Abstract
The molecular process of RNA editing allows changes in RNA transcripts that increase genomic diversity. These highly conserved RNA editing events are catalyzed by a group of enzymes known as adenosine deaminases acting on double-stranded RNA (ADARs). ADARs are necessary for normal development, they bind to over thousands of genes, impact millions of editing sites, and target critical components of the central nervous system (CNS) such as glutamate receptors, serotonin receptors, and potassium channels. Dysfunctional ADARs are known to cause alterations in CNS protein products and therefore play a role in chronic or acute neurodegenerative and psychiatric diseases as well as CNS cancer. Here, we review how RNA editing deficiency impacts CNS function and summarize its role during disease pathogenesis.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- RNA editing
- MARCH
- AMPA
- GluA2
- 5HT receptors
- K channels
- Excitotoxicity
- Neurodegeneration
- Psychiatric diseases
- Cancer
3.1 Introduction: RNA Editing Overview
RNA editing is a molecular process that allows changes in the sequence of specific RNA transcripts to increase the diversity of different RNAs that can be generated from the genome. This can result in translation of different protein variants, but can also alter alternative splicing events and micro RNA-binding efficiencies [1]. RNA editing occurs during or after transcription through two distinct mechanisms: (1) chemically modifying a nucleotide, and therefore, altering the nucleotide sequence; (2) inserting or deleting nucleotides and changing the length of the mRNA. This chapter will focus on the most common form of RNA editing, the Adenosine to Inosine (A/I) nucleotide modification of RNA catalyzed by a family of enzymes known as adenosine deaminase acting on double-stranded RNA (ADARs). We will summarize how the dysfunction of these RNA editing enzymes and the subsequent substrate alterations contributes to central nervous system (CNS) diseases [2,3,4,5].
While other RNA editing events such as C-to-U or G-to-A exist, the catalytic deamination of Adenosine into Inosine is the most prevalent [6, 7]. There are three ADAR gene family members in mammals : ADAR1, ADAR2, and ADAR3 (These enzymes have also been referred to as ADAR, ADARB1, and ADARB2 respectively). All ADAR proteins contain double-stranded RNA-binding domains (dsRBDs ), a nuclear localization sequence (NLS), and a C terminal deaminase domain [8] (see Fig. 3.1). There are two ADAR1 isoforms, ADAR1 p150 containing two additional Z DNA-binding domains and an NES and ADAR1 p110 a truncation isoform maintaining one Z DNA-binding domain and no NES [9]. ADAR1 is widely expressed throughout the body and to a lesser extent in the CNS. It has been shown to bind to over 10,000 genes and is necessary for normal development [10, 11]. ADAR1−/− mice are embryonic lethal and die around day E11.5 [10]. The mouse embryos undergo widespread apoptosis and show severe liver disintegration due to the loss of ADAR1 [10, 11]. ADAR2 is highly expressed in the CNS, and to a lesser extent in peripheral tissues [12]. ADAR2 has been shown to be responsible for the A/I editing of transcripts that are most actively edited. Knockout mouse models have shown that ADAR2 is required for normal development and ADAR2−/− mice die by P20 and become progressively seizure prone [13]. The third and final member of the family, ADAR3, is thought to have no RNA editing activity [14]. ADAR3 contains an additional arginine-rich domain [14]. Unlike its family members, ADAR3 is expressed exclusively in the brain [15]. Because no ADAR3 editing activity has been reported, the function of the enzyme is still an area of debate. There is a growing amount of evidence to suggest that ADAR3 acts as a negative regulator of overall RNA editing by binding and sequestering editing substrates of ADAR1 and ADAR2 [15, 16].
ADAR domain structures . ADAR family members do share certain domain structures, including a C-terminal Deaminase Domain, dsRNA-binding domains (dsRBD), and a nuclear localization domain (NLS). ADAR1 comes in two isoforms, ADAR1 p150 and p110. ADAR1 p150 has two Z-DNA-binding domains, Zα and Zβ, in addition to a nuclear export sequence, which explains why ADAR1 p150 can be found both in the nucleus and in the cytoplasm. ADAR1 p110 only has a Zβ domain, and is only expressed in the nucleus. ADAR3 differs from ADAR1 and ADAR2 by the existence of a N-terminal RG-rich region. ADAR1 and ADAR2 are ubiquitously expressed throughout the body, while ADAR3 is CNS specific. The chromosomal locations of the ADAR1–3 genes are 1q21.3, 21q22.3, and 10p15.3, respectively
How does RNA editing work? The hydrolytic deamination of adenosine by the catalytic activity of ADAR1 and 2 disrupts the canonical Watson and crick base pairing of adenosine and as a result the edited inosine will be interpreted by the translational machinery as a guanosine (see Fig. 3.2). Therefore, RNA A/I editing events that fall within protein-coding regions can potentially alter the codon and allow the translational machinery to introduce amino acid changes into the protein that were not encoded by the genome. This can allow for important variation of protein products produced by a single strand of RNA (e.g. serotonin receptor [17]). Editing also occurs in noncoding regions of the transcriptome where the location of the edited nucleotide can regulate splicing, retain edited mRNA in the nucleus, or prevent micro-RNA processing [7, 18,19,20,21,22,23,24,25]. Historically, the estimation of total RNA editing sites was difficult and RNA editing was studied utilizing the serendipitous discovery of A/I sites [26]. With the ever-increasing capabilities of sequencing technologies, it is now possible to analyze RNA editing sites with far greater detail [19, 26, 27]. There are conflicting reports on the total number of RNA editing events in the human genome with reports claiming over one hundred million editing sites spanning the majority of the transcriptome [7, 18, 19, 26, 27]. The majority of these RNA editing events are found within Alu repetitive elements. These genomic elements are approximately 300 bp in lengths and are primate-specific transposable elements that comprise approximately 10% of the human genome [28]. These repetitive elements form long dsRNA secondary structures that make them ideal targets for ADARs. ADARs edited sites and levels of RNA editing, as well as ADAR proteins themselves are thought to be evolutionary conserved and play a role in environmental adaptation [29].
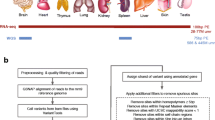
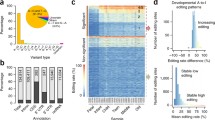
A/I RNA editing has been recognized as a significant event during CNS cortical development [30]. An increasing RNA editing pattern is observed during deep cortical layer formation suggesting these events occur at a critical period in neuronal maturation. As previously stated knockout mouse models of both ADAR1 and ADAR2 have shown that mice deficient in these deaminases form severe developmental phenotypes, emphasizing the importance of A/I RNA editing during CNS development [13]. At mature states, neurons show higher ADAR expression and editing activity than non-neuronal cells suggesting a limited involvement of other brain cells in RNA editing [4, 30]. Editing events may occur in response to environmental factors or to maintain normal CNS physiology. It can alter the function of target genes such as α-Amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptors for fast excitatory neurotransmission, serotonin-5HT2C receptors, or potassium channels Kv1.1 for modulation of neuronal excitability [31, 32]. Due to the regulation of these ion channels by ADARs, RNA A/I editing is considered crucial for proper neuronal function.
3.1.1 Major CNS RNA Editing Targets
To illustrate the importance of RNA editing in the CNS, we decided to introduce briefly three major RNA editing targets, which have shown to play a role in disease pathogenesis of several of the CNS disorders discussed below.
3.1.1.1 AMPA Receptors
AMPA receptors are ionotropic glutamate receptors responsible for fast synaptic transmission in the CNS [33]. The functional properties of AMPA receptors are greatly dependent on its subunit composition, GluA1–4, determining its role in synapse formation, stabilization, and synaptic plasticity [34]. The GluA2 subunit has the ability to regulate the calcium (Ca2+)-permeability of AMPA receptors [35,36,37]. Most AMPA receptors become permeable to Ca2+ by lacking the GluA2 subunit, and these GluA2-lacking receptors are thought to contribute to normal brain function, especially synaptic plasticity [33, 37,38,39,40,41]. However, there are numerous reports suggesting that GluA2-containing AMPA receptors become Ca2+-permeable due to a lack of editing of the GluA2 Q/R site, although in the brain, almost 100% of GluA2 mRNA is present in its edited form [42,43,44,45,46] (see Fig. 3.3). This unique element of GluA2 is regulated by ADAR2-mediated A/I RNA editing [31]. Mice lacking ADAR2 can be rescued by expression of a forced edited GluA2 subunit [13]. This provides evidence that this single editing event is essential for normal development and survival. It further supports the idea that unedited Ca2+-permeable GluA2-containing AMPA receptors do not have a physiological role similar to GluA2-lacking AMPA receptors. In this chapter, we will discuss the role of AMPA receptor GluA2 Q/R editing in the context of the role of glutamate excitotoxicity in neurodegenerative diseases, especially Amyotrophic lateral sclerosis (ALS; see below).
ADAR A/I RNA editing. (a) ADAR enzyme (light green structure) acting on double-stranded RNA. (b) ADAR dsRNA-binding domains act on dsRNA editing sites and its catalytic domain converts adenine to inosine. Within the catalytic domain an amino group on the adenine base is replaced by an oxygen and converted to inosine
3.1.1.2 Serotonin Receptors
Serotonin 5-hydroxytryptamine (5-HT) receptors are a family of chemical messengers that produce a wide variety of physiological responses including circadian rhythms, mood, memory, cognition, and possibly peristalsis in the gastrointestinal tract [47,48,49]. There are 15 unique receptors divided into seven subgroups (5-HT1–7), all subgroups are classified as G-protein coupled receptors with the exception being the 5-HT3 receptors that are ionotropic [50,51,52]. The 5-HT2C receptor subtype is expressed throughout the CNS [53, 54]. There are five ADAR-meditated RNA editing sites on the 5-HT2C mRNA, designated sites A through E [17]. These five editing sites are located within 13 base pairs and are responsible for three codons allowing for significant variation in the protein isoforms [17, 55]. With only 7% of 5-HT2C mRNA lack editing at any of the five sites, the majority of transcripts are exposed to ADAR-mediated A/I editing, the most prevalent showing editing at the ABC and D sites [17]. Editing of this receptor alters binding affinity and functional potency of receptor agonists, and thereby affection receptor function during synaptic signaling. The fully edited 5-HT2C receptor isoforms have been shown to have a 40-fold decrease in serotonergic potency, decreasing inositol phosphate accumulation and calcium release [56,57,58]. The role of serotonin receptor editing is mostly relevant for neuropsychiatric disorders, such as schizophrenia and depression (see below).
Role of GluA2 in AMPA receiver Ca2+ permeability . (a) AMPA receptors containing fully edited GluA2 (R) are impermeable to calcium due to the positively charged arginine in the channel pore. (b) When GluA2 (Q) is unedited, this positive charge is removed with the presence of the glutamine, and AMPA receptors become permeable to calcium. (c) Calcium permeability is also present when AMPA receptors lack GluA2 (Q) altogether and are composed of other AMPA receptor subunits instead
3.1.1.3 Voltage Gated Potassium Channels
Voltage gated potassium channels (Kv channels) are the largest subgroup of potassium channels [59, 60]. Comprised of 12 subgroups (Kv1–12) these six transmembrane domain subunits form tetrameric Kv channels containing an inner pore and external voltage sensor domains allowing for the conversion of voltage across the membrane to be transferred into mechanical work [60]. The Kv1 family (Kv 1.1,2, and 4) has been shown to localize to soma, axons, synaptic terminals, and proximal dendrites [59, 61]. The Kv1.1 channel plays an important role in the regulation of neuronal excitability [62]. An ADAR2-mediated A/I editing site lies within the ion pore of the Kv1.1 subunit, mediating an isoleucine to valine substitution [63]. No differences were observed between the voltage-dependent activation of edited and unedited Kv1.1 channels [63]. In contrast A/I editing at this site has been proposed to target the process of fast inactivation [63]. Fast inactivation of Kv1 channels is mediated by the inactivating ball domain on the Kvβ1 subunit [64]. Regulation of this mechanism by RNA editing will have profound effects on regulation of neuronal excitability.
3.2 RNA Editing Deficits in Neurodegeneration
As summarized above, the post-transcriptional modification of RNA transcripts by ADARs through RNA editing generates protein diversity regulating many critical aspects of CNS function. Therefore, if the RNA editing process fails it could lead to CNS diseases, or exacerbate acute injury and chronic disorders. In the following sections, we will discuss RNA editing deficits for chronic and acute neurodegenerative disorders, neuropsychiatric diseases, and brain cancers.
3.2.1 RNA Editing in Chronic Neurodegenerative Diseases
Alzheimer’s disease (AD) accounts for 60–80% of dementia cases [65]. It is the most prevalent form of dementia characterized by a progressive loss of memory and cognitive dysfunction. The neuropathological hallmarks comprise of plaques and tangles known to play a critical role in neurodegeneration [65, 66]. Areas of the hippocampus, pre-frontal, and temporal cortex play a significant role in AD pathophysiology [67]. Studies done by Akbarian et al. associated deficits in RNA editing of the α-Amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptor subunit GluA2 in the pre-frontal cortex of AD patients with changes in intracellular Ca2+ which could lead to neuronal dysfunction and neurodegeneration due to excessive Ca2+ permeability [68]. The authors showed that the pre-frontal cortex of Alzheimer’s patients has approximately 1.0% of all GluA2 RNA molecules unedited. In healthy states the pre-frontal cortex shows less than 0.1% of all GluA2 RNA molecules are unedited and more than 99.9% are edited. Other studies found lower RNA editing levels at the GluA2 Q/R site in the hippocampus and caudate of sporadic AD patients and Apo E4 carriers, independent of clinical diagnosis. Interestingly, ADAR levels were decreased only in the caudate region of the patient’s brains [69]. The E4 allele of the apolipoprotein ApoE gene has been recognized as a major genetic risk factor for AD and it has been suggested that ApoE plays a role in hippocampus AMPA receptor dynamics and glutamate regulation [70,71,72]. Interestingly, studies performed in the triple-transgenic AD mouse model (3×Tg-AD, PS1(M146 V); APP(Swe); tau (P301L)), a widely used transgenic mouse model for AD which exhibits both plaques and tau pathology, showed decreased levels of all AMPA receptor subunits, except for GluA2, while no editing deficiencies were detected [73]. A study aimed at analyzing the hippocampal transcriptome of normal aged mice using RNA sequencing, also examined age-related RNA editing changes as a mechanism to generate alternative transcripts [74]. In 29 months old mice, 41 out of 682 editing sites were significantly changed, which corresponded to 35 genes. One of the genes exhibiting increased editing was the serotonin receptor 2c, which has previously been found showing altered RNA editing in a mouse model of impaired memory function [75]. A comprehensive study on RNA editing in postmortem AD patient tissue revealed significant loss of RNA editing in the hippocampus, and to a lesser extent in the temporal and frontal lobes [76]. Most of the editing changes showed hypo-editing, including the serotonin receptor 2c, which in contrast to what was found in the aging mice discussed above showed less RNA editing in the hippocampus, temporal and frontal lobes. Surprisingly, the authors were unable to find a true correlation between the editing deficits and the expression levels of neither ADAR1 nor ADAR2, suggesting that ADAR dysfunction could be caused by mechanisms other than decreased transcription.
Amyotrophic lateral sclerosis (ALS) is a fatal neurodegenerative disorder where the progressive death of both the upper and lower motor neurons leads to atrophy of skeletal muscles and ultimately death due to respiratory failure [77]. The known genetic contribution to ALS is relatively little, only 10% of the ALS cases are believed to be familial. The remaining 90% of ALS is designated as sporadic ALS in which there is no familial history of the disease [78]. While the etiology remains largely unknown there have been great strides in understanding the pathology of the disease attributed to the advances in genomic sequencing capabilities [79].
Early studies identified the dysregulation of astrocytic glutamate transporters in ALS as the leading cause for increased levels of glutamate at the synapse [80]. Pyramidal tract projection into the spinal cord uses glutamate as the excitatory neurotransmitter and motor neurons expressing abundant glutamate receptors are most vulnerable to exaggerated glutamate stimulation, supporting excitotoxicity as a major mechanism for motor neuron loss in ALS [81]. A likely contributor to the mechanism behind neuronal excitotoxicity is the dysfunction of the AMPA receptor leading to exaggerated calcium influx and slow neuronal death [82, 83]. As mentioned previously, elevated calcium influx through the AMPA receptors can occur through the absence of GluA2 from the receptor complex or through RNA editing of the GluA2 Q/R site. Initial studies addressing the role of AMPA receptors in motor neuron cell death supported both of these mechanisms [84,85,86,87,88]. Over the years, Kwak and colleagues provided accumulating evidence that spinal motor neurons from sporadic ALS patients showed reduced GluA2 Q/R editing efficiencies, leading to increased Ca2+ permeability of AMPA receptors and subsequent excitotoxic motor neuron cell death [84, 89, 90]. The group further showed that these editing deficits are accompanied by a downregulation of ADAR2 [91], and transgenic mice with specific motor neuron knockdown of ADAR2 exhibited inefficient GluA2 Q/R editing and decreased motor function, which was rescued when the mice were crossed with transgenic mice overexpressing a fully edited version of GluA2 [92]. Interestingly, the oculomotor neurons, which are generally not affected in ALS patients, of these mice were not degenerated despite a loss of ADAR2 and a decrease in GluA2 Q/R editing. Also, the motor neurons of the ADAR2 conditional knockout mice exhibited classical TDP-43 pathology, and similar co-pathologies were found in sporadic ALS patient spinal cord motor neurons [93]. The authors propose that Ca2+ influx via unedited GluA2 containing AMPA receptors leads to activation of calpain, which in turn triggers TDP-43 pathology and nucleo-cytoplasmic transport deficits, in addition to excitotoxicity [94, 95].
The loss of GluA2 Q/R editing efficiency has not been demonstrated in other subgroups of ALS, while decreased ADAR2 levels were reported in spinal motor neurons of a single patient carrying a FUS mutation [96]. A recent transcriptome study using deep RNA sequencing technology reported that while spinal cord tissue shows decreased GluA2 Q/R editing efficiencies compared to other brain regions, there was no detectable difference of GluA2 Q/R editing deficits between control spinal cord patient tissue and sporadic ALS patient tissue [97]. One explanation for this discrepancy could be the use of spinal cord tissue lysate versus laser-captured motor neuron analysis, or, the use of RNA sequencing versus a restriction digest-based RNA editing technique. Future studies are required to address these conflicting results. Finally, Donnelly et al. described sequestration of ADAR3 to C9orf72 repeat RNAs in postmortem C9orf72 ALS patient tissue and patient-derived human-induced pluripotent stem cells differentiated into motor neurons (hiPSC-MNs) [98]. Additionally, the hiPSC-MNs showed increased susceptibility to glutamate toxicity, which was mimicked by siRNA knockdown of ADAR3. Ongoing studies in our laboratory are aimed at understanding how ADAR3 dysfunction could regulate ADAR2 function and subsequent excitotoxicity in C9orf72 ALS/Frontotemporal Dementia (FTD), and whether ADAR2 function itself is altered in C9orf72 ALS/FTD patients.
Huntington’s disease (HD) and Parkinson’s disease (PD) have not been investigated much in regards to RNA editing deficits. HD is an autosomal dominant mutation caused by an abnormal trinucleotide CAG repeat expansion in the huntingtin gene (HTT). Carriers of this mutation produce an unusual polyglutamine sequence that causes disease by a toxic gain of function of the protein huntingtin. Even though HD impacts the entire brain, the most affected regions are the basal ganglia and striatum composed of the caudate nucleus and putamen. To a lesser extent areas of the cerebellum, substantia nigra, hippocampus, and layer III, V and IV of the cerebral cortex are affected [99]. Very early research in HD, often referred to as Huntington’s chorea, suggested that aberrant glutamate homeostasis might be involved in HD disease pathogenesis [100]. As an example, researchers used intrastriatial injections of glutamate or kainic acid to mimic biochemical changes observed in HD [101, 102]. With the cloning and discovery of glutamate receptors, the role of glutamate and excitotoxicity becomes a major disease mechanism for HD [103] and the first study examining RNA editing of glutamate receptors subunits GluA2, 5 and 6 noted no difference in the RNA editing efficiency between healthy control and HD patient brain tissue samples [104]. A later study provided the first evidence to support little, yet significant changes in GluA2 editing in the striatum on HD patient tissue [68]. Nearly 5% of GluA2 Q/R was unedited, which still leaves a large percentage of edited GluA2, but could nevertheless contribute to increased Ca2+ permeability and neuronal death. Interestingly, a more recent study decreased immunostaining for GluA2 in the striatum of HD patient tissue when compared to control tissue, suggesting that an overall lack of GluA2 might further contribute to glutamate excitotoxicity in HD [105].
PD is the second most common neurodegenerative disorder affecting nearly 1% of the population [106]. Patients with PD exhibit crippling motor deficits or bradykinesia (or slowness of movement), rigidity, resting tremor, and postural instability also known as the four cardinal manifestations of PD [107]. These symptoms arise due to the degeneration of dopaminergic neurons in the substantia nigra [106, 108]. Similar to HD, among the many proposed cellular dysfunctions [108] excitotoxicity has been suggested to play a role in the degeneration of the dopaminergic neurons [109]. However, despite the proposed role of excitotoxicity there has been little evidence that suggests any known RNA editing deficits [110]. With the increase in RNA sequencing capabilities the ability to study RNA editing events by whole transcriptome sequencing is allowing for more complex analysis of A/I editing sites in disease. One whole transcriptome study associated Parkinson’s disease with changes in Alu insertions the largest target of the ADAR family of proteins [111]. Due to ADARs RNA editing of micro RNAs and Long noncoding RNA and alterations in these RNAs in PD, RNA editing is hypothesized to play a role in disease pathogenesis [110], but only future studies will prove whether this hypothesis is correct. An intriguing new concept has just been proposed in regards to utilizing endogenous ADAR2 editing activity to repair a PD disease causing mutation in PINK1 [112]. A G-to-A mutation in PINK1 introduces a premature stop codon and shortens the protein’s C-terminus including its kinase domain. The authors designed guideRNAs to enable endogenous ADAR2 to edit and recode the user-defined mRNA target. This was successfully achieved in mammalian cell lines and showed a functional rescue of PINK1/Parkin-mediated mitophagy [112].
3.2.2 RNA Editing in Acute Neurodegeneration
Epilepsy is a neurological disorder characterized by abnormal neuronal hyperexcitability of a subpopulation of cells resulting in unprovoked recurrent seizures [113, 114]. The mechanisms responsible for this neuronal hyperexcitability are multifaceted and include genetic predispositions, acute brain injuries, as well as epigenetic changes alterations. Overstimulated cells have a prolonged increase in intracellular Ca2+ concentrations, which has been suggested to contribute to the mechanisms of hyperexcitability seen in epilepsy. AMPA receptors are involved in fast excitatory neurotransmission and are therefore thought to play a key role in the generation of seizures. Various studies present evidence that connects deficits in AMPA receptor editing with seizure vulnerability. Transgenic mice expressing a fully unedited GluA2 Q/R site die around 3 weeks of age and develop severe seizures [115]. Interestingly, GluA2 knockout mice, while similarly showing premature death, do not show signs of seizures, but instead show increased susceptibility to absence seizures [116]. ADAR2 knockout mice behave very similar to the GluA2 Q/R unedited mice and develop seizures before prematurely dying at 21 days of age [13]. These mice are rescued by crossing the ADAR2 KO mice with transgenic mice overexpressing a fully edited GluA2 Q/R site [13]. RNA editing analyses of epileptic brain tissue resulted in contradictory results, with studies showing no altered RNA editing at the GluA2 Q/R site (while there were RNA editing changes in GluA5 and GluA6) [117]. Only one study examined ADAR2 expression from needle biopsy samples obtained from hypothalamic hamartoma tissue and found loss of nuclear immunostaining of ADAR2 concomitant with lower RNA editing efficiency at the GluA2 Q/R site [118]. A recent genome-wide analysis of epileptic and healthy mouse hippocampus revealed a correlation between seizure frequency and differential RNA editing [119]. Functional enrichment analysis revealed that pathways relevant for epilepsy showed the highest degree of differential RNA editing, e.g., neuron projection, synapse, seizures. More work needs to be done to fully understand whether RNA editing plays a significant role in this disorder.
Stroke patients suffer from a spontaneously disrupted blood supply to the brain resulting in a loss of oxygen and nutrients to affected regions. Accounting for 85% of all strokes an ischemic stroke occurs when blood flow to part of the brain is obstructed. After an ischemic attack and loss of blood supply, cells are immediately unable to sustain normal homeostasis leading to massive irreversible cell death [120]. Because of the rapid neuronal loss in stroke victims immediate and effective treatment is crucial to minimize damage [121, 122]. Post-ischemic excitotoxicity results from consumption of ATP, failure of ATP synthesis, and dysregulation of the ionic concentration across the plasma membrane leading to rapid rise in intracellular calcium concentrations and death of the cell [123]. Historically, the increase in calcium permeability of neurons affected by ischemia was thought to be due the downregulation of GluA2 following ischemia commonly referred to as “The GluA2 hypothesis” [36]. However, in 2006 unedited GluA2 was found in the CA1 pyramidal neurons of rats following ischemia [124]. The calcium permeability of AMPA receptors in the CA1 pyramidal neurons is 18-fold higher following ischemia when compared to control groups [125]. In addition, loss of ADAR2 expression increases neuronal sensitivity to ischemia and can be rescued by expression of a fully edited GluA2(R) [124, 125]. These studies suggest that loss of RNA editing contributes to the disruption in neuronal homeostasis following ischemic stroke and immediate prevention of these deficits may protect against neuronal damage.
Spinal Cord Injury (SCI) is defined as damage to the spinal cord causing reduced or complete loss of motor function [126, 127]. It generally affects glutamatergic tracts descending from varying brain regions and serotonergic tracts descending from the brainstem. Serotonin signaling is critical in the spinal cord by providing neuromodulation to motor neuron and recent studies showed reduced A → I RNA editing of the 5HT2cR serotonin receptor after SCI, which was suggested to contribute to loss of motor neuron function [126,127,128,129]. These studies demonstrated that RNA editing deficiency for 5HT2cR was due to a decrease in the ADAR2 expression suggested to be caused by a continuous inflammatory response during injury. In addition to 5HT2cR, the authors also found reduced RNA editing of potassium channel Kv1.1, an additional ADAR2 target. Additional studies strongly support the fact that microglial cells and immune infiltrating cells are involved in the dysfunction of A → I RNA editing in SCI [4, 128, 129], suggesting that at least during spinal cord injury, RNA deficits of neuronal targets are triggered by non-cell autonomous mechanisms. Future studies are needed to test the hypothesis that these non-cell autonomous mechanisms also occur in other neurodegenerative diseases characterized by RNA editing deficits.
3.3 A → I RNA Editing Dysfunction in Psychiatric Diseases
Depression and Schizophrenia. Depression is a long term mood disorder that affects a person’s thoughts and feelings as well as daily activities such as working, eating and sleeping [130]. This disorder is caused by a combination of genetic, biological and environmental factors. Serotonin or 5-hydroxytryptamine (5HT), a monoamine neurotransmitter has been implicated in this psychiatric disease [131, 132]. Patients suffering from depression have lower levels of serotonin or an increase in the number of serotonin receptors. Selective serotonin reuptake inhibitors are frequently used to treat depression to maintain serotonin for longer periods at the synapse. Schizophrenia is classified as a chronic mental disorder where the patients lose contact with reality and present psychotic behaviors (positive symptoms), disruption of normal behaviors (negative symptoms), poor executive function and poor working memory (cognitive symptoms) [133]. Similar to depression, schizophrenia is caused by genetic aberrations and environmental factors.
5HT-serotonergic receptors are relevant to mental disorders such as depression, anxiety, and schizophrenia. The 5HT2cR , a G-protein couple receptor, is known to undergo RNA editing post-transcriptional modification [32, 134,135,136]. Altered editing of 5HT2cR pre-mRNA occurs in the pre-frontal cortex of depressive and schizophrenic patients. A/I RNA editing of the 5HT2cR occurs at five sites (A-to-E) causing protein and functional diversity. Previous studies have shown that depressive and schizophrenic patients have reduced expression of ADAR2 with a decrease or increase in RNA editing in some of the five 5HT2cR sites [137, 138] making it difficult to elucidate how RNA editing is associated with these psychiatric disorders. These studies suggest that RNA editing is not only disease-specific, but it may also be determined by the severity of the psychiatric diseases.
Cocaine addiction. An estimated 18.3 million people between the ages of 16–64 used cocaine in 2014 making it one of the most common illicit drugs in the world (National Institute on Drug Abuse 2016; [139]). Numerous health risks are associated with cocaine use such as cognitive impairment, respiratory disease, cardiovascular disease, congenital malformations, and premature mortality [140]. Approximately 20% of recreational users will develop a dependence for cocaine within 5 years [141]. Drug-seeking behavior is thought to be influenced by limbic cortical-ventral striatal circuitry which afferents to the basolateral amygdala and nucleus accumbens providing the circuitry for stimulus-reward pathway that reinforces drug seeking behavior [142]. Increased calcium permeable AMPA receptors in the nucleus accumbens have been associated with drug-seeking behavior [143]. These alterations may be due to increased GluA1 in the nucleus accumbens [144]. However, downregulation of ADAR2 and GluA2 Q/R editing deficits have been identified in the nucleus accumbens shell in rats following cocaine self-administration [145]. Both upregulation of GluA1 and misediting of the GluA2 Q/R site could explain alterations in the nucleus accumbens that leads to the reinforcement of drug-seeking behavior.
Considered a multi-factorial disorder autism spectrum disorder (ASD) is a range of neurological abnormalities affecting one in 68 children in the United States [146]. Children affected by ASD exhibit reduced eye contact, facial expression, and body gestures [147]. Due to the heterogeneity of the classification of the disease the etiology is still widely unknown. Genetic causes have only been identified in 10–20% of individuals. Deep whole transcriptome sequencing of 30 patients with ASD identified RNA A/I editing alterations in 20 of 25 sites analyzed [148]. In contrast to other neurodegenerative disorders discussed in this chapter, RNA editing levels in ASD were found to be significantly higher than control groups [148]. Interestingly, the editing at the GluA2 Q/R site is not altered in ASD [148, 149]. Alterations in RNA A/I editing in ASD have been explained by alterations in ADAR2 self-regulation and loss of fmr1 [148,149,150].
3.4 Brain Cancer
Glioblastoma multiforme (GBM) is a tumor generated from astroglial cells generally localized in the cerebral hemispheres, and to a lesser extent in other regions of the brain or spinal cord. A transcriptome study using RNA sequencing for global A-to-I editing events in human revealed that genes with predicted editing events were significantly enriched for cancer-related genes, suggesting that RNA editing plays a significant role in the development of cancer [151]. This was later confirmed by Hwang and colleagues, who showed via gene ontology analyses that there was a selective change in the pattern of RNA editing in gliobastomas [30] (also recently reviewed in [152]). Indeed, early studies found a significant reduction in the GluA2 Q/R and the serotonin receptor 5-HT(2C) editing efficiency in malignant human brain tumors, which correlated with decreased ADAR2 self-editing activity [153]. These studies were confirmed when significantly reduced editing in Alu sequences was found in brain tissues [154]. All three ADAR genes showed lower RNA levels and the reduced ADAR3 levels correlated with the grade of malignancy of glioblastoma multiforme. Along those lines, high grade astrocytomas equally show lack of ADAR2 editing activity when grown in vitro, as well as in vivo via a flank tumor growth model in nude mice [155, 156].
As previously discussed, A-to-I editing also affects miRNAs, ~22 nucleotide long noncoding RNAs known to silence gene expression by binding to the 3′untranslated region (3’UTR) of mRNAs. miRNAs can undergo A-to-I RNA editing at premature states when the miRNA has a double-stranded structure. Analyses of high grade gliomas revealed reduced editing of miRNA-376 [157]. The authors found a strong correlation between the extent of unedited miRNA-376 and tumor spread, which was measured using magnetic resonance imaging of the patient’s brains. The authors further confirmed these results in xenograft mouse models, showing that unedited miRNA-376 promoted glioma growth and spread, while edited miRNA-376 was protective. Similar results were recently reported on miRNA-589-3p [158]. A more recent study showed that A-to-I miRNA editing is enhanced at the seed region of the miRNA, an area critical to bind its target mRNA [159]. The authors further confirmed by RNA sequencing of GBM patient tissue that a significant reduction of miRNA editing occurs in GBM tissue and is correlated with the reduction of ADAR2 expression [159].
Interestingly, one study found elevated levels of ADAR3 in GBMs when compared to control brain tissue [16]. The authors suggested ADAR3 as a potential regulator of the Q/R editing site by binding to GluA2 subunit pre-mRNA and thereby inhibiting editing by ADAR2 in GBM. They hypothesized that an elevated expression of ADAR3 and reduced GluA2 editing will induce calcium permeability through the glutamate receptor, which in turn accelerates cell migration and tumor invasion into surrounding peri-tumoral tissue.
3.5 Conclusions
RNA editing, with now an estimate of over a million editing sites in primates and humans, has gained increasing interest as an important mechanism of RNA processing, not only during development, but also in disease. Given its ability to contribute to the molecular complexity in the human body, including the brain, it is of importance that we learn more about the regulation of RNA editing and how it can contribute to disease pathogenesis. It will be important to fully understand temporal and spatial regulation, of specific brain regions and likely also cell types, of the individual ADAR editing enzymes. This knowledge will be especially critical if we consider targeting ADAR enzymes for therapeutic purposes in any of the discussed diseases, as well as any non-CNS disorders.
References
Rueter SM, Dawson TR, Emeson RB. Regulation of alternative splicing by RNA editing. Nature. 1999;399(6731):75–80.
Gerber AP, Keller W. RNA editing by base deamination: more enzymes, more targets, new mysteries. Trends Biochem Sci. 2001;26(6):376–84.
Paul MS, Bass BL. Inosine exists in mRNA at tissue-specific levels and is most abundant in brain mRNA. EMBO J. 1998;17(4):1120–7.
Gal-Mark N, et al. Abnormalities in A-to-I RNA editing patterns in CNS injuries correlate with dynamic changes in cell type composition. Sci Rep. 2017;7:43421.
Lee SY, et al. RCARE: RNA sequence comparison and annotation for RNA editing. BMC Med Genomics. 2015;8(Suppl 2):S8.
Gu T, et al. Canonical A-to-I and C-to-U RNA editing is enriched at 3′UTRs and microRNA target sites in multiple mouse tissues. PLoS One. 2012;7(3):e33720.
Kim DD, et al. Widespread RNA editing of embedded alu elements in the human transcriptome. Genome Res. 2004;14(9):1719–25.
Yang JH, et al. Intracellular localization of differentially regulated RNA-specific adenosine deaminase isoforms in inflammation. J Biol Chem. 2003;278(46):45833–42.
Patterson JB, Samuel CE. Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: evidence for two forms of the deaminase. Mol Cell Biol. 1995;15(10):5376–88.
Wang Q, et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J Biol Chem. 2004;279(6):4952–61.
Hartner JC, et al. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J Biol Chem. 2004;279(6):4894–902.
Yao L, et al. Large-scale prediction of ADAR-mediated effective human A-to-I RNA editing. Brief Bioinform. 2017;PMID:28968662.
Higuchi M, et al. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature. 2000;406(6791):78–81.
Chen CX, Cho DS, Wang Q, Lai F, Carter KC, Nishikura K. A third member of the RNA-specific adenosine deaminase gene family, ADAR3, contains both single- and double-stranded RNA binding domains. RNA. 2000;6:755–67.
Galipon J, et al. Differential binding of three major human ADAR isoforms to coding and long non-coding transcripts. Genes (Basel). 2017;8(2):pii:E68.
Oakes E, et al. Adenosine deaminase that acts on RNA 3 (ADAR3) binding to glutamate receptor subunit B pre-mRNA inhibits RNA editing in glioblastoma. J Biol Chem. 2017;292(10):4326–35.
Fitzgerald LW, Iyer G, Conklin DS, Krause CM, Marshall A, Patterson JP, Tran DP, Jonak GJ, Hartig PR. Messenger RNA editing of the human serotonin 5-HT 2C receptor. Europsychopharmacology. 1999;21(2S):82S–90S.
Bazak L, et al. A-to-I RNA editing occurs at over a hundred million genomic sites, located in a majority of human genes. Genome Res. 2014;24(3):365–76.
Ramaswami G, et al. Identifying RNA editing sites using RNA sequencing data alone. Nat Methods. 2013;10(2):128–32.
Ramaswami G, et al. Accurate identification of human Alu and non-Alu RNA editing sites. Nat Methods. 2012;9(6):579–81.
Neeman Y, et al. RNA editing level in the mouse is determined by the genomic repeat repertoire. RNA. 2006;12(10):1802–9.
Rodriguez J, Menet JS, Rosbash M. Nascent-seq indicates widespread cotranscriptional RNA editing in drosophila. Mol Cell. 2012;47(1):27–37.
Levanon EY, et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat Biotechnol. 2004;22(8):1001–5.
Blow M, et al. A survey of RNA editing in human brain. Genome Res. 2004;14(12):2379–87.
Athanasiadis A, Rich A, Maas S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2004;2(12):e391.
Tan MH, et al. Dynamic landscape and regulation of RNA editing in mammals. Nature. 2017;550(7675):249–54.
Bahn JH, et al. Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways. Nat Commun. 2015;6:6355.
Korenberg JR, Rykowski MC. Human genome Orginazation: alu, lines, and the molecular structure of metaphase chromosome bands. Cell. 1988;53:391–400.
Yablonovitch AL, et al. The evolution and adaptation of A-to-I RNA editing. PLoS Genet. 2017;13(11):e1007064.
Hwang T, et al. Dynamic regulation of RNA editing in human brain development and disease. Nat Neurosci. 2016;19(8):1093–9.
Higuchi M, et al. RNA editing of AMPA receptor subunit GluR-B: a base-paired intron-exon structure determines position and efficiency. Cell. 1993;75(7):1361–70.
Burns CM, et al. Regulation of serotonin-2C receptor G-protein coupling by RNA editing. Nature. 1997;387(6630):303–8.
Henley JM, Wilkinson KA. Synaptic AMPA receptor composition in development, plasticity and disease. Nat Rev Neurosci. 2016;17(6):337–50.
Huganir RL, Nicoll RA. AMPARs and synaptic plasticity: the last 25 years. Neuron. 2013;80(3):704–17.
Hollmann M, Hartley M, Heinemann SF. Calcium permeability of KA-AMPA-gated glutamate receptor channels depnds on subunit composition. Science. 1991;252:851–3.
Bennett MV, et al. The GluR2 hypothesis: ca(++)-permeable AMPA receptors in delayed neurodegeneration. Cold Spring Harb Symp Quant Biol. 1996;61:373–84.
Wright A, Vissel B. The essential role of AMPA receptor GluR2 subunit RNA editing in the normal and diseased brain. Front Mol Neurosci. 2012;5:34.
Wenthold RJ, et al. Evidence for multiple AMPA receptor complexes in hippocampal CA1/CA2 neurons. J Neurosci. 1996;16(6):1982–9.
Isaac JT, Ashby MC, McBain CJ. The role of the GluR2 subunit in AMPA receptor function and synaptic plasticity. Neuron. 2007;54(6):859–71.
Cull-Candy S, Kelly L, Farrant M. Regulation of Ca2+-permeable AMPA receptors: synaptic plasticity and beyond. Curr Opin Neurobiol. 2006;16(3):288–97.
Sanderson JL, Gorski JA, Dell'Acqua ML. NMDA receptor-dependent LTD requires transient synaptic incorporation of ca(2+)-permeable AMPARs mediated by AKAP150-anchored PKA and calcineurin. Neuron. 2016;89(5):1000–15.
Nishikura K. Functions and regulation of RNA editing by ADAR deaminases. Annu Rev Biochem. 2010;79:321–49.
Washburn MC, et al. The dsRBP and inactive editor ADR-1 utilizes dsRNA binding to regulate A-to-I RNA editing across the C. elegans transcriptome. Cell Rep. 2014;6(4):599–607.
Melcher T, et al. RED2, a brain-specific member of the RNA-specific adenosine deaminase family. J Biol Chem. 1996;271(50):31795–8.
Melcher T, et al. Editing of alpha-amino-3-hydroxy-5-methylisoxazole-4-propionic acid receptor GluR-B pre-mRNA in vitro reveals site-selective adenosine to inosine conversion. J Biol Chem. 1995;270(15):8566–70.
Sommer B, et al. RNA editing in brain controls a determinant of ion flow in glutamate-gated channels. Cell. 1991;67(1):11–9.
Ray RS, et al. Impaired respiratory and body temperature control upon acute serotonergic neuron inhibition. Science. 2011;333(6042):637–42.
Gershon MD, et al. 5-HT receptor subtypes outside the central nervous system. Roles in the physiology of the gut. Neuropsychopharmacology. 1990;3(5–6):385–95.
Spencer NJ, Keating DJ. Is there a role for endogenous 5-HT in gastrointestinal motility? How recent studies have changed our understanding. Adv Exp Med Biol. 2016;891:113–22.
Hood JL, Emeson RB. Editing of neurotransmitter receptor and ion channel RNAs in the nervous system. Curr Top Microbiol Immunol. 2012;353:61–90.
Bockaert J, et al. Neuronal 5-HT metabotropic receptors: fine-tuning of their structure, signaling, and roles in synaptic modulation. Cell Tissue Res. 2006;326(2):553–72.
Hoyer D, Clarke DE, Fozard JR, Hartig PR, Martin GR, Mylecharane EJ, Saxena PR, Humphrey PP. International Union of Pharmacology classification of receptors for 5-hydroxytryptamine (serotonin). Pharmacol Rev. 1994;46(2):157–203.
McCorvy JD, Roth BL. Structure and function of serotonin G protein-coupled receptors. Pharmacol Ther. 2015;150:129–42.
Helton L, Thor KB, Baez M. 5-Hydroxytryptamine2A, 5-hydroxytryptamine2B, and 5-hydroxytryptamine2C receptor mRNA expression in the spinal cord of rat, cat, monkey and human. Mol Neurosci. 1994;5:2617–20.
Iwamoto K, et al. Measuring RNA editing of serotonin 2C receptor. Biochemistry (Mosc). 2011;76(8):912–4.
Price RD, et al. RNA editing of the human serotonin 5-HT2C receptor alters receptor-mediated activation of G13 protein. J Biol Chem. 2001;276(48):44663–8.
Niswender CM, Copeland SC, Herrick-Davis K, Emeson RB, Sanders-Bush E. RNA editing of the human serotonin 5-hydroxytryptamine 2C receptor silences constitutive activity. J Biol Chem. 1999;274(14):9472–8.
Sanders-Bush E, Price RD. RNA editing of the human serotonin 5-HT2C receptor delays agonist-stimulated calcium release. Mol Pharmacol. 2000;58(4):859–62.
Coetzee WA, Amarillo Y, Chiu J, Chow A, Lau D, McCormack T, Moreno H, Nadal MS, Ozaita A, Pountney D, Saganich M, Vega-Saenz de Miera E, Rudy B. Molecular diversity of K+ channels. Ann N Y Acad Sci. 1999;868:233–85.
Tian C, et al. Potassium channels: structures, diseases, and modulators. Chem Biol Drug Des. 2014;83(1):1–26.
Wang H, Kunkel DD, Schwartzkroin PA, Tempel BL. Localization of Kv1.1 and Kv1.2, two K channel proteins, to synaptic terminals, somata, and dendrites in the mouse brain. J Neurosci. 1994;14(8):4588–99.
Robbins CA, Tempel BL. Kv1.1 and Kv1.2: similar channels, different seizure models. Epilepsia. 2012;53(Suppl 1):134–41.
Bhalla T, et al. Control of human potassium channel inactivation by editing of a small mRNA hairpin. Nat Struct Mol Biol. 2004;11(10):950–6.
Rettig J, Heinemann SH, Wunder F, Lorra C, Parcej DN, Dolly JO, Pongs O. Inactivation properties of voltage-gated K+ channels altered by presence of beta-subunit. Nature. 1994;369(6478):289–94.
Kumar A, Singh A, Ekavali. A review on Alzheimer’s disease pathophysiology and its management: an update. Pharmacol Rep. 2015;67(2):195–203.
Area-Gomez E, Schon EA. Alzheimer disease. Adv Exp Med Biol. 2017;997:149–56.
Calderon-Garciduenas AL, Duyckaerts C. Alzheimer disease. Handb Clin Neurol. 2017;145:325–37.
Akbarian S, Smith MA, Jones EG. Editing for an AMPA receptor subunit RNA in prefrontal cortex and striatum in Alzheimer’s disease, Huntington’s disease and schizophrenia. Brain Res. 1995;699(2):297–304.
Gaisler-Salomon I, et al. Hippocampus-specific deficiency in RNA editing of GluA2 in Alzheimer’s disease. Neurobiol Aging. 2014;35(8):1785–91.
Payami H, et al. Apolipoprotein E genotype and Alzheimer’s disease. Lancet. 1993;342(8873):738.
Saunders AM, et al. Association of apolipoprotein E allele epsilon 4 with late-onset familial and sporadic Alzheimer’s disease. Neurology. 1993;43(8):1467–72.
Valastro B, et al. AMPA receptor regulation and LTP in the hippocampus of young and aged apolipoprotein E-deficient mice. Neurobiol Aging. 2001;22(1):9–15.
Cantanelli P, et al. Age-dependent modifications of AMPA receptor subunit expression levels and related cognitive effects in 3xTg-AD mice. Front Aging Neurosci. 2014;6:200.
Stilling RM, et al. De-regulation of gene expression and alternative splicing affects distinct cellular pathways in the aging hippocampus. Front Cell Neurosci. 2014;8:373.
Stilling RM, et al. K-lysine acetyltransferase 2a regulates a hippocampal gene expression network linked to memory formation. EMBO J. 2014;33(17):1912–27.
Khermesh K, et al. Reduced levels of protein recoding by A-to-I RNA editing in Alzheimer’s disease. RNA. 2016;22(2):290–302.
Vucic S, Rothstein JD, Kiernan MC. Advances in treating amyotrophic lateral sclerosis: insights from pathophysiological studies. Trends Neurosci. 2014;37(8):433–42.
Taylor JP, Brown RH Jr, Cleveland DW. Decoding ALS: from genes to mechanism. Nature. 2016;539(7628):197–206.
Bettencourt C, Houlden H. Exome sequencing uncovers hidden pathways in familial and sporadic ALS. Nat Neurosci. 2015;18(5):611–3.
Chien-Liang Glenn Lin LAB, Lin J, Margaret Dykes-Hoberg TC, Lora-Clawson JDR. 1-s2.0-S0896627300809976-main.Pdf. Neuron. 1998;20:589–602.
Rothstein JD, et al. Abnormal excitatory amino acid metabolism in amyotrophic lateral sclerosis. Ann Neurol. 1990;28(1):18–25.
Lu YM, Yin HZ, Chiang J, Weiss JH. Ca2+-permeable AMPA/Kainate and NMDA channels: high rate of Ca2+ influx underlies potent induction of injury. J Neurosci. 1996;16(17):5457–65.
Carriedo SG, Yin HZ, Weiss JH. Motor neurons are selectively vulnerable to AMPA/kainate receptor-mediated injury in vitro. J Neurosci. 1996;16(13):4069–79.
Takuma H, et al. Reduction of GluR2 RNA editing, a molecular change that increases calcium influx through AMPA receptors, selective in the spinal ventral gray of patients with amyotrophic lateral sclerosis. Ann Neurol. 1999;46(6):806–15.
Kawahara Y, et al. Human spinal motoneurons express low relative abundance of GluR2 mRNA: an implication for excitotoxicity in ALS. J Neurochem. 2003;85(3):680–9.
Williams TL, et al. Calcium-permeable alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid receptors: a molecular determinant of selective vulnerability in amyotrophic lateral sclerosis. Ann Neurol. 1997;42(2):200–7.
Van Den Bosch L, et al. Ca(2+)-permeable AMPA receptors and selective vulnerability of motor neurons. J Neurol Sci. 2000;180(1–2):29–34.
Morrison BM, et al. Light and electron microscopic distribution of the AMPA receptor subunit, GluR2, in the spinal cord of control and G86R mutant superoxide dismutase transgenic mice. J Comp Neurol. 1998;395(4):523–34.
Kawahara Y, et al. Glutamate receptors: RNA editing and death of motor neurons. Nature. 2004;427(6977):801.
Kwak S, Kawahara Y. Deficient RNA editing of GluR2 and neuronal death in amyotropic lateral sclerosis. J Mol Med (Berl). 2005;83(2):110–20.
Hideyama T, et al. Profound downregulation of the RNA editing enzyme ADAR2 in ALS spinal motor neurons. Neurobiol Dis. 2012;45(3):1121–8.
Hideyama T, et al. Induced loss of ADAR2 engenders slow death of motor neurons from Q/R site-unedited GluR2. J Neurosci. 2010;30(36):11917–25.
Aizawa H, et al. TDP-43 pathology in sporadic ALS occurs in motor neurons lacking the RNA editing enzyme ADAR2. Acta Neuropathol. 2010;120(1):75–84.
Yamashita T, et al. Calpain-dependent disruption of nucleo-cytoplasmic transport in ALS motor neurons. Sci Rep. 2017;7:39994.
Yamashita T, Akamatsu M, Kwak S. Altered intracellular milieu of ADAR2-deficient motor neurons in amyotrophic lateral sclerosis. Genes (Basel). 2017;8(2):pii: E60.
Aizawa H, et al. Deficient RNA-editing enzyme ADAR2 in an amyotrophic lateral sclerosis patient with a FUSP525L mutation. J Clin Neurosci. 2016;32:128–9.
D’Erchia AM, et al. Massive transcriptome sequencing of human spinal cord tissues provides new insights into motor neuron degeneration in ALS. Sci Rep. 2017;7(1):10046.
Donnelly CJ, et al. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron. 2013;80(2):415–28.
Bates GP, et al. Huntington disease. Nat Rev Dis Primers. 2015;1:15005.
Olney JW, de Gubareff T. Glutamate neurotoxicity and Huntington’s chorea. Nature. 1978;271(5645):557–9.
McGeer EG, McGeer PL. Duplication of biochemical changes of Huntington’s chorea by intrastriatal injections of glutamic and kainic acids. Nature. 1976;263(5577):517–9.
Coyle JT, Schwarcz R. Lesion of striatal neurones with kainic acid provides a model for Huntington’s chorea. Nature. 1976;263(5574):244–6.
Beal MF. Huntington’s disease, energy, and excitotoxicity. Neurobiol Aging. 1994;15(2):275–6.
Paschen W, Hedreen JC, Ross CA. RNA editing of the glutamate receptor subunits GluR2 and GluR6 in human brain tissue. J Neurochem. 1994;63(5):1596–602.
Fourie C, et al. Differential changes in postsynaptic density proteins in postmortem Huntington’s disease and Parkinson’s disease human brains. J Neurodegener Dis. 2014;2014:938530.
Garrett E, Alexander M. Biology of Parkinson’s disease: pathogenesis and pathophysiology of a multisystem neurodegenerative disorder. Dialogues Clin Neurosci. 2004;6(3):259–80.
Jankovic J. Parkinson's disease: clinical features and diagnosis. J Neurol Neurosurg Psychiatry. 2008;79(4):368–76.
Maiti P, Manna J, Dunbar GL. Current understanding of the molecular mechanisms in Parkinson’s disease: targets for potential treatments. Transl Neurodegener. 2017;6:28.
Dong XX, Wang Y, Qin ZH. Molecular mechanisms of excitotoxicity and their relevance to pathogenesis of neurodegenerative diseases. Acta Pharmacol Sin. 2009;30(4):379–87.
Labbe C, Lorenzo-Betancor O, Ross OA. Epigenetic regulation in Parkinson’s disease. Acta Neuropathol. 2016;132(4):515–30.
Paz-Yaacov N, et al. Adenosine-to-inosine RNA editing shapes transcriptome diversity in primates. Proc Natl Acad Sci U S A. 2010;107(27):12174–9.
Wettengel J, et al. Harnessing human ADAR2 for RNA repair – recoding a PINK1 mutation rescues mitophagy. Nucleic Acids Res. 2017;45(5):2797–808.
Chen T, et al. Genetic and epigenetic mechanisms of epilepsy: a review. Neuropsychiatr Dis Treat. 2017;13:1841–59.
Wang J, et al. Epilepsy-associated genes. Seizure. 2017;44:11–20.
Brusa R, et al. Early-onset epilepsy and postnatal lethality associated with an editing-deficient GluR-B allele in mice. Science. 1995;270(5242):1677–80.
Hu RQ, et al. Gamma-hydroxybutyric acid-induced absence seizures in GluR2 null mutant mice. Brain Res. 2001;897(1–2):27–35.
Kortenbruck G, et al. RNA editing at the Q/R site for the glutamate receptor subunits GLUR2, GLUR5, and GLUR6 in hippocampus and temporal cortex from epileptic patients. Neurobiol Dis. 2001;8(3):459–68.
Kitaura H, et al. Ca(2+)-permeable AMPA receptors associated with epileptogenesis of hypothalamic hamartoma. Epilepsia. 2017;58(4):e59–63.
Srivastava PK, et al. Genome-wide analysis of differential RNA editing in epilepsy. Genome Res. 2017;27(3):440–50.
RAB JN, Allen SG, Fujikawa DG, Wasterlain CG. Hypoxic neuronal necrosis: protein synthesisindependent activation of a cell death program. PNAS. 2003;100(5):2825–30.
Saver JL. Time is brain—quantified. Stroke. 2006;37(1):263–6.
Vilela P, Rowley HA. Brain ischemia: CT and MRI techniques in acute ischemic stroke. Eur J Radiol. 2017;96:162–72.
Khoshnam SE, et al. Pathogenic mechanisms following ischemic stroke. Neurol Sci. 2017;38(7):1167–86.
Peng PL, et al. ADAR2-dependent RNA editing of AMPA receptor subunit GluR2 determines vulnerability of neurons in forebrain ischemia. Neuron. 2006;49(5):719–33.
Liu S, et al. Expression of ca(2+)-permeable AMPA receptor channels primes cell death in transient forebrain ischemia. Neuron. 2004;43(1):43–55.
Ahuja CS, et al. Traumatic spinal cord injury. Nat Rev Dis Primers. 2017;3:17018.
Ahuja CS, et al. Traumatic spinal cord injury-repair and regeneration. Neurosurgery. 2017;80(3S):S9–S22.
Di Narzo AF, et al. Decrease of mRNA editing after spinal cord injury is caused by down-regulation of ADAR2 that is triggered by inflammatory response. Sci Rep. 2015;5:12615.
Murray KC, et al. Recovery of motoneuron and locomotor function after spinal cord injury depends on constitutive activity in 5-HT2C receptors. Nat Med. 2010;16(6):694–700.
Huang YJ, Lane HY, Lin CH. New treatment strategies of depression: based on mechanisms related to neuroplasticity. Neural Plast. 2017;2017:4605971.
Palacios JM, Pazos A, Hoyer D. A short history of the 5-HT2C receptor: from the choroid plexus to depression, obesity and addiction treatment. Psychopharmacology. 2017;234(9–10):1395–418.
Lin SH, Lee LT, Yang YK. Serotonin and mental disorders: a concise review on molecular neuroimaging evidence. Clin Psychopharmacol Neurosci. 2014;12(3):196–202.
Owen MJ, Sawa A, Mortensen PB. Schizophrenia. Lancet. 2016;388(10039):86–97.
Baxter G, et al. 5-HT2 receptor subtypes: a family re-united? Trends Pharmacol Sci. 1995;16(3):105–10.
Sergeeva OA, Amberger BT, Haas HL. Editing of AMPA and serotonin 2C receptors in individual central neurons, controlling wakefulness. Cell Mol Neurobiol. 2007;27(5):669–80.
Barnes NM, Sharp T. A review of central 5-HT receptors and their function. Neuropharmacology. 1999;38(8):1083–152.
Lyddon R, et al. Serotonin 2c receptor RNA editing in major depression and suicide. World J Biol Psychiatry. 2013;14(8):590–601.
Kubota-Sakashita M, et al. A role of ADAR2 and RNA editing of glutamate receptors in mood disorders and schizophrenia. Mol Brain. 2014;7:5.
Park TM, Haning WF III. Stimulant use disorders. Child Adolesc Psychiatr Clin N Am. 2016;25(3):461–71.
Mathuru AS. A little rein on addiction. Semin Cell Dev Biol. 2017. https://doi.org/10.1016/j.semcdb.2017.09.030.
Lopez-Quintero C, et al. Probability and predictors of transition from first use to dependence on nicotine, alcohol, cannabis, and cocaine: results of the National Epidemiologic Survey on alcohol and related conditions (NESARC). Drug Alcohol Depend. 2011;115(1–2):120–30.
Everitt BJ. Neural and psychological mechanisms underlying compulsive drug seeking habits and drug memories—indications for novel treatments of addiction. Eur J Neurosci. 2014;40(1):2163–82.
Carr KD, et al. AMPA receptor subunit GluR1 downstream of D-1 dopamine receptor stimulation in nucleus accumbens shell mediates increased drug reward magnitude in food-restricted rats. Neuroscience. 2010;165(4):1074–86.
Zheng D, et al. Nucleus accumbens AMPA receptor involvement in cocaine-conditioned place preference under different dietary conditions in rats. Psychopharmacology. 2015;232(13):2313–22.
Schmidt HD, et al. ADAR2-dependent GluA2 editing regulates cocaine seeking. Mol Psychiatry. 2015;20(11):1460–6.
Deborah L, Christensen P, et al. Prevalence and characteristics of autism spectrum disorder among children aged 8 years – autism and developmental disabilities monitoring network, 11 sites, United States, 2012. MMWR. 2016;65(3):1–23.
Park HR, et al. A short review on the current understanding of autism spectrum disorders. Exp Neurobiol. 2016;25(1):1–13.
Eran A, et al. Comparative RNA editing in autistic and neurotypical cerebella. Mol Psychiatry. 2013;18(9):1041–8.
Shamay-Ramot A, et al. Fmrp interacts with adar and regulates RNA editing, synaptic density and locomotor activity in zebrafish. PLoS Genet. 2015;11(12):e1005702.
Feng Y, et al. Altered RNA editing in mice lacking ADAR2 autoregulation. Mol Cell Biol. 2006;26(2):480–8.
Bahn JH, et al. Accurate identification of A-to-I RNA editing in human by transcriptome sequencing. Genome Res. 2012;22(1):142–50.
Wang C, et al. Mechanisms and implications of ADAR-mediated RNA editing in cancer. Cancer Lett. 2017;411:27–34.
Maas S, et al. Underediting of glutamate receptor GluR-B mRNA in malignant gliomas. Proc Natl Acad Sci U S A. 2001;98(25):14687–92.
Paz N, et al. Altered adenosine-to-inosine RNA editing in human cancer. Genome Res. 2007;17(11):1586–95.
Galeano F, et al. ADAR2-editing activity inhibits glioblastoma growth through the modulation of the CDC14B/Skp2/p21/p27 axis. Oncogene. 2013;32(8):998–1009.
Cenci C, et al. Down-regulation of RNA editing in pediatric astrocytomas: ADAR2 editing activity inhibits cell migration and proliferation. J Biol Chem. 2008;283(11):7251–60.
Choudhury Y, et al. Attenuated adenosine-to-inosine editing of microRNA-376a* promotes invasiveness of glioblastoma cells. J Clin Invest. 2012;122(11):4059–76.
Cesarini V, et al. ADAR2/miR-589-3p axis controls glioblastoma cell migration/invasion. Nucleic Acids Res. 2017;46(4):2045–59.
Paul D, et al. A-to-I editing in human miRNAs is enriched in seed sequence, influenced by sequence contexts and significantly hypoedited in glioblastoma multiforme. Sci Rep. 2017;7(1):2466.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Lorenzini, I., Moore, S., Sattler, R. (2018). RNA Editing Deficiency in Neurodegeneration. In: Sattler, R., Donnelly, C. (eds) RNA Metabolism in Neurodegenerative Diseases. Advances in Neurobiology, vol 20. Springer, Cham. https://doi.org/10.1007/978-3-319-89689-2_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-89689-2_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-89688-5
Online ISBN: 978-3-319-89689-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)