound [NiFe]-hydrogenase of Ralstonia eutropha. ChemPhysChem 11:1107–1119
Leroux F, Dementin S, Burlat B et al (2008) Experimental approaches to kinetics of gas diffusion in hydrogenase. Proc Natl Acad Sci U S A 105:11188–11193
Liang Y, Wu X, Gan L, Xu H, Hu Z, Long M (2009) Increased biological hydrogen production by deletion of hydrogen-uptake system in photosynthetic bacteria. Microbiol Res 164:674–679
Liebgott PP, Leroux F, Burlat B, Dementin S, Baffert C, Lautier T, Fourmond V, Ceccaldi P, Cavazza C, Meynial-Salles I, Soucaille P, Fontecilla-Camps JC, Guigliarelli B, Bertrand P, Rousset M, Léger C (2010) Relating diffusion along the substrate tunnel and oxygen sensitivity in hydrogenase. Nat Chem Biol 6:63–70
Liebgott PP, De Lacey AL, Burlat B, Cournac L, Richaud P, Brugna M, Fernandez VM, Guigliarelli B, Rousset M, Léger C, Dementin S (2011) Original design of an oxygen-tolerant [NiFe] hydrogenase: major effect of a valine-to-cysteine mutation near the active site. J Am Chem Soc 133:986–997
Liu ZP, Hu P (2002) A density functional theory study on the active center of Fe-only hydrogenase: characterization and electronic structure of the redox states. J Am Chem Soc 124:5175–5182
Liu X, Zhu Y, Yang ST (2006) Construction and characterization of ack deleted mutant of Clostridium tyrobutyricum for enhanced butyric acid and hydrogen production. Biotechnol Prog 22:1265–1275
Lubitz W, Reijerse E, van Gastel M (2007) [NiFe] and [FeFe] hydrogenases studied by advanced magnetic resonance techniques. Chem Rev 107:4331–4365
Lubner CE, Knoïzer P, Silva PJN, Vincent KA, Happe T, Bryant DA, Golbeck JH (2010) Wiring an [FeFe]-hydrogenase with photosystem I for light-induced hydrogen production. Biochemistry 49:10264–10266
Ludwig M, Cracknell JA, Vincent KA, Armstrong FA, Lenz O (2009a) Oxygen-tolerant H2 oxidation by membrane-bound [NiFe] hydrogenases of Ralstonia species. Coping with low level H2 in air. J Biol Chem 284:465–477
Ludwig M, Schubert T, Zebger I, Wisitruangsakul N, Saggu M, Strack A, Lenz O, Hildebrandt P, Friedrich B (2009b) Concerted action of two novel auxiliary proteins in assembly of the active site in a membrane-bound [NiFe] hydrogenase. J Biol Chem 284:2159–2168
Lukey MJ, Parkin A, Roessler MM, Murphy BJ, Harmer J, Palmer T, Sargent F, Armstrong FA (2010) How Escherichia coli is equipped to oxidize hydrogen under different redox conditions. J Biol Chem 285:3928–3938
Lukey MJ, Roessler MM, Parkin A, Evans RM, Davies RA, Lenz O, Friedrich B, Sargent F, Armstrong FA (2011) Oxygen-tolerant [NiFe]-hydrogenases: the individual and collective importance of supernumerary cysteines at the proximal Fe-S cluster. J Am Chem Soc 133:16881–16892
Lyon EJ, Shima S, Buurman G, Chowdhuri S, Batschauer A, Steinbach K, Thauer RK (2004) UV-A/blue-light inactivation of the ‘metal-free’ hydrogenase (Hmd) from methanogenic archaea. Eur J Biochem 271:195–204
Maeda T, Sanchez-Torres V, Wood TK (2007) Enhanced hydrogen production from glucose by metabolically engineered Escherichia coli. Appl Microbiol Biotechnol 77:879–890
Maier T, Binder U, Bock A (1996) Analysis of the hydA locus of Escherichia coli: two genes (hydN and hypF) involved in formate and hydrogen metabolism. Arch Microbiol 165:333–341
Manyani H, Rey L, Palacios JM, Imperial J, Ruiz-Argueso T (2005) Gene products of the hupGHIJ operon are involved in maturation of the iron-sulfur subunit of the [NiFe] hydrogenase from Rhizobium leguminosarum bv. viciae. J Bacteriol 187:7018–7026
Maroti G, Fodor BD, Rakhely G, Kovacs AT, Arvani S, Kovacs KL (2003) Accessory proteins functioning selectively and pleiotropically in the biosynthesis of [NiFe] hydrogenases in Thiocapsa roseopersicina. Eur J Biochem 270:2218–2227
Maroti G, Tong Y, Yooseph S, Baden-Tillson H, Smith HO, Kovàcs KL, Frazier M, Venter JC, Xu Q (2009) Discovery of [NiFe] hydrogenase genes in metagenomic DNA: cloning and heterologous expression in Thiocapsa roseopersicina. Appl Environ Microbiol 75:5821–5830
Mason JE (2007) World energy analysis: H2 now or later? Energy Policy 35:1315–1329
Matias PM, Soares CM, Saraiva LM, Coelho R, Morais J, Le Gall J, Carrondo MA (2001) [NiFe] hydrogenase from Desulfovibrio desulfuricans ATCC 27774: gene sequencing, three-dimensional structure determination and refinement at 1.8 A and modelling studies of its interaction with the tetrahaem cytochrome c3. J Biol Inorg Chem 6:63–81
McKinlay JB, Harwood CS (2010) Photobiological production of hydrogen gas as a biofuel. Curr Opin Biotechnol 21:244–251
Menon NK, Chatelus CY, Dervartanian M, Wendt JC, Shanmugam KT, Peck HD Jr, Przybyla AE (1994) Cloning, sequencing, and mutational analysis of the hyb operon encoding Escherichia coli hydrogenase 2. J Bacteriol 176:4416–4423
Meyer J (2007) [FeFe] hydrogenases and their evolution: a genomic perspective. Cell Mol Life Sci 64:1063–1084
Montet Y, Amara P, Volbeda A, Vernede X, Hatchikian EC, Field MJ, Frey M, Fontecilla-Camps JC (1997) Gas access to the active site of Ni-Fe hydrogenases probed by X-ray crystallography and molecular dynamics. Nat Struct Biol 4:523–526
Morimoto K, Kimura T, Sakka K, Ohmiya K (2005) Overexpression of a hydrogenase gene in Clostridium paraputrificum to enhance hydrogen gas production. FEMS Microbiol Lett 246:229–234
Morozov SV, Karyakina EE, Zorin NA, Varfolomeyev SD, Cosnier S, Karyakin AA (2002) Direct and electrically wired bioelectrocatalysis by hydrogenase from Thiocapsa roseopersicina. Bioelectrochemistry 55:169–171
Mulder DW, Shepard EM, Meuser JE, Joshi N, King PW, Posewitz MC, Broderick JB, Peters JW (2011) Insights into [FeFe]-hydrogenase structure, mechanism, and maturation. Structure 19:1038–1052
Nicolet Y, Fontecilla-Camps JC (2012) Structure-function relationships in [FeFe]-hydrogenase active site maturation. J Biol Chem 287:13532–13540
Nicolet Y, Piras C, Legrand P, Hatchikian CE, Fontecilla-Camps JC (1999) Desulfovibrio desulfuricans iron hydrogenase: the structure shows unusual coordination to an active site Fe binuclear center. Structure 7:13–23
Nicolet Y, Lemon BJ, Fontecilla-Camps JC, Peters JW (2000) A novel FeS cluster in Fe-only hydrogenases. Trends Biochem Sci 25:138–143
Nicolet Y, Cavazza C, Fontecilla-Camps JC (2002) Fe-only hydrogenases: structure, function and evolution. J Inorg Biochem 91:1–8
Nienhaus K, Deng P, Kriegl JM, Nienhaus GU (2003) Structural dynamics of myoglobin: effect of internal cavities on ligand migration and binding Biochem 42:9647–9658
Ogata H, Lubitz W, Higuchi Y (2009) [NiFe] hydrogenases: structural and spectroscopic studies of the reaction mechanism. Dalton Trans 37:7577–7587
Ogata H, Kellers P, Lubitz W (2010) The crystal structure of the [NiFe] hydrogenase from the photosynthetic bacterium Allochromatium vinosum: characterization of the oxidized enzyme (Ni-A State). J Mol Biol 402:428–444
Oh Y-K, Raj SM, Jung GY, Park S (2011) Current status of the metabolic engineering of microorganisms for biohydrogen production. Bioresour Technol 102:8357–8367
Okura I, Otsuka K, Nakada N, Hasumi F (1990) Regeneration of NADH and ketone hydrogenation by hydrogen with the combination of hydrogenase and alcohol dehydrogenase. Appl Biochem Biotechnol 24–25:425–430
Pandelia M-E, Fourmond V, Tron-Infossi P, Lojou E, Bertrand P, Léger C, Giudici-Orticoni M-T, Lubitz W (2010a) Membrane-bound hydrogenase I from the hyperthermophilic bacterium Aquifex aeolicus: enzyme activation, redox intermediates and oxygen tolerance. J Am Chem Soc 132:6991–7004
Pandelia M-E, Ogata H, Lubitz W (2010b) Intermediates in the catalytic cycle of [NiFe] hydrogenase: functional spectroscopy of the active site. ChemPhysChem 11:1127–1140
Pandelia M-E, Nitschke W, Infossi P, M-Trs G-O, Bill E, Lubitz W (2011) Characterization of a unique [FeS] cluster in the electron transfer chain of the oxygen tolerant [NiFe] hydrogenase from Aquifex aeolicus. Proc Natl Acad Sci U S A 108:6097–6102
Pardo A, De Lacey AL, Fernandez VM, Fan HJ, Fan Y, Hall MB (2006) Density functional study of the catalytic cycle of nickel-iron [NiFe] hydrogenases and the involvement of high-spin nickel(II). J Biol Inorg Chem 11:286–306
Parish D, Benach J, Liu G, Singarapu KK, Xiao R, Acton T, Su M, Bansal S, Prestegard JH, Hunt J, Montelione GT, Szyperski T (2008) Protein chaperones Q8ZP25_SALTY from Salmonella typhimurium and HYAE_ECOLI from Escherichia coli exhibit thioredoxin-like structures despite lack of canonical thioredoxin active site sequence motif. J Struct Funct Genomics 9:41–49
Parkin A, Sargent F (2012) The how and whys of aerobic hydrogen metabolism. Curr Opin Chem Biol 16:26–34
Paschos A, Bauer A, Zimmermann A, Zehelein E, Bock A (2002) HypF, a carbamoyl phosphate-converting enzyme involved in [NiFe] hydrogenase maturation. J Biol Chem 277:49945–49951
Perez-Arellano JL, Martin T, Lopez-Novoa JM, Sanchez ML, Montero A, Jimenez A (1998) BN 52021 (a platelet activating factor-receptor antagonist) decreases alveolar macrophage-mediated lung injury in experimental extrinsic allergic alveolitis. Mediators Inflamm 7:201–210
Peters JW, Lanzilotta WN, Lemon BJ, Seefeldt LC (1998) X-ray crystal structure of the Fe-only hydrogenase (CpI) from Clostridium pasteurianum to 1.8 angstrom resolution. Science 282:1853–1858
Petkun S, Shi R, Li Y, Asinas A, Munger C, Zhang L, Waclawek M, Soboh B, Sawers RG, Cygler M (2011) Structure of hydrogenase maturation protein HypF with reaction intermediates shows two active sites. Structure 19:1773–1783
Pierik AJ, Roseboom W, Happe RP, Bagley KA, Albracht SP (1999) Carbon monoxide and cyanide as intrinsic ligands to iron in the active site of [NiFe]-hydrogenases. NiFe(CN)2CO, biology’s way to activate H2. J Biol Chem 274:3331–3337
Pohlmann A, Fricke WF, Reinecke F, Kusian B, Liesegang H, Cramm R, Eitinger T, Ewering C, Pötter M, Schwartz E, Strittmatter A, Voss I, Gottschalk G, Steinbüchel A, Friedrich B, Bowien B (2006) Genome sequence of the bioplastic-producing Knallgas bacterium Ralstonia eutropha H16. Nat Biotechnol 24:1257–1262
Porthun A, Bernhard M, Friedrich B (2002) Expression of a functional NAD-reducing [NiFe] hydrogenase from the gram-positive Rhodococcus opacus in the gram-negative Ralstonia eutropha. Arch Microbiol 177:159–166
Posewitz MC, Smolinski SL, Kanakagiri S, Melis A, Seibert M, Ghirardi ML (2004) Hydrogen photoproduction is attenuated by disruption of an isoamylase gene in Chlamydomonas reinhardtii. Plant Cell 16:2151–2163
Prince RC, Kheshgi HS (2005) The photobiological production of hydrogen: potential efficiency and effectiveness as a renewable fuel. Crit Rev Microbiol 31:19–31
Ratzka J, Lauterbach L, Lenz O, Ansorge-Schumacher MB (2011) Systematic evaluation of the dihydrogen-oxidising and NAD+-reducing soluble [NiFe]-hydrogenase from Ralstonia eutropha H16 as a cofactor regeneration catalyst. Biocatal Biotransfor 29:246–252
Reddie KG, Carroll KS (2008) Expanding the functional diversity of proteins through cysteine oxidation. Curr Opin Chem Biol 12:746–754
Richardson AJ, Stewart CS (1990) Hydrogen transfer between Neocallimastix frontalis and Selenomonas ruminantium grown in mixed culture. In: Bélaich JP, Bruschi M, Garcia JL (eds) Microbiology and biochemistry of strict anaerobes involved in interspecies hydrogen transfer. Plenum Publishing Corporation, New York, pp 463–466
Rousset M, Montet Y, Guigliarelli B, Forget N, Asso M, Bertrand P, Fontecilla-Camps JC, Hatchikian EC (1998) [3Fe-4S] to [4Fe-4S] cluster conversion in Desulfovibrio fructosovorans [NiFe] hydrogenase by site-directed mutagenesis. Proc Natl Acad Sci U S A 95:11625–11630
Rupprecht J, Hankamer B, Mussgnug JH, Ananyev G, Dismukes C, Kruse O (2006) Perspectives and advances of biological H2 production in microorganisms. Appl Microbiol Biotechnol 72:442–449
Sauter M, Bohm R, Bock A (1992) Mutational analysis of the operon (hyc) determining hydrogenase 3 formation in Escherichia coli. Mol Microbiol 6:1523–1532
Schmitz O, Boison G, Salzmann H, Bothe H, Schutz K, Wang SH, Happe T (2002) HoxE–a subunit specific for the pentameric bidirectional hydrogenase complex (HoxEFUYH) of cyanobacteria. Biochim Biophys Acta 1554:66–74
Schubert T, Lenz O, Krause E, Volkmer R, Friedrich B (2007) Chaperones specific for the membrane-bound [NiFe]-hydrogenase interact with the Tat signal peptide of the small subunit precursor in Ralstonia eutropha H16. Mol Microbiol 66:453–467
Schwartz E, Henne A, Cramm R, Eitinger T, Friedrich B, Gottschalk G (2003) Complete nucleotide sequence of pHG1: a Ralstonia eutropha H16 megaplasmid encoding key enzymes of H(2)-based lithoautotrophy and anaerobiosis. J Mol Biol 332:369–383
Self WT, Hasona A, Shanmugam KT (2004) Expression and regulation of a silent operon, hyf, coding for hydrogenase 4 isoenzyme in Escherichia coli. J Bacteriol 186:580–587
Shima S (2008) Structure of [Fe]-hydrogenase and the convergent evolution of the active site of hydrogenases. Seikagaku 80:846–849
Shomura Y, Yoon K-S, Nishihara H, Higuchi Y (2011) Structural basis for a [4Fe-3S] cluster in the oxygen-tolerant membrane-bound [NiFe]-hydrogenase. Nature 479:253–256
Silakov A, Wenk B, Reijerse E, Albracht SP, Lubitz W (2009) Spin distribution of the H-cluster in the H(ox)-CO state of the [FeFe] hydrogenase from Desulfovibrio desulfuricans: HYSCORE and ENDOR study of (14)N and (13)C nuclear interactions. J Biol Inorg Chem 14:301–313
Soboh B, Kuhns M, Braussemann M, Waclawek M, Muhr E, Pierik AJ, Sawers RG (2012) Evidence for an oxygen-sensitive iron-sulfur cluster in an immature large subunit species of Escherichia coli [NiFe]-hydrogenase 2. Biochem Biophys Res Commun 424:158–163
Stadler C, de Lacey AL, Montet Y, Volbeda A, Fontecilla-Camps JC, Conesa JC, Fernandez VM (2002) Density functional calculations for modeling the active site of nickel-iron hydrogenases. 2. Predictions for the unready and ready states and the corresponding activation processes. Inorg Chem 41:4424–4434
Stadtman ER (2004) Cyclic oxidation and reduction of methionine residues of proteins in antioxidant defense and cellular regulation. Arch Biochem Biophys 423:2–5
Stadtman ER (2006) Protein oxidation and aging. Free Radic Res 40:1250–1258
Stadtman ER, Moskovitz J, Berlett BS, Levine RL (2002) Cyclic oxidation and reduction of protein methionine residues is an important antioxidant mechanism. Mol Cell Biochem 234–235:3–9
Stein M, van Lenthe E, Baerends EJ, Lubitz W (2001) Relativistic DFT calculations of the paramagnetic intermediates of [NiFe] hydrogenase. Implications for the enzymatic mechanism. J Am Chem Soc 123:5839–5840
Stripp ST, Goldet G, Brandmayr C, Sanganas O, Vincent KA, Haumann M, Armstrong FA, Happe T (2009) How oxygen attacks [FeFe] hydrogenases from photosynthetic organisms. Proc Natl Acad Sci U S A 106:17331–17336
Strnad H, Lapidus A, Paces J, Ulbrich P, Vlcek C, Paces V, Haselkorn R (2010) Complete genome sequence of the photosynthetic purple nonsulfur bacterium Rhodobacter capsulatus SB 1003. J Bacteriol 192:3545–3546
Sun J, Hopkins RC, Jenney FE Jr, McTernan PM, Adams MWW (2010) Heterologous expression and maturation of an NADP-dependent [NiFe]-hydrogenase: a key enzyme in biofuel production. PLoS One 5:e10526
Sybirna K, Antoine T, Lindberg P, Fourmond V, Rousset M, Mejean V, Bottin H (2008) Shewanella oneidensis: a new and efficient system for expression and maturation of heterologous [Fe-Fe] hydrogenase from Chlamydomonas reinhardtii. BMC Biotechnol 8:73
Teixeira M, Fauque G, Moura I, Lespinat PA, Berlier Y, Prickril B, Peck HD Jr, Xavier AV, Le Gall J, Moura JJ (1987) Nickel-[iron-sulfur]-selenium-containing hydrogenases from Desulfovibrio baculatus (DSM 1743). Redox centers and catalytic properties. Eur J Biochem 167:47–58
Teixeira VH, Baptista AM, Soares CM (2006) Pathways of H2 toward the active site of [NiFe]-hydrogenase. Biophys J 91:2035–2045
Topin J, Rousset M, Antonczak S, Golebiowski J (2012) Kinetics and thermodynamics of gas diffusion in a NiFe hydrogenase. Proteins 80:677–682
Tremblay P-L, Lovley DR (2012) Role of the NiFe hydrogenase Hya in oxidative stress defense in Geobacter sulfurreducens. J Bacteriol 194:2248–2253
van der Zwaan JW, Coremans JM, Bouwens EC, Albracht SP (1990) Effect of 17O2 and 13CO on EPR spectra of nickel in hydrogenase from Chromatium vinosum. Biochim Biophys Acta 1041:101–110
van Gastel M, Stein M, Brecht M, Schroder O, Lendzian F, Bittl R, Ogata H, Higuchi Y, Lubitz W (2006) A single-crystal ENDOR and density functional theory study of the oxidized states of the [NiFe] hydrogenase from Desulfovibrio vulgaris Miyazaki F. J Biol Inorg Chem 11:41–51
van Gastel M, Shaw JL, Blake AJ, Flores M, Schroder M, McMaster J, Lubitz W (2008) Electronic structure of a binuclear nickel complex of relevance to [NiFe] hydrogenase. Inorg Chem 47:11688–11697
Vargas WA, Weyman PD, Tong Y, Smith HO, Xu Q (2011) [NiFe] hydrogenase from Alteromonas macleodii with unusual stability in the presence of oxygen and high temperature. Appl Environ Microbiol 77:1990–1998
Vignais PM, Billoud B (2007) Occurrence, classification, and biological function of hydrogenases: an overview. Chem Rev 107:4206–4272
Vignais PM, Billoud B, Meyer J (2001) Classification and phylogeny of hydrogenases. FEMS Microbiol Rev 25:455–501
Vincent KA, Parkin A, Armstrong FA (2007) Investigating and exploiting the electrocatalytic properties of hydrogenases. Chem Rev 107:4366–4413
Volbeda A, Fontecilla-Camps JC (2004) Crystallographic evidence for a CO/CO(2) tunnel gating mechanism in the bifunctional carbon monoxide dehydrogenase/acetyl coenzyme A synthase from Moorella thermoacetica. J Biol Inorg Chem 9:525–532
Volbeda A, Charon MH, Piras C, Hatchikian EC, Frey M, Fontecilla-Camps JC (1995) Crystal structure of the nickel-iron hydrogenase from Desulfovibrio gigas. Nature 373:580–587
Volbeda A, Garcin E, Piras C, De Lacey AL, Fernandez VM, Hatchikian CE, Frey M, Fontecilla-Camps JC (1996) Structure of the [NiFe] hydrogenase active site: evidence for biologically uncommon Fe ligands. J Am Chem Soc 118:12989–12996
Volbeda A, Montet Y, Vernede X, Hatchikian CE, Fontecilla-Camps JC (2002) High-resolution crystallographic analysis of Desulfovibrio fructosovorans [NiFe] hydrogenase. Int J Hydrog Energy 27:1449–1461
Volbeda A, Martin L, Cavazza C, Matho M, Faber BW, Roseboom W, Albracht SP, Garcin E, Rousset M, Fontecilla-Camps JC (2005) Structural differences between the ready and unready oxidized states of [NiFe] hydrogenases. J Biol Inorg Chem 10:239–249
Volbeda A, Fontecilla-Camps JC, Sazanov L (2012) The evolutionary relationship between complex I and [NiFe]-hydrogenase. In: Sazanov L (ed) A structural perspective on respiratory Complex I: structure and function of NADH:ubiquinone oxidoreductase. Springer, Dordrecht, pp 109–121
Voordouw G, Hagen WR, Kruse-Wolters KM, van Berkel-Arts A, Veeger C (1987) Purification and characterization of Desulfovibrio vulgaris (Hildenborough) hydrogenase expressed in Escherichia coli. Eur J Biochem 162:31–36
Wells MA, Mercer J, Mott RA, Pereira-Medrano AG, Burja AM, Radianingtyas H, Wright PC (2011) Engineering a non-native hydrogen production pathway into Escherichia coli via a cyanobacterial [NiFe] hydrogenase. Metab Eng 13:445–453
Weyman PD, Vargas WA, Tong Y, Yu J, Maness P-C, Smith HO, Xu Q (2011) Heterologous expression of Alteromonas macleodii and Thiocapsa roseopersicina [NiFe] hydrogenases in Synechococcus elongatus. PLoS One 6:e20126
Wolf I, Buhrke T, Dernedde J, Pohlmann A, Friedrich B (1998) Duplication of hyp genes involved in maturation of [NiFe] hydrogenases in Alcaligenes eutrophus H16. Arch Microbiol 170:451–459
Yacoby I, Pochekailov S, Toporik H, Ghirardi ML, King PW, Zhang S (2011) Photosynthetic electron partitioning between [FeFe]-hydrogenase and ferredoxin:NADP+-oxidoreductase (FNR) enzymes in vitro. Proc Natl Acad Sci U S A 108:9396–9401
Yacoby I, Tegler LT, Pochekailov S, Zhang S, King PW (2012) Optimized expression and purification for high-activity preparations of algal [FeFe]-hydrogenase. PLoS One 7:e35886
Zhao J, Song W, Cheng J, Zhang C (2010) Cloning of Enterobacter aerogenes fh1A gene and overexpression of hydrogen production. Wei Sheng Wu Xue Bao 50:736–742
Zheng H, Zhang C, Lu Y, Jiang P-X, Xing X-H (2012) Alteration of anaerobic metabolism in Escherichia coli for enhanced hydrogen production by heterologous expression of hydrogenase genes originating from Synechocystis sp. Biochem Eng J 60:81–86
Zirngibl C, Van Dongen W, Schworer B, Von Bunau R, Richter M, Klein A, Thauer RK (1992) H2-forming methylenetetrahydromethanopterin dehydrogenase, a novel type of hydrogenase without iron-sulfur clusters in methanogenic archaea. Eur J Biochem 208:511–520
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Rousset, M., Liebgott, PP. (2014). Engineering Hydrogenases for H2 Production: Bolts and Goals. In: Zannoni, D., De Philippis, R. (eds) Microbial BioEnergy: Hydrogen Production. Advances in Photosynthesis and Respiration, vol 38. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-8554-9_3
Download citation
DOI: https://doi.org/10.1007/978-94-017-8554-9_3
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-8553-2
Online ISBN: 978-94-017-8554-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)