Abstract
Sequence-specific transcription factors (TFs) play a central role in regulating transcription initiation by directing the recruitment and activity of the general transcription machinery and accessory factors. It is now well established that many of the effects exerted by TFs in eukaryotes are mediated through interactions with a host of coregulators that modify the chromatin state, resulting in a more open (in case of activation) or closed conformation (in case of repression). The relationship between TFs and chromatin is a two-way street, however, as chromatin can in turn influence the recognition and binding of target sequences by TFs. The aim of this chapter is to highlight how this dynamic interplay between TF-directed remodelling of chromatin and chromatin-adjusted targeting of TF binding determines where and how transcription is initiated, and to what degree it is productive.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
11.1 Introduction
The basic principles of transcriptional regulation are similar between prokaryotes and eukaryotes and involve the binding of TFs to specific DNA sequences at target genes, where they recruit and stabilize the general transcriptional machinery required for gene expression [1, 2]. Despite these general similarities, transcription initiation in eukaryotes is considerably more complex, which is likely related to the increased genome size and greater need for organization compared to prokaryotes. One key difference is that DNA in eukaryotes is not readily accessible, but tightly packaged by architectural proteins into chromatin. The basic unit of this packaging is the nucleosome, which consists of ~147 bp of DNA wrapped around an octamer of histone proteins [3, 4]. Nucleosomes play an important role in condensing DNA, thereby allowing the large eukaryotic genome to fit into the nucleus. Perhaps not surprisingly, this compaction also negatively affects transcription initiation in vitro [5, 6] and in vivo [7], as it forms an impediment to the binding of TFs and the formation of a preinitiation complex (PIC) [8, 9]. To initiate transcription, TFs and the PIC must first overcome the physical barrier posed by nucleosomes; however, the stability of nucleosomes means that direct competition for DNA access is inefficient. A host of coactivators therefore exist that can be recruited to regulatory regions by TFs to facilitate transcription initiation. These coactivators typically consist of (or recruit) chromatin modifier (CM) complexes that either displace or evict nucleosomes or covalently modify histones to loosen their interactions with DNA. CMs can also function as corepressors by effecting a more closed chromatin conformation. Consequently, the recruitment of coregulators that affect chromatin structure is now recognized as a major mechanism by which TFs can regulate gene expression.
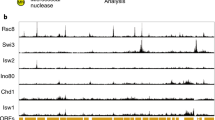
Knowledge of general chromatin architecture has greatly expanded in recent years due to the broad application of classical and novel techniques to map TF binding sites, histone modifications, and chromatin accessibility. Mapping of TF binding sites and histone modifications is typically done using chromatin immunoprecipitation (ChIP) or related techniques such as DamID, which are discussed in more detail in Chapter 8. Most of the techniques to map chromatin accessibility make use of the fact that regulatory sites and the short DNA linkers connecting nucleosomes are more sensitive to nuclease digestion by micrococcal nuclease (MNase) or DNase I, each of which has distinct cleavage patterns that provide a different view of chromatin structure [10]. MNase cuts preferentially in linker regions between nucleosome and it is therefore typically used to map the positions of nucleosomes. On the other hand, DNaseI also cuts DNA associated with nucleosomes, when used at higher concentrations, and its cleavage pattern therefore typifies general chromatin accessibility. Another approach to identify regions of open chromatin, formaldehyde-assisted isolation of regulatory elements (FAIRE), has also been described [11]. This method exploits the property that fragmented DNA that is highly crosslinked to histones after formaldehyde treatment (i.e. closed chromatin) can be separated from DNA with a low degree of crosslinking (i.e. open chromatin) by phenol extraction.
Advances in microarray and sequencing technology have made it possible to apply these various methods to create genome-wide maps of nucleosome occupancy [12–15], potential regulatory sites [16, 17], as well as patterns of histone modifications and TF binding [18–23]. A common observation in these studies is that active promoters and distal regulatory elements such as enhancers are associated with regions of open chromatin and enriched for bound TFs and their coregulators, underscoring that transcriptional regulation is universally linked to chromatin remodelling. These studies have also provided an unprecedented view of the higher-order structure of the genome, where broad domains of more accessible chromatin (i.e. euchromatin) alternate with regions that are less accessible to the transcription machinery (i.e. heterochromatin). It should be noted, though, that these techniques provide only a snapshot of the chromatin structure at the time of fixation and while many regulatory regions may appear stable, several lines of evidence suggest that remodelling is in fact a highly dynamic and continuously ongoing process. For example, nucleosomes found in yeast promoters exchange more rapidly than nucleosomes located in gene bodies [24, 25] and FRAP (fluorescence recovery after photobleaching) studies suggest that many TFs only transiently interact with DNA in vivo, even at active promoters [26–29]. Thus, chromosomal domains and regulatory regions with apparently stable chromatin are likely in a dynamic equilibrium between competing forces, the balance of which ultimately determines the degree of DNA accessibility [8].
Following a brief introduction into the types of CM involved in chromatin remodelling, this chapter will highlight how TFs can regulate gene expression by recruiting these coregulators to orchestrate changes in the chromatin state, and in turn, how chromatin can affect TF target recognition and binding. Then, I will discuss how these dynamic and antagonistic forces may be coordinated to organize chromatin and direct transcription at specific locations in the genome. Other recent reviews that consider these and related topics include [30–33], as well as Chapters 10 and 12 in this volume, which specifically consider TF–nucleosome interactions, and the auxiliary domains of TFs that mediate many of these functions, respectively. This chapter also contains a Glossary at the end which provides an overview of key terminology used throughout.
11.2 An Overview of Coregulators that Effect Changes in Chromatin Structure
A broad distinction can be made between two types of CMs, based on their mechanism of action: histone modifiers and ATPase nucleosome remodelling complexes. Histone modifiers are responsible for the wide variety of covalent modifications found on histone proteins, in particular on their unstructured N-terminal tails (Reviewed in [34, 35]). At least eight different types of histone modifications and their associated enzymes have been identified, with the number of distinctly modified residues currently standing at well over a hundred [34]. It has been proposed that combinations of these modifications constitute a “histone code” that is read by proteins that interact with specific modifications [36], allowing for an organized association of proteins with different stages of transcription. Indeed, the different modifications can serve as interaction sites for other coregulators, such as ATPase remodelers, that can direct further changes to chromatin structure (see examples below). The ultimate effect of histone modifications on chromatin structure – be it compacting or unwrapping – is therefore presumably to a large degree determined by the type of proteins that interact with them. Another way that histone modifications can affect chromatin structure is by changing the electrostatic properties of nucleosomes. For example, the acetylation of histone tails by histone acetyl transferases (HATs) neutralizes positive charges that would otherwise interact with negatively charged DNA [37], facilitating nucleosome unwrapping and mobility (Fig. 11.1a). It is unclear whether other modifications similarly affect chromatin through effects on the chemical properties of nucleosomes, but it has been suggested that phosphorylation may, like acetylation, reduce chromatin compaction through its effects on nucleosome charge [34].
Effects of chromatin modifiers on chromatin structure. a Acetylation of histone tails by histone acetyl transferases (HATs) results in a more open chromatin conformation. b Model for nucleosome sliding by ATPase remodelers based on studies of the ACF complex [273]. In this model, the ATPase remodeler draws in DNA from the linker region (bottom arrow), resulting in the formation of a small DNA loop at the nucleosome entry site, which then propagates over the nucleosome, resulting in a lateral displacement along the DNA. The illustration shows one possible effect of remodelling at regulatory regions, namely the exposure of TF binding sites that would otherwise be rendered inaccessible by nucleosomes
Genome-wide studies have revealed that the occurrence of most modifications is tightly coupled to the location and activity of genes and their regulatory regions, in a manner that reflects their effects on chromatin structure. For example, acetylation marks are predominantly found at the beginning of active genes in yeast [22, 38–41] and at promoters and CpG islands in higher eukaryotes [42–45], although activation has also been linked to decreased acetylation of lysine residue 16 on histone H4 (i.e. H4K16ac) [38, 46, 47]. In contrast, methylation patterns differ depending on the residue that is modified, and distinct methylation states can be associated with either repression or activation [31, 34]. Classical examples include H3K4me and H3K27me, which mark regions of active and silent chromatin, respectively. The difference between acetylation and methylation patterns is mirrored in the specificity of their enzymes: HATs typically act indiscriminantly on multiple histone residues [34], whereas methyltransferases are restricted to a single residue on one histone type [48]. Some effects of HATs on chromatin may also be mediated through other targets, as it has become increasingly clear that they can acetylate many non-histone proteins, including TFs [49–51]. For other modifications, the relation to the transcriptional state is less well characterized, but in general, phosphorylation appears to correlate with activation [52, 53], while sumoylation has been associated with repression [54, 55]. Ubiquitination, like methylation, can be associated with either transcriptional state [56–58]. Extensive crosstalk between modifications presumably contributes to these complex patterns. For example, phosphorylation of H3S10 can stimulate acetylation of H3K14 [59, 60] and inhibit H3K19 methylation [61], while repression by sumoylation may be directly related to the fact that it competes for the same residues as acetylation and ubiquitination.
The second class of CMs, ATPase remodelers, can directly affect the degree of chromatin packing by repositioning or sliding nucleosomes along the DNA (Reviewed in [62]) (Fig. 11.1b). The primary driving force behind this motion comes from a central catalytic subunit, which contains a conserved ATPase domain that provides the energy to move nucleosomes by rewinding the DNA around them. This process involves breaking and reforming most histone–DNA interactions, which likely explains the broad effects that remodelers can have on nucleosomal DNA accessibility [63, 64], nucleosome eviction [65–67] and histone exchange [68, 69]. Besides the ATPase domain, the catalytic subunits contain various additional domains that have been used to classify these remodelers into four major families: SWI/SNF, ISWI, CHD and INO80. Interestingly, with the exception of INO80 subunits, many of these additional domains mediate affinity to distinct histone modifications [70, 71], which are thought to confer different preferences for specifically modified chromatin structures to each family [72, 73]. SWI/SNF remodelers contain a bromodomain which binds acetylated histones [74], while the CHD family possesses chromodomains that can interact with methylated histone tails [75–78]. ISWI family proteins have a pair of SANT and SLIDE domains that are believed to form a module with affinity for unmodified histones [79], though it is as yet unclear to what degree this interaction may be affected by specific modifications.
The diversity of CMs is further increased through the association of the core catalytic subunits with different complements of additional proteins, which can vary even within families [62, 70, 80]. These accessory subunits can play a structural role, and can also contribute a variety of additional interaction domains and catalytic activities. Some complexes, such as NURD (nucleosome-remodelling and histone deacytelase), even combine ATPase remodeler and histone modifier activities [81]. As in the case of histone modifications and their associated enzymes, a broad classification can be made regarding the effects of the ATPase remodelers on gene expression. For example, recruitment of SWI/SNF complexes is predominantly associated with transcriptional activation, consistent with its preference for acetylated histones, while ISWI complexes typically function as repressors [82]. This distinction is by no means sharply defined, though, and most ATPase remodelers have been found to function as activators at some promoters and repressors at others. Thus the ultimate effect of remodelling can vary depending on the context in which this remodelling takes place.
11.3 TFs Play a Central Role in Targeting Chromatin Remodelling
Exactly how chromatin remodelling complexes are guided to their target regions remains an active area of investigation. One clearly established pathway is direct recruitment by TFs, with TFs providing the targeting component through their sequence-specific DNA binding domains. This recruitment typically involves transient interactions with the transactivation or effector domains of TFs, which are discussed in more detail in Chapter 12 of this volume. The intrinsic preferences for specific histone modifications found in many CMs, discussed above, do indicate that there are also alternative routes that do not involve direct recruitment by TFs. For example, the bromodomains in the yeast Swi2/Snf2 remodelers and Gcn5 HAT are sufficient to anchor their respective complexes to acetylated promoters in the absence of transcriptional activators [74]. Individual histone binding domains may in general not be sufficient for effective targeting, however, given the low binding affinities of the domains characterized to date [62]. Instead, the interaction domains could serve other purposes that do not involve recruitment, such as regulating remodeler ATPase activity [62]. Regardless, even if histone modifications indeed provide important targeting cues for CMs, the question remains as to how these modifications are established in the first place, given that histone-modifying enzymes generally do not posses intrinsic DNA sequence preferences. One possible answer comes from detailed studies of model genes in yeast (Reviewed in [83]), which have shown that the actions of histone modifiers in the early stages of transcription initiation are primarily guided by sequence-specific TFs. It is therefore likely that TFs play a central role in targeting chromatin remodelling, whether this is through direct interactions with remodelling complexes, or by guiding initial histone modifications and/or other coregulators that mediate these interactions indirectly. An overview of some of the key features of TF-mediated recruitment of CMs and their implications for gene regulation will be given in the following paragraphs; readers are referred to Chapter 12 for more details.
Individual TFs can interact with a surprisingly wide variety of modifier complexes and other coregulators. This promiscuity is in part due to the intrinsic characteristics of the TF transactivation domains (also discussed in greater detail in Chapter 12), which are generally unstructured and only become stabilized upon interacting with their binding partners [84, 85] property that may allow for some degree of flexibility in the selection of binding partners [86]. The diversity of TF partners is also increased through interactions with subunits that are shared between different CM complexes. For example, acidic activation domains such as those found in the yeast Gal4 TF can recruit both the SAGA and NuA4 HATs through interactions with the Tra1 subunit that is present in both these complexes [87–89] (Fig. 11.2a). The great diversity of TF binding partners may serve multiple purposes. First, it enables the same TF to participate in distinct mechanisms of transcription initiation at different genes, as has been described for the activation of transcription by Pho2 and Pho4 at the PHO5 and PHO8 promoters in budding yeast [83]. Second, the transient nature of TF interactions at individual regulatory regions [26–29] could allow for repeated cycles of TF binding to the same target site with different coregulators, enabling a TF to affect initiation in more than one way. The particular coregulator(s) recruited at each site likely depends on other elements such as local chromatin structure and interactions with other TFs.
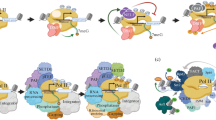
Targeting of chromatin remodelling by TFs. a The diversity of TF interactions with CMs is increased through shared subunits in remodeler complexes, as illustrated here by the interaction between the Gal4 TF and the Tra1 subunit in the SAGA and NuA4 complexes. b Targeting of the RSC complex in S. cerevisiae by the Rsc3 TF subunit. c CBP hub function at the IFN-β enhanceosome. CBP interacts with the enhanceosome TFs, resulting in recruitment of the RNA polymerase II holoenzyme, PIC assembly and the initiation of transcription [274]
In addition to mediating targeting through transient interactions, TFs can be integrated into CM complexes as stable components (Fig. 11.2b). The budding yeast TF Rsc3 is a subunit of the RSC chromatin remodelling complex [90], and was shown to promote nucleosome exclusion at promoters containing Rsc3 binding motifs [91], suggesting that it directs the RSC complex to these locations. Likewise, the Iec1 TF subunit of the INO80 complex is required for recruitment to target genes in fission yeast, and for associated histone remodelling [92]. Numerous putative DNA binding domains have also been identified in subunits of SWI/SNF remodelers in higher eukaryotes, including high mobility group (HMG) domains, C2H2 zinc fingers, and AT–rich interaction domains (ARIDs) [93]. The function of these domains is still largely uncharacterized and some, such as the HMG and ARID domains, are known to predominantly bind DNA in a sequence-independent manner and likely have structural roles [94, 95]. Nevertheless, it is possible that others will turn out to be important for targeting. Interestingly, the integration of sequence-specific TFs in remodelling complexes does not appear to be highly conserved between species. The RSC complex in higher eukaryotes lacks the specific DNA-binding determinants found in yeast [93, 96]; similarly, the INO80 component Iec1 is fungal-specific and has no ortholog in budding yeast. The stable integration of these particular TFs in remodelling complexes may therefore be the result of adaptations to specific selective pressures during evolution.
The multitude of subunits found in CMs means that they too can have many binding partners, greatly increasing their potential to regulate diverse targets. The subunit composition of complexes associated with each CM can also vary, such that different versions can pair with distinct sets of TFs. This enables individual complexes to be involved in gene- and cell type-specific functions, as exemplified by the mammalian SWI/SNF-type ATPases Brahma (BRM) and its paralog Brahma related gene 1 (BRG1), which are part of numerous chromatin remodelling complexes that target specific promoters to control gene expression [97]. BRG1 can be associated with WINAC (WSTF including nucleosome assembly complex), which can inhibit or activate target gene expression through subunit–specific interactions with the Vitamin D receptor [98]. Alternatively, when incorporated in the NUMAC (nucleosomal methylation activation) complex it can associate with estrogen receptor-responsive promoters to activate transcription [99]. Dynamic changes in CM subunit composition during development have also been shown to result in alterations in targeting by TFs. For example, the BRG1/BRM associated factors (BAFs) BAF45A and BAF53A in the SWI/SNF-type neuronal-progenitor-specific BAF complex (npBAF) are replaced by BAF45B and BAF53B upon differentiation, to form a neuron-specific complex (nBAF) [100]. The inclusion of BAF53B allows the nBAF complex to interact with the calcium-responsive transactivator (CREST) to regulate genes that are essential for dendritic outgrowth in the differentiated cells [101]. A similar requirement for specific BAF complex components has been observed in the differentiation of cardiomyocytes, where ectopic expression of the GATA4 and TBX5 TFs in combination with the BAF60C but not BAF60A subunits can induce the differentiation of mesoderm into contracting cardiomyocytes in developing mouse embryos [102]. Together, these observations indicate that TF binding can be interpreted differently in distinct cell types, depending on the complement of coregulators that is expressed. This modularity underscores the importance of combinatorial subunit assembly in establishing gene regulatory networks and reveals an additional layer of complexity that must be considered in our attempts to reconstruct these networks.
CM complexes can also be used as scaffolds for the assembly of different components of the transcriptional machinery. Indeed, the main catalytic function of CMs is sometimes dispensable altogether, as illustrated by the fact that SAGA-mediated activation of GAL genes does not require its HAT activity [103–105]. Instead, SAGA is believed to serve as a platform for the assembly of the PIC at GAL promoters. Similar functions have also been demonstrated for the general transcriptional coactivators CREB binding protein (CBP) and P300, two highly similar HATs with homologs in most multicellular organisms. In addition to the HAT domain, P300/CBP proteins contain other domains that mediate interactions with RNA polymerase II and a multitude of basal and gene-specific TFs [106, 107], allowing P300/CBP proteins to operate as hubs that can integrate signals from multiple TFs. This function has been most clearly described at the IFN-β enhanceosome, a stable complex of TFs and other nucleoproteins directly upstream of the IFN-β core promoter [108]. In this complex, CBP simultaneously interacts with multiple TFs bound across a 55 bp region, acting as a mediator for their synergistic activation of IFN-β transcription [108, 109] (Fig. 11.2c).
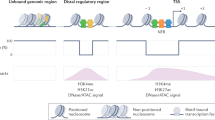
Consistent with their numerous interaction partners, P300/CBP have been linked to regulation of many genes, often acting at enhancers. Indeed, recent ChIP studies have identified P300/CBP binding as a key component of a wider signature of histone modifications and trans-acting factors that distinguish distal enhancers from gene promoters [20, 110–114]. Another component of this signature is H3K4 monomethylation, which peaks at enhancers but not promoters. Nevertheless, despite the predominance of P300/CBP at distal enhancers, both proteins can also be associated with proximal promoters and genes [115], underscoring their versatile roles in gene regulation.
11.4 Determinants of TF Access to Chromatin
A complicating factor for any model of chromatin remodelling based primarily on targeting by TFs is that they typically recognize small DNA motifs (~6–12 bp) that can occur randomly at high frequencies. For example, an 8-bp recognition sequence will appear 45,000 times in a human-sized genome with random sequence composition, and in reality this number will be dozens of times greater considering that TFs typically bind degenerate motifs in vitro [116]. Chromatin is believed to significantly increase TF specificity by reducing the accessibility of many spurious binding sites [117, 118]. This central role of chromatin in restricting where transcription initiation takes place is underscored by observations that failure to properly reconstitute nucleosomes in the body of transcribed yeast genes results in the appearance of cryptic transcripts, presumably initiated from exposed sequences that resemble promoters [119, 120]. Nonetheless, the packaging of DNA by nucleosomes is not the only means by which TF specificity is achieved in vivo. For example, TF–TF interactions, direct or indirect (e.g. through scaffold proteins or by outcompeting nucleosomes), can decrease the number of potential target sites due to the larger size of the combined binding specificity. Moreover, recognition sites are often clustered together in regulatory regions, allowing for further synergistic interactions between TFs [121–123]. A more in-depth overview of the various factors that play a role in TF target site selection can be found in Chapters 8 and 9.
The fact that nucleosomes can restrict access to DNA to prevent spurious transcription raises an important question: how can TFs bind their bona fide target sites to initiate the remodelling required for active transcription, given that much of the genome is covered by nucleosomes? Part of the answer to this question lies in the aforementioned fact that regulatory sites tend to be associated with open chromatin and nucleosome depleted regions (NDRs) [12–15, 124]. In yeast and C. elegans, there is strong evidence that the intrinsic DNA sequence preferences of nucleosomes play a key role in establishing these regions, and that these preferences are encoded in the genome sequence [125, 126]. Rigid DNA sequences such as poly(dA:dT) tracts are common in many eukaryotic promoters and have long been known to disrupt nucleosome–DNA interactions, increasing accessibility of nearby TF binding sequences [12, 127–130] (Fig. 11.3a). For example, the presence of a poly(dA:dT) tract in the Candida glabrata AFT1 promoter destabilizes a well-positioned nucleosome containing a metal responsive element, enabling Aft1 to bind and autoactivate its gene expression [131–133]. Poly(dA:dT) tracts were also found to be major determinants of nucleosome exclusion in studies aimed at predicting in vivo nucleosome positions from DNA sequence features in a range of species [12, 134, 135]. Perhaps the most direct indication of the importance of intrinsic nucleosome sequence preferences in the establishment of NDRs at promoters has come from comparisons of in vivo yeast nucleosome occupancy patterns to those of nucleosomes reconstituted in vitro on purified yeast genomic DNA, which showed a high correlation between the two profiles [125, 136]. The importance of nucleosome disfavouring sequences in establishing NDRs is now widely accepted, though there is still some debate about the degree in which intrinsic sequence preferences dictate nucleosome positions outside these regions [137–140].
Mechanisms of TF access to chromatin. a Rigid Poly(dA:dT) elements (red) are refractory to nucleosome assembly, allowing TFs to access nearby binding sites. b A model for progressive opening of chromatin by sequential binding of multiple TFs, as proposed by Polach and Widom [153]. c Presentation of the Gal4 UAS by RSC (Reproduced, with permission, from [63]). The model shows two exposed binding sites in the Gal4 UAS in the RSC/UASg/nucleosome complex and that Gal4 (red and purple) can access these sites without disrupting the complex. The structure of RSC/nucleosome complex (yellow) was determined by cryo-electronmicroscopy [275] and the position of the DNA helix is indicated in green and blue
Despite their general applicability, models based on intrinsic nucleosome sequence preferences alone cannot fully explain the architecture of promoters and other regulatory sequences observed in living cells, even in yeast. An assessment of the influence of a wide range of sequence features on in vivo nucleosome positioning in budding yeast revealed additional strong nucleosome excluding elements that corresponded to binding motifs of sequence-specific TFs such as Reb1 and Abf1 [12]. The role of these factors in establishing NDRs was confirmed in Reb1 and Abf1 loss-of-function mutants that showed greatly increased nucleosome occupancy at hundreds of promoters containing their binding motifs [91, 141]. Moreover, the in vitro reconstituted nucleosome occupancy at Abf1 and Reb1 binding sites was higher than that measured in vivo [125]. Taken together, these data clearly indicate that TFs are capable of establishing NDRs at yeast promoters that lack intrinsic nucleosome-disfavouring sequences. Correspondingly, the concept of a universally encoded open promoter structure does not appear to apply to all genes: a subset of yeast genes that display highly variable expression levels have increased nucleosome occupancy in their promoters, consistent with predictions based on intrinsic sequence preferences [142]. It was proposed that the positioning of nucleosomes in these promoters plays a key role in the variable regulation of these genes.
The degree of basal nucleosome occupancy at promoters and other regulatory sequences also appears to vary between species. When applied to the human genome, models based on intrinsic nucleosome sequence preferences actually predict an overall increased occupancy at regulatory sites, in sharp contrast to most yeast promoters [143]. One explanation that was offered for this difference is that higher eukaryotes have greater requirements for variable gene expression, such as in the case of cell-type specific genes, and a constitutive open state might therefore not be desired [143]. Examples of TF binding to regions with high nucleosome occupancy have been described for the CCCTC-binding factor (CTCF) [144] and p53 [145], suggesting that the predicted increased nucleosome binding preferences in regulatory regions are relevant in vivo. Given these various observations it is evident that other mechanisms must exist to ensure TF access to DNA in regulatory regions that are occupied by nucleosomes. One model of TF binding to nucleosomal DNA that does not depend on external factors is based on in vitro observations that compacted DNA can undergo spontaneous transitions to more open states, allowing for brief windows of opportunity for TF access [146–148]. These movements can affect relatively small regions of DNA near the nucleosome entry sites, a process referred to as “nucleosome breathing”, or involve the unwinding of DNA over longer stretches [147, 149]. The increased accessibility of DNA at nucleosome entry sites is consistent with observations that TF binding sites are, on average, enriched at these locations in vivo [150–152]. Given the need to prevent cryptic transcription initiation, the thermodynamic balance in cells is likely such that individual TF binding events are not sufficient to prevent rapid rewrapping of nucleosomal DNA; however, cooperative binding of multiple TFs may overcome this barrier. Polach and Widom proposed that the binding of one TF could lead to further unwinding of the DNA on a nucleosome, enabling other factors to bind to nearby sites in a stepwise process that could ultimately result in a stable TF-DNA complex [153] (Fig. 11.3b). This cooperative model of TF access to nucleosomal DNA has two major additional benefits. First, it enables TFs to interact with each other without direct protein-protein contacts, creating new opportunities for coregulated gene expression. Second, the requirement for multiple closely spaced TF binding sites ensures regulatory site specificity. Cooperative binding of TFs to nucleosomal DNA has been demonstrated both in vitro [154] and in vivo [154–157], though it remains difficult to assess how widespread this mode of regulation is across the genome.
There is also evidence that TFs can interact with DNA in a manner that involves additional direct contacts with nucleosomes. For example, FOXA1 (HNF3A) binds more strongly to nucleosomal DNA than to naked DNA [158]. The source of this unique behaviour can be traced to the protein structure of the FoxA family members. FOXA1-3 contain a C-terminal domain that interacts with the core histones H3 and H4, as well as a winged helix N-terminal forkhead DNA binding domain that structurally resembles that of linker histone H1 [159]. In stark contrast to H1 linker histones, which are known for their ability to stabilize nucleosomes and higher order chromatin structures [160, 161], FoxA factors have intrinsic chromatin opening activity [159, 162]. Interestingly, this activity does not require the action of CMs such as SWI/SNF. Because of their ability to open condensed chromatin, FoxA proteins have been proposed to function as “pioneer” TFs that facilitate the binding of other factors [159]. A similar pioneer function has also been described for the RAR and RXR members of the nuclear receptor family, due to their ability to bind a highly compacted chromatin fibre containing a PEPCK promoter in an in vitro system that recaptured the chromatin dynamics observed at this promoter in vivo [163]. In this system, the action of the RAR/RXR heterodimer together with CMs was required to disrupt the chromatin for subsequent binding of nuclear factor 1 (NF1), an essential coregulator for transcriptional activation of PEPCK. The requirement for additional coregulators in transcriptional activation by both FoxA and RAR/RXR may be essential to ensure that their actions do not result in spurious transcription at non-specific sites in the genome. In the case of FoxA, methylation patterns associated with repressive or active chromatin domains also further guide recruitment to specific sites [164].
Other TFs that are able to access condensed chromatin include the CAAT-box/enhancer binding protein (C/EBP), though its pioneering role may be limited to a subset of genes [165]. In yeast, the Reb1 and Abf1 TFs can clearly function as pioneers as well, as evidenced by their aforementioned ability to direct the formation of NDRs [91, 141]. Finally, Gal4 upstream activating sequences (UAS) are able form mini-promoters regardless of their location in the genome [166], indicating that Gal4 binding can also disrupt chromatin. The Gal4 UAS used in this study contained multiple Gal4 binding sites, suggesting cooperative binding as a possible mechanism underlying this effect. Alternatively, Gal4 access to nucleosomal DNA can also be aided by the actions of CMs in a manner that does not involve displacing nucleosomes away from binding sites, as it was recently shown that the RSC complex can envelop and partially unwind a nucleosome in the GAL1/GAL10 promoter, with RSC essentially “presenting” this element for Gal4 binding [63] (Fig. 11.3c).
11.5 A Dynamic Regulatory Role for Chromatin
Up to this point, the relationship between TFs and chromatin has mainly been explored in terms of how TFs overcome the chromatin barrier to access DNA and facilitate further remodelling. However, the involvement of chromatin in gene expression goes beyond merely forming a passive impediment to TF binding. Indeed, there are many indications that CMs are causative for gene expression outputs, so presumably they must be both regulated and regulatory. In the remainder of this chapter I will examine some of the other roles of chromatin remodelling, such as effecting transcriptional repression and controlling the accessibility and activity of regulatory regions, as well as establishing higher-order chromatin organization. In all these cases the role of TFs will be highlighted in particular.
CMs are essential coregulators in TF-mediated repression of many target genes. A large number of these coregulators belong to a family of histone deacetylases (HDACs) [167, 168], which catalyze the removal of acetyl groups that are closely associated with a relaxed chromatin structure. Accordingly, they prevent initiation by maintaining chromatin in a condensed state that is inaccessible to the transcription machinery. Some of the effects of HDACs may also be mediated by deacetylation of proteins other than histones, such as TFs [169]. Like their HAT counterparts, HDACs typically operate as part of larger corepressor complexes that include other chromatin binding or remodelling activities, as has been described for the NURD [81] and NCoR (nuclear receptor corepressor) complexes [168, 170]. The importance of HDACs in transcriptional repression is reflected in the size of their family, which includes as many as 6 different members in yeast and 18 in human, distributed over four main classes [171]. In addition to HDACs, other CMs such as ATPase nucleosome remodelers have also been implicated in the formation of repressive chromatin structures. For example, the ISW2 complex can be recruited to a large variety of promoters by the Ume6 repressor in budding yeast, where it establishes a repressive chromatin environment as evidenced by decreased nuclease sensitivity [172]. SWI/SNF remodelers can also effect transcriptional repression, either directly [173–175], or as part of larger corepressor complexes that include deacetylase activities [81, 170, 176]. In contrast to HDACs, the mechanisms by which ATPase remodelers act to repress transcription are less well understood, but presumably involve chromatin compaction [172, 173] and/or the repositioning of nucleosomes to block important TF binding sites [177].
By condensing chromatin at promoters of repressed genes, CMs can place important restrictions on the actions of TFs, as illustrated by the effects of the Tup1-Cyc8 corepressor on Rap1-mediated gene activation in budding yeast [178]. The Tup1-Cyc8 complex was one of the first corepressors to be identified [179] and is targeted to promoters by a variety of sequence-specific TFs [180–183] where it recruits HDACs and the Isw2 remodeler complex to induce chromatin condensation [184, 185]. Among the Tup1-Cyc8 targets are promoters of genes that are bound by Rap1 in low- but not high-glucose conditions, despite the fact that Rap1 directs the expression of other genes encoding glycolytic enzymes and ribosomal protein subunits when glucose is present [186, 187]. The increased number of Rap1 targets in low-glucose is even more surprising given that global Rap1 levels actually decrease during a shift to low glucose medium [178]. The contradictory behaviour of Rap1 binding was explained by the actions of Tup1-Cyc8, which prevent Rap1 binding to low-glucose specific genes when glucose is present. The Tup1-Cyc8-mediated promoter compaction is only released upon glucose depletion, presumably through a mechanism that involves the release or inactivation of the TFs responsible for recruiting Tup1-Cyc8, allowing Rap1 to bind [178]. This example shows that chromatin remodelling can provide an additional level of regulation of gene expression by preventing activators from recognizing their binding sites in target promoters.
An unexpected finding has been that the actions of chromatin-targeting corepressors are not just limited to transcriptionally silent regions. Genome-wide ChIP experiments have revealed that HDACs are also associated with active promoters [188, 189]. Even more surprising, the degree of HDAC recruitment was positively correlated with transcription levels. To explain this paradox, it was proposed that the presence of HDACs at active promoters was needed to reset the chromatin state between subsequent rounds of initiation [189, 190], which suggests that histone acetylation – like TF and nucleosome interactions – may be inherently transient. Indeed, the dynamic nature of TFs interactions with DNA in vivo may well be directly connected to negative feedback from CMs. For example, the human glucocorticoid receptor can be actively removed from promoter templates by SWI/SNF remodelers [26, 191] and Rsc2 can speed up the release of Ace1 from non-specific binding sites in yeast [27]. Nevertheless, the presence of remodelling complexes associated with repression at active promoters does not necessarily have to be associated with returning these promoters to their basal state. The yeast SWI/SNF ATPase Mot1 is a global repressor known for its role in removing TBP from DNA [192], and like HDACs, its presence at promoters is positively correlated with transcript levels [193]. However, in this particular case it was shown that Mot1 can actually make a positive contribution to PIC assembly at active promoters by releasing a transcriptionally inert TBP complexed with the NC2 inhibitor, thereby allowing entry of free TBP and productive initiation [193].
The precise positioning of nucleosomes at promoters may also be important for establishing regulated gene expression, as illustrated by the actions of the RSC complex at the CHA1 promoter in budding yeast. In uninduced conditions, RSC represses CHA1 expression by placing a nucleosome over the TATA box, resulting in a decreased level of TBP binding [177, 194]. Crucially, in the absence of two key RSC components (Swh3 and Sth1), the expression levels of CHA1 in uninduced cells are approximately equal to those observed in fully induced cells. Thus, the presence of an inhibitory nucleosome over binding motifs recognized by the basal transcription machinery is vital for maintaining activator-regulated expression of CHA1. Similar regulation mechanisms are likely far more widespread, given the aforementioned observation that yeast genes with variable expression levels tend to have increased nucleosome occupancy within their promoter regions, often overlapping TATA boxes [142]. Taken together, these various observations show that the complex interplay between chromatin, CMs and TFs affects all aspects of transcription regulation.
11.6 TFs and Higher Order Chromatin Organization
In addition to the localized organization at the level of individual regulatory regions, chromatin is also arranged into higher-order structures that can span broad regions and affect multiple genes. These domains typically share a common chromatin environment that is characterized by a specific signature of histone marks and associated proteins. Classic examples of such domains include the condensed heterochromatin regions found at telomeres and in the pericentric regions surrounding centromeres in most organisms, as well as the mating-type loci in yeasts [195]. The heterochromatin in these regions is characterized by the presence of heterochromatin protein 1 (HP1) [196], histone hypoacetylation and H3K9 methylation (H3K9me) [197]. The co-occurrence of these marks is no coincidence, as H3K9me serves as an anchor point for the chromodomain that is present in HP1 [75]. Homologues of HP1 have been identified in Drosophila, vertebrates and fission yeast and its loss invariably leads to defects in telomere and centromere function. Additional domains marked by HP1 and H3K9me have also been associated with silencing of a number of genes dispersed throughout the genome [198–200].
A second important type of chromatin domain involved in gene silencing is established by Polycomb group (PcG) proteins. PcG proteins were initially identified as key developmental regulators of the Hox gene cluster in Drosophila (Reviewed in [201]), and two main PcG protein complexes have since been characterized with distinct roles in silencing in plants, vertebrates and flies. Polycomb repressive complex 2 (PRC2) has histone modifier activity and trimethylates H3K27, a characteristic signature of PcG chromatin domains, which can span up to 100 kb [202–204]. This methylation mark can be read by PRC1, which possesses ubiquitination activity. The specific mechanisms underlying HP1 and PcG silencing have been discussed in great detail elsewhere [195, 205–207]. Here, I will use these two domain types to illustrate the role of TFs in establishing higher order chromatin structure.
Heterochromatin typically originates at specific nucleation sites from which chromatin condensation spreads along the chromatin fibre. At telomeres, pericentric regions and yeast mating type loci, these nucleation sites often consist of highly repetitive DNA elements [208–210]. Studies in fission yeast have shown that repeat-based silencing depends on transcription of the repetitive regions and RNAi pathways [211, 212], and similar mechanisms have since been found to operate in fly, plants and vertebrates (Reviewed in [213]). There are also many examples where silencing is nucleated by TF binding, however. In fission yeast, the Pcr1 and Atf1 TFs can bind a heptamer sequence in the REIII element at the mating-type locus [214] and recruit the Clr4 histone methylase, the HP1 homolog Swi6, and the histone deacetylase Clr3 silencing factors [215, 216]. Budding yeast lacks HP1 homologs, but possesses silent information regulator (SIR) proteins that perform similar functions and which can be recruited to telomeres and mating-type loci by the synergistic actions of Rap1, Abf1 and Orc1 [217]. In tetrapods (four-limbed vertebrates), a large family of kruppel-associated box domain zinc finger TFs (KRAB-ZF) has also been implicated in silencing. The KRAB domain that characterizes this family interacts with KRAB associated protein 1 (KAP1) [218, 219], which acts as a scaffold for several heterochromatin-associated proteins, including HP1 [220–222]. Synthetic TF constructs with KRAB domains have been shown to induce heterochromatin silencing over broad regions, up to 12 kb away from their binding site [223, 224]. Natural KRAB-ZF proteins have been linked to the autoregulation of large clusters of KRAB-ZF genes [199, 200], but given that KRAB domains are present in more than 200 human TFs, they likely play a much wider role in chromatin metabolism. The KRAB domain is also discussed in Chapters 4 and 12 of this volume.
In contrast to HP1-associated heterochromatin, the origins of Polycomb domains are less well understood. In Drosophila, silencing by PcG proteins is driven by Polycomb response elements (PREs), which contain binding sites for the Pleiohomeotic (PHO) and PHO-like zinc finger TFs [225, 226], the only PcG proteins identified to date with DNA sequence specificity. The importance of PHO and PHO-like for PRE function is firmly established, as their disruption results in silencing defects at Hox genes [225, 227, 228] and a loss of PRC1 and PRC2 components [228]; however, PHO binding sites alone are insufficient to confer PRE-mediated silencing [225, 226, 229]. Many other TFs have been shown to bind PREs in Drosophila, including Pipsqueak, Zeste and GAGA factor (GAF) (Reviewed in [72]), but their role in silencing is unclear, given that null mutants for many of these genes do not show obvious PcG phenotypes. One possible explanation is that these TFs act synergistically at PREs, which is consistent with computational analyses that show that clusters of TF binding motifs – but not individual sites – can distinguish PRE from non-PRE sequences [230]. Redundancy between factors may explain why some null mutants do not show phenotypes.
Even less is known about PRC recruitment in vertebrates, where it has proved challenging to identify PREs because PcG proteins are often distributed over broad regions [202, 204, 231, 232]. A 3 kb DNA fragment in the MafB gene region that possesses activities consistent with a PRE was recently identified in mouse [233]. This fragment, named PRE-kr, was shown to bind PcG proteins and contains conserved binding sites for the mammalian PHO homolog YY1, as well as GAGAG motifs that are known to be bound by GAF and Pipsqueak in Drosophila. Another PRE with conserved YY1 binding sites has since been characterized in the human HOXD cluster, and disruption of these sites negatively affected binding of the PRC1 component BMI1 [234]. The role of YY1 in PcG silencing is consistent with earlier observations that YY1 knockdown results in loss of recruitment of the PRC2 component Ezh2 and H3K27me [235], as well as with other studies that have shown that YY1 interacts with PcG components [236–238]. Taken together, these data suggest that at least some of the PcG-targeting mechanisms are conserved between flies and mammals. Nonetheless, other TFs such as the embryonic stem cell regulators OCT4 and NANOG may also be involved in targeting PcG proteins in mammals, based on their high degree of overlap with PcG proteins in ChIP studies [202, 231, 239]. Moreover, the discovery of the HOTAIR transcript, which targets PRC2 to the human HOXD locus, indicates that ncRNAs also play a role in directing Polycomb silencing [240]. Future studies will undoubtedly reveal whether this latter mechanism is more widespread.
Several mechanisms are believed to operate to expand chromatin domains beyond their initial nucleation sites (Reviewed in [241]). One model of spreading described for HP1 family members depends on a self-sustaining wave of silencing complex assembly, which is based in the ability of HP1 to bind both H3K9 methylated histones as well as the methyltransferase responsible for this modification (Fig. 11.4a) [75, 77, 242]. Starting at the nucleation site, H3K9 methylation of neighboring nucleosomes by HP1-recruited methyltransferases creates new HP1 binding sites, resulting in more HP1 binding and further propagation of the signal. A similar mechanism involving repeated cycles of deacetylation has also been described for SIR proteins in budding yeast [243, 244]. Recurrent assembly cannot completely account for all observations of spreading from a nucleation site, however, as indicated by the following examples. In budding yeast, individual Rap1 and Abf1 binding sites that are unable to direct silencing independently can enhance the actions of a silencer that is 4 kb away [245], suggesting long–range interactions between these sites. Another signal spreading from a subtelomeric silencer was shown to “skip over” an active reporter gene flanked by subtelomeric antisilencing regions (STARS), but still affected a second distal reporter gene [246]. Finally, ChIP studies of PcG proteins in Drosophila have revealed distribution patterns that seem inconsistent with a progressive spreading of Polycomb complexes. For example, while the H3K27me3 mark is consistently found in large domains [203, 247–250], the PRC1 components Ph and Psc and the PRC2 methyltransferase E(z) are concentrated in much smaller peaks [203, 247]. Currently, the most favoured model to explain these various observations involves folding of the DNA in a manner that allows nucleation sites to contact and modify the surrounding chromatin (Fig. 11.4b), and has been proposed to explain the difference in distribution patterns of PcG components and H3K27me3 [251]. Several cases of long–range interactions between PREs and distant regulatory sites have also been described, forming higher order chromatin loop configurations that may facilitate gene silencing across broad domains [252, 253]. The relationship of TFs to higher-order chromatin structure is described in more detail in Chapter 13.
Formation of chromatin domains. a Mechanism of spreading for HP1 heterochromatin at the S. pombe mating type locus from TF nucleation sites (Modified from [214]). Atf1 and Pcr1 binding results in the recruitment of the Clr3 histone deacetylase, which subsequently cooperates with heterochromatin proteins (HP) such as the HP1 homolog Swi6 to promote H3K9me of neighbouring nucleosomes. This creates additional HP1 binding sites, which form the basis for the spreading process. b Schematic representation of spreading of chromatin domains by looping interactions between the nucleation site and the surrounding DNA. c Model for the enhancer-blocker function of CTCF. Interactions between distant CTCF binding sites can form looped domains, thereby isolating genes from the actions of upstream enhancers
Given that silencing can propagate autonomously along the chromatin fibre, and that distal regulatory elements such as PREs and enhancers can operate over large distances, how are their effects on one region of the genome kept from spilling over to nearby genes? The answer to this question lies in yet another group of regulatory elements called insulators [254–256], which possess one of two distinct characteristics: (1) they can block enhancers from activating genes when placed between the enhancer and the gene or (2) they can act as boundary elements to prevent the spread of the silencing effects of heterochromatin. These two activities are separate and measured in different assays, though many insulators can perform both functions in vivo, such as the 5′HS4 insulator in the chicken β-globin locus [257, 258]. Once again, TFs play a central role in establishing insulator regions, and at least five different insulator-binding TFs have been identified in Drosophila to date: ZW5, Su(Hw), dCTCF, BEAF, and GAGA (Reviewed in [259]). In contrast, most vertebrate insulators appear to depend on only a single TF, the CCCTC-binding factor (CTCF) [257]. CTCF is considered to mainly function as an enhancer blocker rather than as a boundary protein, as evidenced by the fact that it is dispensable for blocking the spread of heterochromatin at the chicken β-globin locus [260]. Instead, this latter function depends on the USF1 TF, which binds boundary elements in the 5′HS4 insulator as a heterodimer with USF2 [258, 261]. The USF1/USF2 heterodimer recruits HATs and the SET 7/9 methyltransferase, which establish a region of open chromatin that is thought to prevent the progression of silencing analogous to the manner in which firewalls prevent forest fires from spreading. In contrast, enhancer-blocking insulators such as those bound by Su(Hw) in Drosophila (Reviewed in [262]) or CTCF in vertebrates (Reviewed in [263]) have been suggested to operate by organizing chromatin into looped domains, isolating the genes contained inside from their distant regulatory elements (Fig. 11.4c). In addition, CTCF has also been implicated in anchoring DNA to the nuclear periphery, an area that is typically associated with a repressive chromatin environment, as it was found to be enriched at the boundaries of domains that are linked to the nuclear lamina [264].
11.7 Concluding Remarks
The complexity of chromatin–TF interactions is reflected in the considerable variability in initiation mechanisms for the few genes studied in great detail [83] suggesting that there are many routes leading to productive transcription. Indeed, considering that the requirement for coregulators at a single gene can vary depending on external conditions, and that promoters are typically unique in a genome, the number of transcriptional activation mechanisms may yet prove to be larger than the number of genes. Nonetheless, the number of possibilities is clearly not unlimited, since at any given regulatory region only a subset of TFs and their coregulators play a dominant role. Thus, it should be possible to build a catalogue of the proteins most commonly bound to these elements in specific cell types, and eventually decode the mechanisms that control gene expression. ChIP in combination with either microarrays or next-generation sequencing is currently the most widely used method for the identification of the proteins and histone modifications associated with DNA [265, 266]; however, this technique has several drawbacks. First, it can only identify the location of a handful of proteins at the same time, and second, it requires advance knowledge of the factor(s) to study. An alternative approach called proteomics of isolated chromatin segments (PICh) was recently developed that does not suffer from these limitations, and uses mass-spectrometry to detect proteins associated with a chromatin segment [267]. If this approach were to be applied to the large collections of regulatory regions that are now being identified in genome-wide nuclease hypersensitivity assays such as those undertaken by the ENCODE and modENCODE consortia [268], it might greatly expand our knowledge of the interplay between TFs and chromatin at these locations.
Simply knowing which proteins are associated with a given genomic region will not be enough to understand how these proteins operate to regulate transcription, since they generally do not work in isolation. Protein–protein interaction maps should also greatly facilitate mapping gene regulatory mechanisms, since they reveal interactions between and among TFs and CMs [269]. Moreover, maps of long range interactions between regulatory regions are needed to understand the interplay between promoters, enhancers, silencers and insulators. The advent of new technologies such as the numerous derivatives of chromosome conformation capture (3C) [270, 271] now make such approaches possible at a genome-wide level (see Chapter 13). Finally, detailed knowledge of the affinities of TFs and their coregulators for DNA, as well as for their protein binding partners will also be essential. This will require the application of techniques that can assess both the intrinsic DNA sequence specificities of TFs (see Chapter 8) and the binding kinetics of proteins, in a high-throughput and quantitative fashion. Potential strategies for the latter have been outlined by Segal and Widom [272]. Together, these various types of data will provide valuable insight into the ground rules that govern the interactions between DNA, chromatin and the transcription machinery. These rules can then form the basis for in silico modeling of these processes, which will be essential if we are to fully understand the intricate relationships between TFs and chromatin.
References
Lee TI, Young RA (2000) Transcription of eukaryotic protein-coding genes. Annu Rev Genet 34:77–137
van Hijum SA, Medema MH, Kuipers OP (2009) Mechanisms and evolution of control logic in prokaryotic transcriptional regulation. Microbiol Mol Biol Rev 73 (3):481–509
Kornberg RD, Thomas JO (1974) Chromatin structure; oligomers of the histones. Science 184 (139):865–868
Richmond TJ, Finch JT, Rushton B, Rhodes D, Klug A (1984) Structure of the nucleosome core particle at 7 A resolution. Nature 311 (5986):532–537
Knezetic JA, Luse DS (1986) The presence of nucleosomes on a DNA template prevents initiation by RNA polymerase II in vitro. Cell 45 (1):95–104
Lorch Y, LaPointe JW, Kornberg RD (1987) Nucleosomes inhibit the initiation of transcription but allow chain elongation with the displacement of histones. Cell 49 (2):203–210
Han M, Grunstein M (1988) Nucleosome loss activates yeast downstream promoters in vivo. Cell 55 (6):1137–1145
Hager GL, McNally JG, Misteli T (2009) Transcription dynamics. Mol Cell 35 (6):741–753
Segal E, Widom J (2009a) What controls nucleosome positions? Trends Genet 25 (8):335–343
Wu C, Bingham PM, Livak KJ, Holmgren R, Elgin SC (1979) The chromatin structure of specific genes: I. Evidence for higher order domains of defined DNA sequence. Cell 16 (4):797–806
Giresi PG, Kim J, McDaniell RM, Iyer VR, Lieb JD (2007) FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin. Genome Res 17 (6):877–885
Lee W, Tillo D, Bray N, Morse RH, Davis RW, Hughes TR, Nislow C (2007) A high-resolution atlas of nucleosome occupancy in yeast. Nat Genet 39 (10):1235–1244
Mavrich TN, Jiang C, Ioshikhes IP, Li X, Venters BJ, Zanton SJ, Tomsho LP, Qi J, Glaser RL, Schuster SC, Gilmour DS, Albert I, Pugh BF (2008a) Nucleosome organization in the Drosophila genome. Nature 453 (7193):358–362
Schones DE, Cui K, Cuddapah S, Roh TY, Barski A, Wang Z, Wei G, Zhao K (2008) Dynamic regulation of nucleosome positioning in the human genome. Cell 132 (5):887–898
Yuan GC, Liu YJ, Dion MF, Slack MD, Wu LF, Altschuler SJ, Rando OJ (2005) Genome-scale identification of nucleosome positions in S. cerevisiae. Science 309 (5734):626–630
Boyle AP, Davis S, Shulha HP, Meltzer P, Margulies EH, Weng Z, Furey TS, Crawford GE (2008) High-resolution mapping and characterization of open chromatin across the genome. Cell 132 (2):311–322
Hesselberth JR, Chen X, Zhang Z, Sabo PJ, Sandstrom R, Reynolds AP, Thurman RE, Neph S, Kuehn MS, Noble WS, Fields S, Stamatoyannopoulos JA (2009) Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat Methods 6 (4):283–289
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K (2007) High-resolution profiling of histone methylations in the human genome. Cell 129 (4):823–837
Harbison CT, Gordon DB, Lee TI, Rinaldi NJ, Macisaac KD, Danford TW, Hannett NM, Tagne JB, Reynolds DB, Yoo J, Jennings EG, Zeitlinger J, Pokholok DK, Kellis M, Rolfe PA, Takusagawa KT, Lander ES, Gifford DK, Fraenkel E, Young RA (2004) Transcriptional regulatory code of a eukaryotic genome. Nature 431 (7004):99–104
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, Wang W, Weng Z, Green RD, Crawford GE, Ren B (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39 (3):311–318
Kim TH, Barrera LO, Zheng M, Qu C, Singer MA, Richmond TA, Wu Y, Green RD, Ren B (2005) A high-resolution map of active promoters in the human genome. Nature 436 (7052):876–880
Pokholok DK, Harbison CT, Levine S, Cole M, Hannett NM, Lee TI, Bell GW, Walker K, Rolfe PA, Herbolsheimer E, Zeitlinger J, Lewitter F, Gifford DK, Young RA (2005) Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122 (4):517–527
Wang Z, Zang C, Rosenfeld JA, Schones DE, Barski A, Cuddapah S, Cui K, Roh TY, Peng W, Zhang MQ, Zhao K (2008) Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40 (7):897–903
Dion MF, Kaplan T, Kim M, Buratowski S, Friedman N, Rando OJ (2007) Dynamics of replication-independent histone turnover in budding yeast. Science 315 (5817):1405–1408
Linger J, Tyler JK (2006) Global replication-independent histone H4 exchange in budding yeast. Eukaryot Cell 5 (10):1780–1787
Fletcher TM, Xiao N, Mautino G, Baumann CT, Wolford R, Warren BS, Hager GL (2002) ATP-dependent mobilization of the glucocorticoid receptor during chromatin remodeling. Mol Cell Biol 22 (10):3255–3263
Karpova TS, Chen TY, Sprague BL, McNally JG (2004) Dynamic interactions of a transcription factor with DNA are accelerated by a chromatin remodeller. EMBO Rep 5 (11):1064–1070
McNally JG, Muller WG, Walker D, Wolford R, Hager GL (2000) The glucocorticoid receptor: rapid exchange with regulatory sites in living cells. Science 287 (5456):1262–1265
Sharp ZD, Mancini MG, Hinojos CA, Dai F, Berno V, Szafran AT, Smith KP, Lele TP, Ingber DE, Mancini MA (2006) Estrogen-receptor-alpha exchange and chromatin dynamics are ligand- and domain-dependent. J Cell Sci 119 (Pt 19):4101–4116
Farnham PJ (2009) Insights from genomic profiling of transcription factors. Nat Rev Genet 10 (9):605–616
Li B, Carey M, Workman JL (2007) The role of chromatin during transcription. Cell 128 (4):707–719
Venters BJ, Pugh BF (2009) How eukaryotic genes are transcribed. Crit Rev Biochem Mol Biol 44 (2-3):117–141
Weake VM, Workman JL (2010) Inducible gene expression: diverse regulatory mechanisms. Nat Rev Genet 11 (6):426–437
Kouzarides T (2007) Chromatin modifications and their function. Cell 128 (4):693–705
Reid G, Gallais R, Metivier R (2009) Marking time: the dynamic role of chromatin and covalent modification in transcription. Int J Biochem Cell Biol 41 (1):155–163
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403 (6765):41–45
Rice JC, Allis CD (2001) Histone methylation versus histone acetylation: new insights into epigenetic regulation. Curr Opin Cell Biol 13 (3):263–273
Kurdistani SK, Tavazoie S, Grunstein M (2004) Mapping global histone acetylation patterns to gene expression. Cell 117 (6):721–733
Liu CL, Kaplan T, Kim M, Buratowski S, Schreiber SL, Friedman N, Rando OJ (2005) Single-nucleosome mapping of histone modifications in S. cerevisiae. PLoS Biol 3 (10):e328
Roh TY, Ngau WC, Cui K, Landsman D, Zhao K (2004) High-resolution genome-wide mapping of histone modifications. Nat Biotechnol 22 (8):1013–1016
Sinha I, Wiren M, Ekwall K (2006) Genome-wide patterns of histone modifications in fission yeast. Chromosome Res 14 (1):95–105
Bernstein BE, Kamal M, Lindblad-Toh K, Bekiranov S, Bailey DK, Huebert DJ, McMahon S, Karlsson EK, Kulbokas EJ, 3rd, Gingeras TR, Schreiber SL, Lander ES (2005) Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120 (2):169–181
Liang G, Lin JC, Wei V, Yoo C, Cheng JC, Nguyen CT, Weisenberger DJ, Egger G, Takai D, Gonzales FA, Jones PA (2004) Distinct localization of histone H3 acetylation and H3-K4 methylation to the transcription start sites in the human genome. Proc Natl Acad Sci U S A 101 (19):7357–7362
Roh TY, Cuddapah S, Zhao K (2005) Active chromatin domains are defined by acetylation islands revealed by genome-wide mapping. Genes Dev 19 (5):542–552
Schubeler D, MacAlpine DM, Scalzo D, Wirbelauer C, Kooperberg C, van Leeuwen F, Gottschling DE, O’Neill LP, Turner BM, Delrow J, Bell SP, Groudine M (2004) The histone modification pattern of active genes revealed through genome-wide chromatin analysis of a higher eukaryote. Genes Dev 18 (11):1263–1271
Shahbazian MD, Grunstein M (2007) Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem 76:75–100
Wiren M, Silverstein RA, Sinha I, Walfridsson J, Lee HM, Laurenson P, Pillus L, Robyr D, Grunstein M, Ekwall K (2005) Genomewide analysis of nucleosome density histone acetylation and HDAC function in fission yeast. Embo J 24 (16):2906–2918
Bannister AJ, Kouzarides T (2005) Reversing histone methylation. Nature 436 (7054):1103–1106
Kouzarides T (2000) Acetylation: a regulatory modification to rival phosphorylation? Embo J 19 (6):1176–1179
Spange S, Wagner T, Heinzel T, Kramer OH (2009) Acetylation of non-histone proteins modulates cellular signalling at multiple levels. Int J Biochem Cell Biol 41 (1):185–198
Yang XJ (2004) The diverse superfamily of lysine acetyltransferases and their roles in leukemia and other diseases. Nucleic Acids Res 32 (3):959–976
Ge Z, Liu C, Bjorkholm M, Gruber A, Xu D (2006) Mitogen-activated protein kinase cascade-mediated histone H3 phosphorylation is critical for telomerase reverse transcriptase expression/telomerase activation induced by proliferation. Mol Cell Biol 26 (1):230–237
Mahadevan LC, Willis AC, Barratt MJ (1991) Rapid histone H3 phosphorylation in response to growth factors, phorbol esters, okadaic acid, and protein synthesis inhibitors. Cell 65 (5):775–783
Nathan D, Ingvarsdottir K, Sterner DE, Bylebyl GR, Dokmanovic M, Dorsey JA, Whelan KA, Krsmanovic M, Lane WS, Meluh PB, Johnson ES, Berger SL (2006) Histone sumoylation is a negative regulator in Saccharomyces cerevisiae and shows dynamic interplay with positive-acting histone modifications. Genes Dev 20 (8):966–976
Van Rechem C, Boulay G, Pinte S, Stankovic-Valentin N, Guerardel C, Leprince D (2010) Differential regulation of HIC1 target genes by CtBP and NuRD, via an acetylation/SUMOylation switch, in quiescent versus proliferating cells. Mol Cell Biol 30 (16):4045–4059
Chandrasekharan MB, Huang F, Sun ZW (2010) Histone H2B ubiquitination and beyond: Regulation of nucleosome stability, chromatin dynamics and the trans-histone H3 methylation. Epigenetics 5 (6)
Wang H, Zhai L, Xu J, Joo HY, Jackson S, Erdjument-Bromage H, Tempst P, Xiong Y, Zhang Y (2006) Histone H3 and H4 ubiquitylation by the CUL4-DDB-ROC1 ubiquitin ligase facilitates cellular response to DNA damage. Mol Cell 22 (3):383–394
Zhu B, Zheng Y, Pham AD, Mandal SS, Erdjument-Bromage H, Tempst P, Reinberg D (2005) Monoubiquitination of human histone H2B: the factors involved and their roles in HOX gene regulation. Mol Cell 20 (4):601–611
Cheung P, Tanner KG, Cheung WL, Sassone-Corsi P, Denu JM, Allis CD (2000) Synergistic coupling of histone H3 phosphorylation and acetylation in response to epidermal growth factor stimulation. Mol Cell 5 (6):905–915
Lo WS, Trievel RC, Rojas JR, Duggan L, Hsu JY, Allis CD, Marmorstein R, Berger SL (2000) Phosphorylation of serine 10 in histone H3 is functionally linked in vitro and in vivo to Gcn5-mediated acetylation at lysine 14. Mol Cell 5 (6):917–926
Rea S, Eisenhaber F, O’Carroll D, Strahl BD, Sun ZW, Schmid M, Opravil S, Mechtler K, Ponting CP, Allis CD, Jenuwein T (2000) Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406 (6796):593–599
Clapier CR, Cairns BR (2009) The biology of chromatin remodeling complexes. Annu Rev Biochem 78:273–304
Floer M, Wang X, Prabhu V, Berrozpe G, Narayan S, Spagna D, Alvarez D, Kendall J, Krasnitz A, Stepansky A, Hicks J, Bryant GO, Ptashne M (2010) A RSC/nucleosome complex determines chromatin architecture and facilitates activator binding. Cell 141 (3):407–418
Narlikar GJ, Phelan ML, Kingston RE (2001) Generation and interconversion of multiple distinct nucleosomal states as a mechanism for catalyzing chromatin fluidity. Mol Cell 8 (6):1219–1230
Dechassa ML, Sabri A, Pondugula S, Kassabov SR, Chatterjee N, Kladde MP, Bartholomew B (2010) SWI/SNF has intrinsic nucleosome disassembly activity that is dependent on adjacent nucleosomes. Mol Cell 38 (4):590–602
Lorch Y, Maier-Davis B, Kornberg RD (2006) Chromatin remodeling by nucleosome disassembly in vitro. Proc Natl Acad Sci U S A 103 (9):3090–3093
Vicent GP, Nacht AS, Smith CL, Peterson CL, Dimitrov S, Beato M (2004) DNA instructed displacement of histones H2A and H2B at an inducible promoter. Mol Cell 16 (3):439–452
Bruno M, Flaus A, Stockdale C, Rencurel C, Ferreira H, Owen-Hughes T (2003) Histone H2A/H2B dimer exchange by ATP-dependent chromatin remodeling activities. Mol Cell 12 (6):1599–1606
Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C (2004) ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303 (5656):343–348
Bao Y, Shen X (2007) SnapShot: chromatin remodeling complexes. Cell 129 (3):632
Eisen JA, Sweder KS, Hanawalt PC (1995) Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res 23 (14):2715–2723
Bottomley MJ (2004) Structures of protein domains that create or recognize histone modifications. EMBO Rep 5 (5):464–469
de la Cruz X, Lois S, Sanchez-Molina S, Martinez-Balbas MA (2005) Do protein motifs read the histone code? Bioessays 27 (2):164–175
Hassan AH, Prochasson P, Neely KE, Galasinski SC, Chandy M, Carrozza MJ, Workman JL (2002) Function and selectivity of bromodomains in anchoring chromatin-modifying complexes to promoter nucleosomes. Cell 111 (3):369–379
Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410 (6824):120–124
Flanagan JF, Mi LZ, Chruszcz M, Cymborowski M, Clines KL, Kim Y, Minor W, Rastinejad F, Khorasanizadeh S (2005) Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature 438 (7071):1181–1185
Lachner M, O’Carroll D, Rea S, Mechtler K, Jenuwein T (2001) Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410 (6824):116–120
Sims RJ, 3rd, Chen CF, Santos-Rosa H, Kouzarides T, Patel SS, Reinberg D (2005) Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J Biol Chem 280 (51):41789–41792
Boyer LA, Latek RR, Peterson CL (2004) The SANT domain: a unique histone-tail-binding module? Nat Rev Mol Cell Biol 5 (2):158–163
Ho L, Crabtree GR (2010) Chromatin remodelling during development. Nature 463 (7280):474–484
Bowen NJ, Fujita N, Kajita M, Wade PA (2004) Mi-2/NuRD: multiple complexes for many purposes. Biochim Biophys Acta 1677 (1-3):52–57
Dirscherl SS, Krebs JE (2004) Functional diversity of ISWI complexes. Biochem Cell Biol 82 (4):482–489
Biddick R, Young ET (2009) The disorderly study of ordered recruitment. Yeast 26 (4):205–220
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 6 (3):197–208
Liu J, Perumal NB, Oldfield CJ, Su EW, Uversky VN, Dunker AK (2006a) Intrinsic disorder in transcription factors. Biochemistry 45 (22):6873–6888
Dunker AK, Lawson JD, Brown CJ, Williams RM, Romero P, Oh JS, Oldfield CJ, Campen AM, Ratliff CM, Hipps KW, Ausio J, Nissen MS, Reeves R, Kang C, Kissinger CR, Bailey RW, Griswold MD, Chiu W, Garner EC, Obradovic Z (2001) Intrinsically disordered protein. J Mol Graph Model 19 (1):26–59
Hassan AH, Neely KE, Vignali M, Reese JC, Workman JL (2001) Promoter targeting of chromatin-modifying complexes. Front Biosci 6:D1054–1064
Narlikar GJ, Fan HY, Kingston RE (2002) Cooperation between complexes that regulate chromatin structure and transcription. Cell 108 (4):475–487
Peterson CL, Workman JL (2000) Promoter targeting and chromatin remodeling by the SWI/SNF complex. Curr Opin Genet Dev 10 (2):187–192
Cairns BR, Lorch Y, Li Y, Zhang M, Lacomis L, Erdjument-Bromage H, Tempst P, Du J, Laurent B, Kornberg RD (1996) RSC, an essential, abundant chromatin-remodeling complex. Cell 87 (7):1249–1260
Badis G, Chan ET, van Bakel H, Pena-Castillo L, Tillo D, Tsui K, Carlson CD, Gossett AJ, Hasinoff MJ, Warren CL, Gebbia M, Talukder S, Yang A, Mnaimneh S, Terterov D, Coburn D, Li Yeo A, Yeo ZX, Clarke ND, Lieb JD, Ansari AZ, Nislow C, Hughes TR (2008) A library of yeast transcription factor motifs reveals a widespread function for Rsc3 in targeting nucleosome exclusion at promoters. Mol Cell 32 (6):878–887
Hogan CJ, Aligianni S, Durand-Dubief M, Persson J, Will WR, Webster J, Wheeler L, Mathews CK, Elderkin S, Oxley D, Ekwall K, Varga-Weisz PD (2009) Fission yeast Iec1-ino80-mediated nucleosome eviction regulates nucleotide and phosphate metabolism. Mol Cell Biol 30 (3):657–674
Mohrmann L, Verrijzer CP (2005) Composition and functional specificity of SWI2/SNF2 class chromatin remodeling complexes. Biochim Biophys Acta 1681 (2-3):59–73
Patsialou A, Wilsker D, Moran E (2005) DNA-binding properties of ARID family proteins. Nucleic Acids Res 33 (1):66–80
Thomas JO, Travers AA (2001) HMG1 and 2, and related ‘architectural’ DNA-binding proteins. Trends Biochem Sci 26 (3):167–174
Wilson B, Erdjument-Bromage H, Tempst P, Cairns BR (2006) The RSC chromatin remodeling complex bears an essential fungal-specific protein module with broad functional roles. Genetics 172 (2):795–809
Trotter KW, Archer TK (2008) The BRG1 transcriptional coregulator. Nucl Recept Signal 6:e004
Kitagawa H, Fujiki R, Yoshimura K, Mezaki Y, Uematsu Y, Matsui D, Ogawa S, Unno K, Okubo M, Tokita A, Nakagawa T, Ito T, Ishimi Y, Nagasawa H, Matsumoto T, Yanagisawa J, Kato S (2003) The chromatin-remodeling complex WINAC targets a nuclear receptor to promoters and is impaired in Williams syndrome. Cell 113 (7):905–917
Xu W, Cho H, Kadam S, Banayo EM, Anderson S, Yates JR, 3rd, Emerson BM, Evans RM (2004) A methylation-mediator complex in hormone signaling. Genes Dev 18 (2):144–156
Lessard J, Wu JI, Ranish JA, Wan M, Winslow MM, Staahl BT, Wu H, Aebersold R, Graef IA, Crabtree GR (2007) An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron 55 (2):201–215
Wu JI, Lessard J, Olave IA, Qiu Z, Ghosh A, Graef IA, Crabtree GR (2007) Regulation of dendritic development by neuron-specific chromatin remodeling complexes. Neuron 56 (1):94–108
Takeuchi JK, Bruneau BG (2009) Directed transdifferentiation of mouse mesoderm to heart tissue by defined factors. Nature 459 (7247):708–711
Bhaumik SR, Green MR (2001) SAGA is an essential in vivo target of the yeast acidic activator Gal4p. Genes Dev 15 (15):1935–1945
Bhaumik SR, Green MR (2002) Differential requirement of SAGA components for recruitment of TATA-box-binding protein to promoters in vivo. Mol Cell Biol 22 (21):7365–7371
Larschan E, Winston F (2001) The S. cerevisiae SAGA complex functions in vivo as a coactivator for transcriptional activation by Gal4. Genes Dev 15 (15):1946–1956
Chan HM, La Thangue NB (2001) p300/CBP proteins: HATs for transcriptional bridges and scaffolds. J Cell Sci 114 (Pt 13):2363–2373
Kalkhoven E (2004) CBP and p300: HATs for different occasions. Biochem Pharmacol 68 (6):1145–1155
Panne D, Maniatis T, Harrison SC (2007) An atomic model of the interferon-beta enhanceosome. Cell 129 (6):1111–1123
Merika M, Williams AJ, Chen G, Collins T, Thanos D (1998) Recruitment of CBP/p300 by the IFN beta enhanceosome is required for synergistic activation of transcription. Mol Cell 1 (2):277–287
Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, Kuehn MS, Taylor CM, Neph S, Koch CM, Asthana S, Malhotra A, Adzhubei I, Greenbaum JA, Andrews RM, Flicek P, Boyle PJ, Cao H, Carter NP, Clelland GK, Davis S, Day N, Dhami P, Dillon SC, Dorschner MO, Fiegler H, Giresi PG, Goldy J, Hawrylycz M, Haydock A, Humbert R, James KD, Johnson BE, Johnson EM, Frum TT, Rosenzweig ER, Karnani N, Lee K, Lefebvre GC, Navas PA, Neri F, Parker SC, Sabo PJ, Sandstrom R, Shafer A, Vetrie D, Weaver M, Wilcox S, Yu M, Collins FS, Dekker J, Lieb JD, Tullius TD, Crawford GE, Sunyaev S, Noble WS, Dunham I, Denoeud F, Reymond A, Kapranov P, Rozowsky J, Zheng D, Castelo R, Frankish A, Harrow J, Ghosh S, Sandelin A, Hofacker IL, Baertsch R, Keefe D, Dike S, Cheng J, Hirsch HA, Sekinger EA, Lagarde J, Abril JF, Shahab A, Flamm C, Fried C, Hackermuller J, Hertel J, Lindemeyer M, Missal K, Tanzer A, Washietl S, Korbel J, Emanuelsson O, Pedersen JS, Holroyd N, Taylor R, Swarbreck D, Matthews N, Dickson MC, Thomas DJ, Weirauch MT, Gilbert J, Drenkow J, Bell I, Zhao X, Srinivasan KG, Sung WK, Ooi HS, Chiu KP, Foissac S, Alioto T, Brent M, Pachter L, Tress ML, Valencia A, Choo SW, Choo CY, Ucla C, Manzano C, Wyss C, Cheung E, Clark TG, Brown JB, Ganesh M, Patel S, Tammana H, Chrast J, Henrichsen CN, Kai C, Kawai J, Nagalakshmi U, Wu J, Lian Z, Lian J, Newburger P, Zhang X, Bickel P, Mattick JS, Carninci P, Hayashizaki Y, Weissman S, Hubbard T, Myers RM, Rogers J, Stadler PF, Lowe TM, Wei CL, Ruan Y, Struhl K, Gerstein M, Antonarakis SE, Fu Y, Green ED, Karaoz U, Siepel A, Taylor J, Liefer LA, Wetterstrand KA, Good PJ, Feingold EA, Guyer MS, Cooper GM, Asimenos G, Dewey CN, Hou M, Nikolaev S, Montoya-Burgos JI, Loytynoja A, Whelan S, Pardi F, Massingham T, Huang H, Zhang NR, Holmes I, Mullikin JC, Ureta-Vidal A, Paten B, Seringhaus M, Church D, Rosenbloom K, Kent WJ, Stone EA, Batzoglou S, Goldman N, Hardison RC, Haussler D, Miller W, Sidow A, Trinklein ND, Zhang ZD, Barrera L, Stuart R, King DC, Ameur A, Enroth S, Bieda MC, Kim J, Bhinge AA, Jiang N, Liu J, Yao F, Vega VB, Lee CW, Ng P, Shahab A, Yang A, Moqtaderi Z, Zhu Z, Xu X, Squazzo S, Oberley MJ, Inman D, Singer MA, Richmond TA, Munn KJ, Rada-Iglesias A, Wallerman O, Komorowski J, Fowler JC, Couttet P, Bruce AW, Dovey OM, Ellis PD, Langford CF, Nix DA, Euskirchen G, Hartman S, Urban AE, Kraus P, Van Calcar S, Heintzman N, Kim TH, Wang K, Qu C, Hon G, Luna R, Glass CK, Rosenfeld MG, Aldred SF, Cooper SJ, Halees A, Lin JM, Shulha HP, Zhang X, Xu M, Haidar JN, Yu Y, Ruan Y, Iyer VR, Green RD, Wadelius C, Farnham PJ, Ren B, Harte RA, Hinrichs AS, Trumbower H, Clawson H, Hillman-Jackson J, Zweig AS, Smith K, Thakkapallayil A, Barber G, Kuhn RM, Karolchik D, Armengol L, Bird CP, de Bakker PI, Kern AD, Lopez-Bigas N, Martin JD, Stranger BE, Woodroffe A, Davydov E, Dimas A, Eyras E, Hallgrimsdottir IB, Huppert J, Zody MC, Abecasis GR, Estivill X, Bouffard GG, Guan X, Hansen NF, Idol JR, Maduro VV, Maskeri B, McDowell JC, Park M, Thomas PJ, Young AC, Blakesley RW, Muzny DM, Sodergren E, Wheeler DA, Worley KC, Jiang H, Weinstock GM, Gibbs RA, Graves T, Fulton R, Mardis ER, Wilson RK, Clamp M, Cuff J, Gnerre S, Jaffe DB, Chang JL, Lindblad-Toh K, Lander ES, Koriabine M, Nefedov M, Osoegawa K, Yoshinaga Y, Zhu B, de Jong PJ (2007) Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447 (7146):799–816
Heintzman ND, Hon GC, Hawkins RD, Kheradpour P, Stark A, Harp LF, Ye Z, Lee LK, Stuart RK, Ching CW, Ching KA, Antosiewicz-Bourget JE, Liu H, Zhang X, Green RD, Lobanenkov VV, Stewart R, Thomson JA, Crawford GE, Kellis M, Ren B (2009) Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459 (7243):108–112
Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, Harmin DA, Laptewicz M, Barbara-Haley K, Kuersten S, Markenscoff-Papadimitriou E, Kuhl D, Bito H, Worley PF, Kreiman G, Greenberg ME (2010) Widespread transcription at neuronal activity-regulated enhancers. Nature 465 (7295):182–187
Koch CM, Andrews RM, Flicek P, Dillon SC, Karaoz U, Clelland GK, Wilcox S, Beare DM, Fowler JC, Couttet P, James KD, Lefebvre GC, Bruce AW, Dovey OM, Ellis PD, Dhami P, Langford CF, Weng Z, Birney E, Carter NP, Vetrie D, Dunham I (2007) The landscape of histone modifications across 1% of the human genome in five human cell lines. Genome Res 17 (6):691–707
Visel A, Blow MJ, Li Z, Zhang T, Akiyama JA, Holt A, Plajzer-Frick I, Shoukry M, Wright C, Chen F, Afzal V, Ren B, Rubin EM, Pennacchio LA (2009) ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457 (7231):854–858
Ramos YF, Hestand MS, Verlaan M, Krabbendam E, Ariyurek Y, van Galen M, van Dam H, van Ommen GJ, den Dunnen JT, Zantema A, t Hoen PA (2010) Genome-wide assessment of differential roles for p300 and CBP in transcription regulation. Nucleic Acids Res 36 (16):5396–5408
Badis G, Berger MF, Philippakis AA, Talukder S, Gehrke AR, Jaeger SA, Chan ET, Metzler G, Vedenko A, Chen X, Kuznetsov H, Wang CF, Coburn D, Newburger DE, Morris Q, Hughes TR, Bulyk ML (2009) Diversity and complexity in DNA recognition by transcription factors. Science 324 (5935):1720–1723
Liu X, Lee CK, Granek JA, Clarke ND, Lieb JD (2006b) Whole-genome comparison of Leu3 binding in vitro and in vivo reveals the importance of nucleosome occupancy in target site selection. Genome Res 16 (12):1517–1528
Struhl K (1999) Fundamentally different logic of gene regulation in eukaryotes and prokaryotes. Cell 98 (1):1–4
Cheung V, Chua G, Batada NN, Landry CR, Michnick SW, Hughes TR, Winston F (2008) Chromatin- and transcription-related factors repress transcription from within coding regions throughout the Saccharomyces cerevisiae genome. PLoS Biol 6 (11):e277
Kaplan CD, Laprade L, Winston F (2003) Transcription elongation factors repress transcription initiation from cryptic sites. Science 301 (5636):1096–1099
Berman BP, Nibu Y, Pfeiffer BD, Tomancak P, Celniker SE, Levine M, Rubin GM, Eisen MB (2002) Exploiting transcription factor binding site clustering to identify cis-regulatory modules involved in pattern formation in the Drosophila genome. Proc Natl Acad Sci U S A 99 (2):757–762
Georges AB, Benayoun BA, Caburet S, Veitia RA (2010) Generic binding sites, generic DNA-binding domains: where does specific promoter recognition come from? Faseb J 24 (2):346–356
Papatsenko DA, Makeev VJ, Lifanov AP, Regnier M, Nazina AG, Desplan C (2002) Extraction of functional binding sites from unique regulatory regions: the Drosophila early developmental enhancers. Genome Res 12 (3):470–481
Valouev A, Ichikawa J, Tonthat T, Stuart J, Ranade S, Peckham H, Zeng K, Malek JA, Costa G, McKernan K, Sidow A, Fire A, Johnson SM (2008) A high-resolution, nucleosome position map of C. elegans reveals a lack of universal sequence-dictated positioning. Genome Res 18 (7):1051–1063
Kaplan N, Moore IK, Fondufe-Mittendorf Y, Gossett AJ, Tillo D, Field Y, LeProust EM, Hughes TR, Lieb JD, Widom J, Segal E (2009) The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458 (7236):362–366
Sekinger EA, Moqtaderi Z, Struhl K (2005) Intrinsic histone-DNA interactions and low nucleosome density are important for preferential accessibility of promoter regions in yeast. Mol Cell 18 (6):735–748
Anderson JD, Widom J (2001) Poly(dA-dT) promoter elements increase the equilibrium accessibility of nucleosomal DNA target sites. Mol Cell Biol 21 (11):3830–3839
Bao Y, White CL, Luger K (2006) Nucleosome core particles containing a poly(dA.dT) sequence element exhibit a locally distorted DNA structure. J Mol Biol 361 (4):617–624
Iyer V, Struhl K (1995) Poly(dA:dT), a ubiquitous promoter element that stimulates transcription via its intrinsic DNA structure. Embo J 14 (11):2570–2579
Suter B, Schnappauf G, Thoma F (2000) Poly(dA.dT) sequences exist as rigid DNA structures in nucleosome-free yeast promoters in vivo. Nucleic Acids Res 28 (21):4083–4089
Shimizu M, Mori T, Sakurai T, Shindo H (2000) Destabilization of nucleosomes by an unusual DNA conformation adopted by poly(dA) small middle dotpoly(dT) tracts in vivo. Embo J 19 (13):3358–3365
White CL, Luger K (2004) Defined structural changes occur in a nucleosome upon Amt1 transcription factor binding. J Mol Biol 342 (5):1391–1402
Zhu Z, Thiele DJ (1996) A specialized nucleosome modulates transcription factor access to a C. glabrata metal responsive promoter. Cell 87 (3):459–470
Field Y, Kaplan N, Fondufe-Mittendorf Y, Moore IK, Sharon E, Lubling Y, Widom J, Segal E (2008) Distinct modes of regulation by chromatin encoded through nucleosome positioning signals. PLoS Comput Biol 4 (11):e1000216
Peckham HE, Thurman RE, Fu Y, Stamatoyannopoulos JA, Noble WS, Struhl K, Weng Z (2007) Nucleosome positioning signals in genomic DNA. Genome Res 17 (8):1170–1177
Segal E, Fondufe-Mittendorf Y, Chen L, Thastrom A, Field Y, Moore IK, Wang JP, Widom J (2006) A genomic code for nucleosome positioning. Nature 442 (7104):772–778
Kaplan N, Moore I, Fondufe-Mittendorf Y, Gossett AJ, Tillo D, Field Y, Hughes TR, Lieb JD, Widom J, Segal E (2010) Nucleosome sequence preferences influence in vivo nucleosome organization. Nat Struct Mol Biol 17 (8):918–920
Mavrich TN, Ioshikhes IP, Venters BJ, Jiang C, Tomsho LP, Qi J, Schuster SC, Albert I, Pugh BF (2008b) A barrier nucleosome model for statistical positioning of nucleosomes throughout the yeast genome. Genome Res 18 (7):1073–1083
Zhang Y, Moqtaderi Z, Rattner BP, Euskirchen G, Snyder M, Kadonaga JT, Liu XS, Struhl K (2009) Intrinsic histone-DNA interactions are not the major determinant of nucleosome positions in vivo. Nat Struct Mol Biol 16 (8):847–852
Zhang Y, Moqtaderi Z, Rattner BP, Euskirchen G, Snyder M, Kadonaga JT, Liu XS, Struhl K (2010) Reply to “Evidence against a genomic code for nucleosome positioning”. Nat Struct Mol Biol 17 (8):920–923
Hartley PD, Madhani HD (2009) Mechanisms that specify promoter nucleosome location and identity. Cell 137 (3):445–458
Choi JK, Kim YJ (2009) Intrinsic variability of gene expression encoded in nucleosome positioning sequences. Nat Genet 41 (4):498–503
Tillo D, Kaplan N, Moore IK, Fondufe-Mittendorf Y, Gossett AJ, Field Y, Lieb JD, Widom J, Segal E, Hughes TR (2010) High nucleosome occupancy is encoded at human regulatory sequences. PLoS One 5 (2):e9129
Fu Y, Sinha M, Peterson CL, Weng Z (2008) The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome. PLoS Genet 4 (7):e1000138
Lidor Nili E, Field Y, Lubling Y, Widom J, Oren M, Segal E (2010) p53 binds preferentially to genomic regions with high DNA-encoded nucleosome occupancy. Genome Res 20 (10):1361–1368
Li G, Levitus M, Bustamante C, Widom J (2005) Rapid spontaneous accessibility of nucleosomal DNA. Nat Struct Mol Biol 12 (1):46–53
Li G, Widom J (2004) Nucleosomes facilitate their own invasion. Nat Struct Mol Biol 11 (8):763–769
Zlatanova J, Seebart C, Tomschik M (2008) The linker-protein network: control of nucleosomal DNA accessibility. Trends Biochem Sci 33 (6):247–253
Tomschik M, Zheng H, van Holde K, Zlatanova J, Leuba SH (2005) Fast, long-range, reversible conformational fluctuations in nucleosomes revealed by single-pair fluorescence resonance energy transfer. Proc Natl Acad Sci U S A 102 (9):3278–3283
Albert I, Mavrich TN, Tomsho LP, Qi J, Zanton SJ, Schuster SC, Pugh BF (2007) Translational and rotational settings of H2A.Z nucleosomes across the Saccharomyces cerevisiae genome. Nature 446 (7135):572–576
Jiang C, Pugh BF (2009) Nucleosome positioning and gene regulation: advances through genomics. Nat Rev Genet 10 (3):161–172
Koerber RT, Rhee HS, Jiang C, Pugh BF (2009) Interaction of transcriptional regulators with specific nucleosomes across the Saccharomyces genome. Mol Cell 35 (6):889–902
Polach KJ, Widom J (1995) Mechanism of protein access to specific DNA sequences in chromatin: a dynamic equilibrium model for gene regulation. J Mol Biol 254 (2):130–149
Adams CC, Workman JL (1995) Binding of disparate transcriptional activators to nucleosomal DNA is inherently cooperative. Mol Cell Biol 15 (3):1405–1421
Miller JA, Widom J (2003) Collaborative competition mechanism for gene activation in vivo. Mol Cell Biol 23 (5):1623–1632
Pettersson M, Schaffner W (1990) Synergistic activation of transcription by multiple binding sites for NF-kappa B even in absence of co-operative factor binding to DNA. J Mol Biol 214 (2):373–380
Vashee S, Melcher K, Ding WV, Johnston SA, Kodadek T (1998) Evidence for two modes of cooperative DNA binding in vivo that do not involve direct protein-protein interactions. Curr Biol 8 (8):452–458
Cirillo LA, Zaret KS (1999) An early developmental transcription factor complex that is more stable on nucleosome core particles than on free DNA. Mol Cell 4 (6):961–969
Cirillo LA, Lin FR, Cuesta I, Friedman D, Jarnik M, Zaret KS (2002) Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol Cell 9 (2):279–289
Pennings S, Meersseman G, Bradbury EM (1994) Linker histones H1 and H5 prevent the mobility of positioned nucleosomes. Proc Natl Acad Sci U S A 91 (22):10275–10279
Ura K, Hayes JJ, Wolffe AP (1995) A positive role for nucleosome mobility in the transcriptional activity of chromatin templates: restriction by linker histones. Embo J 14 (15):3752–3765
Holmqvist PH, Belikov S, Zaret KS, Wrange O (2005) FoxA1 binding to the MMTV LTR modulates chromatin structure and transcription. Exp Cell Res 304 (2):593–603
Li G, Margueron R, Hu G, Stokes D, Wang YH, Reinberg D (2010) Highly compacted chromatin formed in vitro reflects the dynamics of transcription activation in vivo. Mol Cell 38 (1):41–53
Lupien M, Eeckhoute J, Meyer CA, Wang Q, Zhang Y, Li W, Carroll JS, Liu XS, Brown M (2008) FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 132 (6):958–970
Plachetka A, Chayka O, Wilczek C, Melnik S, Bonifer C, Klempnauer KH (2008) C/EBPbeta induces chromatin opening at a cell-type-specific enhancer. Mol Cell Biol 28 (6):2102–2112
Dobi KC, Winston F (2007) Analysis of transcriptional activation at a distance in Saccharomyces cerevisiae. Mol Cell Biol 27 (15):5575–5586
Lee KK, Workman JL (2007) Histone acetyltransferase complexes: one size doesn’t fit all. Nat Rev Mol Cell Biol 8 (4):284–295
Perissi V, Jepsen K, Glass CK, Rosenfeld MG (2010) Deconstructing repression: evolving models of co-repressor action. Nat Rev Genet 11 (2):109–123
Glozak MA, Sengupta N, Zhang X, Seto E (2005) Acetylation and deacetylation of non-histone proteins. Gene 363:15–23
Underhill C, Qutob MS, Yee SP, Torchia J (2000) A novel nuclear receptor corepressor complex, N-CoR, contains components of the mammalian SWI/SNF complex and the corepressor KAP-1. J Biol Chem 275 (51):40463–40470
Dokmanovic M, Clarke C, Marks PA (2007) Histone deacetylase inhibitors: overview and perspectives. Mol Cancer Res 5 (10):981–989
Goldmark JP, Fazzio TG, Estep PW, Church GM, Tsukiyama T (2000) The Isw2 chromatin remodeling complex represses early meiotic genes upon recruitment by Ume6p. Cell 103 (3):423–433
Inayoshi Y, Kaneoka H, Machida Y, Terajima M, Dohda T, Miyake K, Iijima S (2005) Repression of GR-mediated expression of the tryptophan oxygenase gene by the SWI/SNF complex during liver development. J Biochem 138 (4):457–465
Murphy DJ, Hardy S, Engel DA (1999) Human SWI-SNF component BRG1 represses transcription of the c-fos gene. Mol Cell Biol 19 (4):2724–2733
Ooi L, Belyaev ND, Miyake K, Wood IC, Buckley NJ (2006) BRG1 chromatin remodeling activity is required for efficient chromatin binding by repressor element 1-silencing transcription factor (REST) and facilitates REST-mediated repression. J Biol Chem 281 (51):38974–38980
Sif S, Saurin AJ, Imbalzano AN, Kingston RE (2001) Purification and characterization of mSin3A-containing Brg1 and hBrm chromatin remodeling complexes. Genes Dev 15 (5):603–618
Moreira JM, Holmberg S (1999) Transcriptional repression of the yeast CHA1 gene requires the chromatin-remodeling complex RSC. Embo J 18 (10):2836–2844
Buck MJ, Lieb JD (2006) A chromatin-mediated mechanism for specification of conditional transcription factor targets. Nat Genet 38 (12):1446–1451
Keleher CA, Redd MJ, Schultz J, Carlson M, Johnson AD (1992) Ssn6-Tup1 is a general repressor of transcription in yeast. Cell 68 (4):709–719
De Vit MJ, Waddle JA, Johnston M (1997) Regulated nuclear translocation of the Mig1 glucose repressor. Mol Biol Cell 8 (8):1603–1618
Park SH, Koh SS, Chun JH, Hwang HJ, Kang HS (1999) Nrg1 is a transcriptional repressor for glucose repression of STA1 gene expression in Saccharomyces cerevisiae. Mol Cell Biol 19 (3):2044–2050
Proft M, Serrano R (1999) Repressors and upstream repressing sequences of the stress-regulated ENA1 gene in Saccharomyces cerevisiae: bZIP protein Sko1p confers HOG-dependent osmotic regulation. Mol Cell Biol 19 (1):537–546
Tzamarias D, Struhl K (1994) Functional dissection of the yeast Cyc8-Tup1 transcriptional co-repressor complex. Nature 369 (6483):758–761
Davie JK, Edmondson DG, Coco CB, Dent SY (2003) Tup1-Ssn6 interacts with multiple class I histone deacetylases in vivo. J Biol Chem 278 (50):50158–50162
Watson AD, Edmondson DG, Bone JR, Mukai Y, Yu Y, Du W, Stillman DJ, Roth SY (2000) Ssn6-Tup1 interacts with class I histone deacetylases required for repression. Genes Dev 14 (21):2737–2744
Lieb JD, Liu X, Botstein D, Brown PO (2001) Promoter-specific binding of Rap1 revealed by genome-wide maps of protein-DNA association. Nat Genet 28 (4):327–334
Shore D (1994) RAP1: a protean regulator in yeast. Trends Genet 10 (11):408–412
Kurdistani SK, Robyr D, Tavazoie S, Grunstein M (2002) Genome-wide binding map of the histone deacetylase Rpd3 in yeast. Nat Genet 31 (3):248–254
Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W, Zhao K (2009) Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 138 (5):1019–1031
Wang A, Kurdistani SK, Grunstein M (2002) Requirement of Hos2 histone deacetylase for gene activity in yeast. Science 298 (5597):1412–1414
Nagaich AK, Walker DA, Wolford R, Hager GL (2004) Rapid periodic binding and displacement of the glucocorticoid receptor during chromatin remodeling. Mol Cell 14 (2):163–174
Auble DT, Hansen KE, Mueller CG, Lane WS, Thorner J, Hahn S (1994) Mot1, a global repressor of RNA polymerase II transcription, inhibits TBP binding to DNA by an ATP-dependent mechanism. Genes Dev 8 (16):1920–1934
van Werven FJ, van Bakel H, van Teeffelen HA, Altelaar AF, Koerkamp MG, Heck AJ, Holstege FC, Timmers HT (2008) Cooperative action of NC2 and Mot1p to regulate TATA-binding protein function across the genome. Genes Dev 22 (17):2359–2369
Li G, Chandler SP, Wolffe AP, Hall TC (1998) Architectural specificity in chromatin structure at the TATA box in vivo: nucleosome displacement upon beta-phaseolin gene activation. Proc Natl Acad Sci U S A 95 (8):4772–4777
Grewal SI, Jia S (2007) Heterochromatin revisited. Nat Rev Genet 8 (1):35–46
James TC, Elgin SC (1986) Identification of a nonhistone chromosomal protein associated with heterochromatin in Drosophila melanogaster and its gene. Mol Cell Biol 6 (11):3862–3872
Ebert A, Lein S, Schotta G, Reuter G (2006) Histone modification and the control of heterochromatic gene silencing in Drosophila. Chromosome Res 14 (4):377–392
Nielsen SJ, Schneider R, Bauer UM, Bannister AJ, Morrison A, O’Carroll D, Firestein R, Cleary M, Jenuwein T, Herrera RE, Kouzarides T (2001) Rb targets histone H3 methylation and HP1 to promoters. Nature 412 (6846):561–565
O’Geen H, Squazzo SL, Iyengar S, Blahnik K, Rinn JL, Chang HY, Green R, Farnham PJ (2007) Genome-wide analysis of KAP1 binding suggests autoregulation of KRAB-ZNFs. PLoS Genet 3 (6):e89
Vogel MJ, Guelen L, de Wit E, Peric-Hupkes D, Loden M, Talhout W, Feenstra M, Abbas B, Classen AK, van Steensel B (2006) Human heterochromatin proteins form large domains containing KRAB-ZNF genes. Genome Res 16 (12):1493–1504
Schwartz YB, Pirrotta V (2007) Polycomb silencing mechanisms and the management of genomic programmes. Nat Rev Genet 8 (1):9–22
Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS, Kumar RM, Chevalier B, Johnstone SE, Cole MF, Isono K, Koseki H, Fuchikami T, Abe K, Murray HL, Zucker JP, Yuan B, Bell GW, Herbolsheimer E, Hannett NM, Sun K, Odom DT, Otte AP, Volkert TL, Bartel DP, Melton DA, Gifford DK, Jaenisch R, Young RA (2006) Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 125 (2):301–313
Schwartz YB, Kahn TG, Nix DA, Li XY, Bourgon R, Biggin M, Pirrotta V (2006) Genome-wide analysis of Polycomb targets in Drosophila melanogaster. Nat Genet 38 (6):700–705
Squazzo SL, O’Geen H, Komashko VM, Krig SR, Jin VX, Jang SW, Margueron R, Reinberg D, Green R, Farnham PJ (2006) Suz12 binds to silenced regions of the genome in a cell-type-specific manner. Genome Res 16 (7):890–900
Fanti L, Pimpinelli S (2008) HP1: a functionally multifaceted protein. Curr Opin Genet Dev 18 (2):169–174
Muller J, Verrijzer P (2009) Biochemical mechanisms of gene regulation by polycomb group protein complexes. Curr Opin Genet Dev 19 (2):150–158
Simon JA, Kingston RE (2009) Mechanisms of polycomb gene silencing: knowns and unknowns. Nat Rev Mol Cell Biol 10 (10):697–708
Dorer DR, Henikoff S (1994) Expansions of transgene repeats cause heterochromatin formation and gene silencing in Drosophila. Cell 77 (7):993–1002
Luff B, Pawlowski L, Bender J (1999) An inverted repeat triggers cytosine methylation of identical sequences in Arabidopsis. Mol Cell 3 (4):505–511
Selker EU (2002) Repeat-induced gene silencing in fungi. Adv Genet 46:439–450
Hall IM, Shankaranarayana GD, Noma K, Ayoub N, Cohen A, Grewal SI (2002) Establishment and maintenance of a heterochromatin domain. Science 297 (5590):2232–2237
Volpe TA, Kidner C, Hall IM, Teng G, Grewal SI, Martienssen RA (2002) Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297 (5588):1833–1837
Grewal SI (2010) RNAi-dependent formation of heterochromatin and its diverse functions. Curr Opin Genet Dev 20 (2):134–141
Yamada T, Fischle W, Sugiyama T, Allis CD, Grewal SI (2005) The nucleation and maintenance of heterochromatin by a histone deacetylase in fission yeast. Mol Cell 20 (2):173–185
Jia S, Noma K, Grewal SI (2004) RNAi-independent heterochromatin nucleation by the stress-activated ATF/CREB family proteins. Science 304 (5679):1971–1976
Kim HS, Choi ES, Shin JA, Jang YK, Park SD (2004) Regulation of Swi6/HP1-dependent heterochromatin assembly by cooperation of components of the mitogen-activated protein kinase pathway and a histone deacetylase Clr6. J Biol Chem 279 (41):42850–42859
Rusche LN, Kirchmaier AL, Rine J (2003) The establishment, inheritance, and function of silenced chromatin in Saccharomyces cerevisiae. Annu Rev Biochem 72:481–516
Abrink M, Ortiz JA, Mark C, Sanchez C, Looman C, Hellman L, Chambon P, Losson R (2001) Conserved interaction between distinct Kruppel-associated box domains and the transcriptional intermediary factor 1 beta. Proc Natl Acad Sci U S A 98 (4):1422–1426
Peng H, Begg GE, Schultz DC, Friedman JR, Jensen DE, Speicher DW, Rauscher FJ, 3rd (2000) Reconstitution of the KRAB-KAP-1 repressor complex: a model system for defining the molecular anatomy of RING-B box-coiled-coil domain-mediated protein-protein interactions. J Mol Biol 295 (5):1139–1162
Lechner MS, Begg GE, Speicher DW, Rauscher FJ, 3rd (2000) Molecular determinants for targeting heterochromatin protein 1-mediated gene silencing: direct chromoshadow domain-KAP-1 corepressor interaction is essential. Mol Cell Biol 20 (17):6449–6465
Schultz DC, Ayyanathan K, Negorev D, Maul GG, Rauscher FJ, 3rd (2002) SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16 (8):919–932
Schultz DC, Friedman JR, Rauscher FJ, 3rd (2001) Targeting histone deacetylase complexes via KRAB-zinc finger proteins: the PHD and bromodomains of KAP-1 form a cooperative unit that recruits a novel isoform of the Mi-2alpha subunit of NuRD. Genes Dev 15 (4):428–443
Ayyanathan K, Lechner MS, Bell P, Maul GG, Schultz DC, Yamada Y, Tanaka K, Torigoe K, Rauscher FJ, 3rd (2003) Regulated recruitment of HP1 to a euchromatic gene induces mitotically heritable, epigenetic gene silencing: a mammalian cell culture model of gene variegation. Genes Dev 17 (15):1855–1869
Groner AC, Meylan S, Ciuffi A, Zangger N, Ambrosini G, Denervaud N, Bucher P, Trono D (2010) KRAB-zinc finger proteins and KAP1 can mediate long-range transcriptional repression through heterochromatin spreading. PLoS Genet 6 (3):e1000869
Brown JL, Fritsch C, Mueller J, Kassis JA (2003) The Drosophila pho-like gene encodes a YY1-related DNA binding protein that is redundant with pleiohomeotic in homeotic gene silencing. Development 130 (2):285–294
Fritsch C, Brown JL, Kassis JA, Muller J (1999) The DNA-binding polycomb group protein pleiohomeotic mediates silencing of a Drosophila homeotic gene. Development 126 (17):3905–3913
Klymenko T, Papp B, Fischle W, Kocher T, Schelder M, Fritsch C, Wild B, Wilm M, Muller J (2006) A Polycomb group protein complex with sequence-specific DNA-binding and selective methyl-lysine-binding activities. Genes Dev 20 (9):1110–1122
Wang L, Brown JL, Cao R, Zhang Y, Kassis JA, Jones RS (2004) Hierarchical recruitment of polycomb group silencing complexes. Mol Cell 14 (5):637–646
Dejardin J, Rappailles A, Cuvier O, Grimaud C, Decoville M, Locker D, Cavalli G (2005) Recruitment of Drosophila Polycomb group proteins to chromatin by DSP1. Nature 434 (7032):533–538
Ringrose L, Rehmsmeier M, Dura JM, Paro R (2003) Genome-wide prediction of Polycomb/Trithorax response elements in Drosophila melanogaster. Dev Cell 5 (5):759–771
Boyer LA, Plath K, Zeitlinger J, Brambrink T, Medeiros LA, Lee TI, Levine SS, Wernig M, Tajonar A, Ray MK, Bell GW, Otte AP, Vidal M, Gifford DK, Young RA, Jaenisch R (2006) Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441 (7091):349–353
Ku M, Koche RP, Rheinbay E, Mendenhall EM, Endoh M, Mikkelsen TS, Presser A, Nusbaum C, Xie X, Chi AS, Adli M, Kasif S, Ptaszek LM, Cowan CA, Lander ES, Koseki H, Bernstein BE (2008) Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet 4 (10):e1000242
Sing A, Pannell D, Karaiskakis A, Sturgeon K, Djabali M, Ellis J, Lipshitz HD, Cordes SP (2009) A vertebrate Polycomb response element governs segmentation of the posterior hindbrain. Cell 138 (5):885–897
Woo CJ, Kharchenko PV, Daheron L, Park PJ, Kingston RE (2010) A region of the human HOXD cluster that confers polycomb-group responsiveness. Cell 140 (1):99–110
Caretti G, Di Padova M, Micales B, Lyons GE, Sartorelli V (2004) The Polycomb Ezh2 methyltransferase regulates muscle gene expression and skeletal muscle differentiation. Genes Dev 18 (21):2627–2638
Garcia E, Marcos-Gutierrez C, del Mar Lorente M, Moreno JC, Vidal M (1999) RYBP, a new repressor protein that interacts with components of the mammalian Polycomb complex, and with the transcription factor YY1. Embo J 18 (12):3404–3418
Kim SY, Paylor SW, Magnuson T, Schumacher A (2006) Juxtaposed Polycomb complexes co-regulate vertebral identity. Development 133 (24):4957–4968
Satijn DP, Hamer KM, den Blaauwen J, Otte AP (2001) The polycomb group protein EED interacts with YY1, and both proteins induce neural tissue in Xenopus embryos. Mol Cell Biol 21 (4):1360–1369
Endoh M, Endo TA, Endoh T, Fujimura Y, Ohara O, Toyoda T, Otte AP, Okano M, Brockdorff N, Vidal M, Koseki H (2008) Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development 135 (8):1513–1524
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY (2007) Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129 (7):1311–1323
Talbert PB, Henikoff S (2006) Spreading of silent chromatin: inaction at a distance. Nat Rev Genet 7 (10):793–803
Nakayama J, Rice JC, Strahl BD, Allis CD, Grewal SI (2001) Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292 (5514):110–113
Hoppe GJ, Tanny JC, Rudner AD, Gerber SA, Danaie S, Gygi SP, Moazed D (2002) Steps in assembly of silent chromatin in yeast: Sir3-independent binding of a Sir2/Sir4 complex to silencers and role for Sir2-dependent deacetylation. Mol Cell Biol 22 (12):4167–4180
Rusche LN, Kirchmaier AL, Rine J (2002) Ordered nucleation and spreading of silenced chromatin in Saccharomyces cerevisiae. Mol Biol Cell 13 (7):2207–2222
Boscheron C, Maillet L, Marcand S, Tsai-Pflugfelder M, Gasser SM, Gilson E (1996) Cooperation at a distance between silencers and proto-silencers at the yeast HML locus. Embo J 15 (9):2184–2195
Fourel G, Revardel E, Koering CE, Gilson E (1999) Cohabitation of insulators and silencing elements in yeast subtelomeric regions. Embo J 18 (9):2522–2537
Beisel C, Buness A, Roustan-Espinosa IM, Koch B, Schmitt S, Haas SA, Hild M, Katsuyama T, Paro R (2007) Comparing active and repressed expression states of genes controlled by the Polycomb/Trithorax group proteins. Proc Natl Acad Sci U S A 104 (42):16615–16620
Holohan EE, Kwong C, Adryan B, Bartkuhn M, Herold M, Renkawitz R, Russell S, White R (2007) CTCF genomic binding sites in Drosophila and the organisation of the bithorax complex. PLoS Genet 3 (7):e112
Negre N, Hennetin J, Sun LV, Lavrov S, Bellis M, White KP, Cavalli G (2006) Chromosomal distribution of PcG proteins during Drosophila development. PLoS Biol 4 (6):e170
Tolhuis B, de Wit E, Muijrers I, Teunissen H, Talhout W, van Steensel B, van Lohuizen M (2006) Genome-wide profiling of PRC1 and PRC2 Polycomb chromatin binding in Drosophila melanogaster. Nat Genet 38 (6):694–699
Kahn TG, Schwartz YB, Dellino GI, Pirrotta V (2006) Polycomb complexes and the propagation of the methylation mark at the Drosophila Ubx gene. J Biol Chem 281 (39):29064–29075
Lanzuolo C, Roure V, Dekker J, Bantignies F, Orlando V (2007) Polycomb response elements mediate the formation of chromosome higher-order structures in the bithorax complex. Nat Cell Biol 9 (10):1167–1174
Tiwari VK, Cope L, McGarvey KM, Ohm JE, Baylin SB (2008) A novel 6C assay uncovers Polycomb-mediated higher order chromatin conformations. Genome Res 18 (7):1171–1179
Molto E, Fernandez A, Montoliu L (2009) Boundaries in vertebrate genomes: different solutions to adequately insulate gene expression domains. Brief Funct Genomic Proteomic 8 (4):283–296
Wallace JA, Felsenfeld G (2007) We gather together: insulators and genome organization. Curr Opin Genet Dev 17 (5):400–407
West AG, Gaszner M, Felsenfeld G (2002) Insulators: many functions, many mechanisms. Genes Dev 16 (3):271–288
Bell AC, West AG, Felsenfeld G (1999) The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98 (3):387–396
Huang S, Li X, Yusufzai TM, Qiu Y, Felsenfeld G (2007) USF1 recruits histone modification complexes and is critical for maintenance of a chromatin barrier. Mol Cell Biol 27 (22):7991–8002
Gurudatta BV, Corces VG (2009) Chromatin insulators: lessons from the fly. Brief Funct Genomic Proteomic 8 (4):276–282
Recillas-Targa F, Pikaart MJ, Burgess-Beusse B, Bell AC, Litt MD, West AG, Gaszner M, Felsenfeld G (2002) Position-effect protection and enhancer blocking by the chicken beta-globin insulator are separable activities. Proc Natl Acad Sci U S A 99 (10):6883–6888
West AG, Huang S, Gaszner M, Litt MD, Felsenfeld G (2004) Recruitment of histone modifications by USF proteins at a vertebrate barrier element. Mol Cell 16 (3):453–463
Gaszner M, Felsenfeld G (2006) Insulators: exploiting transcriptional and epigenetic mechanisms. Nat Rev Genet 7 (9):703–713
Phillips JE, Corces VG (2009) CTCF: master weaver of the genome. Cell 137 (7):1194–1211
Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, de Klein A, Wessels L, de Laat W, van Steensel B (2008) Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453 (7197):948–951
Park PJ (2009) ChIP-seq: advantages and challenges of a maturing technology. Nat Rev Genet 10 (10):669–680
Ren B, Robert F, Wyrick JJ, Aparicio O, Jennings EG, Simon I, Zeitlinger J, Schreiber J, Hannett N, Kanin E, Volkert TL, Wilson CJ, Bell SP, Young RA (2000) Genome-wide location and function of DNA binding proteins. Science 290 (5500):2306–2309
Dejardin J, Kingston RE (2009) Purification of proteins associated with specific genomic Loci. Cell 136 (1):175–186
Celniker SE, Dillon LA, Gerstein MB, Gunsalus KC, Henikoff S, Karpen GH, Kellis M, Lai EC, Lieb JD, MacAlpine DM, Micklem G, Piano F, Snyder M, Stein L, White KP, Waterston RH (2009) Unlocking the secrets of the genome. Nature 459 (7249):927–930
Ravasi T, Suzuki H, Cannistraci CV, Katayama S, Bajic VB, Tan K, Akalin A, Schmeier S, Kanamori-Katayama M, Bertin N, Carninci P, Daub CO, Forrest AR, Gough J, Grimmond S, Han JH, Hashimoto T, Hide W, Hofmann O, Kamburov A, Kaur M, Kawaji H, Kubosaki A, Lassmann T, van Nimwegen E, MacPherson CR, Ogawa C, Radovanovic A, Schwartz A, Teasdale RD, Tegner J, Lenhard B, Teichmann SA, Arakawa T, Ninomiya N, Murakami K, Tagami M, Fukuda S, Imamura K, Kai C, Ishihara R, Kitazume Y, Kawai J, Hume DA, Ideker T, Hayashizaki Y (2010) An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140 (5):744–752
van Berkum NL, Lieberman-Aiden E, Williams L, Imakaev M, Gnirke A, Mirny LA, Dekker J, Lander ES (2010) Hi-C: a method to study the three-dimensional architecture of genomes. J Vis Exp (39):1869
Vassetzky Y, Gavrilov A, Eivazova E, Priozhkova I, Lipinski M, Razin S (2009) Chromosome conformation capture (from 3C to 5C) and its ChIP-based modification. Methods Mol Biol 567:171–188
Segal E, Widom J (2009b) From DNA sequence to transcriptional behaviour: a quantitative approach. Nat Rev Genet 10 (7):443–456
Strohner R, Wachsmuth M, Dachauer K, Mazurkiewicz J, Hochstatter J, Rippe K, Langst G (2005) A ‘loop recapture’ mechanism for ACF-dependent nucleosome remodeling. Nat Struct Mol Biol 12 (8):683–690
Agalioti T, Lomvardas S, Parekh B, Yie J, Maniatis T, Thanos D (2000) Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 103 (4):667–678
Chaban Y, Ezeokonkwo C, Chung WH, Zhang F, Kornberg RD, Maier-Davis B, Lorch Y, Asturias FJ (2008) Structure of a RSC-nucleosome complex and insights into chromatin remodeling. Nat Struct Mol Biol 15 (12):1272–1277
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Glossary
- Chromatin
-
The combination of DNA and accessory proteins, such as histones, that together constitute chromosomes.
- Transcriptional coregulator
-
An accessory factor recruited by transcription factors to modulate gene expression. Cofactors typically lack intrinsic DNA binding specificity and rely on transcription factors for targeting. Most cofactors excert their effects by locally modifying chromatin structure.
- Transcriptional coactivator
-
A coregulator that positively affects gene expression.
- Transcriptional corepressor
-
A coregulator that negatively affects gene expression.
- Chromatin modifiers
-
Proteins or protein complexes that can effect changes in chromatin structure by covalently modifying histones or moving nucleosomes. In this chapter the term chromatin modifier is used generally to refer to histone modifiers and ATPase nucleosome remodelers.
- Histone modifiers
-
The enzymes responsible for adding or removing covalent modifications on histones, the majority of which are are found on the flexible histone tails. Some histone modifiers, such as HDACs and HATs can also have non-histone targets.
- ATPase nucleosome remodelers
-
Protein complexes that use the energy generated by ATP hydrolysis to alter nucleosome-DNA interactions and displace nucleosomes.
- Heterochromatin
-
A tightly packed form of chromatin where DNA is typically rendered inaccessible to the transcriptional machinery. Different types of heterochromatin are associated with distinct chromatin marks, such as HP1 heterochromatin (HP1 binding and H3K9me) or Polycomb domains (H3K27me).
- Euchromatin
-
An open chromatin conformation in which DNA is easily accessible. This type of chromatin is often, but not exclusively, associated with active transcription.
- Histone code
-
Distinct patterns of histone modifications are believed to constitute a code that is used to direct specific activities on DNA, such as during transcriptional silencing or during the various stages of the transcriptional cycle. For example, the initiation, elongation and termination of transcription are each associated with different patterns of histone modifications that are believed to contribute to the recruitment and regulation of the proteins required in each stage.
- Epigenetics
-
Inherited changes in phenotypes or expression profiles that are not due to changes in the underlying DNA sequence. Examples of epigenetic modifications include DNA methylation and covalent histone modifications, which play an important role in a variety of processes, including cell differentiation, X chromosome inactivation and imprinting.
- Polycomb-group proteins
-
A family of proteins, initially discovered in Drosophila, that are involved in epigenetic silencing of genes by inducing a repressive chromatin structure. Polycomb group proteins are predominantly found as part of two main protein complexes: Polycomb-group Repressive Complex 1 and 2 (PRC1 and PRC2).
- Nucleosome
-
The basic building block of chromatin, consisting of ~147 bp of DNA wrapped around an octamer of two of each of the histones H2A, H2B, H3 and H4.
- Effector domains
-
The domains in transcription factors that are responsible for mediating their effects on gene expression. These effects can be activating or inhibitory and involve a variety of mechanisms, including recruitment of chromatin modifiers, or interactions with components of the basal transcriptional machinery and other transcription factors.
- DNA binding domain
-
A protein domain with DNA binding activity. In the case of transcription factors, these domains typically possess specificity affinity for a limited number of DNA sequences.
- Enhancer
-
A DNA element bound by transcription factors that can operate over long distances (up to thousands of basepairs) to stimulate transcription of its target gene(s). Enhancers are thought to operate through looping interactions with promoter regions. In addition to their distance to genes, enhancers can also be distinguished from promoters by a unique chromatin profile. Though most enhancers act in cis, they can also be located on different chromosomes.
- Silencer
-
Like enhancers, silencers are DNA elements that can be located far away from the genes they control, but their effect on gene expression is negative. Silencers can also act as nucleation sites for repressive chromatin domains.
- Insulator
-
A DNA element that either prevents an enhancer from activating target genes, or acts as a boundary element to delineate different chromatin domains. Insulators are distinct from from silencer regions in that an insulator needs to be located between an enhancer and a gene to affect expression, while silencers can typically operate in any orientation relative to a gene.
- Chromatin domain
-
A relatively uniform region of chromatin characterized by distinct histone and/or DNA modifications. Examples include Polycomb domains as well as telomeric- and pericentromeric heterochromatin.
- Preinitiation complex
-
Large complex of proteins required for successful transcription initiation by RNA Polymerase II. Major components include the basal transcription factors TFIIA, TFIIB, TFIID, TFIIE, TFIIF, and TFIIH. The preinitiation complex plays a role in positioning polymerase and melting the DNA so that it is properly configured to fit in the active site. Positioning is aided by motifs recognized by the general transcription factors.
- CpG island
-
Sequence elements rich in CG dinucleotides that are found at a large number of mammalian promoters.
- General transcription factors
-
Transcription factors that are universally required for RNA polymerase II transcription. Most GTFs are part of the preinitiation complex.
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media B.V.
About this chapter
Cite this chapter
van Bakel, H. (2011). Interactions of Transcription Factors with Chromatin. In: Hughes, T. (eds) A Handbook of Transcription Factors. Subcellular Biochemistry, vol 52. Springer, Dordrecht. https://doi.org/10.1007/978-90-481-9069-0_11
Download citation
DOI: https://doi.org/10.1007/978-90-481-9069-0_11
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-90-481-9068-3
Online ISBN: 978-90-481-9069-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)