Abstract
Friedreich ataxia (FRDA) is an autosomal recessive progressively debilitating degenerative disease that principally affects the nervous system and the heart. Although FRDA is considered a rare disease, is the most common inherited ataxia. It is caused by loss-of-function mutations in the FXN gene, mainly an expanded GAA triplet repeat in the intron 1. The genetic defect results in the reduction of frataxin levels, a protein targeted to the mitochondria. Frataxin deficiency leads to mitochondrial dysfunction, oxidative damage and iron accumulation. Studies of the yeast and animal models of the disease have led to propose several different roles for frataxin. Animal models have also been important for dissecting the steps of pathogenesis in FRDA and they are essential for the development of effective therapies. Currently, antioxidant and iron chelation therapies are under evaluation in clinical trials. Gene reactivation, gene therapy and protein replacement strategies for FRDA are promising approaches.
This review focuses on the current models developed for FRDA, the different roles proposed for frataxin and the progress of potential treatment strategies for the disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Friedreich ataxia
- Frataxin
- Mitochondria
- Iron-sulfur clusters
- Oxidative stress
- Oxidative phosphorylation
- Antioxidant therapy
- Iron chelators
- Recombinant human erythropoietin
- Histone deacetylase inhibitors
17.1 Introduction
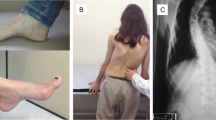
In 1863, Nicholaus Friedreich described a new disease that was given the name of Friedreich Ataxia (FRDA, MIM 229300) [1, 2]. FRDA is the most common hereditary ataxia in the Caucasian population. Specifically, in Spain the prevalence is about 4.7 cases per 100,000 habitants [3, 4]. FRDA is characterized by progressive sensory ataxia with onset typically before 25 years of age (cases with later onset are not uncommon), areflexia, dysarthria, extensor plantar responses and loss of position and vibration sense [5]. Patients have primary degeneration of dorsal root ganglia associated with axonal degeneration of the posterior columns, spinocerebellar tracts, and corticospinal tracts, and large myelinated fibers in peripheral nerves. Most individuals with FRDA develop a cardiomyopathy and diabetes mellitus.
17.2 Friedreich Ataxia Gene and Molecular Pathology
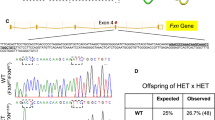
Friedreich ataxia is an autosomal recessive degenerative disorder caused by a GAA triplet repeat expansion or point mutations in the FXN gene on chromosome 9q13 [6, 7]. The FXN gene product, frataxin, is a widely expressed mitocondrial protein that is severely reduced in FRDA patients [6].
Most patients with FRDA (approximately 97%) have expansions of a GAA repeat in the first intron of both FXN alleles [6, 8, 9]. The GAA repeat expansions reduce the levels of messenger RNA by the formation of non-B DNA structures (triplexes or sticky DNA), the formation of a persistent DNA × RNA hybrid, or heterochromatin formation [10–14]. Individuals with FRDA have also been identified (aprox. 3%) who carry one allele with a trinucleotide repeat expansion and the other allele with a point mutation [8, 10, 11, 15–26].
The nuclear encoded human frataxin protein is synthesized as a 210 amino acid precursor with a N-terminal mitochondrial targeting sequence where it is proteolytically cleaved by the mitochondrial processing peptidase (MPP) to the mature form via a processing intermediate [27, 28]. The initial characterization demonstrated that the endogenous mature form (m-FXN) was encoded by amino acids 56-210. However, there are evidences that endogenous mature frataxin corresponds to amino acids 81-210 [29].
The structure of human frataxin reveals a novel protein fold. A five-stranded, antiparallel β sheets provides a flat platform, which supports a pair of parallel α-helix, to form a compact αβ sandwich [30]. A cluster of 12 acidic residues from the first helix and the first strand of the large sheet form a contiguous anionic surface on the protein. Many of these amino acids are necessary for stabilizing the structure and can not be replaced, being conserved in the different species and appear mutated in several patients. These structural findings predict potential modes of protein-protein and protein iron binding.
17.3 Animal Models in Friedreich Ataxia
Frataxin is a mitochondrial protein which shows high conservation throughout evolution, with orthologs in essentially all eukaryotes and some prokaryotes [31]. Many studies in model organisms as yeast, mice, Drosophila and C. elegans contribute extensively to our current understanding.
In yeast, the deletion of the YFH1 gene, showed a strain unable to grow on a non-fermentable source of carbon suggesting that it could not carry out oxidative phosphorylation. The strain yfh1Δ, showed mitochondrial iron accumulation (up to 10 times more than the wild strain) and exhibited sensitivity to oxidative stress [32–36]. Frataxin can substitute for Yfh1p in yeast, indicating that the two proteins are functional as well as structural homologs [36, 37].
Generation of fly models of FRDA has been possible after the identification of the Drosophila frataxin homolog, fh, [38]. Systemic and tissue-specific down-regulation of fh has been carried out combining the UAS-GAL4 system and RNA interference (RNAi). Moderate systemic reduction of fh expression to levels of 30% of normal fh-mRNA, a situation which strikingly parallel with Friedreich ataxia, is compatible with normal embryonic development in fly [39]. However in such conditions, adults show a decrease of mean and maximum life span of 60 and 32% respectively, and a reduction in climbing abilities. Meanwhile, widespread fh silencing to undetectable levels results in lethality during development and larvae are incapable of reaching the adult phase [39, 40]. Silencing of fh in early mesoderm also provokes lethality in agreement with the essential role for frataxin during development, previously proposed in a mouse model [41]. At the biochemical level, almost completely depletion of Drosophila frataxin reduces activities of mitochondrial Fe-S containing enzymes, and confers hypersensitivity to iron and to hyperoxia [39, 40]. However aconitase is the only Fe-S enzyme which shows a dramatic reduction in its activity under hyperoxia in flies with moderate reduction of frataxin. Meanwhile the respiratory chain complexes operate normally in this condition. Therefore the decay in aconitase activity by reactive oxygen species (ROS) inactivation would be one of the primary events in frataxin deficient condition. In turn, aconitase inactivation would induce further oxidative stress by different mechanisms [42, 43] that might affect other macromolecules, such as the Fe-S enzymes of the respiratory chain. Frataxin would act as an aconitase chaperone protecting this enzyme against ROS-mediated inactivation supporting a causative role of oxidative stress in Friedreich ataxia. In addition, overexpression of the H2O2-scavenging enzymes (peroxisomal and mitochondrial catalases and a mitochondrial peroxiredoxin) rescues the shortened life span in frataxin deficient flies [44]. Expressions of these enzymes also alleviate the hypersensibility to H2O2 shown by flies showing deficiency of frataxin in the peripheral nervous system. These data suggest that H2O2 is an important pathogenic substrate in FRDA with promising therapeutic implications.
As expected overexpression of Drosophila frataxin improve antioxidant capability, resistance to oxidative stress insults, and longevity in flies [45]. However a 9-fold of normal level of Drosophila frataxin is lethal in flies [39]. Moreover, the excess of frataxin induces defects in Drosophila by oxidative-mediated inactivation of mitochondrial aconitase. Frataxin overload might induce new in vivo function for this protein, by limiting iron availability conducting to aconitase dysfunction. Frataxin has been shown to form in vitro iron-binding multimers as ferritin [46]. Alternatively, an excess of frataxin might saturate interactions with its partners leading to mitochondrial dysfunction. The pathogenic effect of frataxin excess in Drosophila has essential repercussion in gene therapy.
Several transient knock-down models of C. elegans frataxin homologue gene, frh-1, by RNAi have been reported. Some authors have observed increase in lifespan when frataxin was knocking-down by RNAi [47]. However, microinjection in L4 larvae showed lifespan significantly reduced and worms have increased sensitivity to oxidative stress that, in turn, might explain the reduction of worms’ longevity [48]. These worms showed a consistent pleiotropic phenotype that includes lethargic behaviour, reduced broodsize, egg laying defects, slow growth, abnormal pharyngeal pumping and altered defecation. In a third study, reduction of life expectancy in knock-down worms was confirmed and lifespan was positively correlated with impaired respiration [49]. This frataxin deficient model in C. elegans may be a useful biological tool for drug screening in FRDA and to investigate the role of frataxin in biochemical pathways.
Generation of a representative mouse model of FRDA is considered important for the further understanding of disease pathology and the testing of potential therapeutic strategies. Initially, a homozygous knockout of the murine FRDA gene leads to early embryonic lethality without evidence of iron accumulation [41]. This mouse has shown that frataxin plays an important role during early embryonic development. As an attempt to generate a mouse model of the disease, it was generated a striated muscle frataxin deficient line and a neuron/cardiac muscle frataxin deficient line through a conditional gene targeting approach [50]. These mice reproduce the most important pathophysiological and biochemical features of the human disease. To obtain specific and progressive neurological models for FRDA, Puccio and collegues generated inducible knock-out mouse models [51]. They showed that ablation of frataxin in adult mice led to progressive neurological symptoms resembling FRDA. These mutants have specific damage to the large sensory neuron cell bodies in the dorsal root ganglia, and they have degeneration of the posterior columns of spinal cord that appear translucent, because of demyelination and loss of fibers, and severe lesions of neurons in Clarke’s columns. Because FRDA is typically caused by a large GAA repeat expansion within the first intron of the FXN gene, several mouse models harbouring this mutation have been created. One of them is a knockin mouse that carries a (GAA)230 repeat expansion in the first intron of the mouse frataxin homolog gene (Frda) [52]. GAA repeat knockin mice were crossed with frataxin knockout mice to obtain double heterozygous mice expressing 25–36% of wild-type frataxin levels. These mice were viable and did not develop anomalies of motor coordination, iron metabolism or response to iron loading. In addition, repeats were meiotically and mitotically stable. As well, it was generated two lines of human FXN YAC transgenic mice containing GAA repeat expansions derived from FRDA patient DNA [53]. Offspring from crosses between these transgenic mice and heterozygous Fxn-knockout mice [54] express comparatively reduced levels of human frataxin and rescue the homozygous Fxn-knockout embryonic lethality. These mice represent the first GAA repeat expansion-based FRDA mouse models that exhibit progressive FRDA-like pathology. Moreover, transgenic mice carrying expanded human FXN GAA repeats (190 or 82 triplets) showed tissue-specific and age-dependent somatic instability specifically in the cerebellum and dorsal root ganglia [55].
17.4 The Function of Frataxin
Frataxin is a soluble protein with no previously known function. No specific domains related to protein families are represented in its polypeptide sequences. Experiments in cell systems, specially in yeast Saccharomyces cerevisiae, have provided relevant information about the possible function of frataxin in mitochondria and its role in the pathogenesis of the disease. At least four hypotheses for the primary mitochondrial function of frataxin have been proposed.
17.4.1 Frataxin and the Homeostasis of Mitochondrial Iron
The first biochemical data came from the experiments in yeast and from clinical observations in FRDA patients. They shown a dysregulation of the iron metabolism, had an injury caused by free radicals and mitochondrial dysfunction [56, 57]. Therefore, frataxin was related with mitochondrial iron homeostasis. Yeast lacking Yfh1p accumulates mitochondrial iron at the expense of cytosolic iron. Excess iron in mitochondria when reacting with oxygen generates toxic reactive oxygen species (ROS) by the Fenton reaction. These ROS causes the oxidation of cellular components: damage to proteins and nucleic acids, and lipid peroxidation [32]. Some of the proteins that are more easily affected by free radicals are those containing Fe-S clusters, such as complex I, II and III of the mitochondrial transport chain and aconitase, showing a decrease of activity in human fibroblasts and yeast [58, 59]. In contrast, data from frataxin Knock-out mouse model has raised some questions about the role of iron in the disease pathogenesis. Iron deposits were not detected suggesting that iron accumulation might not be the primary consequence of frataxin deficiency, but a secondary effect [41].
17.4.2 Frataxin as an Iron-Storage Protein
A second hypothesis suggests that the function of frataxin is to bind iron and keep it in soluble and bioavailable form [46]. Yeast frataxin is a soluble monomer that contains no iron. But, when ferrous iron was added in presence of oxygen, it was induced the formation of frataxin trimers that catalyzed iron oxidation [60, 61]. New results confirm that yeast frataxin is stable as an iron-loaded monomer, and the protein can bind two ferrous iron atoms with micromolar binding affinity [62]. On the other hand, high iron concentration led to the protein assembly reaching to sequestrate more than 2000 iron atoms [63]. The higher molecular weight of this complex reminded us the ferritin structure, which contains a large number of ferric iron atoms. Although human frataxin is not able to form those complexes in vitro, at physiologic iron concentrations in the mitochondria, close to 10% of human frataxin are able to form a spherical complex which contained this metal [64]. Differences founded between yfh1 and mammalian frataxin could be due to the existence of a mitochondrial ferritin of higher eukaryotes (MtF) [65]. MtF expression in frataxin deficient yeast could rescue the respiratory deficiency, thus allowing cell growth in a non-fermentable carbon source and preventing mitochondrial iron accumulation, so protecting the activity of the iron-sulphur enzymes as well [66]. These data support the idea that MtF could replace frataxin functions in yeast, suggesting that frataxin would be directly involved in mitochondrial iron-storage and detoxification.
A common characteristic of yfh1 and mammalian frataxin is that iron storage is easily reversible by the addition of chelating agents, indicating that iron could be biologically available. Interpreting all these data, we would say that frataxin could be a mitochondrial iron chaperone that prevents Fenton reaction by hiding iron to reactive oxygen species and keeping it for other biosynthetic pathways. The problem to accept this hypothesis is that changes in intracellular iron levels do not have an effect on frataxin expression, so this is not consistent with the protein role on iron storage [67].
17.4.3 Frataxin as a Chaperone for Iron
Ferrochelatase, the final enzyme in the heme biosynthetic pathway that incorporate iron into protoporphyrin IX molecule, is one of the protein that frataxin could acts as a Fe(II) chaperone . Multiple studies have demonstrated a physical interaction between frataxin and ferrochelatase [61, 68, 69]. However, decreases in ferrochelatase activity have not been observed in frataxin deficient HeLa cells or patient lymphoblasts [70, 71]. The only other Fe-S enzyme besides ferrochelatase known to participate in the heme biosynthetic pathway is adrenodoxin which carries out the first step of the conversion of heme O to heme a, that is required for cytochrome oxidase activity. Frataxin deficient human oligodendroglial cell line has decreased adrenodoxin activity, decreased heme a and heme c levels and decreased cytochrome oxidase activity [72].
There is evidence in yeast and in mammals that frataxin deletion leads to severe alteration of Fe-S enzymes activities [50, 58, 73]. Fe-S clusters are essential components of respiratory electron transfer complexes as well as the tricarboxylic acid cycle enzymes, aconitase and succinate dehydrogenase. Frataxin is required, but not essential, for Fe-S cluster (ISC) biosynthesis [73, 74]. In eukaryotes, initial ISC biogenesis is mitochondrial [75]. In yeast, [2Fe-2S] cluster assembly is catalyzed by the redundant scaffold proteins Isu1 and Isu2 [76]. Yfh1p has been suggested to be the iron donor in this process [77]. In addition, frataxin interacts with Isu1p in yeast [78, 79]. Sulfur from cysteine is supplied by the cysteine desulfurase Nfs1p in association with essential protein partner Isd11 [76]. New data have shown that human frataxin binds the iron-sulfur biogenesis NFS1/ISCU complex through ISD11 and this interaction is nickel dependent [80]. Recently, studies characterising biophysical and metal binding properties and the abilities to transfer Fe(II) of the fly frataxin, support a frataxin’s role as an iron binding protein that delivers Fe(II) to the Fe-S cluster biosynthesis machinery via Isu in an iron dependent manner [81]. Moreover, Drosophila frataxin enhances the stability of DNA and protect it from oxidative stress because frataxin has the ability to regulate Fenton chemistry, function correlated directly or indirectly to Fe-S cluster formation [81].
Frataxin interacts with aconitase in presence of citrate. Citrate prevents aconitase cluster disassembly and is required for enzyme reactivation. Thus, the citrate requirement for interaction of frataxin and aconitase supports the contention that frataxin can stabilize the [4Fe-4S]2+ cluster and facilitate enzyme reactivation [82].
17.4.4 Frataxin as a Regulatory of Oxidative Phosphorylation
Deletion of the frataxin yeast homologue, YFH1, results in mutant strains that show a growth defect on fermentable carbon sources and a reduction in mitochondrial respiration [32, 34], because this mutant has affected the oxidative phosphorylation [58]. In skeletal muscle of FRDA patients, using phosphorus magnetic resonance spectroscopy, it has been demonstrated a deficit in a mitochondrial ATP production [83]. Ristow et al. hypothesized that frataxin might primarily affect oxidative phosphorylation acting as an activator [84]. They demonstrated that overexpression of frataxin in mammalian cells causes an increased mitochondrial membrane potential and results in an elevated cellular ATP content. In Arabidopsis thaliana it was observed a higher transcript level of the gene AtFH (homologue of frataxin) in flowers, a high energy demanding tissues in plants [85]. Moreover, AtFH expression was further increased in flowers from A. thaliana lines showing a mitochondrial dysfunction induced by the expression of the unedited version of ATP synthase subunit 9. Thus, the induction of AtFH expression in these lines could also be interpreted as a nuclear compensation in response to a decrease in mitochondrial respiration.
Physical interactions between the yeast frataxin Yfh1p and succinate dehydrogenase complex subunits Sdh1p and Sdh2p, which are components of the mitochondrial complex II, have been observed [86]. This result has been also confirmed by functional relationships between Yfh1p and Sdh1p and Sdh2p carrying out synthetic genetic interaction studies of mutated genes. Interaction among human frataxin and the human succinate dehydrogenase subunits SDHA and SDHB have been confirmed [86]. These findings suggest that frataxin is working in the mitochondrial electron transfer chain, perhaps by regulating the delivery of electrons via complex II towards the ubiquinone and also suggest that respiratory chain may have a primary role in the pathogenesis of FRDA. Genetic interaction with a complex II subunit of the respiratory chain, has been observed by synthetic genetic analysis in C. elegans [48], confirming similar findings previously observed in S. cerevisiae [86]. Finally, co-immunoprecipitation results from mitochondria of lymphoblast and COS-cells confirm the study of frataxin’s partners in yeast and C. elegans, in which frataxin was observed to interact with succinate dehydrogenase [80].
17.5 Therapies for Friedreich Ataxia
Pathophysiology of Friedreich ataxia based on cell and animal models and human studies indicates several possible therapeutic approaches: iron-chelation, antioxidant therapy, therapy with metalloporphyrins and molecules to increases frataxin expression.
17.5.1 Antioxidant Therapy
Coenzyme Q10 (CoQ) and vitamin E are important mitochondrial antioxidants. Whereas vitamin E is obtained solely from diet, CoQ is obtained in the diet and also synthesised in every cell via a pathway that shares some of the steps with cholesterol biosynthesis. It is a component of the electron transport chain. Lodi et al. have shown that combined CoQ (400 mg/d) and vitamin E (2100 IU/d) therapy resulted in a significant improvement in cardiac and skeletal muscle energy metabolism in 10 patients with FRDA after 3 months of treatment [87]. A long-term longitudinal follow-up study of these patients with FRDA caused a prolonged improvement in cardiac and skeletal muscle bioenergetics clearly demonstrating its biochemical efficacy [88].
Idebenone is a short-chain analogue of CoQ. It has been demonstrated that idebenone treatment inhibits lipid peroxidation, stimulates mitochondrial functions, and improves the myocardial energy state in cardiac hypertrophy [89]. Most trials demonstrated a positive effect on cardiac hypertrophy [90–92]. The neurological function is in general not modified in adult patients, but a dose-dependent effect was demonstrated in young Friedreich ataxia patients [93]. The treatment with idebenone (5–20 mg/kg/day) for 3–5 years in paediatric and adults patients, prevented progression of cardiomyopathy in both patients, whereas its stabilizing effect on neurological dysfunction was present only in the paediatric population. This suggests that the age at which idebenone treatment is initiated may be an important factor in the effectiveness of the therapy [94].
Mitoquinone (MitoQ) is an antioxidant selectively targeted to mitochondria. This mitochondrially localized antioxidant protects against H2O2-induced apoptosis in cell culture studies, in contrast to untargeted ubiquinone analogues [95]. MitoQ was shown to be 800-fold more potent than idebenone in protecting FRDA fibroblasts from death due to endogenous oxidative stress generated by inhibition of glutathione synthesis [96]. Until now no clinical trials have been reported.
17.5.2 Iron Chelators Therapy
Since excess of mitochondrial iron is likely to play a role in the pathology of FRDA, iron chelators have considered in the treatment of FRDA.
Deferiprone is a chelator specifically targeting mitochondrial iron. This chelator is orally active, membrane permeant, and capable of scavenging iron from mitochondria of specific areas of the brain. Treatment to FRDA patients with deferiprone caused no apparent hematologic or neurologic side effects while reducing neuropathy and ataxic gait in the youngest patients [97]. This was the first clinical demonstration of chelation removing labile iron accumulated in a specific brain area implicated in a neurodegenerative disease. In cell HEK-293 deficient in frataxin, deferiprone restored of impaired mitochondrial membrane, increased ATP production and oxygen consumption and attenuation of mitochondrial DNA damage [98]. However, in skin fibroblasts of FRDA patients and in frataxin depleted neuroblastoma derived cells, a direct consequence of chelation mitochondrial free iron is a concentration and time dependent loss of aconitase activity [99].
A group of iron chelator, 2-pyridylcarboxaldehyde isonicotinoyl hydrazone (PCIH) ligands, has been designed to specifically target mitochondrial iron pools, being the 2-pyridylcarboxaldehyde 2-thiophenecarboxyl hydrazone (PCTH) as the most promising compound. PCTH is highly effective at preventing H2O2-induced cytotoxicity and at preventing oxidative stress. It also increased FRDA fibroblast cell viability by up to 70% [100].
17.5.3 Therapy with Metalloporphyrins
Hemin is an enzyme inhibitor derived from processed red blood cells. It is an iron containing metalloporphyrin. Exogenous heme administration with hemin increases activity of the iron-sulfur cluster enzymes adrenoxin and aconitase in frataxin deficient human oligodendroglial cell line [72].
17.5.4 Therapy to Increase Frataxin Expression
Recombinant human erythropoietin (rhoEPO) is a glycoprotein hormone that controls erythropoiesis, or red blood cell production. rhoEPO has broad neuroprotective and cardioprotective capabilities [101, 102]. In various cell types: lymphocytes from FRDA patients, primary human cardiac cells and in a neuronal cell line, rhoEPO increases frataxin expression significantly [103].
Heber and colleagues described a screening of small molecule DNA ligands for their effects on EGFP expression in the reporter cell lines [104]. While some of the tested compounds were cytotoxic, some compounds such as Hoeschst 33258, pentamidine, DAPI, distamycin and diminazene, were found to increase GFP expression. Pentamidine was also shown to increase frataxin expression in primary FRDA lymphocytes.
Polyamides possess the ability to bind duplex DNA. These molecules are unique in that they can be programmed to bind predetermined DNA sequences [105]. Polyamide FA1 increased FXN transcription approximately 2–3 fold [106].
Butyric acid has been identified to possess the ability to increase frataxin expression or transcription through GAA repeats [107].
Molecules that alter chromatin structure by changing the postsynthetic modification states of the histone proteins, namely histone deacetylase inhibitors, have been screened for their effects on FXN gene expression. HDAC inchibitors have been tested in FRDA patient lymphocytes and lymphoblasts. One compound, BML-210 and related analogues (in particular component 4b), showed a significant increase in frataxin message levels by approximately two-fold [108]. Compound 106, a derivative of 4b, increases histone acetylation in the brain at a dose that causes no apparent toxicity in wild type C57Bl/6 mouse or in KIKI mice (a knock-in mouse carrying a (GAA)230 repeat in the first intron of the endogenous frataxin gene [52]). This compound was able to restore normal frataxin levels in the central nervous system and heart of KIKI mice, tissues that are relevant targets as they are involved in FRDA pathology [109].
References
Friedreich N. Über degenerative Atrophie der spinalen Hinterstránge. Virchows Arch Pathol Anat 1863;27:1–26.
Friedreich N. Über degenerative Atrophie der spinalen Hinterstränge. Virchows Arch Pathol Anat 1863;26(433–459).
Lopez-Arlandis JM, Vilchez JJ, Palau F, et al. Friedreich’s ataxia: an epidemiological study in Valencia, Spain, based on consanguinity analysis. Neuroepidemiology 1995;14(1):14–19.
Polo JM, Calleja J, Combarros O, et al. Hereditary ataxias and paraplegias in Cantabria, Spain. An epidemiological and clinical study. Brain 1991; 114 ( Pt 2):855–866.
Durr A, Cossee M, Agid Y, et al. Clinical and genetic abnormalities in patients with Friedreich’s ataxia. N Engl J Med 1996;335 (16):1169–1175.
Campuzano V, Montermini L, Molto MD, et al. Friedreich’s ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science 1996;271 (5254):1423–1427.
Chamberlain S, Shaw J, Rowland A, et al. Mapping of mutation causing Friedreich’s ataxia to human chromosome 9. Nature 1988;334(6179):248–250.
Cossee M, Schmitt M, Campuzano V, et al. Evolution of the Friedreich’s ataxia trinucleotide repeat expansion: founder effect and premutations. Proc Natl Acad Sci U S A 1997;94(14):7452–7457.
Montermini L, Andermann E, Labuda M, et al. The Friedreich ataxia GAA triplet repeat: premutation and normal alleles. Hum Mol Genet 1997;6(8):1261–1266.
Bidichandani SI, Ashizawa T, Patel PI. Atypical Friedreich ataxia caused by compound heterozygosity for a novel missense mutation and the GAA triplet-repeat expansion. Am J Hum Genet 1997;60(5):1251–1256.
Campuzano V, Montermini L, Lutz Y, et al. Frataxin is reduced in Friedreich ataxia patients and is associated with mitochondrial membranes. Hum Mol Genet 1997;6(11):1771–1780.
Grabczyk E, Mancuso M, Sammarco MC. A persistent RNA.DNA hybrid formed by transcription of the Friedreich ataxia triplet repeat in live bacteria, and by T7 RNAP in vitro. Nucleic Acids Res 2007;35(16):5351–5359.
Grabczyk E, Usdin K. The GAA*TTC triplet repeat expanded in Friedreich’s ataxia impedes transcription elongation by T7 RNA polymerase in a length and supercoil dependent manner. Nucleic Acids Res 2000;28(14):2815–2822.
Soragni E, Herman D, Dent SY, et al. Long intronic GAA*TTC repeats induce epigenetic changes and reporter gene silencing in a molecular model of Friedreich ataxia. Nucleic Acids Res 2008;36(19):6056–6065.
Al-Mahdawi S, Pook M, Chamberlain S. A novel missense mutation (L198R) in the Friedreich’s ataxia gene. Hum Mutat 2000;16(1):95.
Bartolo C, Mendell JR, Prior TW. Identification of a missense mutation in a Friedreich’s ataxia patient: implications for diagnosis and carrier studies. Am J Med Genet 1998;79(5):396–399.
Cossee M, Durr A, Schmitt M, et al. Friedreich’s ataxia: point mutations and clinical presentation of compound heterozygotes. Ann Neurol 1999;45(2):200–206.
De Castro M, Garcia-Planells J, Monros E, et al. Genotype and phenotype analysis of Friedreich’s ataxia compound heterozygous patients. Hum Genet 2000;106(1):86–92.
De Michele G, Filla A, Cavalcanti F, et al. Atypical Friedreich ataxia phenotype associated with a novel missense mutation in the X25 gene. Neurology 2000;54(2):496–499.
Doudney K PM, Al-Mahdawi S, Carvajal J, Hillerman R, Chamberlain S. A novel site mutation (384+1G-A) in the Friedreich’s ataxia gene. Hum Mutat 1997;11:415.
Forrest SM, Knight M, Delatycki MB, et al. The correlation of clinical phenotype in Friedreich ataxia with the site of point mutations in the FRDA gene. Neurogenetics 1998;1(4):253–257.
Labuda M PJaPM. A missense mutation (W155R) in an American patients with Friedreich ataxia. Hum Mutat 1999;13:506–507.
McCormack ML, Guttmann RP, Schumann M, et al. Frataxin point mutations in two patients with Friedreich’s ataxia and unusual clinical features. J Neurol Neurosurg Psychiatry 2000;68(5):661–664.
Pook MA, Al-Mahdawi SA, Thomas NH, et al. Identification of three novel frameshift mutations in patients with Friedreich’s ataxia. J Med Genet 2000;37(11):E38.
Zhu D, Burke C, Leslie A, et al. Friedreich’s ataxia with chorea and myoclonus caused by a compound heterozygosity for a novel deletion and the trinucleotide GAA expansion. Mov Disord 2002;17(3):585–589.
Zuhlke C, Laccone F, Cossee M, et al. Mutation of the start codon in the FRDA1 gene: linkage analysis of three pedigrees with the ATG to ATT transversion points to a unique common ancestor. Hum Genet 1998;103(1):102–105.
Branda SS, Cavadini P, Adamec J, et al. Yeast and human frataxin are processed to mature form in two sequential steps by the mitochondrial processing peptidase. J Biol Chem 1999;274(32):22763–22769.
Koutnikova H, Campuzano V, Koenig M. Maturation of wild-type and mutated frataxin by the mitochondrial processing peptidase. Hum Mol Genet 1998;7(9):1485–1489.
Schmucker S, Argentini M, Carelle-Calmels N, et al. The in vivo mitochondrial two-step maturation of human frataxin. Hum Mol Genet 2008;17(22):3521–35231.
Dhe-Paganon S, Shigeta R, Chi YI, et al. Crystal structure of human frataxin. J Biol Chem 2000;275(40):30753–30756.
Gibson TJ, Koonin EV, Musco G, et al. Friedreich’s ataxia protein: phylogenetic evidence for mitochondrial dysfunction. Trends Neurosci 1996;19(11):465–468.
Babcock M, de Silva D, Oaks R, et al. Regulation of mitochondrial iron accumulation by Yfh1p, a putative homolog of frataxin. Science 1997;276(5319):1709–1712.
Foury F, Cazzalini O. Deletion of the yeast homologue of the human gene associated with Friedreich’s ataxia elicits iron accumulation in mitochondria. FEBS Lett 1997;411(2–3):373–377.
Koutnikova H, Campuzano V, Foury F, et al. Studies of human, mouse and yeast homologues indicate a mitochondrial function for frataxin. Nat Genet 1997;16(4):345–351.
Radisky DC, Babcock MC, Kaplan J. The yeast frataxin homologue mediates mitochondrial iron efflux. Evidence for a mitochondrial iron cycle. J Biol Chem 1999;274(8):4497–4499.
Wilson RB, Roof DM. Respiratory deficiency due to loss of mitochondrial DNA in yeast lacking the frataxin homologue. Nat Genet 1997;16(4):352–357.
Cavadini P, Gellera C, Patel PI, et al. Human frataxin maintains mitochondrial iron homeostasis in Saccharomyces cerevisiae. Hum Mol Genet 2000;9(17):2523–2530.
Canizares J, Blanca JM, Navarro JA, et al. dfh is a Drosophila homolog of the Friedreich’s ataxia disease gene. Gene 2000;256(1–2):35–42.
Llorens JV, Navarro JA, Martinez-Sebastian MJ, et al. Causative role of oxidative stress in a Drosophila model of Friedreich ataxia. Faseb J 2007;21(2):333–344.
Anderson PR, Kirby K, Hilliker AJ, et al. RNAi-mediated suppression of the mitochondrial iron chaperone, frataxin, in Drosophila. Hum Mol Genet 2005;14(22):3397–3405.
Cossee M, Puccio H, Gansmuller A, et al. Inactivation of the Friedreich ataxia mouse gene leads to early embryonic lethality without iron accumulation. Hum Mol Genet 2000;9(8):1219–1226.
Das N, Levine RL, Orr WC, et al. Selectivity of protein oxidative damage during aging in Drosophila melanogaster. Biochem J 2001;360(Pt 1):209–216.
Prabhu HR, Krishnamurthy S. Ascorbate-dependent formation of hydroxyl radicals in the presence of iron chelates. Indian J Biochem Biophys 1993;30(5):289–292.
Anderson PR, Kirby K, Orr WC, et al. Hydrogen peroxide scavenging rescues frataxin deficiency in a Drosophila model of Friedreich’s ataxia. Proc Natl Acad Sci U S A 2008;105(2):611–616.
Runko AP, Griswold AJ, Min KT. Overexpression of frataxin in the mitochondria increases resistance to oxidative stress and extends lifespan in Drosophila. FEBS Lett 2008;582(5):715–719.
Adamec J, Rusnak F, Owen WG, et al. Iron-dependent self-assembly of recombinant yeast frataxin: implications for Friedreich ataxia. Am J Hum Genet 2000;67(3):549–562.
Ventura N, Rea S, Henderson ST, et al. Reduced expression of frataxin extends the lifespan of Caenorhabditis elegans. Aging Cell 2005;4(2):109–112.
Vazquez-Manrique RP, Gonzalez-Cabo P, Ros S, et al. Reduction of Caenorhabditis elegans frataxin increases sensitivity to oxidative stress, reduces lifespan, and causes lethality in a mitochondrial complex II mutant. Faseb J 2006;20(1):172–174.
Zarse K, Schulz TJ, Birringer M, et al. Impaired respiration is positively correlated with decreased life span in Caenorhabditis elegans models of Friedreich Ataxia. Faseb J 2007;21(4):1271–1275.
Puccio H, Simon D, Cossee M, et al. Mouse models for Friedreich ataxia exhibit cardiomyopathy, sensory nerve defect and Fe-S enzyme deficiency followed by intramitochondrial iron deposits. Nat Genet 2001;27(2):181–186.
Simon D, Seznec H, Gansmuller A, et al. Friedreich ataxia mouse models with progressive cerebellar and sensory ataxia reveal autophagic neurodegeneration in dorsal root ganglia. J Neurosci 2004;24(8):1987–1995.
Miranda CJ, Santos MM, Ohshima K, et al. Frataxin knockin mouse. FEBS Lett 2002;512(1–3):291–297.
Al-Mahdawi S, Pinto RM, Ruddle P, et al. GAA repeat instability in Friedreich ataxia YAC transgenic mice. Genomics 2004;84(2):301–310.
Al-Mahdawi S, Pinto RM, Varshney D, et al. GAA repeat expansion mutation mouse models of Friedreich ataxia exhibit oxidative stress leading to progressive neuronal and cardiac pathology. Genomics 2006;88(5):580–590.
Clark RM, De Biase I, Malykhina AP, et al. The GAA triplet-repeat is unstable in the context of the human FXN locus and displays age-dependent expansions in cerebellum and DRG in a transgenic mouse model. Hum Genet 2007;120(5):633–640.
Waldvogel D, van Gelderen P, Hallett M. Increased iron in the dentate nucleus of patients with Friedrich’s ataxia. Ann Neurol 1999;46(1):123–125.
Lamarche JB, Cote M, Lemieux B. The cardiomyopathy of Friedreich’s ataxia morphological observations in 3 cases. Can J Neurol Sci 1980;7(4):389–396.
Rotig A, de Lonlay P, Chretien D, et al. Aconitase and mitochondrial iron-sulphur protein deficiency in Friedreich ataxia. Nat Genet 1997;17(2):215–217.
Foury F. Low iron concentration and aconitase deficiency in a yeast frataxin homologue deficient strain. FEBS Lett 1999;456(2):281–284.
Park S, Gakh O, Mooney SM, et al. The ferroxidase activity of yeast frataxin. J Biol Chem 2002;277(41):38589–38595.
Park S, Gakh O, O’Neill HA, et al. Yeast frataxin sequentially chaperones and stores iron by coupling protein assembly with iron oxidation. J Biol Chem 2003;278(33):31340–31351.
Cook JD, Bencze KZ, Jankovic AD, et al. Monomeric yeast frataxin is an iron-binding protein. Biochemistry 2006;45(25):7767–7777.
Nichol H, Gakh O, O’Neill HA, et al. Structure of frataxin iron cores: an X-ray absorption spectroscopic study. Biochemistry 2003;42(20):5971–5976.
Cavadini P, O’Neill HA, Benada O, et al. Assembly and iron-binding properties of human frataxin, the protein deficient in Friedreich ataxia. Hum Mol Genet 2002;11(3):217–227.
Levi S, Corsi B, Bosisio M, et al. A human mitochondrial ferritin encoded by an intronless gene. J Biol Chem 2001;276(27):24437–24440.
Campanella A, Isaya G, O’Neill HA, et al. The expression of human mitochondrial ferritin rescues respiratory function infrataxin-deficient yeast. Hum Mol Genet 2004;13(19):2279–2288.
Becker EM, Greer JM, Ponka P, et al. Erythroid differentiation and protoporphyrin IX down-regulate frataxin expression in Friend cells: characterization of frataxin expression compared to molecules involved in iron metabolism and hemoglobinization. Blood 2002;99(10):3813–3822.
Yoon T, Cowan JA. Frataxin-mediated iron delivery to ferrochelatase in the final step of heme biosynthesis. J Biol Chem 2004;279(25):25943–25946.
Lesuisse E, Santos R, Matzanke BF, et al. Iron use for haeme synthesis is under control of the yeast frataxin homologue (Yfh1). Hum Mol Genet 2003;12(8):879–889.
Lange H, Muhlenhoff U, Denzel M, et al. The heme synthesis defect of mutants impaired in mitochondrial iron-sulfur protein biogenesis is caused by reversible inhibition of ferrochelatase. J Biol Chem 2004;279(28):29101–29108.
Napoli E, Taroni F, Cortopassi GA. Frataxin, iron-sulfur clusters, heme, ROS, and aging. Antioxid Redox Signal 2006;8(3–4):506–516.
Napoli E, Morin D, Bernhardt R, et al. Hemin rescues adrenodoxin, heme a and cytochrome oxidase activity in frataxin-deficient oligodendroglioma cells. Biochim Biophys Acta 2007;1772(7):773–780.
Muhlenhoff U, Richhardt N, Ristow M, et al. The yeast frataxin homolog Yfh1p plays a specific role in the maturation of cellular Fe/S proteins. Hum Mol Genet 2002;11(17):2025–2036.
Duby G, Foury F, Ramazzotti A, et al. A non-essential function for yeast frataxin in iron-sulfur cluster assembly. Hum Mol Genet 2002;11(21):2635–2643.
Lill R, Diekert K, Kaut A, et al. The essential role of mitochondria in the biogenesis of cellular iron-sulfur proteins. Biol Chem 1999;380(10):1157–1166.
Lill R, Muhlenhoff U. Iron-sulfur protein biogenesis in eukaryotes: components and mechanisms. Annu Rev Cell Dev Biol 2006;22:457–486.
Yoon T, Cowan JA. Iron-sulfur cluster biosynthesis. Characterization of frataxin as an iron donor for assembly of [2Fe-2S] clusters in ISU-type proteins. J Am Chem Soc 2003;125(20):6078–6084.
Gerber J, Muhlenhoff U, Lill R. An interaction between frataxin and Isu1/Nfs1 that is crucial for Fe/S cluster synthesis on Isu1. EMBO Rep 2003;4(9):906–911.
Ramazzotti A, Vanmansart V, Foury F. Mitochondrial functional interactions between frataxin and Isu1p, the iron-sulfur cluster scaffold protein, in Saccharomyces cerevisiae. FEBS Lett 2004;557(1–3):215–220.
Shan Y, Napoli E, Cortopassi G. Mitochondrial frataxin interacts with ISD11 of the NFS1/ISCU complex and multiple mitochondrial chaperones. Hum Mol Genet 2007;16(8):929–941.
Kondapalli KC, Kok NM, Dancis A, et al. Drosophila frataxin: an iron chaperone during cellular Fe-S cluster bioassembly. Biochemistry 2008;47(26):6917–6927.
Bulteau AL, O’Neill HA, Kennedy MC, et al. Frataxin acts as an iron chaperone protein to modulate mitochondrial aconitase activity. Science 2004;305(5681):242–245.
Lodi R, Cooper JM, Bradley JL, et al. Deficit of in vivo mitochondrial ATP production in patients with Friedreich ataxia. Proc Natl Acad Sci U S A 1999;96(20):11492–11495.
Ristow M, Pfister MF, Yee AJ, et al. Frataxin activates mitochondrial energy conversion and oxidative phosphorylation. Proc Natl Acad Sci U S A 2000;97(22):12239–12243.
Busi MV, Zabaleta EJ, Araya A, et al. Functional and molecular characterization of the frataxin homolog from Arabidopsis thaliana. FEBS Lett 2004;576(1–2):141–144.
Gonzalez-Cabo P, Vazquez-Manrique RP, Garcia-Gimeno MA, et al. Frataxin interacts functionally with mitochondrial electron transport chain proteins. Hum Mol Genet 2005;14(15):2091–2098.
Lodi R, Hart PE, Rajagopalan B, et al. Antioxidant treatment improves in vivo cardiac and skeletal muscle bioenergetics in patients with Friedreich’s ataxia. Ann Neurol 2001;49(5):590–596.
Hart PE, Lodi R, Rajagopalan B, et al. Antioxidant treatment of patients with Friedreich ataxia: four-year follow-up. Arch Neurol 2005;62(4):621–626.
Geromel V, Darin N, Chretien D, et al. Coenzyme Q(10) and idebenone in the therapy of respiratory chain diseases: rationale and comparative benefits. Mol Genet Metab 2002;77(1–2):21–30.
Mariotti C, Solari A, Torta D, et al. Idebenone treatment in Friedreich patients: one-year-long randomized placebo-controlled trial. Neurology 2003;60(10):1676–1679.
Ribai P, Pousset F, Tanguy ML, et al. Neurological, cardiological, and oculomotor progression in 104 patients with Friedreich ataxia during long-term follow-up. Arch Neurol 2007;64(4):558–564.
Rustin P, von Kleist-Retzow JC, Chantrel-Groussard K, et al. Effect of idebenone on cardiomyopathy in Friedreich’s ataxia: a preliminary study. Lancet 1999;354(9177):477–479.
Di Prospero NA, Baker A, Jeffries N, et al. Neurological effects of high-dose idebenone in patients with Friedreich’s ataxia: a randomised, placebo-controlled trial. Lancet Neurol 2007;6(10):878–886.
Pineda M, Arpa J, Montero R, et al. Idebenone treatment in paediatric and adult patients with Friedreich ataxia: long-term follow-up. Eur J Paediatr Neurol 2008;12(6):470–475.
Kelso GF, Porteous CM, Coulter CV, et al. Selective targeting of a redox-active ubiquinone to mitochondria within cells: antioxidant and antiapoptotic properties. J Biol Chem 2001;276(7):4588–4596.
Jauslin ML, Meier T, Smith RA, et al. Mitochondria-targeted antioxidants protect Friedreich Ataxia fibroblasts from endogenous oxidative stress more effectively than untargeted antioxidants. Faseb J 2003;17(13):1972–1974.
Boddaert N, Le Quan Sang KH, Rotig A, et al. Selective iron chelation in Friedreich ataxia: biologic and clinical implications. Blood 2007;110(1):401–408.
Kakhlon O, Manning H, Breuer W, et al. Cell functions impaired by frataxin deficiency are restored by drug-mediated iron relocation. Blood 2008;112(13):5219–5227.
Goncalves S, Paupe V, Dassa EP, et al. Deferiprone targets aconitase: implication for Friedreich’s ataxia treatment. BMC Neurol 2008;8:20.
Lim CK, Kalinowski DS, Richardson DR. Protection against hydrogen peroxide-mediated cytotoxicity in Friedreich’s ataxia fibroblasts using novel iron chelators of the 2-pyridylcarboxaldehyde isonicotinoyl hydrazone class. Mol Pharmacol 2008;74(1):225–235.
Siren AL, Ehrenreich H. Erythropoietin–a novel concept for neuroprotection. Eur Arch Psychiatry Clin Neurosci 2001;251(4):179–184.
Smith KJ, Bleyer AJ, Little WC, et al. The cardiovascular effects of erythropoietin. Cardiovasc Res 2003;59(3):538–548.
Sturm B, Stupphann D, Kaun C, et al. Recombinant human erythropoietin: effects on frataxin expression in vitro. Eur J Clin Invest 2005;35(11):711–717.
Grant L, Sun J, Xu H, et al. Rational selection of small molecules that increase transcription through the GAA repeats found in Friedreich’s ataxia. FEBS Lett 2006;580(22):5399–53405.
Dervan PB, Edelson BS. Recognition of the DNA minor groove by pyrrole-imidazole polyamides. Curr Opin Struct Biol 2003;13(3):284–299.
Burnett R, Melander C, Puckett JW, et al. DNA sequence-specific polyamides alleviate transcription inhibition associated with long GAA.TTC repeats in Friedreich’s ataxia. Proc Natl Acad Sci U S A 2006;103(31):11497–11502.
Sarsero JP, Li L, Wardan H, et al. Upregulation of expression from the FRDA genomic locus for the therapy of Friedreich ataxia. J Gene Med 2003;5(1):72–81.
Herman D, Jenssen K, Burnett R, et al. Histone deacetylase inhibitors reverse gene silencing in Friedreich’s ataxia. Nat Chem Biol 2006;2(10):551–8.
Rai M, Soragni E, Jenssen K, et al. HDAC inhibitors correct frataxin deficiency in a Friedreich ataxia mouse model. PLoS ONE 2008;3(4):e1958.
Acknowledgements
This work is supported by the Spanish Ministry of Science and Innovation and the Fondo de Investigación Sanitaria. The CIBER de Enfermedades Raras is an initiative of the Instituto de Salud Carlos III.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer Science+Business Media B.V.
About this chapter
Cite this chapter
González-Cabo, P., Llorens, J.V., Palau, F., Moltó, M.D. (2009). Friedreich Ataxia: An Update on Animal Models, Frataxin Function and Therapies. In: Espinós, C., Felipo, V., Palau, F. (eds) Inherited Neuromuscular Diseases. Advances in Experimental Medicine and Biology, vol 652. Springer, Dordrecht. https://doi.org/10.1007/978-90-481-2813-6_17
Download citation
DOI: https://doi.org/10.1007/978-90-481-2813-6_17
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-90-481-2812-9
Online ISBN: 978-90-481-2813-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)