Abstract
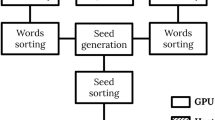
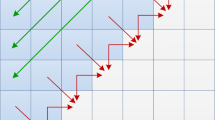
Discovery of motifs that are repeated in groups of biological sequences is a major task in bioinformatics. Iterative methods such as expectation maximization (EM) are used as a common approach to find such patterns. However, corresponding algorithms are highly compute-intensive due to the small size and degenerate nature of biological motifs. Runtime requirements are likely to become even more severe due to the rapid growth of available gene transcription data. In this paper we present a novel approach to accelerate motif discovery based on commodity graphics hardware (GPUs). To derive an efficient mapping onto this type of architecture, we have formulated the compute-intensive parts of the popular MEME tool as streaming algorithms. Our experimental results show that a single GPU allows speedups of one order of magnitude with respect to the sequential MEME implementation. Furthermore, parallelization on a GPU-cluster even improves the speedup to two orders of magnitude.
Chapter PDF
Similar content being viewed by others
References
Bailey, T.L., Elkan, C.: Unsupervised learning of multiple motifs in biopolymers using expectation maximization. Machine Learning 21, 51–80 (1995)
Bailey, T.L., Williams, N., Misleh, C., Li, W.W.: MEME: discovering and analyzing DNA and protein motifs. Nucleic Acid Research 34, W369–W373 (2006)
Grundy, W.N., Bailey, T.L., Elkan, C.P.: ParaMEME: A parallel implementation and a web interface for a DNA and protein motif discovery tool. Computer Applications in the Biological Sciences (CABIOS) 12, 303–310 (1996)
Kessenich, J., Baldwin, D., Rost, R.: The OpenGL Shading Language, Document Revision 8 (2006), http://www.opengl.org/documentation/glsl/
Lawrence, C., Altschul, S., Boguski, M., Liu, J., Neuwald, A., Wootton, J.: Detecting subtle sequence signals: a Gibbs sampling strategy for multiple alignment. Science 262, 208–214 (1993)
Liu, W., Schmidt, B., Voss, G., Muller-Wittig, W.: Streaming Algorithms for Biological Sequence Alignment on GPUs. IEEE Transactions on Parallel and Distributed Systems 18(10), 1270–1281 (2007)
Manavski, S.A., Valle, G.: CUDA compatible GPU cards as efficient hardware accelerators for Smith-Waterman sequence alignment. BMC Bioinformatics 9(Suppl. 2), S10 (2008)
Sabatti, C., Rohlin, L., Lange, K., Liao, J.C.: Vocabulon: a dictionary model approach for reconstruction and localization of transcription factor binding sites. Bioinformatics 21(7), 922–931 (2005)
Sandve, G.K., Nedland, M., Syrstad, B., Eidsheim, L.A., Abul, O., Drablas, F.: Accelerating motif discovery: Motif matching on parallel hardware. In: Bücher, P., Moret, B.M.E. (eds.) WABI 2006. LNCS (LNBI), vol. 4175, pp. 197–206. Springer, Heidelberg (2006)
Schatz, M.C., Trapnell, C., Delcher, A.L., Varshney, A.: High-throughput sequence alignment using Graphics Processing Units. BMC Bioinformatics 8(474) (2007)
Sumazin, P., et al.: DWE: Discriminating Word Enumerator. Bioinformatics 21(1), 31038 (2005)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Chen, C., Schmidt, B., Weiguo, L., Müller-Wittig, W. (2008). GPU-MEME: Using Graphics Hardware to Accelerate Motif Finding in DNA Sequences. In: Chetty, M., Ngom, A., Ahmad, S. (eds) Pattern Recognition in Bioinformatics. PRIB 2008. Lecture Notes in Computer Science(), vol 5265. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-88436-1_38
Download citation
DOI: https://doi.org/10.1007/978-3-540-88436-1_38
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-88434-7
Online ISBN: 978-3-540-88436-1
eBook Packages: Computer ScienceComputer Science (R0)