Abstract
RNA metabolism impacts different steps of mRNA life cycle including splicing, polyadenylation, nucleo-cytoplasmic export, translation, and decay. Growing evidence indicates that defects in any of these steps lead to devastating diseases in humans. This chapter reviews the various RNA metabolic mechanisms that are disrupted in Myotonic Dystrophy—a trinucleotide repeat expansion disease—due to dysregulation of RNA-Binding Proteins. We also compare Myotonic Dystrophy to other microsatellite expansion disorders and describe how some of these mechanisms commonly exert direct versus indirect effects toward disease pathologies.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Microsatellite repeat expansions
- Post-transcriptional gene regulation
- RNA toxicity
- Alternative splicing and polyadenylation
- RNA-binding proteins
8.1 Introduction
Gene expression is a highly coordinated multistep process, which allows organisms to integrate intrinsic and environmental information to exert appropriate cellular functions. The expression of most genes can be regulated at distinct stages of RNA metabolism including synthesis or transcription, post-transcriptional processing or maturation, nucleo-cytoplasmic export, translation, as well as degradation at a rate that is often dictated by transcript- and cell-type-specific cues. Although transcription is a general point of control, many co- and post-transcriptional pre-mRNA processing events add substantial capacity to tune overall gene expression [1]. The typical pre-mRNA processing events comprise 5′ capping, splicing, and 3′ polyadenylation, which are directly linked to the nucleo-cytoplasmic export and eventual fate of mRNAs. RNA-Binding Proteins (RBPs) are essential in carrying out these processing events in both the nucleus and cytoplasm by interacting with RNA sequence or structural elements and forming distinct mRNA-protein (mRNP) complexes [2]. Disruption of RBP function(s), therefore, frequently results in deleterious RNA metabolism defects that in some cases become pathogenic [3, 4].
Neurodegenerative diseases are a heterogeneous group of neurological disorders characterized by progressive degeneration of structure and function of the central or peripheral nervous systems. Aberrant RNA metabolism is increasingly implicated in neurodegenerative diseases, a subset of which are caused by the expansion of short repetitive elements (microsatellites) within particular genes [5]. The causative repeat expansion mutation for this group of disorders is unstable because the repeat size changes through generations and even within an individual, as different tissues have cell populations with variable repeat length and in some cases the repeat length varies within the same tissue [6]. The severity of a repeat expansion disease is dependent on numerous variables, including the length of the repeat, its sequence context, and the native function of the protein-coding gene with which the repeat is associated. A typical pathogenic feature of these diseases is the accumulation of repeat-containing transcripts into aberrant RNA foci, which can sequester RBPs and prevent them from performing their normal functions [7,8,9]. Interestingly, once the repeat length cross a critical number, the repeat-containing RNAs can undergo phase separation—partitioning into granules due to multivalent base-pairing between repeat RNAs—or spontaneous gelation to form RNA foci, explaining why disease symptoms appear to be triggered after the expansions have reached a particular threshold number [10].
8.2 Toxicity of Coding and Noncoding Microsatellite Repeat Expansions
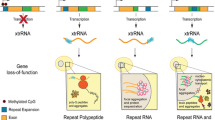
Over 25 human genes with tandem repeat expansions have been identified to date, and these disease-causing repeats can occur in the coding or noncoding regions [6] (Fig. 8.1 and Table 8.1). Majority of the microsatellites arise due to the expansion of trinucleotide repeats. However, expanded tetranucleotide, pentanucleotide, and hexanucleotide repeats are also detected. In the early 1990s, two microsatellites were discovered providing the first evidence that simple repeat expansions are linked to human disease. Fragile X Syndrome (FXS)—an X-linked disorder caused by CGG repeat expansions in the 5′ untranslated region (UTR) of the FMR1 gene—is the most prevalent form of inherited cognitive impairment and mental retardation [11,12,13,14,15,16]. The repeat expansion in FXS causes loss of FMR1 gene product FMRP, a polyribosome-associated RBP that binds ~4% of brain mRNAs and regulates their expression—either enhancing or suppressing translation through unknown mechanisms [17,18,19,20].
Spinal and bulbar muscular atrophy (SBMA)—the other microsatellite disease discovered along with FXS—arises due to a CAG repeat expansion in the coding region of the X chromosome-linked androgen receptor (AR) gene [21]. The discovery of SBMA was soon followed by the elucidation of a similar mutation as the basis for a group of disorders now known as the polyglutamine (polyQ ) neurodegenerative diseases (Table 8.1). Along with SBMA, the polyQ diseases include Huntington disease (HD), dentatorubral-pallidoluysian atrophy, and six spinocerebellar ataxias (SCA ) 1, 2, 3, 6, 7, and 17 [22]. As a group, these nine diseases are among the more common forms of inherited neurodegeneration. The translation of exons containing CAG repeats gives rise to elongated stretches of polyQs in mutant proteins, which aggregate into nuclear or cytoplasmic inclusions in the diseased brain [23,24,25]. Several observations indicate that the CAG repeat-containing RNAs, in the absence of coding for a protein, may also be a source of toxicity in polyQ diseases [26, 27]. GGGGCC hexanucleotide repeat expansion in the C9ORF72 gene has gained much attention in the past few years and is now considered the most frequent inherited cause of Amyotrophic lateral sclerosis (ALS) and Frontotemporal dementia (FTD ) [28, 29]. Pathology occurs due to the toxicity of expanded repeats, which are transcribed in both the sense and antisense directions and give rise to distinct sets of intracellular RNA and protein aggregates [30,31,32,33].
Myotonic Dystrophy (DM) is part of a group of diseases characterized by repeat expansions in noncoding regions of genes. DM is defined in two clinical and molecular forms: myotonic dystrophy type 1 (DM1), and type 2 (DM2), both of which are inherited in an autosomal dominant fashion. The combined worldwide incidence of DM is approximately 1 in 8000 [34, 35]. DM1 is the most prevalent form of adult onset muscular dystrophy [36] and is caused by a CTG repeat expansion in the 3′ UTR of Dystrophia Myotonica Protein Kinase (DMPK) gene [37, 38]. DM2, on the other hand, is caused by a CCTG repeat expansion in an intron of Zinc Finger Protein 9 (ZNF9) gene [39]. While 5–37 repeats are considered normal, DM1 patients can have up to several thousand CTG repeats, which can reduce expression of DMPK [40] (Fig. 8.2a). DMPK is expressed in multiple tissues, and the major symptoms of the disease include muscle hyperexcitability (myotonia), progressive muscle wasting, cardiac defects, insulin resistance, and neuropsychiatric disturbances [41,42,43,44]. Table 8.1 provides further description of tissue-specific symptoms observed in DM and other microsatellite expansion disorders.
Schematic showing different pathological mechanisms for Myotonic Dystrophy type 1 (DM1) and 2 (DM2). (a) Causative mutation for DM1 is CUG repeat expansion in 3′UTR of DMPK gene and for DM2 is CCUG repeat expansion in intron 1 of ZNF9 gene. The severity of the disease is dependent on the number of repeats. Although these mutations are in two different genes, the disease mechanisms for both diseases are surprisingly similar. Most of the pathology is consistent with the toxic RNA gain-of-function mechanism and affects general RNA metabolism in both the nucleus and cytoplasm. (b) After transcription, the repeat-containing transcripts form stable hairpin loop comprising secondary structures (pink), which aggregate to form ribonuclear foci. (c) Members of the Mbnl family of RNA-binding proteins (RBPs) MBNL1/2 (purple) bind the CUG Fig. 8.2 (continued) or CCUG repeats and are sequestered in the ribonuclear foci. (d) Hyperphosphorylation by PKC stabilizes another RBP, CELF1, resulting in its gain-of-function. (e) Both MBNL and CELF proteins regulate various aspects of RNA metabolism during normal development. Alterations in their functional levels due to toxic repeat RNA cause adult-to-fetal reversion of splicing and polyadenylation for many pre-mRNAs in the nucleus. (f) MBNL depletion also leads to cellular mislocalization of many mRNAs. CELF1 gain-of-function further affects (g) miRNA metabolism and (h) mRNA translation. (i) Dysregulation of MBNL and CELF activity in the cytoplasm also affects mRNA stability through various mechanisms. (j) Both sense and antisense CUG/CCUG-containing transcripts are subject to RAN translation in all three frames giving rise to homopolymeric polypeptides that accumulate in the cytoplasm and form pathological intracellular aggregates
8.3 RNA Metabolism Defects in Myotonic Dystrophy
Closely after the discovery of repeats, the DMPK haploinsufficiency model was put forward to explain the DM1 pathology. However, the removal of DMPK gene in mice failed to recapitulate the major neuromuscular symptoms of DM1 [45, 46]. A separate hypothesis proposed that expanded CTG repeats might affect the expression of nearby genes. Although the adjacent gene, SIX5, exhibits reduced expression in DM1 patients [47], Six5 knockout mice also do not reproduce DM1 muscle pathology [48]. Instead, the CTG repeats alone, regardless of the gene context, are sufficient to induce pathogenic features of DM1 [49, 50]. The predominant pathology of DM1 actually stems from the toxic effects of expanded CUG RNA, which disrupts the normal activity of certain RBPs. Further support for the RNA toxicity model comes from the finding that although the repeat expansion in DM2 is on an entirely different gene, both diseases exhibit similar symptoms.
In both DM1 and DM2, the RNAs with expanded repeats (CUG in DM1; and CCUG in DM2) fold into stable hairpin loops that accumulate as ribonuclear foci in the nuclei of affected tissues [9] (Fig. 8.2b). These expanded RNA transcripts directly trap RBPs such as muscleblind-like proteins (MBNLs) and cause upregulation of CUG-binding protein 1 (CELF1) family of alternative splicing factors [51,52,53,54], which results in aberrant splicing of many transcripts and a broad, multi-systemic phenotype (Fig. 8.2c, d). Alternative pre-mRNA splicing generates much of the transcriptome diversity in higher eukaryotes as it enables the production of multiple transcripts with potentially different functions from each individual gene [55]. Alternative splicing decisions are generally influenced by cis-acting regulatory elements within pre-mRNAs that promote or inhibit exon recognition, as well as expression/activity of trans-acting factors (e.g., MBNL and CELF proteins) that bind to these cis elements and regulate the accessibility of the spliceosome to splice sites [56]. The misregulated splicing events in DM are usually developmentally regulated and exhibit an adult-to-embryonic switch in splicing patterns (Fig. 8.2e). Some of these embryonic isoforms fail to meet the adult tissue requirements and thus directly contribute to the overall disease pathology [54].
8.3.1 Misregulation of mRNA Processing
MBNL loss-of-function in DM1 and DM2 is a prominent example of RBP sequestration by disease-associated microsatellite expansion RNAs. The MBNL proteins were initially identified in Drosophila melanogaster for their requirement in muscle development and eye differentiation [57], and they were later shown as direct regulators of alternative splicing [58]. There are three MBNL paralogues in mammals, named MBNL1–3. MBNL1 and MBNL2 are widely expressed across many tissues, including brain, heart, muscle, and liver, whereas MBNL3 expression is restricted to the placenta [59]. In a majority of tissues, MBNL1 and MBNL2 mRNA levels rise during differentiation [60, 61]. Besides their roles in pre-mRNA processing, MBNLs also influence gene expression by regulating cellular mRNA transport, stability as well as microRNA biogenesis [62,63,64,65,66,67]. The high expression of MBNL1 in the heart and skeletal muscle is consistent with the most severe DM phenotypes in these tissues. For instance, independent of the repeat expansion, Mbnl1 deletion in mice reproduces many of the cardinal symptoms of DM1 such as myotonia, myopathy, cataracts, and misregulation of developmentally regulated RNA processing [63, 68].
The expanded repeat-containing RNAs in DM sequester MBNL1, 2, and 3 in nuclear RNA foci [69,70,71], and this protein redistribution explains the inhibition of their normal functions predominantly in alternative splicing and polyadenylation, microRNA processing, and mRNA localization [58, 62, 67, 72,73,74,75]. The MBNL loss-of-function hypothesis is further supported by studies on Mbnl single- and compound-knockout mice, which recapitulate many of the DM phenotypes [68, 76,77,78]. The extent of symptoms, however, varies depending on the tissue context, relative concentrations of MBNL paralogues, and the degree to which they are sequestered [78]. For instance, compared to skeletal muscle, only few splicing defects are observed in the brains of Mbnl1 knockout mice [63, 79]. Alternatively, Mbnl2 knockout mice exhibit a number of DM-related central nervous system abnormalities including irregular REM sleep propensity and deficits in spatial memory [76], which is consistent with the observation that MBNL2 expression in the brain is higher than MBNL1 [59]. MBNL2 is directly sequestered by repeat expansions in the brain tissue of human DM patients resulting in misregulation of alternative splicing and polyadenylation of its normal RNA targets [80]. One of the most misspliced mRNA due to loss of MBNL2 is human microtubule-associated protein tau (MAPT) in the DM1 frontal cortex [80]. RNA toxicity mediated through MBNL2 sequestration leads to abnormal expression of tau isoforms and the progressive appearance of neurofibrillary tangles composed of intraneuronal aggregates of hyper-phosphorylated tau protein [81].
More recently, MBNL proteins were found to serve essential roles in poly(A) site selection for many transcripts (Fig. 8.2e). By integrating HITS-CLIP and RNA-seq from MBNL knockout cells and transgenic DM1 mouse model, along with minigene reporter studies, Swanson and colleagues demonstrated that MBNL proteins directly suppress or activate polyadenylation for thousands of pre-mRNAs [75, 80]. Thus, MBNL proteins coordinate multiple pre-mRNA processing steps and their sequestration in DM depletes them from their normal RNA targets.
Besides MBNL loss-of-function, there is accumulation and aberrant sub-cellular distribution of another splicing factor CELF1 in DM. CELF proteins are normally downregulated during postnatal striated muscle development, which facilitates fetal-to-adult splicing transitions in hundreds of muscle transcripts [61, 82]. CELF1 actually does not colocalize with RNA foci [83], and its upregulation in DM1 occurs through two separate mechanisms. First, CELF1 protein is stabilized through its hyper-phosphorylation [84]; and second, reduced levels of microRNAs in DM1 derepress CELF1 protein translation [85, 86] (Fig. 8.2d, g, h). The situation is less clear in DM2, with conflicting reports of normal [87, 88] and increasing CELF1 protein levels [89] in patient tissues and cells. It is interesting to note that for many pre-mRNAs whose splicing is disrupted in DM1, CELF1 and MBNL1 regulate them in an antagonistic manner [58, 61, 90,91,92]. The antagonism, however, is not due to direct competition for the binding site as both CELF1 and MBNL1 bind and regulate splicing independently via distinct cis-acting RNA motifs.
In addition to MBNL and CELF proteins, other RNA splicing factors are implicated in DM. For instance, hnRNP H binds to DMPK-derived CUG-expanded RNAs in vitro and increased hnRNP H levels may also contribute toward DM pathogenesis [93]. hnRNP H forms a repressor complex with MBNL1 and nine other proteins (hnRNP H2, H3, F, A2/B1, K, L, DDX5, DDX17, and DHX9) in normal myoblast extracts but elevated hnRNP H levels in DM1 disrupt the stoichiometry of these complexes which affects splicing of specific pre-mRNAs [94, 95]. Since expanded CUG repeat RNAs fold into hairpin structures [96], the partial recruitment and colocalization of the RNA helicase p68/DDX5 with RNA foci may also have a contributing role in splicing dysregulation. Moreover, p68/DDX5 can modulate MBNL1-binding activity, and its colocalization with nuclear RNA foci can further stimulate MBNL1 binding to repeat RNAs [97].
8.3.2 Misregulation of mRNA Localization and Stability
Following transcription, newly synthesized and fully processed mRNAs are bound by specific RBPs to form export-competent mRNPs, which help their transport through the nuclear pore complex (NPC). Some pre-mRNAs are processed at the speckle periphery before being exported and repeat-containing nuclear foci can colocalize at the periphery of nuclear speckles, a non-membrane bound nuclear assembly of macromolecules including splicing factors. The presence of expanded CUG repeats may, therefore, prevent entry of other RNAs into the nuclear speckle [98, 99]. However, in DM2, the mutant ZNF9 mRNA is exported normally as the expanded CCUG repeats are removed during splicing. The nuclear foci formed by DM2 intronic repeats are widely dispersed in the nucleoplasm and not associated with nuclear speckles. Also, it is not yet clear whether the DM1 and/or DM2 nuclear foci contain partially degraded fragments of CUG or CCUG repeats or larger intact RNAs respectively.
As discussed above, CELF1 upregulation and MBNL sequestration by the CUG repeats in DM1 cause misprocessing of hundreds of transcripts. Aberrant processing results in nucleocytoplasmic export defects for many of these transcripts. Furthermore, MBNL proteins are localized both in the nucleus and cytoplasm and several studies have demonstrated their direct roles in mRNA localization [62, 100] (Fig. 8.2f). For instance, by interacting with the 3′-UTR of Integrin α3, MBNL2 moves it to the plasma membrane for its local translation [64]. Similarly, MBNL1 also plays major roles in mRNA localization and membrane-associated translation. Transcriptome-wide analyses of subcellular compartments from mouse myoblasts showed widespread defects in mRNA localization upon combined depletion of MBNL1 and MBNL2 [62]. Many of the mislocalized mRNAs encode for secreted proteins, extracellular matrix components, and proteins involved in cell–cell communication. MBNL depletion in DM can thus have a significant impact on mRNA localization potentially affecting proper neuromuscular junction formation.
In the cytoplasm, MBNLs also regulate mRNA stability [101] (Fig. 8.2i). MBNL1 specifically recognizes YGCY-containing motifs within the 3′-UTR regions and destabilizes the target mRNAs through unknown mechanisms [65, 92]. CELF1, on the other hand, induces mRNA decay of short-lived transcripts through interactions with GU-rich elements (GREs) in their 3′-UTR and possibly recruitment of poly(A)-specific ribonuclease, which promotes deadenylation of target transcripts [102,103,104]. Many of the CELF mRNA targets with GREs encode proteins essential for muscle cell development and function [105,106,107,108]. Interestingly, CELF1 binds to the mRNAs coding for SRP protein subunits and promotes their decay [109]. Signal recognition particle (SRP) is a cytoplasmic ribonucleoprotein complex, which regulates the translation of secreted and membrane-associated proteins. It is likely that the CELF1 overexpression contributes to the faster turnover of SRP mRNAs and the reduced SRP levels thereby attenuate the protein secretory pathway in DM1 [109].
8.3.3 Misregulation of mRNA Translation
CELF1 is additionally involved in the regulation of mRNA translation [106, 110,111,112] (Fig. 8.2h). The affinity of CELF1 toward its mRNA targets can be modulated through phosphorylation [113]. For instance, phosphorylated CELF1 interacts with a subunit of initiation factor eIF2, leading to the recruitment of translational machinery to target mRNAs [106]. In myoblasts, AKT phosphorylates CELF1 and increases its affinity for CCND1 mRNA. During myoblast-to-myotube differentiation, cyclinD3-cdk4/6 phosphorylates CELF1, which increases CELF1 interaction with 5′-UTR of p21 mRNA (a cell cycle inhibitor) and enhances its translation. Myoblasts from DM1 patients show an increased interaction between CELF1 and AKT and have reduced cyclinD3-CDK4/6 levels during differentiation [105]. Moreover, DM1 myoblasts during differentiation show a reduced ability to withdraw from cell cycle, which may be due to the altered translation of P21 or myogenic transcription factor MEF2A by CELF1 [111, 112].
mRNA translation in DM1 is also affected due to microRNA deregulation (Fig. 8.2g). A subset of developmentally regulated microRNAs associated with cardiac arrhythmias is downregulated in the hearts of DM1 patients and mice [67, 86]. Downregulation of these microRNAs recapitulates particular gene expression deficits seen in DM1 hearts including enhanced protein levels of miR-1 targets CX43 and Cav1.2 as well as miR-23a/b target CELF1 [67, 86]. In DM1 and DM2 skeletal muscle biopsies, both the levels and cellular distribution of several evolutionarily conserved microRNAs are altered affecting their downstream targets [114,115,116,117]. Furthermore, specific microRNAs are differentially detected in peripheral blood plasma of DM1 patients, which inversely correlate with skeletal muscle strength and may serve as noninvasive biomarkers [118]. More recently, reduced expression of miR-200c/141 tumor suppressor family was shown to correlate with increased oncologic risk in women with DM1 especially for gynecologic, brain, and thyroid cancer [119].
Besides altering cellular translation through misregulation of RBPs and microRNAs, the microsatellite expansions also promote unconventional translation of repeats in multiple reading frames producing homopolymeric peptides that aggregate in both the nucleus and the cytoplasm [120] (Fig. 8.2j). Designated as Repeat Associated Non-AUG Translation (RAN translation), it was first described for the expanded CAG and CTG repeats that cause spinocerebellar ataxia 8 (SCA8) and DM1, respectively [120]. Interestingly, the efficiency of RAN translation increases with the size of repeats and when RNA forms hairpin-like structures [121]. Additionally, the cells making the toxic RAN protein products are prone to apoptosis as detected in tissues of affected patients, indicating a potential contribution of RAN to pathogenesis. In addition to DM1, Zu et al. recently demonstrated that in DM2 the tetranucleotide expansion repeats are bidirectionally transcribed, and the resulting transcripts are RAN translated, producing tetrapeptide expansion proteins with Leu-Pro-Ala-Cys (LPAC) from the sense strand or Gln-Ala-Gly-Arg (QAGR) repeats from the antisense strand [122]. These RAN proteins were readily detected in the DM2 patient brains; however, the specific roles of these RAN proteins regarding toxicity, mechanism of action, and their regulation are yet to be determined.
Since their original discovery, RAN translation has now been observed in many other repeat-expansion diseases, including ALS/FTD, FXTAS, and Huntington’s disease [52, 123]. However, the exact mechanisms initiating translation from these repeats likely differ across diverse sequence contexts [124]. For instance, in case of FMR1, expanded CGG repeats in the 5′-UTR initiate CAP-dependent RAN translation upstream of the canonical AUG start codon, producing FMRpolyGlycine and FMRpolyAlanine in FXTAS [123, 125]. In contrast to FXTAS, the expanded repeats in DM1 exist within the 3′ UTR of DMPK mRNA, which is not in the normal path of ribosome scanning; thus, unconventional ribosome interactions must contribute in their translation. For HTT in Huntington’s disease, the CAG repeats are in the ORF, and canonical translation starts at the native AUG codon upstream of the repeats. But in some instances, HTTpolySerine and HTTpolyAlanine proteins are also produced due to RAN-translation and frame shifting from the normal HTTpolyGlutamine frame of the repeats [126]. Finally, in case of ALS/FTD, the GGGGCC repeats are within C9ORF72 intron, and the RAN-translation generates polyGlycine-Alanine, polyGlycine-Arginine, and polyGlycine-Proline dipeptide products [31, 127]. The RAN translation in this case, however, may occur from the intron retained transcript, spliced lariat, or a 3′ truncated RNA generated due to stalled transcription [124, 128].
8.4 Disrupted Function of RBPs in Other Microsatellite Expansion Disorders
Recent paradigm-shifting advances have established that defective RNA processing through disrupted function of RBPs is central to many other repeat expansion diseases (Table 8.2). For instance, RBP defects occur in both familial and sporadic cases of ALS/FTD [129, 130]. Mutations in TARDBP and FUS genes respectively encoding TDP-43 and FUS/TLS proteins result in abnormal aggregation of these proteins in neurons and are considered pathogenic for ALS/FTD. TDP-43 and FUS/TLS are RNA/DNA-binding proteins, with noticeable structural and functional similarities.
TDP-43 functions in multiple RNA processing steps including pre-mRNA splicing [131,132,133,134], RNA stability [135,136,137], and transport [138]. Similar to TDP-43, FUS interacts with serine-arginine (SR) proteins that serve diverse roles in splicing [139] and regulates transcription by recruiting other RBPs through noncoding RNAs [140]. Hence, the association of TDP43 and FUS/TLS with ALS and FTD is redirecting research efforts toward identifying additional RBPs that are mutated in neurological diseases, defining their normal RNA substrates and determining the misprocessed RNAs that underlie particular disease symptoms. In fact, mutations in several other RBPs that are functionally and structurally similar to FUS/TLS such as TAF15 [141, 142] and EWSR1 [143, 144], as well as the less closely related RBPs—Ataxin 2 [145], hnRNPA2B1 [146], hnRNPA1 [146], and Matrin3 [147] were recently identified. Among these RBPs, TDP-43, FUS, and hnRNPA1 harbor low complexity domains (LCDs), which can polymerize and drive phase separation to form dynamic membrane-less organelles or liquid droplets. For instance, a 57-residue segment within the FUS-LCD was recently shown to assemble into a fibril core that promotes phase-separation and hydrogel formation. Interestingly, phosphorylation of the core-forming residues by DNA-dependent protein kinase dissolves the FUS-LCD liquid droplets providing a molecular basis for the dynamics of LCD polymerization and phase separation [148].
Disease-associated mutations within LCDs of RBPs also enhance prion-like properties and accelerate the shift from liquid to solid phase disturbing proper ribonucleoprotein (RNP) formation [127, 149, 150]. These mutations likely trigger protein aggregation due to aberrant self-assembly of LCDs. The cytoplasmic aggregation of RBPs not only affects their typical functions in RNA metabolism but also diminishes general nucleocytoplasmic trafficking, a common consequence of ALS-initiating mutations [151,152,153]. While the exact reasons impeding nuclear/cytoplasmic transport in ALS are not yet fully established, multiple independent mechanisms have been proposed. For example, nucleocytoplasmic trafficking defects can arise due to proteotoxicity caused by cytoplasmic β-sheet containing protein aggregations [154], direct interactions between repeat RNAs and nuclear import factors [153], or inhibition by RAN translation-products of repeat RNAs [151]. Interestingly, arginine-containing dipeptide repeats produced from RAN translation of hexanucleotide GGGGCC expansions in ALS interact with LCDs of RBPs, which disrupts the dynamics and functions of membrane-less organelle formation by LCDs [155, 156]. Furthermore, subsets of these arginine-containing dipeptides frequently bind to the LCDs encoded by the nuclear pore proteins blocking the transport of macromolecules into and out of the nucleus [157]. Thus, interaction of RAN translation products with LCDs is a yet another pathogenic mechanism that interferes with the normal function of RBPs in microsatellite expansion disorders.
8.5 Conclusions
The past decade has seen remarkable progress in our understanding of the molecular pathogenesis of microsatellite repeat expansion disorders. Although the repeats may vary in terms of their length and location within a gene or the multiple ways through which they cause disease, one commonality of microsatellite expansions is the production of toxic RNA species containing repeats. Mechanistically, the pathology arises either due to loss-of-function of the affected gene, or gain-of-function of the repeat-containing RNAs. Regarding loss-of-function, the repeats can induce transcriptional silencing of the affected gene through epigenetic modifications or produce a non-functional protein that contains a long stretch of homopolymeric amino acids. In case of gain-of-function, the RNAs with expanded repeats often sequester RBPs and thus disrupt their normal activities. Alternatively, the translated protein with a repetitive stretch of homopolymeric peptide sequence can misfold, aggregate, and trap critical cellular proteins causing nucleo-cytoplasmic export defects and further proteotoxicity. For a number of repeat expansion disorders, there is an intricate overlap of such loss- and gain-of-function mechanisms resulting in complex molecular pathologies. We envision that for many repeat expansions, the future investigations will be geared toward determining the unique versus overlapping disease mechanisms, dissecting direct versus indirect RNA metabolism defects, and finally, understanding whether alterations in RNA metabolism occur early or during late stages of the disease.
References
Arif W, Datar G, Kalsotra A. Intersections of post-transcriptional gene regulatory mechanisms with intermediary metabolism. Biochim Biophys Acta. 2017;1860(3):349–62.
Lewis CJ, Pan T, Kalsotra A. RNA modifications and structures cooperate to guide RNA-protein interactions. Nat Rev Mol Cell Biol. 2017;18(3):202–10.
Scotti MM, Swanson MS. RNA mis-splicing in disease. Nat Rev Genet. 2016;17(1):19–32.
Brinegar AE, Cooper TA. Roles for RNA-binding proteins in development and disease. Brain Res. 2016;1647:1–8.
Mirkin SM. Expandable DNA repeats and human disease. Nature. 2007;447(7147):932–40.
La Spada AR, Taylor JP. Repeat expansion disease: progress and puzzles in disease pathogenesis. Nat Rev Genet. 2010;11(4):247–58.
Renoux AJ, Todd PK. Neurodegeneration the RNA way. Prog Neurobiol. 2012;97(2):173–89.
O’Rourke JR, Swanson MS. Mechanisms of RNA-mediated disease. J Biol Chem. 2009;284(12):7419–23.
Zhang N, Ashizawa T. RNA toxicity and foci formation in microsatellite expansion diseases. Curr Opin Genet Dev. 2017;44:17–29.
Jain A, Vale RD. RNA phase transitions in repeat expansion disorders. Nature. 2017;546(7657):243–7.
Heitz D, Rousseau F, Devys D, Saccone S, Abderrahim H, Le Paslier D, Cohen D, Vincent A, Toniolo D, Della Valle G, et al. Isolation of sequences that span the fragile X and identification of a fragile X-related CpG island. Science. 1991;251(4998):1236–9.
Kremer EJ, Pritchard M, Lynch M, Yu S, Holman K, Baker E, Warren ST, Schlessinger D, Sutherland GR, Richards RI. Mapping of DNA instability at the fragile X to a trinucleotide repeat sequence p(CCG)n. Science. 1991;252(5013):1711–4.
Oberle I, Rousseau F, Heitz D, Kretz C, Devys D, Hanauer A, Boue J, Bertheas MF, Mandel JL. Instability of a 550-base pair DNA segment and abnormal methylation in fragile X syndrome. Science. 1991;252(5009):1097–102.
Verkerk AJ, Pieretti M, Sutcliffe JS, Fu YH, Kuhl DP, Pizzuti A, Reiner O, Richards S, Victoria MF, Zhang FP, et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell. 1991;65(5):905–14.
Bhakar AL, Dolen G, Bear MF. The pathophysiology of fragile X (and what it teaches us about synapses). Annu Rev Neurosci. 2012;35:417–43.
Santoro MR, Bray SM, Warren ST. Molecular mechanisms of fragile X syndrome: a twenty-year perspective. Annu Rev Pathol. 2012;7:219–45.
Ashley CT Jr, Wilkinson KD, Reines D, Warren ST. FMR1 protein: conserved RNP family domains and selective RNA binding. Science. 1993;262(5133):563–6.
Brown V, Small K, Lakkis L, Feng Y, Gunter C, Wilkinson KD, Warren ST. Purified recombinant Fmrp exhibits selective RNA binding as an intrinsic property of the fragile X mental retardation protein. J Biol Chem. 1998;273(25):15521–7.
Siomi H, Choi M, Siomi MC, Nussbaum RL, Dreyfuss G. Essential role for KH domains in RNA binding: impaired RNA binding by a mutation in the KH domain of FMR1 that causes fragile X syndrome. Cell. 1994;77(1):33–9.
Siomi H, Siomi MC, Nussbaum RL, Dreyfuss G. The protein product of the fragile X gene, FMR1, has characteristics of an RNA-binding protein. Cell. 1993;74(2):291–8.
La Spada AR, Wilson EM, Lubahn DB, Harding AE, Fischbeck KH. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature. 1991;352(6330):77–9.
Orr HT, Zoghbi HY. Trinucleotide repeat disorders. Annu Rev Neurosci. 2007;30:575–621.
Davies SW, Turmaine M, Cozens BA, DiFiglia M, Sharp AH, Ross CA, Scherzinger E, Wanker EE, Mangiarini L, Bates GP. Formation of neuronal intranuclear inclusions underlies the neurological dysfunction in mice transgenic for the HD mutation. Cell. 1997;90(3):537–48.
DiFiglia M, Sapp E, Chase KO, Davies SW, Bates GP, Vonsattel JP, Aronin N. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science. 1997;277(5334):1990–3.
Paulson HL, Perez MK, Trottier Y, Trojanowski JQ, Subramony SH, Das SS, Vig P, Mandel JL, Fischbeck KH, Pittman RN. Intranuclear inclusions of expanded polyglutamine protein in spinocerebellar ataxia type 3. Neuron. 1997;19(2):333–44.
Li LB, Bonini NM. Roles of trinucleotide-repeat RNA in neurological disease and degeneration. Trends Neurosci. 2010;33(6):292–8.
Gatchel JR, Zoghbi HY. Diseases of unstable repeat expansion: mechanisms and common principles. Nat Rev Genet. 2005;6(10):743–55.
DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ, Nicholson AM, Finch NA, Flynn H, Adamson J, et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron. 2011;72(2):245–56.
Renton AE, Majounie E, Waite A, Simon-Sanchez J, Rollinson S, Gibbs JR, Schymick JC, Laaksovirta H, van Swieten JC, Myllykangas L, et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron. 2011;72(2):257–68.
Ash PE, Bieniek KF, Gendron TF, Caulfield T, Lin WL, Dejesus-Hernandez M, van Blitterswijk MM, Jansen-West K, Paul JW III, Rademakers R, et al. Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron. 2013;77(4):639–46.
Mori K, Weng SM, Arzberger T, May S, Rentzsch K, Kremmer E, Schmid B, Kretzschmar HA, Cruts M, Van Broeckhoven C, et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science. 2013;339(6125):1335–8.
Xu Z, Poidevin M, Li X, Li Y, Shu L, Nelson DL, Li H, Hales CM, Gearing M, Wingo TS, et al. Expanded GGGGCC repeat RNA associated with amyotrophic lateral sclerosis and frontotemporal dementia causes neurodegeneration. Proc Natl Acad Sci U S A. 2013;110(19):7778–83.
Zu T, Liu Y, Banez-Coronel M, Reid T, Pletnikova O, Lewis J, Miller TM, Harms MB, Falchook AE, Subramony SH, et al. RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc Natl Acad Sci U S A. 2013;110(51):E4968–77.
Faustino NA, Cooper TA. Pre-mRNA splicing and human disease. Genes Dev. 2003;17(4):419–37.
Wheeler TM. Myotonic dystrophy: therapeutic strategies for the future. Neurotherapeutics. 2008;5(4):592–600.
Harper P. Myotonic dystrophy. London: W.B. Saunders; 2001.
Brook JD, McCurrach ME, Harley HG, Buckler AJ, Church D, Aburatani H, Hunter K, Stanton VP, Thirion JP, Hudson T, et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell. 1992;68(4):799–808.
Mahadevan M, Tsilfidis C, Sabourin L, Shutler G, Amemiya C, Jansen G, Neville C, Narang M, Barcelo J, O’Hoy K, et al. Myotonic dystrophy mutation: an unstable CTG repeat in the 3′ untranslated region of the gene. Science. 1992;255(5049):1253–5.
Liquori CL, Ricker K, Moseley ML, Jacobsen JF, Kress W, Naylor SL, Day JW, Ranum LP. Myotonic dystrophy type 2 caused by a CCTG expansion in intron 1 of ZNF9. Science. 2001;293(5531):864–7.
Yum K, Wang ET, Kalsotra A. Myotonic dystrophy: disease repeat range, penetrance, age of onset, and relationship between repeat size and phenotypes. Curr Opin Genet Dev. 2017;44:30–7.
Groh WJ, Groh MR, Saha C, Kincaid JC, Simmons Z, Ciafaloni E, Pourmand R, Otten RF, Bhakta D, Nair GV, et al. Electrocardiographic abnormalities and sudden death in myotonic dystrophy type 1. N Engl J Med. 2008;358(25):2688–97.
Heatwole C, Bode R, Johnson N, Quinn C, Martens W, McDermott MP, Rothrock N, Thornton C, Vickrey B, Victorson D, et al. Patient-reported impact of symptoms in myotonic dystrophy type 1 (PRISM-1). Neurology. 2012;79(4):348–57.
Phillips MF, Harper PS. Cardiac disease in myotonic dystrophy. Cardiovasc Res. 1997;33(1):13–22.
Salehi LB, Bonifazi E, Stasio ED, Gennarelli M, Botta A, Vallo L, Iraci R, Massa R, Antonini G, Angelini C, et al. Risk prediction for clinical phenotype in myotonic dystrophy type 1: data from 2,650 patients. Genet Test. 2007;11(1):84–90.
Jansen G, Groenen PJ, Bachner D, Jap PH, Coerwinkel M, Oerlemans F, van den Broek W, Gohlsch B, Pette D, Plomp JJ, et al. Abnormal myotonic dystrophy protein kinase levels produce only mild myopathy in mice. Nat Genet. 1996;13(3):316–24.
Berul CI, Maguire CT, Aronovitz MJ, Greenwood J, Miller C, Gehrmann J, Housman D, Mendelsohn ME, Reddy S. DMPK dosage alterations result in atrioventricular conduction abnormalities in a mouse myotonic dystrophy model. J Clin Invest. 1999;103(4):R1–7.
Thornton CA, Wymer JP, Simmons Z, McClain C, Moxley RT III. Expansion of the myotonic dystrophy CTG repeat reduces expression of the flanking DMAHP gene. Nat Genet. 1997;16(4):407–9.
Klesert TR, Cho DH, Clark JI, Maylie J, Adelman J, Snider L, Yuen EC, Soriano P, Tapscott SJ. Mice deficient in Six5 develop cataracts: implications for myotonic dystrophy. Nat Genet. 2000;25(1):105–9.
Mankodi A, Logigian E, Callahan L, McClain C, White R, Henderson D, Krym M, Thornton CA. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science. 2000;289(5485):1769–73.
Gomes-Pereira M, Cooper TA, Gourdon G. Myotonic dystrophy mouse models: towards rational therapy development. Trends Mol Med. 2011;17(9):506–17.
Echeverria GV, Cooper TA. RNA-binding proteins in microsatellite expansion disorders: mediators of RNA toxicity. Brain Res. 2012;1462:100–11.
Cleary JD, Ranum LP. Repeat associated non-ATG (RAN) translation: new starts in microsatellite expansion disorders. Curr Opin Genet Dev. 2014;26:6–15.
Mohan A, Goodwin M, Swanson MS. RNA-protein interactions in unstable microsatellite diseases. Brain Res. 2014;1584:3–14.
Chau A, Kalsotra A. Developmental insights into the pathology of and therapeutic strategies for DM1: back to the basics. Dev Dyn. 2015;244(3):377–90.
Nilsen TW, Graveley BR. Expansion of the eukaryotic proteome by alternative splicing. Nature. 2010;463(7280):457–63.
Kalsotra A, Cooper TA. Functional consequences of developmentally regulated alternative splicing. Nat Rev Genet. 2011;12(10):715–29.
Begemann G, Paricio N, Artero R, Kiss I, Perez-Alonso M, Mlodzik M. muscleblind, a gene required for photoreceptor differentiation in Drosophila, encodes novel nuclear Cys3His-type zinc-finger-containing proteins. Development. 1997;124(21):4321–31.
Ho TH, Charlet BN, Poulos MG, Singh G, Swanson MS, Cooper TA. Muscleblind proteins regulate alternative splicing. EMBO J. 2004;23(15):3103–12.
Kanadia RN, Urbinati CR, Crusselle VJ, Luo D, Lee YJ, Harrison JK, Oh SP, Swanson MS. Developmental expression of mouse muscleblind genes Mbnl1, Mbnl2 and Mbnl3. Gene Expr Patterns. 2003;3(4):459–62.
Konieczny P, Stepniak-Konieczna E, Sobczak K. MBNL proteins and their target RNAs, interaction and splicing regulation. Nucleic Acids Res. 2014;42(17):10873–87.
Kalsotra A, Xiao X, Ward AJ, Castle JC, Johnson JM, Burge CB, Cooper TA. A postnatal switch of CELF and MBNL proteins reprograms alternative splicing in the developing heart. Proc Natl Acad Sci U S A. 2008;105(51):20333–8.
Wang ET, Cody NA, Jog S, Biancolella M, Wang TT, Treacy DJ, Luo S, Schroth GP, Housman DE, Reddy S, et al. Transcriptome-wide regulation of pre-mRNA splicing and mRNA localization by muscleblind proteins. Cell. 2012;150(4):710–24.
Du H, Cline MS, Osborne RJ, Tuttle DL, Clark TA, Donohue JP, Hall MP, Shiue L, Swanson MS, Thornton CA, et al. Aberrant alternative splicing and extracellular matrix gene expression in mouse models of myotonic dystrophy. Nat Struct Mol Biol. 2010;17(2):187–93.
Adereth Y, Dammai V, Kose N, Li R, Hsu T. RNA-dependent integrin alpha3 protein localization regulated by the Muscleblind-like protein MLP1. Nat Cell Biol. 2005;7(12):1240–7.
Masuda A, Andersen HS, Doktor TK, Okamoto T, Ito M, Andresen BS, Ohno K. CUGBP1 and MBNL1 preferentially bind to 3′ UTRs and facilitate mRNA decay. Sci Rep. 2012;2:209.
Osborne RJ, Lin X, Welle S, Sobczak K, O’Rourke JR, Swanson MS, Thornton CA. Transcriptional and post-transcriptional impact of toxic RNA in myotonic dystrophy. Hum Mol Genet. 2009;18(8):1471–81.
Rau F, Freyermuth F, Fugier C, Villemin JP, Fischer MC, Jost B, Dembele D, Gourdon G, Nicole A, Duboc D, et al. Misregulation of miR-1 processing is associated with heart defects in myotonic dystrophy. Nat Struct Mol Biol. 2011;18(7):840–5.
Kanadia RN, Johnstone KA, Mankodi A, Lungu C, Thornton CA, Esson D, Timmers AM, Hauswirth WW, Swanson MS. A muscleblind knockout model for myotonic dystrophy. Science. 2003;302(5652):1978–80.
Kino Y, Mori D, Oma Y, Takeshita Y, Sasagawa N, Ishiura S. Muscleblind protein, MBNL1/EXP, binds specifically to CHHG repeats. Hum Mol Genet. 2004;13(5):495–507.
Warf MB, Berglund JA. MBNL binds similar RNA structures in the CUG repeats of myotonic dystrophy and its pre-mRNA substrate cardiac troponin T. RNA. 2007;13(12):2238–51.
Yuan Y, Compton SA, Sobczak K, Stenberg MG, Thornton CA, Griffith JD, Swanson MS. Muscleblind-like 1 interacts with RNA hairpins in splicing target and pathogenic RNAs. Nucleic Acids Res. 2007;35(16):5474–86.
Fardaei M, Rogers MT, Thorpe HM, Larkin K, Hamshere MG, Harper PS, Brook JD. Three proteins, MBNL, MBLL and MBXL, co-localize in vivo with nuclear foci of expanded-repeat transcripts in DM1 and DM2 cells. Hum Mol Genet. 2002;11(7):805–14.
Jiang H, Mankodi A, Swanson MS, Moxley RT, Thornton CA. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum Mol Genet. 2004;13(24):3079–88.
Miller JW, Urbinati CR, Teng-Umnuay P, Stenberg MG, Byrne BJ, Thornton CA, Swanson MS. Recruitment of human muscleblind proteins to (CUG)(n) expansions associated with myotonic dystrophy. EMBO J. 2000;19(17):4439–48.
Batra R, Charizanis K, Manchanda M, Mohan A, Li M, Finn DJ, Goodwin M, Zhang C, Sobczak K, Thornton CA, et al. Loss of MBNL leads to disruption of developmentally regulated alternative polyadenylation in RNA-mediated disease. Mol Cell. 2014;56(2):311–22.
Charizanis K, Lee KY, Batra R, Goodwin M, Zhang C, Yuan Y, Shiue L, Cline M, Scotti MM, Xia G, et al. Muscleblind-like 2-mediated alternative splicing in the developing brain and dysregulation in myotonic dystrophy. Neuron. 2012;75(3):437–50.
Poulos MG, Batra R, Li M, Yuan Y, Zhang C, Darnell RB, Swanson MS. Progressive impairment of muscle regeneration in muscleblind-like 3 isoform knockout mice. Hum Mol Genet. 2013;22(17):3547–58.
Lee KY, Li M, Manchanda M, Batra R, Charizanis K, Mohan A, Warren SA, Chamberlain CM, Finn D, Hong H, et al. Compound loss of muscleblind-like function in myotonic dystrophy. EMBO Mol Med. 2013;5(12):1887–900.
Suenaga K, Lee KY, Nakamori M, Tatsumi Y, Takahashi MP, Fujimura H, Jinnai K, Yoshikawa H, Du H, Ares M Jr, et al. Muscleblind-like 1 knockout mice reveal novel splicing defects in the myotonic dystrophy brain. PLoS One. 2012;7(3):e33218.
Goodwin M, Mohan A, Batra R, Lee KY, Charizanis K, Fernandez Gomez FJ, Eddarkaoui S, Sergeant N, Buee L, Kimura T, et al. MBNL sequestration by toxic RNAs and RNA misprocessing in the myotonic dystrophy brain. Cell Rep. 2015;12(7):1159–68.
Sergeant N, Sablonniere B, Schraen-Maschke S, Ghestem A, Maurage CA, Wattez A, Vermersch P, Delacourte A. Dysregulation of human brain microtubule-associated tau mRNA maturation in myotonic dystrophy type 1. Hum Mol Genet. 2001;10(19):2143–55.
Ladd AN, Charlet N, Cooper TA. The CELF family of RNA binding proteins is implicated in cell-specific and developmentally regulated alternative splicing. Mol Cell Biol. 2001;21(4):1285–96.
Fardaei M, Larkin K, Brook JD, Hamshere MG. In vivo co-localisation of MBNL protein with DMPK expanded-repeat transcripts. Nucleic Acids Res. 2001;29(13):2766–71.
Kuyumcu-Martinez NM, Wang GS, Cooper TA. Increased steady-state levels of CUGBP1 in myotonic dystrophy 1 are due to PKC-mediated hyperphosphorylation. Mol Cell. 2007;28(1):68–78.
Kalsotra A, Wang K, Li PF, Cooper TA. MicroRNAs coordinate an alternative splicing network during mouse postnatal heart development. Genes Dev. 2010;24(7):653–8.
Kalsotra A, Singh RK, Gurha P, Ward AJ, Creighton CJ, Cooper TA. The Mef2 transcription network is disrupted in myotonic dystrophy heart tissue, dramatically altering miRNA and mRNA expression. Cell Rep. 2014;6(2):336–45.
Lin X, Miller JW, Mankodi A, Kanadia RN, Yuan Y, Moxley RT, Swanson MS, Thornton CA. Failure of MBNL1-dependent post-natal splicing transitions in myotonic dystrophy. Hum Mol Genet. 2006;15(13):2087–97.
Pelletier R, Hamel F, Beaulieu D, Patry L, Haineault C, Tarnopolsky M, Schoser B, Puymirat J. Absence of a differentiation defect in muscle satellite cells from DM2 patients. Neurobiol Dis. 2009;36(1):181–90.
Salisbury E, Schoser B, Schneider-Gold C, Wang GL, Huichalaf C, Jin B, Sirito M, Sarkar P, Krahe R, Timchenko NA, et al. Expression of RNA CCUG repeats dysregulates translation and degradation of proteins in myotonic dystrophy 2 patients. Am J Pathol. 2009;175(2):748–62.
Savkur RS, Philips AV, Cooper TA. Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat Genet. 2001;29(1):40–7.
Charlet BN, Savkur RS, Singh G, Philips AV, Grice EA, Cooper TA. Loss of the muscle-specific chloride channel in type 1 myotonic dystrophy due to misregulated alternative splicing. Mol Cell. 2002;10(1):45–53.
Wang ET, Ward AJ, Cherone JM, Giudice J, Wang TT, Treacy DJ, Lambert NJ, Freese P, Saxena T, Cooper TA, et al. Antagonistic regulation of mRNA expression and splicing by CELF and MBNL proteins. Genome Res. 2015;25(6):858–71.
Kim DH, Langlois MA, Lee KB, Riggs AD, Puymirat J, Rossi JJ. HnRNP H inhibits nuclear export of mRNA containing expanded CUG repeats and a distal branch point sequence. Nucleic Acids Res. 2005;33(12):3866–74.
Paul S, Dansithong W, Kim D, Rossi J, Webster NJ, Comai L, Reddy S. Interaction of muscleblind, CUG-BP1 and hnRNP H proteins in DM1-associated aberrant IR splicing. EMBO J. 2006;25(18):4271–83.
Paul S, Dansithong W, Jog SP, Holt I, Mittal S, Brook JD, Morris GE, Comai L, Reddy S. Expanded CUG repeats dysregulate RNA splicing by altering the stoichiometry of the muscleblind 1 complex. J Biol Chem. 2011;286(44):38427–38.
Krzyzosiak WJ, Sobczak K, Wojciechowska M, Fiszer A, Mykowska A, Kozlowski P. Triplet repeat RNA structure and its role as pathogenic agent and therapeutic target. Nucleic Acids Res. 2012;40(1):11–26.
Laurent FX, Sureau A, Klein AF, Trouslard F, Gasnier E, Furling D, Marie J. New function for the RNA helicase p68/DDX5 as a modifier of MBNL1 activity on expanded CUG repeats. Nucleic Acids Res. 2012;40(7):3159–71.
Holt I, Mittal S, Furling D, Butler-Browne GS, Brook JD, Morris GE. Defective mRNA in myotonic dystrophy accumulates at the periphery of nuclear splicing speckles. Genes Cells. 2007;12(9):1035–48.
Smith KP, Byron M, Johnson C, Xing Y, Lawrence JB. Defining early steps in mRNA transport: mutant mRNA in myotonic dystrophy type I is blocked at entry into SC-35 domains. J Cell Biol. 2007;178(6):951–64.
Taliaferro JM, Vidaki M, Oliveira R, Olson S, Zhan L, Saxena T, Wang ET, Graveley BR, Gertler FB, Swanson MS, et al. Distal alternative last exons localize mRNAs to neural projections. Mol Cell. 2016;61(6):821–33.
Wang ET, Taliaferro JM, Lee JA, Sudhakaran IP, Rossoll W, Gross C, Moss KR, Bassell GJ. Dysregulation of mRNA localization and translation in genetic disease. J Neurosci. 2016;36(45):11418–26.
Vlasova IA, Tahoe NM, Fan D, Larsson O, Rattenbacher B, Sternjohn JR, Vasdewani J, Karypis G, Reilly CS, Bitterman PB, et al. Conserved GU-rich elements mediate mRNA decay by binding to CUG-binding protein 1. Mol Cell. 2008;29(2):263–70.
Lee JE, Lee JY, Wilusz J, Tian B, Wilusz CJ. Systematic analysis of cis-elements in unstable mRNAs demonstrates that CUGBP1 is a key regulator of mRNA decay in muscle cells. PLoS One. 2010;5(6):e11201.
Moraes KC, Wilusz CJ, Wilusz J. CUG-BP binds to RNA substrates and recruits PARN deadenylase. RNA. 2006;12(6):1084–91.
Timchenko L. Molecular mechanisms of muscle atrophy in myotonic dystrophies. Int J Biochem Cell Biol. 2013;45(10):2280–7.
Dasgupta T, Ladd AN. The importance of CELF control: molecular and biological roles of the CUG-BP, Elav-like family of RNA-binding proteins. Wiley Interdiscip Rev RNA. 2012;3(1):104–21.
Rattenbacher B, Beisang D, Wiesner DL, Jeschke JC, von Hohenberg M, St Louis-Vlasova IA, Bohjanen PR. Analysis of CUGBP1 targets identifies GU-repeat sequences that mediate rapid mRNA decay. Mol Cell Biol. 2010;30(16):3970–80.
Zhang L, Lee JE, Wilusz J, Wilusz CJ. The RNA-binding protein CUGBP1 regulates stability of tumor necrosis factor mRNA in muscle cells: implications for myotonic dystrophy. J Biol Chem. 2008;283(33):22457–63.
Russo J, Lee JE, Lopez CM, Anderson J, Nguyen TP, Heck AM, Wilusz J, Wilusz CJ. The CELF1 RNA-binding protein regulates decay of signal recognition particle mRNAs and limits secretion in mouse myoblasts. PLoS One. 2017;12(1):e0170680.
Vlasova-St Louis I, Dickson AM, Bohjanen PR, Wilusz CJ. CELFish ways to modulate mRNA decay. Biochim Biophys Acta. 2013;1829(6–7):695–707.
Timchenko NA, Iakova P, Cai ZJ, Smith JR, Timchenko LT. Molecular basis for impaired muscle differentiation in myotonic dystrophy. Mol Cell Biol. 2001;21(20):6927–38.
Timchenko NA, Patel R, Iakova P, Cai ZJ, Quan L, Timchenko LT. Overexpression of CUG triplet repeat-binding protein, CUGBP1, in mice inhibits myogenesis. J Biol Chem. 2004;279(13):13129–39.
Salisbury E, Sakai K, Schoser B, Huichalaf C, Schneider-Gold C, Nguyen H, Wang GL, Albrecht JH, Timchenko LT. Ectopic expression of cyclin D3 corrects differentiation of DM1 myoblasts through activation of RNA CUG-binding protein, CUGBP1. Exp Cell Res. 2008;314(11–12):2266–78.
Gambardella S, Rinaldi F, Lepore SM, Viola A, Loro E, Angelini C, Vergani L, Novelli G, Botta A. Overexpression of microRNA-206 in the skeletal muscle from myotonic dystrophy type 1 patients. J Transl Med. 2010;8:48.
Perbellini R, Greco S, Sarra-Ferraris G, Cardani R, Capogrossi MC, Meola G, Martelli F. Dysregulation and cellular mislocalization of specific miRNAs in myotonic dystrophy type 1. Neuromuscul Disord. 2011;21(2):81–8.
Fernandez-Costa JM, Garcia-Lopez A, Zuniga S, Fernandez-Pedrosa V, Felipo-Benavent A, Mata M, Jaka O, Aiastui A, Hernandez-Torres F, Aguado B, et al. Expanded CTG repeats trigger miRNA alterations in Drosophila that are conserved in myotonic dystrophy type 1 patients. Hum Mol Genet. 2013;22(4):704–16.
Greco S, Perfetti A, Fasanaro P, Cardani R, Capogrossi MC, Meola G, Martelli F. Deregulated microRNAs in myotonic dystrophy type 2. PLoS One. 2012;7(6):e39732.
Perfetti A, Greco S, Bugiardini E, Cardani R, Gaia P, Gaetano C, Meola G, Martelli F. Plasma microRNAs as biomarkers for myotonic dystrophy type 1. Neuromuscul Disord. 2014;24(6):509–15.
Fernandez-Torron R, Garcia-Puga M, Emparanza JI, Maneiro M, Cobo AM, Poza JJ, Espinal JB, Zulaica M, Ruiz I, Martorell L, et al. Cancer risk in DM1 is sex-related and linked to miRNA-200/141 downregulation. Neurology. 2016;87(12):1250–7.
Zu T, Gibbens B, Doty NS, Gomes-Pereira M, Huguet A, Stone MD, Margolis J, Peterson M, Markowski TW, Ingram MA, et al. Non-ATG-initiated translation directed by microsatellite expansions. Proc Natl Acad Sci U S A. 2011;108(1):260–5.
Cleary JD, Ranum LP. Repeat-associated non-ATG (RAN) translation in neurological disease. Hum Mol Genet. 2013;22(R1):R45–51.
Zu T, Cleary JD, Liu Y, Banez-Coronel M, Bubenik JL, Ayhan F, Ashizawa T, Xia G, Clark HB, Yachnis AT, et al. RAN translation regulated by Muscleblind proteins in myotonic dystrophy type 2. Neuron. 2017;95(6):1292–1305 e1295.
Kearse MG, Todd PK. Repeat-associated non-AUG translation and its impact in neurodegenerative disease. Neurotherapeutics. 2014;11(4):721–31.
Green KM, Linsalata AE, Todd PK. RAN translation-what makes it run? Brain Res. 2016;1647:30–42.
Todd PK, Oh SY, Krans A, He F, Sellier C, Frazer M, Renoux AJ, Chen KC, Scaglione KM, Basrur V, et al. CGG repeat-associated translation mediates neurodegeneration in fragile X tremor ataxia syndrome. Neuron. 2013;78(3):440–55.
Banez-Coronel M, Ayhan F, Tarabochia AD, Zu T, Perez BA, Tusi SK, Pletnikova O, Borchelt DR, Ross CA, Margolis RL, et al. RAN translation in Huntington disease. Neuron. 2015;88(4):667–77.
Harrison AF, Shorter J. RNA-binding proteins with prion-like domains in health and disease. Biochem J. 2017;474(8):1417–38.
Tran H, Almeida S, Moore J, Gendron TF, Chalasani U, Lu Y, Du X, Nickerson JA, Petrucelli L, Weng Z, et al. Differential toxicity of nuclear RNA foci versus dipeptide repeat proteins in a Drosophila model of C9ORF72 FTD/ALS. Neuron. 2015;87(6):1207–14.
Liu EY, Cali CP, Lee EB. RNA metabolism in neurodegenerative disease. Dis Models Mech. 2017;10(5):509–18.
Ling SC, Polymenidou M, Cleveland DW. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron. 2013;79(3):416–38.
Buratti E, Dork T, Zuccato E, Pagani F, Romano M, Baralle FE. Nuclear factor TDP-43 and SR proteins promote in vitro and in vivo CFTR exon 9 skipping. EMBO J. 2001;20(7):1774–84.
Ling JP, Pletnikova O, Troncoso JC, Wong PC. TDP-43 repression of nonconserved cryptic exons is compromised in ALS-FTD. Science. 2015;349(6248):650–5.
Shiga A, Ishihara T, Miyashita A, Kuwabara M, Kato T, Watanabe N, Yamahira A, Kondo C, Yokoseki A, Takahashi M, et al. Alteration of POLDIP3 splicing associated with loss of function of TDP-43 in tissues affected with ALS. PLoS One. 2012;7(8):e43120.
Tollervey JR, Curk T, Rogelj B, Briese M, Cereda M, Kayikci M, Konig J, Hortobagyi T, Nishimura AL, Zupunski V, et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat Neurosci. 2011;14(4):452–8.
Costessi L, Porro F, Iaconcig A, Muro AF. TDP-43 regulates beta-adducin (Add2) transcript stability. RNA Biol. 2014;11(10):1280–90.
Liu X, Li D, Zhang W, Guo M, Zhan Q. Long non-coding RNA gadd7 interacts with TDP-43 and regulates Cdk6 mRNA decay. EMBO J. 2012;31(23):4415–27.
Strong MJ, Volkening K, Hammond R, Yang W, Strong W, Leystra-Lantz C, Shoesmith C. TDP43 is a human low molecular weight neurofilament (hNFL) mRNA-binding protein. Mol Cell Neurosci. 2007;35(2):320–7.
Alami NH, Smith RB, Carrasco MA, Williams LA, Winborn CS, Han SSW, Kiskinis E, Winborn B, Freibaum BD, Kanagaraj A, et al. Axonal transport of TDP-43 mRNA granules is impaired by ALS-causing mutations. Neuron. 2014;81(3):536–43.
Yang L, Embree LJ, Tsai S, Hickstein DD. Oncoprotein TLS interacts with serine-arginine proteins involved in RNA splicing. J Biol Chem. 1998;273(43):27761–4.
Wang X, Arai S, Song X, Reichart D, Du K, Pascual G, Tempst P, Rosenfeld MG, Glass CK, Kurokawa R. Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature. 2008;454(7200):126–30.
Couthouis J, Hart MP, Shorter J, DeJesus-Hernandez M, Erion R, Oristano R, Liu AX, Ramos D, Jethava N, Hosangadi D, et al. A yeast functional screen predicts new candidate ALS disease genes. Proc Natl Acad Sci U S A. 2011;108(52):20881–90.
Ticozzi N, Vance C, Leclerc AL, Keagle P, Glass JD, McKenna-Yasek D, Sapp PC, Silani V, Bosco DA, Shaw CE, et al. Mutational analysis reveals the FUS homolog TAF15 as a candidate gene for familial amyotrophic lateral sclerosis. Am J Med Genet B Neuropsychiatric Genet. 2011;156B(3):285–90.
Neumann M, Bentmann E, Dormann D, Jawaid A, DeJesus-Hernandez M, Ansorge O, Roeber S, Kretzschmar HA, Munoz DG, Kusaka H, et al. FET proteins TAF15 and EWS are selective markers that distinguish FTLD with FUS pathology from amyotrophic lateral sclerosis with FUS mutations. Brain. 2011;134(Pt 9):2595–609.
Couthouis J, Hart MP, Erion R, King OD, Diaz Z, Nakaya T, Ibrahim F, Kim HJ, Mojsilovic-Petrovic J, Panossian S, et al. Evaluating the role of the FUS/TLS-related gene EWSR1 in amyotrophic lateral sclerosis. Hum Mol Genet. 2012;21(13):2899–911.
Elden AC, Kim HJ, Hart MP, Chen-Plotkin AS, Johnson BS, Fang X, Armakola M, Geser F, Greene R, Lu MM, et al. Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature. 2010;466(7310):1069–75.
Kim HJ, Kim NC, Wang YD, Scarborough EA, Moore J, Diaz Z, MacLea KS, Freibaum B, Li S, Molliex A, et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature. 2013;495(7442):467–73.
Johnson JO, Pioro EP, Boehringer A, Chia R, Feit H, Renton AE, Pliner HA, Abramzon Y, Marangi G, Winborn BJ, et al. Mutations in the Matrin 3 gene cause familial amyotrophic lateral sclerosis. Nat Neurosci. 2014;17(5):664–6.
Murray DT, Kato M, Lin Y, Thurber KR, Hung I, McKnight SL, Tycko R. Structure of FUS protein fibrils and its relevance to self-assembly and phase separation of low-complexity domains. Cell. 2017;171(3):615–627 e616.
Murakami T, Qamar S, Lin JQ, Schierle GS, Rees E, Miyashita A, Costa AR, Dodd RB, Chan FT, Michel CH, et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron. 2015;88(4):678–90.
Taylor JP, Brown RH Jr, Cleveland DW. Decoding ALS: from genes to mechanism. Nature. 2016;539(7628):197–206.
Freibaum BD, Lu Y, Lopez-Gonzalez R, Kim NC, Almeida S, Lee KH, Badders N, Valentine M, Miller BL, Wong PC, et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature. 2015;525(7567):129–33.
Jovicic A, Mertens J, Boeynaems S, Bogaert E, Chai N, Yamada SB, Paul JW III, Sun S, Herdy JR, Bieri G, et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat Neurosci. 2015;18(9):1226–9.
Zhang K, Donnelly CJ, Haeusler AR, Grima JC, Machamer JB, Steinwald P, Daley EL, Miller SJ, Cunningham KM, Vidensky S, et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature. 2015;525(7567):56–61.
Woerner AC, Frottin F, Hornburg D, Feng LR, Meissner F, Patra M, Tatzelt J, Mann M, Winklhofer KF, Hartl FU, et al. Cytoplasmic protein aggregates interfere with nucleocytoplasmic transport of protein and RNA. Science. 2016;351(6269):173–6.
Lee KH, Zhang P, Kim HJ, Mitrea DM, Sarkar M, Freibaum BD, Cika J, Coughlin M, Messing J, Molliex A, et al. C9orf72 dipeptide repeats impair the assembly, dynamics, and function of membrane-less organelles. Cell. 2016;167(3):774–788 e717.
Lin Y, Mori E, Kato M, Xiang S, Wu L, Kwon I, McKnight SL. Toxic PR poly-dipeptides encoded by the C9orf72 repeat expansion target LC domain polymers. Cell. 2016;167(3):789–802 e712.
Shi KY, Mori E, Nizami ZF, Lin Y, Kato M, Xiang S, Wu LC, Ding M, Yu Y, Gall JG, et al. Toxic PRn poly-dipeptides encoded by the C9orf72 repeat expansion block nuclear import and export. Proc Natl Acad Sci U S A. 2017;114(7):E1111–7.
Hagerman RJ, Hull CE, Safanda JF, Carpenter I, Staley LW, O’Connor RA, Seydel C, Mazzocco MM, Snow K, Thibodeau SN, et al. High functioning fragile X males: demonstration of an unmethylated fully expanded FMR-1 mutation associated with protein expression. Am J Med Genet. 1994;51(4):298–308.
Coffey SM, Cook K, Tartaglia N, Tassone F, Nguyen DV, Pan R, Bronsky HE, Yuhas J, Borodyanskaya M, Grigsby J, et al. Expanded clinical phenotype of women with the FMR1 premutation. Am J Med Genet A. 2008;146A(8):1009–16.
Jacquemont S, Hagerman RJ, Leehey M, Grigsby J, Zhang L, Brunberg JA, Greco C, Des Portes V, Jardini T, Levine R, et al. Fragile X premutation tremor/ataxia syndrome: molecular, clinical, and neuroimaging correlates. Am J Hum Genet. 2003;72(4):869–78.
Greco CM, Hagerman RJ, Tassone F, Chudley AE, Del Bigio MR, Jacquemont S, Leehey M, Hagerman PJ. Neuronal intranuclear inclusions in a new cerebellar tremor/ataxia syndrome among fragile X carriers. Brain. 2002;125(Pt 8):1760–71.
Leehey MA. Fragile X-associated tremor/ataxia syndrome: clinical phenotype, diagnosis, and treatment. J Investig Med. 2009;57(8):830–6.
Penagarikano O, Mulle JG, Warren ST. The pathophysiology of fragile x syndrome. Annu Rev Genomics Hum Genet. 2007;8:109–29.
Sreeram N, Wren C, Bhate M, Robertson P, Hunter S. Cardiac abnormalities in the fragile X syndrome. Br Heart J. 1989;61(3):289–91.
Lozano R, Azarang A, Wilaisakditipakorn T, Hagerman RJ. Fragile X syndrome: a review of clinical management. Intractable Rare Dis Res. 2016;5(3):145–57.
Holmes SE, O’Hearn EE, McInnis MG, Gorelick-Feldman DA, Kleiderlein JJ, Callahan C, Kwak NG, Ingersoll-Ashworth RG, Sherr M, Sumner AJ, et al. Expansion of a novel CAG trinucleotide repeat in the 5′ region of PPP2R2B is associated with SCA12. Nat Genet. 1999;23(4):391–2.
O’Hearn E, Holmes SE, Calvert PC, Ross CA, Margolis RL. SCA-12: tremor with cerebellar and cortical atrophy is associated with a CAG repeat expansion. Neurology. 2001;56(3):299–303.
Ranum LP, Rasmussen PF, Benzow KA, Koob MD, Day JW. Genetic mapping of a second myotonic dystrophy locus. Nat Genet. 1998;19(2):196–8.
Peric S, Rakocevic Stojanovic V, Mandic Stojmenovic G, Ilic V, Kovacevic M, Parojcic A, Pesovic J, Mijajlovic M, Savic-Pavicevic D, Meola G. Clusters of cognitive impairment among different phenotypes of myotonic dystrophy type 1 and type 2. Neurol Sci. 2017;38(3):415–23.
Tieleman AA, Knoop H, van de Logt AE, Bleijenberg G, van Engelen BG, Overeem S. Poor sleep quality and fatigue but no excessive daytime sleepiness in myotonic dystrophy type 2. J Neurol Neurosurg Psychiatry. 2010;81(9):963–7.
Hund E, Jansen O, Koch MC, Ricker K, Fogel W, Niedermaier N, Otto M, Kuhn E, Meinck HM. Proximal myotonic myopathy with MRI white matter abnormalities of the brain. Neurology. 1997;48(1):33–7.
Schneider-Gold C, Bellenberg B, Prehn C, Krogias C, Schneider R, Klein J, Gold R, Lukas C. Cortical and subcortical grey and white matter atrophy in myotonic dystrophies Type 1 and 2 is associated with cognitive impairment, depression and daytime sleepiness. PLoS One. 2015;10(6):e0130352.
Meola G, Sansone V, Perani D, Scarone S, Cappa S, Dragoni C, Cattaneo E, Cotelli M, Gobbo C, Fazio F, et al. Executive dysfunction and avoidant personality trait in myotonic dystrophy type 1 (DM-1) and in proximal myotonic myopathy (PROMM/DM-2). Neuromuscul Disord. 2003;13(10):813–21.
Meola G, Cardani R. Myotonic dystrophy type 2: an update on clinical aspects, genetic and pathomolecular mechanism. J Neuromuscul Dis. 2015;2(s2):S59–71.
Giordana MT, Ferrero P, Grifoni S, Pellerino A, Naldi A, Montuschi A. Dementia and cognitive impairment in amyotrophic lateral sclerosis: a review. Neurol Sci. 2011;32(1):9–16.
Graff-Radford NR, Woodruff BK. Frontotemporal dementia. Semin Neurol. 2007;27(1):48–57.
Kirshner HS. Frontotemporal dementia and primary progressive aphasia, a review. Neuropsychiatr Dis Treat. 2014;10:1045–55.
Thomas PS Jr, Fraley GS, Damian V, Woodke LB, Zapata F, Sopher BL, Plymate SR, La Spada AR. Loss of endogenous androgen receptor protein accelerates motor neuron degeneration and accentuates androgen insensitivity in a mouse model of X-linked spinal and bulbar muscular atrophy. Hum Mol Genet. 2006;15(14):2225–38.
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell. 1993;72(6):971–83.
Louis ED, Lee P, Quinn L, Marder K. Dystonia in Huntington’s disease: prevalence and clinical characteristics. Mov Disord. 1999;14(1):95–101.
Kim SD, Fung VS. An update on Huntington’s disease: from the gene to the clinic. Curr Opin Neurol. 2014;27(4):477–83.
Koide R, Ikeuchi T, Onodera O, Tanaka H, Igarashi S, Endo K, Takahashi H, Kondo R, Ishikawa A, Hayashi T, et al. Unstable expansion of CAG repeat in hereditary dentatorubral-pallidoluysian atrophy (DRPLA). Nat Genet. 1994;6(1):9–13.
Nagafuchi S, Yanagisawa H, Sato K, Shirayama T, Ohsaki E, Bundo M, Takeda T, Tadokoro K, Kondo I, Murayama N, et al. Dentatorubral and pallidoluysian atrophy expansion of an unstable CAG trinucleotide on chromosome 12p. Nat Genet. 1994;6(1):14–8.
Ikeuchi T, Koide R, Tanaka H, Onodera O, Igarashi S, Takahashi H, Kondo R, Ishikawa A, Tomoda A, Miike T, et al. Dentatorubral-pallidoluysian atrophy: clinical features are closely related to unstable expansions of trinucleotide (CAG) repeat. Ann Neurol. 1995;37(6):769–75.
Nucifora FC Jr, Ellerby LM, Wellington CL, Wood JD, Herring WJ, Sawa A, Hayden MR, Dawson VL, Dawson TM, Ross CA. Nuclear localization of a non-caspase truncation product of atrophin-1, with an expanded polyglutamine repeat, increases cellular toxicity. J Biol Chem. 2003;278(15):13047–55.
Klement IA, Skinner PJ, Kaytor MD, Yi H, Hersch SM, Clark HB, Zoghbi HY, Orr HT. Ataxin-1 nuclear localization and aggregation: role in polyglutamine-induced disease in SCA1 transgenic mice. Cell. 1998;95(1):41–53.
Skinner PJ, Vierra-Green CA, Emamian E, Zoghbi HY, Orr HT. Amino acids in a region of ataxin-1 outside of the polyglutamine tract influence the course of disease in SCA1 transgenic mice. NeuroMolecular Med. 2002;1(1):33–42.
Sanpei K, Takano H, Igarashi S, Sato T, Oyake M, Sasaki H, Wakisaka A, Tashiro K, Ishida Y, Ikeuchi T, et al. Identification of the spinocerebellar ataxia type 2 gene using a direct identification of repeat expansion and cloning technique, DIRECT. Nat Genet. 1996;14(3):277–84.
Pulst SM, Nechiporuk A, Nechiporuk T, Gispert S, Chen XN, Lopes-Cendes I, Pearlman S, Starkman S, Orozco-Diaz G, Lunkes A, et al. Moderate expansion of a normally biallelic trinucleotide repeat in spinocerebellar ataxia type 2. Nat Genet. 1996;14(3):269–76.
Imbert G, Saudou F, Yvert G, Devys D, Trottier Y, Garnier JM, Weber C, Mandel JL, Cancel G, Abbas N, et al. Cloning of the gene for spinocerebellar ataxia 2 reveals a locus with high sensitivity to expanded CAG/glutamine repeats. Nat Genet. 1996;14(3):285–91.
Kawaguchi Y, Okamoto T, Taniwaki M, Aizawa M, Inoue M, Katayama S, Kawakami H, Nakamura S, Nishimura M, Akiguchi I, et al. CAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nat Genet. 1994;8(3):221–8.
Zhuchenko O, Bailey J, Bonnen P, Ashizawa T, Stockton DW, Amos C, Dobyns WB, Subramony SH, Zoghbi HY, Lee CC. Autosomal dominant cerebellar ataxia (SCA6) associated with small polyglutamine expansions in the alpha 1A-voltage-dependent calcium channel. Nat Genet. 1997;15(1):62–9.
David G, Abbas N, Stevanin G, Durr A, Yvert G, Cancel G, Weber C, Imbert G, Saudou F, Antoniou E, et al. Cloning of the SCA7 gene reveals a highly unstable CAG repeat expansion. Nat Genet. 1997;17(1):65–70.
Koide R, Kobayashi S, Shimohata T, Ikeuchi T, Maruyama M, Saito M, Yamada M, Takahashi H, Tsuji S. A neurological disease caused by an expanded CAG trinucleotide repeat in the TATA-binding protein gene: a new polyglutamine disease? Hum Mol Genet. 1999;8(11):2047–53.
Brais B, Bouchard JP, Xie YG, Rochefort DL, Chretien N, Tome FM, Lafreniere RG, Rommens JM, Uyama E, Nohira O, et al. Short GCG expansions in the PABP2 gene cause oculopharyngeal muscular dystrophy. Nat Genet. 1998;18(2):164–7.
Anvar SY, Raz Y, Verway N, van der Sluijs B, Venema A, Goeman JJ, Vissing J, van der Maarel SM, t Hoen PA, van Engelen BG, et al. A decline in PABPN1 induces progressive muscle weakness in oculopharyngeal muscle dystrophy and in muscle aging. Aging. 2013;5(6):412–26.
Stromme P, Mangelsdorf ME, Shaw MA, Lower KM, Lewis SM, Bruyere H, Lutcherath V, Gedeon AK, Wallace RH, Scheffer IE, et al. Mutations in the human ortholog of Aristaless cause X-linked mental retardation and epilepsy. Nat Genet. 2002;30(4):441–5.
Ropers HH, Hamel BC. X-linked mental retardation. Nat Rev Genet. 2005;6(1):46–57.
Nawara M, Szczaluba K, Poirier K, Chrzanowska K, Pilch J, Bal J, Chelly J, Mazurczak T. The ARX mutations: a frequent cause of X-linked mental retardation. Am J Med Genet A. 2006;140(7):727–32.
Messaed C, Rouleau GA. Molecular mechanisms underlying polyalanine diseases. Neurobiol Dis. 2009;34(3):397–405.
Winblad S, Lindberg C, Hansen S. Cognitive deficits and CTG repeat expansion size in classical myotonic dystrophy type 1 (DM1). Behav Brain Funct. 2006;2:16.
Koob MD, Moseley ML, Schut LJ, Benzow KA, Bird TD, Day JW, Ranum LP. An untranslated CTG expansion causes a novel form of spinocerebellar ataxia (SCA8). Nat Genet. 1999;21(4):379–84.
Ikeda Y, Dalton JC, Moseley ML, Gardner KL, Bird TD, Ashizawa T, Seltzer WK, Pandolfo M, Milunsky A, Potter NT, et al. Spinocerebellar ataxia type 8: molecular genetic comparisons and haplotype analysis of 37 families with ataxia. Am J Hum Genet. 2004;75(1):3–16.
Paulson HL. The spinocerebellar ataxias. J Neuroophthalmol. 2009;29(3):227–37.
Jin P, Duan R, Qurashi A, Qin Y, Tian D, Rosser TC, Liu H, Feng Y, Warren ST. Pur alpha binds to rCGG repeats and modulates repeat-mediated neurodegeneration in a Drosophila model of fragile X tremor/ataxia syndrome. Neuron. 2007;55(4):556–64.
Sofola OA, Jin P, Qin Y, Duan R, Liu H, de Haro M, Nelson DL, Botas J. RNA-binding proteins hnRNP A2/B1 and CUGBP1 suppress fragile X CGG premutation repeat-induced neurodegeneration in a Drosophila model of FXTAS. Neuron. 2007;55(4):565–71.
Timchenko LT, Miller JW, Timchenko NA, DeVore DR, Datar KV, Lin L, Roberts R, Caskey CT, Swanson MS. Identification of a (CUG)n triplet repeat RNA-binding protein and its expression in myotonic dystrophy. Nucleic Acids Res. 1996;24(22):4407–14.
Sellier C, Rau F, Liu Y, Tassone F, Hukema RK, Gattoni R, Schneider A, Richard S, Willemsen R, Elliott DJ, et al. Sam68 sequestration and partial loss of function are associated with splicing alterations in FXTAS patients. EMBO J. 2010;29(7):1248–61.
Pieretti M, Zhang FP, Fu YH, Warren ST, Oostra BA, Caskey CT, Nelson DL. Absence of expression of the FMR-1 gene in fragile X syndrome. Cell. 1991;66(4):817–22.
Feng Y, Gutekunst CA, Eberhart DE, Yi H, Warren ST, Hersch SM, Fragile X. mental retardation protein: nucleocytoplasmic shuttling and association with somatodendritic ribosomes. J Neurosci. 1997;17(5):1539–47.
Khandjian EW, Huot ME, Tremblay S, Davidovic L, Mazroui R, Bardoni B. Biochemical evidence for the association of fragile X mental retardation protein with brain polyribosomal ribonucleoparticles. Proc Natl Acad Sci U S A. 2004;101(36):13357–62.
Stefani G, Fraser CE, Darnell JC, Darnell RB. Fragile X mental retardation protein is associated with translating polyribosomes in neuronal cells. J Neurosci. 2004;24(33):7272–6.
Greenough WT, Klintsova AY, Irwin SA, Galvez R, Bates KE, Weiler IJ. Synaptic regulation of protein synthesis and the fragile X protein. Proc Natl Acad Sci U S A. 2001;98(13):7101–6.
Hokkanen S, Feldmann HM, Ding H, Jung CK, Bojarski L, Renner-Muller I, Schuller U, Kretzschmar H, Wolf E, Herms J. Lack of Pur-alpha alters postnatal brain development and causes megalencephaly. Hum Mol Genet. 2012;21(3):473–84.
Qurashi A, Li W, Zhou JY, Peng J, Jin P. Nuclear accumulation of stress response mRNAs contributes to the neurodegeneration caused by Fragile X premutation rCGG repeats. PLoS Genet. 2011;7(6):e1002102.
Li Y, Jin P. RNA-mediated neurodegeneration in fragile X-associated tremor/ataxia syndrome. Brain Res. 2012;1462:112–7.
Wang GS, Kearney DL, De Biasi M, Taffet G, Cooper TA. Elevation of RNA-binding protein CUGBP1 is an early event in an inducible heart-specific mouse model of myotonic dystrophy. J Clin Invest. 2007;117(10):2802–11.
Ho TH, Savkur RS, Poulos MG, Mancini MA, Swanson MS, Cooper TA. Colocalization of muscleblind with RNA foci is separable from mis-regulation of alternative splicing in myotonic dystrophy. J Cell Sci. 2005;118(Pt 13):2923–33.
Ladd AN, Stenberg MG, Swanson MS, Cooper TA. Dynamic balance between activation and repression regulates pre-mRNA alternative splicing during heart development. Dev Dyn. 2005;233(3):783–93.
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, Bruce J, Schuck T, Grossman M, Clark CM, et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science. 2006;314(5796):130–3.
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, Mann D, Tsuchiya K, Yoshida M, Hashizume Y, et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun. 2006;351(3):602–11.
Kwiatkowski TJ Jr, Bosco DA, Leclerc AL, Tamrazian E, Vanderburg CR, Russ C, Davis A, Gilchrist J, Kasarskis EJ, Munsat T, et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science. 2009;323(5918):1205–8.
Vance C, Rogelj B, Hortobagyi T, De Vos KJ, Nishimura AL, Sreedharan J, Hu X, Smith B, Ruddy D, Wright P, et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science. 2009;323(5918):1208–11.
Mackenzie IR, Nicholson AM, Sarkar M, Messing J, Purice MD, Pottier C, Annu K, Baker M, Perkerson RB, Kurti A, et al. TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron. 2017;95(4):808–16. e809
Todd PK. Making sense of the antisense transcripts in C9FTD/ALS. Acta Neuropathol. 2013;126(6):785–7.
Polymenidou M, Lagier-Tourenne C, Hutt KR, Huelga SC, Moran J, Liang TY, Ling SC, Sun E, Wancewicz E, Mazur C, et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci. 2011;14(4):459–68.
Ito D, Suzuki N. Conjoint pathologic cascades mediated by ALS/FTLD-U linked RNA-binding proteins TDP-43 and FUS. Neurology. 2011;77(17):1636–43.
Ito D, Seki M, Tsunoda Y, Uchiyama H, Suzuki N. Nuclear transport impairment of amyotrophic lateral sclerosis-linked mutations in FUS/TLS. Ann Neurol. 2011;69(1):152–62.
Dormann D, Rodde R, Edbauer D, Bentmann E, Fischer I, Hruscha A, Than ME, Mackenzie IR, Capell A, Schmid B, et al. ALS-associated fused in sarcoma (FUS) mutations disrupt Transportin-mediated nuclear import. EMBO J. 2010;29(16):2841–57.
King OD, Gitler AD, Shorter J. The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease. Brain Res. 2012;1462:61–80.
Kato M, Han TW, Xie S, Shi K, Du X, Wu LC, Mirzaei H, Goldsmith EJ, Longgood J, Pei J, et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell. 2012;149(4):753–67.
Lim L, Wei Y, Lu Y, Song J. ALS-causing mutations significantly perturb the self-assembly and interaction with nucleic acid of the intrinsically disordered prion-like domain of TDP-43. PLoS Biol. 2016;14(1):e1002338.
Molliex A, Temirov J, Lee J, Coughlin M, Kanagaraj AP, Kim HJ, Mittag T, Taylor JP. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell. 2015;163(1):123–33.
Daughters RS, Tuttle DL, Gao W, Ikeda Y, Moseley ML, Ebner TJ, Swanson MS, Ranum LP. RNA gain-of-function in spinocerebellar ataxia type 8. PLoS Genet. 2009;5(8):e1000600.
Mutsuddi M, Marshall CM, Benzow KA, Koob MD, Rebay I. The spinocerebellar ataxia 8 noncoding RNA causes neurodegeneration and associates with staufen in Drosophila. Curr Biol. 2004;14(4):302–8.
Acknowledgments
A.K. is supported by grants from the US National Institute of Health (R01HL126845), Muscular Dystrophy Association (MDA514335), and the Center for Advanced Study at the University of Illinois. C.M. is supported by the American Heart Association post-doctoral fellowship (16POST29950018).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Misra, C., Lin, F., Kalsotra, A. (2018). Deregulation of RNA Metabolism in Microsatellite Expansion Diseases. In: Sattler, R., Donnelly, C. (eds) RNA Metabolism in Neurodegenerative Diseases. Advances in Neurobiology, vol 20. Springer, Cham. https://doi.org/10.1007/978-3-319-89689-2_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-89689-2_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-89688-5
Online ISBN: 978-3-319-89689-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)