Abstract
In contrast to plant mitochondrial genomes, which possess a high degree of plasticity, plastid genomes (plastomes) across photosynthetically active species are characterized by a remarkable conservation of structure, coding capacity, gene order, and intron content. Exceptions to the general pattern, however, are seen in parasitic species that have abandoned photosynthesis as the main energy source and live on other organisms, mainly other plants or algae. The relief from the selective pressure normally exerted by photosynthesis is reflected in a “plastomic striptease” that concerns the loss of coding sequences, introns, RNA-editing sites, and regulatory sequences from the plastid genome in higher plants and has, in extreme cases of nonphotosynthetic algal descendents, resulted in the loss of the entire organelle. This review attempts to give an overview over the trail of plastomic losses and reductions in parasitic species of the plant kingdom, with a main focus on higher plants.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
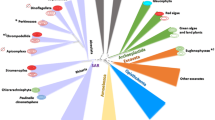
The view that plastids have evolved from cyanobacteria by endosymbiosis and that this event took place originally only once is nowadays widely accepted (see Chap. 2). This primary endosymbiosis gave rise to the common ancestor of today’s three primary plastid-containing lineages (together known as Archaeplastida, Table 4.1): (1) green algae and plants, (2) red algae, and (3) glaucophyte algae (Reyes-Prieto and Bhattacharya 2007; Lane and Archibald 2008). From these lineages, plastids have spread laterally by secondary endosymbiosis to form further lineages (see Chap. 2). Thus, two unrelated lineages containing plastids of green algal ancestry (euglenoids and chlorarachniophytes) and several lineages containing plastids of red algal ancestry (haptophytes, cryptophytes, stramenopiles, dinoflagellates, and apicomplexa) have evolved (Cavalier-Smith 1999, 2002; Lane and Archibald 2008). In addition, the occurrence of serial secondary endosymbiosis and tertiary plastids that replaced the original secondary plastid (Yoon et al. 2005) is under discussion (Lane and Archibald 2008; Janouskovec et al. 2010). Most of the resulting species gain their energy through photoautotrophic carbon fixation. However, in probably every land plant and algal lineage, parasitic species have evolved that have abandoned photoautotrophic growth and rather live by parasitizing on other plants, fungi, or even animals.
Species of colorless unicellular algae with reduced plastid genomes have been described among the green algae (de Koning and Keeling 2004; Tartar and Boucias 2004; Borza et al. 2005; de Koning and Keeling 2006), the euglenoid algae (Gockel and Hachtel 2000), and the dinoflagellates (Sanchez-Puerta et al. 2007; Matsuzaki et al. 2008). The apicomplexan group of unicellular parasitic organisms, last but not least, provides the most prominent example that plastid-containing parasites have been able to utilize a very wide host range (Lim and McFadden 2010). More recently, claims of a photosynthetic/red algal (and therefore, possibly, plastid) history of the oomycete Phytophthora (Tyler et al. 2006) and of ciliates (Reyes-Prieto et al. 2008) have been made (Janouskovec et al. 2010), taking the discussion around adaptations to a nonphotosynthetic life style another step ahead.
Parasitic land plants are – in contrast to their algal counterparts – quite eye catching (Fig. 4.1) and have received much attention not only due to their special lifestyle but also due to the damage they can inflict on agricultural land use. It is estimated that approximately 1% of all angiosperm species from at least 11 different lineages have resorted to a parasitic lifestyle (Barkman et al. 2007). Based on their attachment sites on their hosts, root parasites and shoot parasites are being distinguished (Fig. 4.1). The liverwort Aneura mirabilis (formerly known as Cryptothallus mirabilis) is one example – and as a matter of fact, to date, the only known example – of a nonvascular parasitic land plant that has evolved into a completely nonphotosynthetic lifestyle (Bidartondo et al. 2003).
The following chapters will summarize our current knowledge on the structure and function of plastid genomes from algae to higher plants and will focus on the degeneration of plastid genomes in species that have resorted to a parasitic lifestyle.
2 The Plastid Genome
2.1 The Organization of Plastid DNA
Every plastid possesses its own genetic information that was inherited from the cyanobacterial ancestors of these organelles. Reflecting this ancestry, the plastid genome has retained many features of prokaryotic genomes, including the overall structure, physical properties, gene organization, and regulatory features necessary for gene expression (Bock 2007). Plastid DNA is present in multiple copies per plastid and is compacted by DNA-binding proteins (Kuroiwa 1991). Microscopic studies in combination with fluorescent staining of these high-molecular-weight DNA–protein complexes generally known as nucleoids, revealed that their number, shape, size, and distribution can vary significantly between species and depending on the developmental stage (Kuroiwa 1991). Plastid genome maps of all plant lineages traditionally show the organelle’s genetic information as circular molecules. The only striking exception to this seem to be the dinoflagellates where the normal single circle has been replaced by minicircles, which contain one or a few genes each (Zhang et al. 1999; Barbrook and Howe 2000). Electron microscopic pictures later revealed that the circular conformation is only one of many that the plastid DNA of higher plants can appear in. In addition to circular monomeric molecules, a variety of linear and branched monomers and oligomers (Deng et al. 1989; Lilly et al. 2001; Oldenburg and Bendich 2004; Scharff and Koop 2006) and possibly also shorter fragments (Kolodner and Tewari 1972) have been observed.
In contrast to the structural variability of the nucleoids, plastid coding capacity and gene organization have remained remarkably conserved across the plant kingdom, with higher plants, in particular, showing very little variation in their gene content and plastid genome sizes (Jansen et al. 2007). The selective pressure on the maintenance of photosynthesis-related genes and genes for subunits of the plastid gene-expression machinery seems to have been instrumental for maintaining the plastid genome and conserving a core set of plastid genes (Bock 2007). It is this selective pressure exerted by photosynthesis or, rather opposite, the obvious lack of it in parasitic plants that has drawn the attention to parasitic plant plastid genomes and their particular evolution.
2.2 Structure and Coding Capacity of Plastid Genomes
The first complete plastid genome sequences, namely that of the bryophyte Marchantia polymorpha and of the seed plant Nicotiana tabacum, were reported as early as 1986 (Ohyama et al. 1986; Shinozaki et al. 1986). Since then, more than 230 plastid genomes have been completely sequenced. The vast majority of these plastid genome sequences are from land plants and green algae, while other big groups, such as the red algae, dinoflagellates, or stramenopiles, are still strongly underrepresented (Table 4.1).
A common feature of plastid genomes are two inverted repeat regions (IRA and IRB) that split the remainder of the chromosome into a large and a small single-copy region (LSC and SSC, respectively). The inverted repeats can vary significantly in size, from 75 kbp in Pelargonium (Chumley et al. 2006) to 0.5 kbp in Pinus (Wakasugi et al. 1994). The functional role of this tetrapartite structure is unclear and examples of species without this organization are known from green (Hallick et al. 1993; Wakasugi et al. 1997; Jansen et al. 2008) and red plastid lineages (Glockner et al. 2000; Ohta et al. 2003; Hagopian et al. 2004), suggesting that it could be dispensable.
Compared to the ancestral prokaryotic genome, the plastome of all plants and algae is more or less drastically reduced. The genetic information for the majority of plastid-localized proteins either has been lost altogether in the course of evolution or has been transferred to the nucleus from where the gene products are imported back into the organelle (see Chap. 7). With a few exceptions, plastid genomes of the green lineage range between 120 and 160 kbp in size and code for 100 ± 20 proteins as well as about 40 genes encoding stable RNA species (rRNAs and tRNAs) (Palmer 1990). Even the plastome of the hypothetical ancestor to all green lineages was estimated to have contained a total of only 137 protein-coding genes (Turmel et al. 1999). Free-living descendents of the red plastid lineage have, on average, retained larger plastid genomes compared to the “standard plastome” of the green lineage. Nevertheless, gene numbers and sizes in these genomes are still in the same order of magnitude and feature around 200 protein-coding genes on approximately 160–180 kbp (Reith and Munholland 1993; Ohta et al. 2003). Extreme deviations from the average sizes, on the other hand, do exist in single cases and have been reported, for example, for Acetabularia species whose plastid genomes can be up to 1.5 Mbp in size (Simpson and Stern 2002), while probably the smallest plastid genome of a photosynthetic organism with only 72 kbp is that of the green alga Ostreococcus tauri (Robbens et al. 2007).
The information content of the plastid genome can be roughly divided into three large groups (see also Table 4.2): (1) genetic system genes comprising the RNA and protein components of the transcription and translation machineries as well as a few proteins involved in post-transcriptional and post-translational steps, (2) photosynthesis genes for subunits of the light and dark reactions and the ATPase, and (3) conserved open reading frames and genes with miscellaneous functions.
Genetic system genes required for transcription, translation, and processing steps represent a large portion of all plastid genomes and also represent the major fraction of strongly reduced plastid genomes, such as that of the apicomplexan parasite Toxoplasma gondii (Fig. 4.2). The number of photosynthesis and genetic system genes is slightly higher in red algae and most lineages derived from them, indicating specific losses in the green plastids, whereas a constant number of six genes (equaling 4–5% of the coding potential) on almost all plastid genomes is dedicated to the subunits of the plastid ATPase. In green plastid-derived lineages, the third group that comprises genes with other functions makes up around 20% of the genes, the majority of which are conserved reading frames of unknown function (YCFs) (Fig. 4.2). In most rhodophytes, glaucophytes, and chromalveolates, this gene group is considerably expanded and contains genes for amino acid, lipid, pigment, and cofactor metabolism (Fig. 4.2) that are absent from land plants and green algae. The only exception is the chromalveolate group of apicomplexans, where many typical gene groups of the red plastid lineage have been lost (Wilson et al. 1996). This reduction in genome-coding capacity is related partly to the parasitic life that members of this group are leading and partly to losses that must have occurred independent of the parasitic lifestyle.
Plastome coding capacity expressed in proportion of the various functional categories in the different lineages of the plant kingdom. Genomes used as examples were published as follows: N. tabacum (Shinozaki et al. 1986; Wakasugi et al. 1998); C. rheinhardtii (Maul et al. 2002); Cyanidium caldarium (Glockner et al. 2000); Cyanophora paradoxa (Genbank Acc # U30821); Euglena gracilis (Hallick et al. 1993); Bigelowiella natans (Rogers et al. 2007); Emiliania huxleyi (Sanchez Puerta et al. 2005); Toxoplasma gondii (Genbank Acc # U87145); Vaucheria litorea (Rumpho et al. 2008); and Guillardia theta (Douglas and Penny 1999). Background colors behind the pies indicate whether the plastids are of green algal origin (green), red algal origin (pink) or of glaucophyte origin (blue)
3 Plastid Genomes from Parasitic Species
The angiosperm holoparasite Epifagus virginiana (Beechdrops) has been one of the first parasites to be thoroughly investigated with respect to its plastid genome sequence (Wolfe et al. 1992b); and these analyses have revealed a number of drastic changes that involve mainly gene losses and pseudogenizations. The significant reductions of the coding potential that embrace, among others, all photosynthesis genes in E. virginiana (Table 4.3 and Fig. 4.4), have been explained with the relaxation of the selective pressure exerted otherwise by photosynthesis. The discovery that, among the more than 150 species that are assembled within the holoparasitic genus Cuscuta, not all exhibit the same severe physiological reductions found in Epifagus but that some have, in fact, retained some basal photosynthetic activity (Hibberd et al. 1998; van der Kooij et al. 2000) has more recently enabled glimpses into the transition from photoautotrophy via intermediate mixotrophic states to complete heterotrophy (Krause 2008). A likewise gradient has, so far, not been found anywhere else.
3.1 Structural Changes
The typical organization of plastid chromosomes with a large single-copy region (LSC) and a small single-copy region (SSC) separated by two inverted repeat regions (IRA and IRB) has been retained by all parasitic angiosperms (Wolfe et al. 1992b; Funk et al. 2007; McNeal et al. 2007) (Fig. 4.3). For two Cuscuta species, C. reflexa and C. gronovii, overlapping PCR products have indicated the existence of a circular form of the plastid chromosomes (Funk et al. 2007), suggesting overall structural similarities between parasitic and nonparasitic plastid genomes. The same holds true for parasitic algae. Divergences from the standard pattern, for example, in Euglena longa are also present in the photosynthetic relative E. gracilis and are, therefore, unrelated to parasitism (Hallick et al. 1993; Gockel and Hachtel 2000).
The inverted repeats have been assigned a role as a stabilizing factor that limits genome rearrangements in chloroplasts. Nevertheless, the boundaries of inverted repeats were found to be hot spots for gene duplications or deletions (Yue et al. 2008). In line with this, the IRA–LSC junction (JLA) in C. reflexa and C. exaltata was found within the ycf2 gene, leaving one copy of this gene truncated. As a result of this reduction in IR size, there is only one copy each of rpl2 and trnI-CAU and one complete ycf2 gene (Funk et al. 2007; McNeal et al. 2007). Generally, the size of the inverted repeats in Cuscuta was reduced proportionally to the size of the plastid genome (Fig. 4.3). In E. virginiana, in contrast, the inverted repeats have suffered much less reductions so that their sizes relative to the single-copy regions are much larger. The main reason for this is that the ribosomal genes that are located on the IRs have been retained while many genes that are normally part of the single-copy regions (such as the photosynthesis genes) have been lost. However, differences between IR length and IR gene content are fairly common in higher plants, anyway, as exemplified by the comparison between tobacco and Pelargonium (Fig. 4.3), and even between species of the same genus (Goulding et al. 1996), so inverted repeat sizes cannot be correlated with a particular lifestyle.
Insertions and deletions (“indels”) of larger fragments and inversions that can affect the order of genes on the plastid genome are considered to be almost as important for the evolution of genomes as nucleotide substitutions (Yue et al. 2008). Compared to tobacco and other angiosperm plastid genomes, species of the Cuscuta subgenus Monogyna exhibit three typical sequence inversions within the plastid chromosome, two in the large single-copy region ~2 kb and ~13 kb in length and one of ~1.5 kb length in the small single-copy region (Stefanovic and Olmstead 2005; Funk et al. 2007). Otherwise, the gene order in parasitic plants was not found to be different. Whether the three inversions are of any functional significance is, however, unclear.
A high ratio of coding versus noncoding sequences was found for all Cuscuta species and some parasitic algae, such as Helicosporidium and Euglena longa, as well as for aplicoplast genomes. In contrast, Epifagus exhibits almost the same coding to noncoding sequence ratio as photosynthetic plants (Krause 2008). It has been speculated that an early reaction of the plastid genome to the parasitic lifestyle was the condensation of the genome by loss of many noncoding and possibly unimportant parts of the plastid DNA (Funk et al. 2007). As the adaptations to parasitism became more pronounced, pseudogenization and loss of functional reading frames occurred (Fig. 4.4), with the result that the relative amount of noncoding areas have increased again, as observed in Epifagus. The observation of highly compact genomes in parasitic algae and in Cuscuta might indicate, however, that the Epifagus plastid genome represents an exception.
3.2 Gene Losses in Parasitic Plant Plastomes
A recent study of 64 nonparasitic higher plant plastid genomes has revealed that 66 individual losses of genes have occurred in different species during evolution (Jansen et al. 2007). As many as 62 of these losses were confined to the more derived monocot and eudicot clades, and only four genes (chlB, L, and N as well as trnP-GGG) have disappeared from the plastid genome at a very early stage (Jansen et al. 2007). Two genes were lost particularly often: infA for which 11 independent losses have been recorded in eudicots and that is missing, among others from all Solanaceae (Table 4.3), and accD with a total of six independent losses in monocots and some eudicots. The reported losses seem to follow no specific pattern.
The adaptation to parasitism, on the other hand, has resulted in a loss of genes that is the more pronounced, the lesser the photosynthetic capacity of the parasite has become. Here, functional correlations between gene losses and metabolic activities can be drawn more easily.
3.2.1 Ndh Genes
Eleven ndh genes on a “normal” plastid genome code for a chloroplast NAD(P)H dehydrogenase (Table 4.2). These genes were reported missing in several photosynthetic genera, such as Pinus, Phalaenopsis, and Chlamydomonas (Wakasugi et al. 1994; Maul et al. 2002; Chang et al. 2006), which was interpreted as evidence that these genes are nonessential. This question seems, however, far from being resolved, as a recent report defends the view that the Ndh complex plays an essential role in photosynthesis and discusses evidence for a possible transfer of ndh genes to the nucleus in conifers and Gnetales (Martin and Sabater 2010).
A loss of ndh genes was also reported for all sequenced plastid genomes of parasitic dicots [C. reflexa, C. exaltata, C. gronovii, C. obtusiflora, and E. virginiana (Wolfe et al. 1992b; Funk et al. 2007; McNeal et al. 2007)] (Table 4.3) and the recently published underground orchid Rhizantella gardneri (Delannoy et al. 2011) and besides also for the parasitic liverwort Aneura mirabilis (Wickett et al. 2008a, b). Like in all other cases where these genes were reported missing, the entire set consisting of 11 genes has been lost without exception, since these species do no longer rely on photosynthesis.
In E. longa, the ndh genes are also reported missing. However, in this case, the loss is shared with the photosynthetic species E. gracilis.
3.2.2 Photosystem Genes
Genes for photosystem I and photosystem II as well as the cytochrome b6f complex are encoded on all nonparasitic species. Most photosystem genes were retained in the four species of Cuscuta whose plastid genome sequences are available. C. reflexa exhibits no losses, which coincides with comparatively mild reductions in the rate of photosynthesis (van der Kooij et al. 2000), while photosynthesis rates are more severely compromised in C. gronovii. Since all genes were found to be transcribed (Berg et al. 2003) in both species, the finding that the psaI gene was lost in C. gronovii (Funk et al. 2007) could be of significance in that context. The retention of photosynthesis genes is the biggest difference to the plastid genome of E. virginiana (Table 4.3). All genes associated with the bioenergetic processes of photosynthetic electron transport and ATP synthesis were either lost or pseudogenized (Table 4.3 and Fig. 4.4). The condensation grade of the plastid genome of E. virginiana with only 70 kbp and a coding capacity of just 42 genes (Wolfe et al. 1992b) is in higher plants only outmatched by the underground orchid Rhizantella (Delannoy et al. 2011).
Analyses of nucleotide substitution rates showed that the psaI gene that is missing in, for example, C. gronovii, showed significantly increased KA/KS values in C. reflexa and C. exaltata (Krause 2010). PsaI is a small subunit of photosystem I that has only one transmembrane domain and is involved in the docking of the PsaL subunit to this photosystem (Yu et al. 2008; Vanselow et al. 2009). The high KA/KS values suggest that this protein is obviously not evolving under selective constraint, and that this lack of selective pressure is what presumably leads to its eventual loss. It has, however, not yet been determined whether a copy of this gene has been transferred to the nucleus and can functionally replace the lost plastid gene.
3.2.3 The rbcL Gene
The large subunit of Rubisco, rbcL, is encoded by the plastid genome. In accordance with the loss of photosynthetic activity, this gene was lost in some aphotosynthetic species, such as E. virginiana (Wolfe et al. 1992b), C. odorata (van der Kooij et al. 2000), and R. gardneri (Delannoy et al. 2011). However, many reports on other parasitic plant families, among them holoparasitic Scrophulariaceae, showed that rbcL was surprisingly conserved as a functional plastid gene independent of whether the photosynthetic capacity was retained or not. Open reading frames for rbcL were detected, for example, in Lathrea clandestina, a parasite of alder (Alnus glutinosa), where it seems to be expressed, despite the fact that this plant lacks chlorophyll (Lusson et al. 1998). Similar situations have been described for other holoparasites (Thalouarn et al. 1994; Wolfe and dePamphilis 1997, 1998; Delavault and Thalouarn 2002).
Likewise, the parasitic liverwort Aneura mirabilis and the euglenoid alga E. longa have retained seemingly functional rbcL genes, while all photosystem genes were deleted (Gockel and Hachtel 2000; Wickett et al. 2008a). The fact that rbcL has been retained in many but not all parasitic plant plastomes makes it difficult to associate a particular meaning to this, but it has been discussed that Rubisco could have a separate metabolic function independent of photosynthesis (Krause 2008).
3.2.4 Ribosomal Protein Genes
A total of 25 genes of the ribosomal small and large subunits are encoded by most higher plant plastomes (Table 4.2). Although no tendency toward enhanced nucleotide substitution rates in ribosomal protein genes was observed in Cuscuta species (Krause 2010), parasitic plant genomes exhibit several losses of rpl and rps genes (Morden et al. 1991; Funk et al. 2007). Like with the photosynthesis genes and other gene groups, the number of losses roughly follows the gradient of dependency on heterotrophic growth. In C. reflexa, only two genes, rpl23 and rps16, have nonfunctional reading frames and behave as pseudogenes. Along with a third gene, rpl32, both genes have been completely lost in C. gronovii. E. virginiana has even suffered four losses (rpl22, rpl32, rps15, and rps16) in addition to two pseudogenizations (rpl14 and rps23) (Table 4.3).
The rpl32 gene is not only missing from some parasitic plant plastid genomes but was also lost in a number of photosynthetic angiosperms (Jansen et al. 2007), among them Populus alba (Table 4.3). In P. alba, the corresponding chloroplast gene appears to have been transferred to the nucleus. The transfer of chloroplast genes to the nucleus is a process that requires many steps, including the removal of possible introns, the gain of suitable regulatory elements, as well as the acquisition of a transit peptide that can direct the nuclear gene product back to the plastids (Bungard 2004; Ravi et al. 2008). In case of the P. alba rpl32 gene, it could be shown that it acquired the transit peptide from another plastid targeted gene, cp sod-1 (Ueda et al. 2007), thereby paving the way for the Rpl32 protein’s return into the chloroplast.
Another example for a ribosomal gene whose loss has also been observed in P. alba is rps16 (Ueda et al. 2008). In this case, however, the gene was not simply transferred to the nucleus. Rather, the original plastid rps16 gene has been lost, and this loss has been compensated for by import of the mitochondrial rps16 gene. Apparently, the nuclear gene for the mitochondrial rps16 gene has acquired a dual-targeting signal that is able to direct it to the plastids in addition to mitochondria, rendering the plastid’s own gene dispensable.
This functional replacement of a ribosomal gene from one organelle by a dually targeted counterpart from the other organelle is not a unique case. A recent report has shown that, for example, the rpl10 gene in several plant mitochondrial genomes has been replaced by a dually targeted copy of the original “chloroplast-only” targeted rpl10 isoform (Kubo and Arimura 2010). It is possible that some gene losses from parasitic plant plastomes have been compensated for in a similar manner.
3.2.5 Rpo Genes
Higher plants possess two RNA polymerases, PEP and NEP, which share the responsibility of transcribing the plastid genetic information (Hess and Börner 1999). The PEP is a multi-subunit enzyme and is encoded by four plastid genes: rpoA, rpoB, rpoC1, and rpoC2 (Table 4.2). The encoded subunits display a similarity to the eubacterial multisubunit RNA polymerase, and PEP recognizes promoters with a structure similar to bacterial promoters (Hess and Börner 1999). PEP is predominantly responsible for the transcription of photosynthesis-related genes. The generation of homoplastomic rpo knock-out mutants in tobacco, consequently, leads to a loss of photosynthetic capacity that is accompanied by an off-white phenotype (DeSantis-Maciossek et al. 1999) and a reduction in the amount of transcripts of photosynthesis genes (Krause et al. 2000; Legen et al. 2002).
The entire set of rpo genes was reported missing in E. virginiana, which has lost any photosynthetic activity and with that also any of the PEP-dependent genes. Losses of the rpoA subunit were both previously and subsequently reported for photosynthetic species, such as Euglena gracilis (Hallick et al. 1993), Pelargonium (Chumley et al. 2006), and Passiflora (Jansen et al. 2007) (Table 4.3). The operon containing rpoB, C1, and C2, on the other hand, has generally been retained on plastid genomes. To date, the only exceptions are the plastid genomes of C. gronovii and C. obtusiflora, which were the first and only plastid genomes with a confirmed loss of the entire rpo gene set in a plastid genome where photosynthesis genes are not only present (Funk et al. 2007) but, moreover, actively expressed (Berg et al. 2003). It has been shown that transcription of these genes was taken over by the nuclear-encoded RNA polymerase, NEP, which is highly homologous to the mitochondrial phage-type RNA polymerase and might have evolved from it by gene duplication (see Chap. 12).
Unlike parasitic angiosperms, apicomplexans, Helicosporidium and E. longa have retained the rpo genes in their reduced plastid genomes (Wilson et al. 1996; Gockel and Hachtel 2000; Cai et al. 2003; de Koning and Keeling 2006; Janouskovec et al. 2010). In contrast to higher plants, algae appear to possess only a single nuclear gene for a phage-type RNA polymerase and its gene product seems to be exclusively localized to the mitochondria. A second nuclear-encoded but plastid-localized RNA polymerase, NEP, seems to be missing in algae (Smith and Purton 2002). A loss of the PEP subunits encoded by the plastid rpo genes would therefore deprive the plastids of the possibility to transcribe their genetic information, making these genes essential.
3.2.6 Hypothetical Conserved Reading Frames (Ycfs)
Plastid genomes contain a number of conserved open reading frames of unknown function. The conservation of some of these sequences within plastomes from higher plants down to algae or even cyanobacteria is interpreted as strong indication for their functional importance (Ravi et al. 2008). Attempts to uncover the function of the ycf gene products have been successful in some cases, and the use of aliases to the ycf designations is becoming more common. For example, ycf5 has been renamed ccsA [for c-type cytochrome synthesis, (Xie and Merchant 1996)], ycf9 is psbZ (Swiatek et al. 2001), and ycf10 is now called cemA [for chloroplast envelope membrane protein A, (Rolland et al. 1997)]. For ease of reference, the older ycf designations will be used here in this chapter.
Nicotiana species contain nine conserved reading frames in their plastid genome sequence (ycf1-5, ycf9,10, and 15 and orf350) (Wakasugi et al. 1998; Yukawa et al. 2006), six of which have been lost in E. virginiana (ycf3, 4, 5, 9, 10, and 15). Only the pseudogenization of ycf15 is shared with the Cuscuta species (Table 4.3). The retention of ycf1 and ycf2 on all of the parasitic plant plastid genomes indicates that a function of these genes in photosynthesis can be most likely excluded and that rather, a possible role in gene expression or a photosynthesis-unrelated metabolic function can be envisaged. This observation corroborates previous findings that knockout mutants of ycf1 and ycf2 never yielded viable homoplastomic lines and that these genes are therefore essential for the survival of higher plants (Drescher et al. 2000).
3.2.7 tRNA Genes
The set of tRNA genes encoded by plastomes of photosynthetic plants encompasses around 30 genes and it has been argued that a transfer of tRNA genes to the nucleus and re-import of the RNAs is impossible (Barbrook et al. 2006; Howe and Purton 2007). Nevertheless, some parasitic species do exhibit extensive losses of tRNA genes.
In C. reflexa, only a single tRNA, that for lysine (trnK-UUU) is missing. More extended losses of tRNA genes were observed in C. gronovii, where in addition to trnK-UUU the sequence for trnV-UAC was completely eliminated from the plastid DNA, and four tRNA genes (trnA-UGC, trnG-UCC, trnI-GAU, and trnR-AGC) have been reduced to pseudogenes. In E. virginiana, a total of five tRNAs (trnA-UGC, trnC-GCA, trnI-GAU, trnR-UCU, and trnS-GGA) were pseudogenized and eight tRNAs (trnG-GCC, trnG-UCC, trnK-UUU, trnL-UAA, trnT-GGU, trnT-UGU, trnV-GAC, and trnV-UAC) lost.
The extensive loss of tRNA genes from parasitic plastids has, therefore, raised the question whether the missing tRNAs have been replaced by nuclear-encoded ones or whether the codon usage was adapted to these losses. An analysis of codon usages in two Cuscuta species has shown that all 61 sense codons were found in the coding regions of plastid genes in a similar proportion as in nonparasitic plants that possess a “full” plastid tRNA set (Funk et al. 2007). A similar picture emerges from the Epifagus plastid genome. This was interpreted as circumstantial evidence for an import of cytosolic tRNAs into the chloroplasts (Wolfe et al. 1992a; Funk et al. 2007). It is, however, also possible that an “extended wobble” behavior of the remaining tRNAs partly compensates for the losses.
3.2.8 trnK-UUU and matK
Among the tRNA genes, trnK-UUU has a special role as this tRNA gene harbors in its intron the only RNA maturase gene found on the plastid genome, matK. The trnK-UUU gene was found missing on all parasitic angiosperm plastomes, but neither the gene nor its intron and the matK gene contained within have been reported missing in any nonparasitic plants. In C. reflexa, C. exaltata, and E. virginiana, surprisingly, matK has been retained as a free-standing gene, confirming the probably essential function in plastid intron maturation that has been attributed earlier to its gene product (Hess et al. 1994; Hubschmann et al. 1996; Jenkins et al. 1997; Vogel et al. 1997). Unprecedented in higher plants, however, was the complete loss of matK from C. gronovii and C. obtusiflora (Funk et al. 2007; McNeal et al. 2007).
The exceptional loss of matK from the plastid genomes of C. gronovii and C. obtusiflora is probably closely related to the loss of group IIa introns from a whole suite of genes in parasitic plant plastids. Group IIa introns have been discussed as targets of MatK (Liere and Link 1995; Hubschmann et al. 1996; Jenkins et al. 1997; Vogel et al. 1997). Out of originally eight group IIa introns, only a single group IIa intron, namely intron 2 of clpP, is found in C. gronovii and C. obtusiflora, where the matK gene was deleted. This intron was shown to be faithfully spliced and might therefore not be a target of MatK action (Funk et al. 2007). This conclusion was supported by recent biochemical and molecular evidence linking MatK to all group IIa introns, except the clpP intron 2 (Zoschke et al. 2010).
In Apicomplexa and in E. longa, an absence of the gene matK along with all group II introns was observed as well but this does not seem to be the result of a parasitic lifestyle here. The loss of matK in these species is shared with their photosynthetic relatives since matK was reported to be already missing in Chromera velia (Janouskovec et al. 2010) and E. gracilis (Gockel and Hachtel 2000).
3.3 Reduction of RNA-Editing Sites
The loss of introns was not the only posttranscriptional processing step that has experienced a reduction. A surprise was the reduction of RNA-editing sites that was not only the result of the loss of genes that are typically richer in editing sites. Of the 30–40 editing sites of photosynthetic seed plants (Tsudzuki et al. 2001), only 17 potential sites remain in C. reflexa, while 12 have been found in C. gronovii. Several sites are only partially edited or not edited at all. The loss of editing competence as well as the reduction in the number of introns has been discussed in a very recent review (Tillich and Krause 2010) and the reader is referred to this review for further details.
3.4 Loss of PEP Promoters
Each of the two enzymes sharing the responsibility for plastid transcription (PEP and NEP) differs with respect to the promoters they bind to (Hess and Börner 1999). PEP promoters resemble prokaryotic promoters and occur mainly upstream of the photosynthetic genes, whereas the phage-type NEP promoters can be found upstream of genes for the genetic system. Many genes and operons, such as the gene cluster for the ribosomal RNAs, even possess both promoter types.
The loss of the rpo genes from two Cuscuta plastid genomes where photosynthesis genes are still present (Krause et al. 2003; Berg et al. 2004) raised the question of how the corresponding promoters have developed. The analysis of promoter motifs upstream of photosynthesis genes revealed that the consensus −10 and −35 boxes of PEP promoters have been so severely changed that they must be considered to be nonfunctional (Funk et al. 2007). For the rrn operon and the rbcL gene it could be, moreover, demonstrated that the start sites of transcription have been shifted relative to those of tobacco and that the 5′ region of the novel transcription start site revealed striking similarities to the consensus sequence recognized by the NEP polymerase, indicating a shift from PEP- to NEP-driven transcription of these genes. This shift obviously enables the plastids to transcribe the previous PEP genes with high enough efficiency to allow for low photosynthetic activity.
4 Gene Retentions and the “Raison d’être” of Reduced Plastid Genomes
Overall, many of the changes that were seen in connection with a parasitic lifestyle seem to be shared between higher plant and algal parasites. The postulated forces that must exist, according to deKoning and Keeling, for algal parasites and that shape plastid genomes even after relaxation of photosynthetic selection pressure (de Koning and Keeling 2006) do seemingly also apply to higher plants. The most reduced plastid genomes in both groups (i.e., Epifagus for seed plants and Apicomplexa for algae) are characterized by a domination of genetic system genes and only two to four genes with functions outside of gene expression are present.
One question that has repeatedly been asked but so far never satisfactorily been answered is the mystery why plastid genomes were retained in nonphotosynthetic organisms. In this context, genes that encode for subunits of the gene-expression machinery, such as ribosomal RNAs and proteins, are hardly of much interest, since they are presumed to only secure the expression of the “key” plastid gene(s). Consequently, the answers why the plastid genome has not been lost altogether have been sought in the other retained genes.
In many plastid genomes of parasitic plant species, intact reading frames of the rbcL gene that codes for the photosynthesis protein Rubisco have been retained, despite the loss of photosynthetic activity (Krause 2008). It has been suggested that Rubisco could assume a second function in lipid biosynthesis (Schwender et al. 2004). An argument that strengthens the “essential lipid biosynthesis” hypothesis is the retention of the accD gene even in strongly reduced genomes, such as that from E. virginiana. Tobacco plastome mutants, where accD was interrupted, did never reach a homoplastomic state, underlining the essentiality of this gene for plants (Kode et al. 2005). However, in Apicomplexa there is no accD gene and rbcL is missing in apicomplexan parasites as well as in Epifagus and some Cuscuta species, just to name some (see Sect. 4.3.2).
Another set of genes that appear essential for plastid development independent of photosynthetic capacity are those that encode the subunits of the Clp Protease, clpP and clpC. Clp most likely performs chaperone functions and is engaged in protein import into the plastid. While in seed plants, the clpP gene was retained even in reduced plastomes, such as that of Epifagus, Apicomplexa have retained the gene clpC. However, exceptions are found also here (e.g., Helicosporidium), weakening any hypothesis that was tentatively built up around the clp genes.
Among the protein-coding genes, the two hypothetical reading frames ycf1 and ycf2 remain. Knockouts of each gene resulted persistently in heteroplasmy (Drescher et al. 2000; Shikanai et al. 2001; Kuroda and Maliga 2003) and both belong to the reduced gene set of extremely reduced plastid genomes, such as that of E. virginiana. However, their function might well be found to be associated with gene expression (Bock 2007), which would eliminate also these genes from being candidates for the “raison d’être” of plastid genomes.
Some recent alternative attempts at an explanation for the retention of plastid genomes circle around two transfer RNA genes. The first is the gene for the glutamyl-tRNA (trnE). This (trnE) gene fulfills three tasks in plastids: besides its role in protein biosynthesis, it plays a well-known role in the synthesis of δ-aminolaevulinic acid and, thereby, in heme biosynthesis (Jahn et al. 1992), and may regulate transcriptional activity of the NEP (Hanaoka et al. 2005), although this last function has been later challenged (Bohne et al. 2009). The essentiality of heme biosynthesis and the belief that a functional transfer of tRNA genes from the plastid to the nuclear genome is unlikely if not impossible (Barbrook et al. 2006) has been used as an argument for why a plastid genome has been retained, however much reduced it is. It has even been predicted some years ago that trnE may be the only gene that is found in all genomes, regardless of the degree of reductions (Barbrook et al. 2006). So far, all sequenced plastomes fulfill this prediction. However, also this hypothesis has a caveat, since in Plasmodium, at least, heme was found to be synthesized by an exclusively mitochondrial-located pathway, and it is therefore independent of plastid trnE. A second hypothesis was brought up recently, where the formylmethionyl-tRNA (tRNA fM) plays the main role (Howe and Purton 2007). tRNA fM is needed for translation initiation in plastids and mitochondria, but the only tRNA fM gene in Plasmodium is the one that is located in the apicoplast. Therefore, the formylmethionyl-tRNA pool of the plastids was proposed to be shared by the mitochondria, rendering this particular tRNA indispensable (Howe and Purton 2007). Whether this holds true for further parasitic species still awaits confirmation.
5 From Loss of Photosynthesis to Loss of Plastids?
As described in the previous chapters, all nonphotosynthetic land plants without exception have retained a more or less cryptic plastid with a plastome that exhibits a set of typical losses and also of typical gene retentions. A number of nonphotosynthetically living algae and descendents thereof have likewise retained a cryptic plastid that apparently fulfills some essential functions for the parasites.
For a long time, the debate has been going on why these plastids with their “crippled” plastid genomes have remained so steadfast in the nonphotosynthetic species. The discovery that the apicomplexan species Cryptosporidium has lost its plastid (Huang et al. 2004) has given the debate a new spin. Evidence for plastid-derived genes in the nonphotosynthetic oomycete Phytophthora (Tyler et al. 2006), in Ciliates (Reyes-Prieto et al. 2008), and in trypanosomatid parasites (Bodyl et al. 2010) that has recently been presented, has nourished the discussion of whether these lineages have evolved from a photosynthetic, plastid-bearing ancestor (Janouskovec et al. 2010). This scenario would imply that the loss of photosynthesis genes and the reduction of other plastome features could be just one intermediate step and that there are no evolutionary restrictions that would preclude the total loss of the plastid genome or the entire plastid. It is surprising, however, that so far no species has been found where the plastid DNA but not the plastid compartment as a whole were lost.
6 Conclusion and Perspectives
Analyses of nucleotide substitution rates have revealed that the mutation rates in plastid genomes are considerably lower than in their nuclear counterparts (Wolfe et al. 1987). The hypothesis that the selective pressure exerted by the photoautotrophic lifestyle has contributed significantly to this conservation of the plastid genome can best be tested by analyzing species that have evolved under a different type or level of selective constraint. Such species are present in the various groups of parasitic plants and algae. In the “omics” age, tools for high-throughput analysis and annotation of genomes are not only available, they are, more importantly, also affordable and require very small amounts of plant material. Consequently, not only the number of published chloroplast genomes from agriculturally important plants have increased considerably over the last 5 years, but also the number of genomes from “cryptic” plastids of parasitic species. While this information has been instrumental in getting an insight into the evolution of plastid genomes, a number of questions await answers in the future. Among those is the extent of a possible nuclear transfer of genes that are regarded as essential and that are as of now reported missing from the plastid genome. Another question concerns the coordination of cryptic plastid and nuclear gene expression in a nonphotosynthetic setting. To answer these questions, it will be necessary to concentrate on the nuclear genomes of some of these species in the future.
Abbreviations
- bp:
-
Base pairs
- NEP:
-
Nuclear-encoded RNA polymerase
- PEP:
-
Plastid encoded RNA polymerase
- Rubisco:
-
Ribulose-1,5-bisphosphate carboxylase/oxygenase
References
Barbrook AC, Howe CJ (2000) Minicircular plastid DNA in the dinoflagellate Amphidinium operculatum. Mol Gen Genet 263:152–158
Barbrook AC, Howe CJ, Purton S (2006) Why are plastid genomes retained in non-photosynthetic organisms? Trends Plant Sci 11:101–108
Barkman TJ, McNeal JR, Lim SH, Coat G, Croom HB, Young ND, Depamphilis CW (2007) Mitochondrial DNA suggests at least 11 origins of parasitism in angiosperms and reveals genomic chimerism in parasitic plants. BMC Evol Biol 7:248
Berg S, Krupinska K, Krause K (2003) Plastids of three Cuscuta species differing in plastid coding capacity have a common parasite-specific RNA composition. Planta 218:135–142
Berg S, Krause K, Krupinska K (2004) The rbcL genes of two Cuscuta species, C. gronovii and C. subinclusa, are transcribed by the nuclear-encoded plastid RNA polymerase (NEP). Planta 219:541–546
Bidartondo MI, Bruns TD, Weiss M, Sergio C, Read DJ (2003) Specialized cheating of the ectomycorrhizal symbiosis by an epiparasitic liverwort. Proc Biol Sci 270:835–842
Bock R (2007) Structure, function, and inheritance of plastid genomes. In: Bock R (ed) Cell and molecular biology of plastids, vol 19, Topics in current genetics. Springer, Heidelberg
Bodyl A, Mackiewicz P, Milanowski R (2010) Did trypanosomatid parasites contain a eukaryotic alga-derived plastid in their evolutionary past? J Parasitol 96:465–475
Bohne A-V, Weihe A, Börner T (2009) Transfer RNAs inhibit Arabidopsis phage-type RNA polymerases. Endocytobiosis Cell Res 19:63–69
Borza T, Popescu CE, Lee RW (2005) Multiple metabolic roles for the nonphotosynthetic plastid of the green alga Prototheca wickerhamii. Eukaryot Cell 4:253–261
Bungard RA (2004) Photosynthetic evolution in parasitic plants: insight from the chloroplast genome. Bioessays 26:235–247
Cai X, Fuller AL, McDougald LR, Zhu G (2003) Apicoplast genome of the coccidian Eimeria tenella. Gene 321:39–46
Cavalier-Smith T (1999) Principles of protein and lipid targeting in secondary symbiogenesis: euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J Eukaryot Microbiol 46:347–366
Cavalier-Smith T (2002) The phagotrophic origin of eukaryotes and phylogenetic classification of Protozoa. Int J Syst Evol Microbiol 52:297–354
Chang CC, Lin HC, Lin IP, Chow TY, Chen HH, Chen WH, Cheng CH, Lin CY, Liu SM, Chaw SM (2006) The chloroplast genome of Phalaenopsis aphrodite (Orchidaceae): comparative analysis of evolutionary rate with that of grasses and its phylogenetic implications. Mol Biol Evol 23:279–291
Chumley TW, Palmer JD, Mower JP, Fourcade HM, Calie PJ, Boore JL, Jansen RK (2006) The complete chloroplast genome sequence of Pelargonium x hortorum: organization and evolution of the largest and most highly rearranged chloroplast genome of land plants. Mol Biol Evol 23:2175–2190
de Koning AP, Keeling PJ (2004) Nucleus-encoded genes for plastid-targeted proteins in Helicosporidium: functional diversity of a cryptic plastid in a parasitic alga. Eukaryot Cell 3:1198–1205
de Koning AP, Keeling PJ (2006) The complete plastid genome sequence of the parasitic green alga Helicosporidium sp. is highly reduced and structured. BMC Biol 4:12
Delannoy E, Fujii S, des Francs CC, Brundrett M, Small I (2011) Rampant gene loss in the underground orchid Rhizanthella gardneri highlights evolutionary constraints on plastid genomes. Mol Biol Evol 28(7):2077–2086
Delavault P, Thalouarn P (2002) The obligate root parasite Orobanche cumana exhibits several rbcL sequences. Gene 297:85–92
Deng XW, Wing RA, Gruissem W (1989) The chloroplast genome exists in multimeric forms. Proc Natl Acad Sci U S A 86:4156–4160
DeSantis-Maciossek G, Kofer W, Bock A, Schoch S, Maier RM, Wanner G, Rüdiger W, Koop HU, Herrmann RG (1999) Targeted disruption of the plastid RNA polymerase genes rpoA, B and C1: molecular biology, biochemistry and ultrastructure. Plant J 18:477–489
Douglas SE, Penny SL (1999) The plastid genome of the cryptophyte alga, Guillardia theta: complete sequence and conserved synteny groups confirm its common ancestry with red algae. J Mol Evol 48:236–244
Drescher A, Ruf S, Calsa T Jr, Carrer H, Bock R (2000) The two largest chloroplast genome-encoded open reading frames of higher plants are essential genes. Plant J 22:97–104
Funk HT, Berg S, Krupinska K, Maier UG, Krause K (2007) Complete DNA sequences of the plastid genomes of two parasitic flowering plant species, Cuscuta reflexa and Cuscuta gronovii. BMC Plant Biol 7:45
Glockner G, Rosenthal A, Valentin K (2000) The structure and gene repertoire of an ancient red algal plastid genome. J Mol Evol 51:382–390
Gockel G, Hachtel W (2000) Complete gene map of the plastid genome of the nonphotosynthetic euglenoid flagellate Astasia longa. Protist 151:347–351
Goulding SE, Olmstead RG, Morden CW, Wolfe KH (1996) Ebb and flow of the chloroplast inverted repeat. Mol Gen Genet 252:195–206
Hagopian JC, Reis M, Kitajima JP, Bhattacharya D, de Oliveira MC (2004) Comparative analysis of the complete plastid genome sequence of the red alga Gracilaria tenuistipitata var. liui provides insights into the evolution of rhodoplasts and their relationship to other plastids. J Mol Evol 59:464–477
Hallick RB, Hong L, Drager RG, Favreau MR, Monfort A, Orsat B, Spielmann A, Stutz E (1993) Complete sequence of Euglena gracilis chloroplast DNA. Nucleic Acids Res 21:3537–3544
Hanaoka H, Kanamaru K, Fujiwara M, Takahashi H, Tanaka K (2005) Glutamyl-tRNA mediates a switch in RNA polymerase use during chloroplast biogenesis. EMBO Rep 6:545–550
Hess W, Börner T (1999) Organellar RNA polymerases of higher plants. Int Rev Cytol 190:1–59
Hess W, Müller A, Nagy F, Börner T (1994) Ribosome-deficient plastids affect transcription of light-induced nuclear genes: genetic evidence for a plastid-derived signal. Mol Gen Genet 242:305–312
Hibberd JM, Bungard RA, Press MC, Jeschke WD, Scholes JD, Quick WP (1998) Localization of photosynthetic metabolism in the parasitic angiosperm Cuscuta reflexa. Planta 205:506–513
Howe CJ, Purton S (2007) The little genome of apicomplexan plastids: its raison d’etre and a possible explanation for the ‘delayed death’ phenomenon. Protist 158:121–133
Huang J, Mullapudi N, Lancto CA, Scott M, Abrahamsen MS, Kissinger JC (2004) Phylogenomic evidence supports past endosymbiosis, intracellular and horizontal gene transfer in Cryptosporidium parvum. Genome Biol 5:R88
Hubschmann T, Hess WR, Borner T (1996) Impaired splicing of the rps12 transcript in ribosome-deficient plastids. Plant Mol Biol 30:109–123
Jahn D, Verkamp E, Soll D (1992) Glutamyl-transfer RNA: a precursor of heme and chlorophyll biosynthesis. Trends Biochem Sci 17:215–218
Janouskovec J, Horak A, Obornik M, Lukes J, Keeling PJ (2010) A common red algal origin of the apicomplexan, dinoflagellate, and heterokont plastids. Proc Natl Acad Sci U S A 107:10949–10954
Jansen RK, Cai Z, Raubeson LA, Daniell H, Depamphilis CW, Leebens-Mack J, Muller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee SB, Peery R, McNeal JR, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci U S A 104:19369–19374
Jansen RK, Wojciechowski MF, Sanniyasi E, Lee SB, Daniell H (2008) Complete plastid genome sequence of the chickpea (Cicer arietinum) and the phylogenetic distribution of rps12 and clpP intron losses among legumes (Leguminosae). Mol Phylogenet Evol 48:1204–1217
Jenkins BD, Kulhanek DJ, Barkan A (1997) Nuclear mutations that block group II RNA splicing in maize chloroplasts reveal several intron classes with distinct requirements for splicing factors. Plant Cell 9:283–296
Kode V, Mudd EA, Iamtham S, Day A (2005) The tobacco plastid accD gene is essential and is required for leaf development. Plant J 44:237–244
Kolodner R, Tewari KK (1972) Molecular size and conformation of chloroplast deoxyribonucleic acid from pea leaves. J Biol Chem 247:6355–6364
Krause K (2008) From chloroplasts to “cryptic” plastids: evolution of plastid genomes in parasitic plants. Curr Genet 54:111–121
Krause K (2010) Plastid genome variation and selective constraint in species of the parasitic plant genus Cuscuta. In: Urbano KV (ed) Advances in genetics research, vol. 2, chapter 15. Nova Science Publishers, New York, pp 1–15
Krause K, Maier RM, Kofer W, Krupinska K, Herrmann RG (2000) Disruption of plastid-encoded RNA polymerase genes in tobacco: expression of only a distinct set of genes is not based on selective transcription of the plastid chromosome. Mol Gen Genet 263:1022–1030
Krause K, Berg S, Krupinska K (2003) Plastid transcription in the holoparasitic plant genus Cuscuta: parallel loss of the rrn16 PEP-promoter and of the rpoA and rpoB genes coding for the plastid-encoded RNA polymerase. Planta 216:815–823
Kubo N, Arimura S (2010) Discovery of the rpl10 gene in diverse plant mitochondrial genomes and its probable replacement by the nuclear gene for chloroplast RPL10 in two lineages of angiosperms. DNA Res 17:1–9
Kuroda H, Maliga P (2003) The plastid clpP1 protease gene is essential for plant development. Nature 425:86–89
Kuroiwa T (1991) The replication, differentiation, and inheritance of plastids with emphasis on the concept of organelle nuclei. Int Rev Cytol 128:1–62
Lane CE, Archibald JM (2008) The eukaryotic tree of life: endosymbiosis takes its TOL. Trends Ecol Evol 23:268–275
Legen J, Kemp S, Krause K, Profanter B, Herrmann RG, Maier RM (2002) Comparative analysis of plastid transcription profiles of entire plastid chromosomes from tobacco attributed to wild-type and PEP-deficient transcription machineries. Plant J 31:171–188
Liere K, Link G (1995) RNA-binding activity of the matK protein encoded by the chloroplast trnK intron from mustard (Sinapis alba L.). Nucleic Acids Res 23:917–921
Lilly JW, Havey MJ, Jackson SA, Jiang J (2001) Cytogenomic analyses reveal the structural plasticity of the chloroplast genome in higher plants. Plant Cell 13:245–254
Lim L, McFadden GI (2010) The evolution, metabolism and functions of the apicoplast. Philos Trans R Soc Lond B Biol Sci 365:749–763
Lusson NA, Delavault PM, Thalouarn PA (1998) The rbcL gene from the non-photosynthetic parasite Lathraea clandestina is not transcribed by a plastid-encoded RNA polymerase. Curr Genet 34:212–215
Martin M, Sabater B (2010) Plastid ndh genes in plant evolution. Plant Physiol Biochem 48:636–645
Matsuzaki M, Kuroiwa H, Kuroiwa T, Kita K, Nozaki H (2008) A cryptic algal group unveiled: a plastid biosynthesis pathway in the oyster parasite Perkinsus marinus. Mol Biol Evol 25:1167–1179
Maul JE, Lilly JW, Cui L, dePamphilis CW, Miller W, Harris EH, Stern DB (2002) The Chlamydomonas reinhardtii plastid chromosome: islands of genes in a sea of repeats. Plant Cell 14:2659–2679
McNeal JR, Kuehl JV, Boore JL, de Pamphilis CW (2007) Complete plastid genome sequences suggest strong selection for retention of photosynthetic genes in the parasitic plant genus Cuscuta. BMC Plant Biol 7:57
Morden CW, Wolfe KH, dePamphilis CW, Palmer JD (1991) Plastid translation and transcription genes in a non-photosynthetic plant: intact, missing and pseudo genes. EMBO J 10:3281–3288
Ohta N, Matsuzaki M, Misumi O, Miyagishima SY, Nozaki H, Tanaka K, Shin IT, Kohara Y, Kuroiwa T (2003) Complete sequence and analysis of the plastid genome of the unicellular red alga Cyanidioschyzon merolae. DNA Res 10:67–77
Ohyama K, Fukuzawa H, Kohchi T, Shirai H, Sano T, Sano S, Takeuchi M, Chang Z, Aota S-I, Inokuchi H, Ozeki H (1986) Chloroplast gene organization deduced form complete sequence of liverwort Marchantia polymorpha chloroplast DNA. Nature 322:572–574
Oldenburg DJ, Bendich AJ (2004) Most chloroplast DNA of maize seedlings in linear molecules with defined ends and branched forms. J Mol Biol 335:953–970
Palmer JD (1990) Contrasting modes and tempos of genome evolution in land plant organelles. Trends Genet 6:115–120
Ravi V, Khurana JP, Tyagi AK, Khurana P (2008) An update on chloroplast genomes. Plant Syst Evol 271:101–122
Reith M, Munholland J (1993) A high-resolution gene map of the chloroplast genome of the red alga Porphyra purpurea. Plant Cell 5:465–475
Reyes-Prieto A, Bhattacharya D (2007) Phylogeny of nuclear-encoded plastid-targeted proteins supports an early divergence of glaucophytes within Plantae. Mol Biol Evol 24:2358–2361
Reyes-Prieto A, Moustafa A, Bhattacharya D (2008) Multiple genes of apparent algal origin suggest ciliates may once have been photosynthetic. Curr Biol 18:956–962
Robbens S, Derelle E, Ferraz C, Wuyts J, Moreau H, Van de Peer Y (2007) The complete chloroplast and mitochondrial DNA sequence of Ostreococcus tauri: organelle genomes of the smallest eukaryote are examples of compaction. Mol Biol Evol 24:956–968
Rogers MB, Gilson PR, Su V, McFadden GI, Keeling PJ (2007) The complete chloroplast genome of the chlorarachniophyte Bigelowiella natans: evidence for independent origins of chlorarachniophyte and euglenid secondary endosymbionts. Mol Biol Evol 24:54–62
Rolland N, Dorne AJ, Amoroso G, Sultemeyer DF, Joyard J, Rochaix JD (1997) Disruption of the plastid ycf10 open reading frame affects uptake of inorganic carbon in the chloroplast of Chlamydomonas. EMBO J 16:6713–6726
Rumpho ME, Worful JM, Lee J, Kannan K, Tyler MS, Bhattacharya D, Moustafa A, Manhart JR (2008) Horizontal gene transfer of the algal nuclear gene psbO to the photosynthetic sea slug Elysia chlorotica. Proc Natl Acad Sci U S A 105:17867–17871
Sanchez Puerta MV, Bachvaroff TR, Delwiche CF (2005) The complete plastid genome sequence of the haptophyte Emiliania huxleyi: a comparison to other plastid genomes. DNA Res 12:151–156
Sanchez-Puerta MV, Lippmeier JC, Apt KE, Delwiche CF (2007) Plastid genes in a non-photosynthetic dinoflagellate. Protist 158:105–117
Scharff LB, Koop HU (2006) Linear molecules of tobacco ptDNA end at known replication origins and additional loci. Plant Mol Biol 62:611–621
Schwender J, Goffman F, Ohlrogge JB, Shachar-Hill Y (2004) Rubisco without the Calvin cycle improves the carbon efficiency of developing green seeds. Nature 432:779–782
Shikanai T, Shimizu K, Ueda K, Nishimura Y, Kuroiwa T, Hashimoto T (2001) The chloroplast clp Gene, encoding a proteolytic subunit of ATP-dependent protease, is indispensable for chloroplast development in tobacco. Plant and Cell Physiol 42:264–273
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchi-Shinozaki K, Ohto C, Torazawa K, Meng BY, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Sugiura M (1986) The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J 5:2043–2049
Simpson CL, Stern DB (2002) The treasure trove of algal chloroplast genomes. Surprises in architecture and gene content, and their functional implications. Plant Physiol 129:957–966
Smith AC, Purton S (2002) The transcriptional apparatus of algal plastids. Eur J Phycol 37:301–311
Stefanovic S, Olmstead RG (2005) Down the slippery slope: plastid genome evolution in Convolvulaceae. J Mol Evol 61:292–305
Swiatek M, Kuras R, Sokolenko A, Higgs D, Olive J, Cinque G, Muller B, Eichacker LA, Stern DB, Bassi R, Herrmann RG, Wollman FA (2001) The chloroplast gene ycf9 encodes a photosystem II (PSII) core subunit, PsbZ, that participates in PSII supramolecular architecture. Plant Cell 13:1347–1367
Tartar A, Boucias DG (2004) The non-photosynthetic, pathogenic green alga Helicosporidium sp. has retained a modified, functional plastid genome. FEMS Microbiol Lett 233:153–157
Thalouarn PA, Theodet C, Russo NM, Delavault P (1994) The reduced plastid genome of a non-photosynthetic angiosperm Orobanche hederae has retained the rbcL gene. Plant Physiol Biochem 32:233–242
Tillich M, Krause K (2010) The ins and outs of editing and splicing of plastid RNAs: lessons from parasitic plants. N Biotechnol 27:256–266
Tsudzuki T, Wakasugi T, Sugiura M (2001) Comparative analysis of RNA editing sites in higher plant chloroplasts. J Mol Evol 53:327–332
Turmel M, Otis C, Lemieux C (1999) The complete chloroplast DNA sequence of the green alga Nephroselmis olivacea: insights into the architecture of ancestral chloroplast genomes. Proc Natl Acad Sci U S A 96:10248–10253
Tyler BM, Tripathy S, Zhang X, Dehal P, Jiang RH, Aerts A, Arredondo FD, Baxter L, Bensasson D, Beynon JL, Chapman J, Damasceno CM, Dorrance AE, Dou D, Dickerman AW, Dubchak IL, Garbelotto M, Gijzen M, Gordon SG, Govers F, Grunwald NJ, Huang W, Ivors KL, Jones RW, Kamoun S, Krampis K, Lamour KH, Lee MK, McDonald WH, Medina M, Meijer HJ, Nordberg EK, Maclean DJ, Ospina-Giraldo MD, Morris PF, Phuntumart V, Putnam NH, Rash S, Rose JK, Sakihama Y, Salamov AA, Savidor A, Scheuring CF, Smith BM, Sobral BW, Terry A, Torto-Alalibo TA, Win J, Xu Z, Zhang H, Grigoriev IV, Rokhsar DS, Boore JL (2006) Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313:1261–1266
Ueda M, Fujimoto M, Arimura S, Murata J, Tsutsumi N, Kadowaki K (2007) Loss of the rpl32 gene from the chloroplast genome and subsequent acquisition of a preexisting transit peptide within the nuclear gene in Populus. Gene 402:51–56
Ueda M, Nishikawa T, Fujimoto M, Takanashi H, Arimura S, Tsutsumi N, Kadowaki K (2008) Substitution of the gene for chloroplast RPS16 was assisted by generation of a dual targeting signal. Mol Biol Evol 25:1566–1575
van der Kooij TA, Krause K, Dorr I, Krupinska K (2000) Molecular, functional and ultrastructural characterisation of plastids from six species of the parasitic flowering plant genus Cuscuta. Planta 210:701–707
Vanselow C, Weber AP, Krause K, Fromme P (2009) Genetic analysis of the Photosystem I subunits from the red alga, Galdieria sulphuraria. Biochim Biophys Acta 1787:46–59
Vogel J, Hubschmann T, Borner T, Hess WR (1997) Splicing and intron-internal RNA editing of trnK-matK transcripts in barley plastids: support for MatK as an essential splice factor. J Mol Biol 270:179–187
Wakasugi T, Tsudzuki J, Ito S, Nakashima K, Tsudzuki T, Sugiura M (1994) Loss of all ndh genes as determined by sequencing the entire chloroplast genome of the black pine Pinus thunbergii. Proc Natl Acad Sci U S A 91:9794–9798
Wakasugi T, Nagai T, Kapoor M, Sugita M, Ito M, Ito S, Tsudzuki J, Nakashima K, Tsudzuki T, Suzuki Y, Hamada A, Ohta T, Inamura A, Yoshinaga K, Sugiura M (1997) Complete nucleotide sequence of the chloroplast genome from the green alga Chlorella vulgaris: the existence of genes possibly involved in chloroplast division. Proc Natl Acad Sci U S A 94:5967–5972
Wakasugi T, Sugita M, Tsudzuki T, Sugiura M (1998) Updated map of tobacco chloroplast DNA. Plant Mol Biol Rep 16:231–241
Wickett NJ, Fan Y, Lewis PO, Goffinet B (2008a) Distribution and evolution of pseudogenes, gene losses, and a gene rearrangement in the plastid genome of the nonphotosynthetic liverwort, Aneura mirabilis (Metzgeriales, Jungermanniopsida). J Mol Evol 67:111–122
Wickett NJ, Zhang Y, Hansen SK, Roper JM, Kuehl JV, Plock SA, Wolf PG, DePamphilis CW, Boore JL, Goffinet B (2008b) Functional gene losses occur with minimal size reduction in the plastid genome of the parasitic liverwort Aneura mirabilis. Mol Biol Evol 25:393–401
Wilson RJ, Denny PW, Preiser PR, Rangachari K, Roberts K, Roy A, Whyte A, Strath M, Moore DJ, Moore PW, Williamson DH (1996) Complete gene map of the plastid-like DNA of the malaria parasite Plasmodium falciparum. J Mol Biol 261:155–172
Wolfe AD, dePamphilis CW (1997) Alternate paths of evolution for the photosynthetic gene rbcL in four nonphotosynthetic species of Orobanche. Plant Mol Biol 33:965–977
Wolfe AD, dePamphilis CW (1998) The effect of relaxed functional constraints on the photosynthetic gene rbcL in photosynthetic and nonphotosynthetic parasitic plants. Mol Biol Evol 15:1243–1258
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci U S A 84:9054–9058
Wolfe KH, Morden CW, Ems SC, Palmer JD (1992a) Rapid evolution of the plastid translational apparatus in a nonphotosynthetic plant: loss or accelerated sequence evolution of tRNA and ribosomal protein genes. J Mol Evol 35:304–317
Wolfe KH, Morden CW, Palmer JD (1992b) Function and evolution of a minimal plastid genome from a nonphotosynthetic parasitic plant. Proc Natl Acad Sci U S A 89:10648–10652
Xie Z, Merchant S (1996) The plastid-encoded ccsA gene is required for heme attachment to chloroplast c-type cytochromes. J Biol Chem 271:4632–4639
Yoon HS, Hackett JD, Van Dolah FM, Nosenko T, Lidie KL, Bhattacharya D (2005) Tertiary endosymbiosis driven genome evolution in dinoflagellate algae. Mol Biol Evol 22:1299–1308
Yu J, Ma PJ, Shi DJ, Li SM, Wang CL (2008) Homologous comparisons of photosynthetic system I genes among cyanobacteria and chloroplasts. J Integr Plant Biol 50:929–940
Yue F, Cui L, dePamphilis CW, Moret BM, Tang J (2008) Gene rearrangement analysis and ancestral order inference from chloroplast genomes with inverted repeat. BMC Genomics 9(Suppl 1):S25
Yukawa M, Tsudzuki T, Sugiura M (2006) The chloroplast genome of Nicotiana sylvestris and Nicotiana tomentosiformis: complete sequencing confirms that the Nicotiana sylvestris progenitor is the maternal genome donor of Nicotiana tabacum. Mol Genet Genomics 275:367–373
Zhang Z, Green BR, Cavalier-Smith T (1999) Single gene circles in dinoflagellate chloroplast genomes. Nature 400:155–159
Zoschke R, Nakamura M, Liere K, Sugiura M, Borner T, Schmitz-Linneweber C (2010) An organellar maturase associates with multiple group II introns. Proc Natl Acad Sci U S A 107:3245–3250
Acknowledgments
Critical comments by Prof. Karsten Fischer, University of Tromsø, during the preparation of this manuscript are gratefully acknowledged. Prof. Thomas Börner is thanked for reviewing the final version of this chapter.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Krause, K. (2012). Plastid Genomes of Parasitic Plants: A Trail of Reductions and Losses. In: Bullerwell, C. (eds) Organelle Genetics. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-22380-8_4
Download citation
DOI: https://doi.org/10.1007/978-3-642-22380-8_4
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-22379-2
Online ISBN: 978-3-642-22380-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)