Abstract
Recent efforts have shed new light on the epigenetic mechanisms driving gene expression alterations associated with Parkinson’s disease (PD) pathogenesis. Changes in gene expression are a well-established cause of PD, and epigenetic mechanisms likely play a pivotal role in regulation. Studies in families with PD harboring duplications and triplications of the SNCA gene have demonstrated that gene dosage is associated with increased expression of both SNCA mRNA and protein, and correlates with a fulminant disease course. Furthermore, it is postulated that even subtle changes in SNCA expression caused by common variation is associated with disease risk. Of note, genome-wide association studies have identified over 30 loci associated with PD with most signals located in non-coding regions of the genome, thus likely influencing transcript expression levels. In health, epigenetic mechanisms tightly regulate gene expression, turning genes on and off to balance homeostasis and this, in part, explains why two cells with the same DNA sequence will have different RNA expression profiles. Understanding this phenomenon will be crucial to our interpretation of the selective vulnerability observed in neurodegeneration and specifically dopaminergic neurons in the PD brain. In this review, we discuss epigenetic mechanisms, such as DNA methylation and histone modifications, involved in regulating the expression of genes relevant to PD, RNA-based mechanisms, as well as the effect of toxins and potential epigenetic-based treatments for PD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As the second-most common neurodegenerative disorder after Alzheimer’s disease, Parkinson’s disease (PD) presents with motor symptoms, including tremor, bradykinesia, rigidity, postural instability, and non-motor symptoms, such as REM sleep behavior disorder, autonomic dysfunction, and cognitive impairment [45]. Cytoplasmic aggregates of the α-synuclein protein, termed Lewy bodies, form within neurons, which in combination with the loss of dopaminergic neurons in the substantia nigra pars compacta [20], represent the hallmarks of the disease.
PD heritability, from twin and familial studies, is estimated to be around 34–60 % [33, 120], yet the proportion of phenotypic variance based on known PD genetic loci is approximately 27 % [52] suggesting most of the causal variation is still unknown. The bulk of this missing heritability probably lies within the understudied variation, such as rare variants, structural variants, and variants located in regulatory non-coding regions (e.g., long non-coding RNAs and microRNAs). Furthermore, it is becoming clear that the link between DNA variation and phenotype is not necessarily direct: in general, all somatic cells of an organism have the same DNA, and yet, they can present with very diverse expression profiles, which suggests an additional level of control. An understanding of the precise PD gene expression patterns and regulatory mechanisms is essential to fully decipher the sequence of molecular events leading to PD.
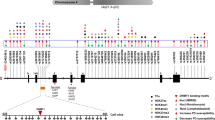
Mounting evidence supports the hypothesis that mechanisms which drive changes in gene expression and protein levels are associated with PD. One such phenomenon is epigenetic modification, which alters the functionality of a locus or chromosome without changing the underlying DNA sequence [32], and may account for some of the fundamental differences in expression patterns between PD patients and controls. Genomic DNA variation is now well-established to drive both familial and sporadic forms of PD. SNCA, which encodes the α-synuclein protein, was identified as the first PD risk gene in 1997 [88], and mutations in six additional genes (PARK2, PINK1, PARK7, LRRK2, VPS35, and CHCHD2) have been linked to familial forms of PD. In addition, genome-wide association studies (GWAS) have identified more than 30 loci associated with the modulation of PD risk at the population level (Fig. 1) [79]. Of note, most of the GWAS hits appear to be driven by non-coding variation and are thus likely associated with the regulation of gene expression [79].
Network of Parkinson’s disease (PD) related genes, and microRNAs that regulate them. Human genome Circos plot [59] showing PD-related genes and regulatory microRNAs. PD-related genes are displayed in the outer part of the plot, and microRNAs are displayed in the inner part of the plot. Green PD genes that have been found using GWAS studies; orange PD genes that cause familial PD; purple PD genes that have been found in GWAS studies and that also cause familial forms of PD. Solid grey lines show direct repression of targeted genes by microRNAs; dotted grey lines represent indirect repression by acting on an intermediate mRNA that causes downstream PD gene repression; dotted red lines represent indirect increase of SNCA-related brain inflammatory response
Differences in gene expression and protein levels of the SNCA gene have been shown to play a critical role in PD susceptibility, as demonstrated in families with SNCA triplications and duplications. SNCA gene/protein levels correlate with the number of extra genomic copies found in PD patients carrying duplications or triplications [73, 78]. The carriers present with widespread cortical Lewy body pathology [56], and SNCA dosage correlates with earlier onset, faster progression, and more severe disease presentation [95]. In addition, even subtle changes in expression due to common SNCA variants can influence disease susceptibility. Indeed, the SNCA locus is a major GWAS peak of association with PD risk. Specific variation including a dinucleotide repeat sequence REP1 within the SNCA promoter, rs356168 in intron 4, and rs356210 in the 3′ UTR region have all been proposed to influence both SNCA expression and disease risk [28, 66, 103].
Gene expression and PD
Changes in gene expression and protein levels are the main consequence of epigenetic mechanisms. Fundamental differences in expression patterns between PD patients and controls have been highlighted by expression arrays and RNA-sequencing. It is also known that within PD patients, gene expression levels are different between different brain regions (i.e., substantia nigra vs less affected brain regions) [64]. One of the caveats of expression studies in PD is that several different platforms and technologies have been used resulting in variability across studies; no specific expression pattern has consistently been replicated yet.
Gene expression can be modulated via a number of ways and can provide mechanistic insight into disease pathogenesis. Recently, the combination of RNA-sequencing, transcriptomics, and mass spectrometry proteomics data from prefrontal cortex allowed Dumitriu et al. to suggest that the main pathways involved in PD development are related to mitochondrial processes, protein folding mechanisms, and metallothioneins [22]. Moreover, the study identified ten genes overlapping with GWAS loci including SNCA which was strongly implicated in these pathways via proteomics. Hoss et al. proposed to differentiate PD brains from control brains by analyzing expression levels of a set of 125 microRNAs (miRNA) in prefrontal cortex [41]. The study used a subset of 29 miRNAs to differentiate PD brains from control brains at a genome-wide level of significance with 93.9 % specificity and 96.6 % sensitivity. The same study highlighted the differences between PD with dementia brains and PD without dementia brains using a subset of 36 miRNAs. These results strongly support a direct link between gene and protein expression and disease state.
In this review, we will present epigenetic regulatory mechanisms in the context of PD (“Appendix 1”), specifically events affecting the expression of SNCA and other PD genes. We will discuss: DNA methylation, histone modifications, and RNA-based mechanisms as well as the effects of neurotoxins and drugs on epigenetic mechanisms.
Epigenetics and the genome: DNA methylation
DNA methylation is the modification by which a methyl group is added to cytosine converting it to 5-methylcytosine. This process typically occurs on cytosines adjacent to guanines (the so-called CpG islands) and is crucial for the regulation of many cellular processes, such as gene expression, cellular differentiation, and development [89]. DNA methylation is analogous to an on/off switch: heavily methylated areas of the genome are usually less active at the transcriptional level (gene expression turned off), whereas areas with less methylation are more active (gene expression turned on). DNA methylation has been extensively studied in cancer research and new research focused on identifying differentially methylated genes and the process in which methylation is maintained or lost has recently become a major area of interest in neurodegenerative diseases research. In this section, we discuss the role of DNA methylation in PD, and we start with the methylation of the SNCA gene and its key regulatory enzyme DNA methyltransferase 1 (Dnmt1). We present the findings of large-scale methylation studies in PD, and we describe the issues with using RNA extracted from blood leukocytes. We then describe the use of S-adenosyl methionine (SAM)/S-adenosyl homocysteine (SAH) ratios as a biomarker and the epigenetic clock, a tool created by Horvath et al., to measure the age of tissues based on methylation markers. We end by discussing the methylation of mRNA and mitochondrial DNA.
Given the known influence of α-synuclein expression levels in disease, the initial PD methylation studies focused on the SNCA gene. In 2010, two groups studied the DNA methylation of SNCA using bisulfide treated cell models [49, 71]. Both groups observed the increased expression of SNCA when a CpG island in intron 1 was de-methylated. Lower methylation levels in several brain regions (substantia nigra, putamen, and cortex) in PD patients compared to controls were also observed. Interestingly, the methylation of the SNCA promoter polymorphism in PD patients was found to be correlated with the amount of L-dopa administered to the patient: patients receiving higher therapy dose have higher methylation levels and thus lower SNCA expression [99].
The Dnmt1 enzyme, one of the key regulators of DNA methylation in mammalian cells that preserves the methylation patterns established in development, has been recently examined in the context of PD. Desplats et al., of the University of California San Diego (UCSD), showed that α-synuclein interacts with Dmnt1, thereby keeping it in the cytoplasm and preventing its action on DNA. Dmnt1 levels are also lower in PD patients compared to controls, and Dmnt1 mislocalization directly alters the methylation of the SNCA gene [19]. Epigenetic regulatory mechanisms surrounding the SNCA gene and the α-synuclein protein are presented in Fig. 2.
Epigenetic mechanisms involved in α-synuclein pathology. a Nucleus. Methylation of histones promotes histone compression and the formation of condensed heterochromatin. Conversely, acetylation of histones decreases their affinity for DNA allowing nucleosome spacing and heterochromatin to transform into its relaxed form, euchromatin, which is conducive to the transcription of genes such as SNCA. Histone deacetylases (HDAC) remove acetyl group from histone and, as a result, repress gene expression. When DNA methylation occurs at the CpG island in the first intron of SNCA, transcription is also repressed. The enzyme Dnmt1 is involved in maintaining methylation patterns. When intron 1 is de-methylated, transcription of SNCA can proceed. SNCA demethylation can be due to Dnmt1 being sequestered in the cytosplasm by α-synuclein. Histone methylation–acetylation status dynamically changes by complex mechanisms that promote or repress gene transcription depending on cellular conditions and stress. Primary miRNAs are processed by the microprocessor complex, which consists of Drosha and DGCR8. Resulting precursor miRNAs (pre-miR) are transported to the cytoplasm by XPO5. b Cytoplasm. SNCA mRNA is translated in the cytoplasm into α-synuclein. In their pathological form α-synuclein monomers are assembled in oligomers and fibrils rich in β-sheets; such fibrils form the basis of the mature Lewy bodies. Pre-miRs are processed by the DICER/TRBP complex into 22 bp-miRNA duplexes. The functional strand of a miRNA duplex is incorporated to the AGO2/GW182 complex to generate mature miRISC. MiRISCs containing either miR-7 or miR-153 can bind to the 3′UTR of SNCA mRNA, destabilize the mRNA and induce translational repression. AAA poly(A) tail; Ac acetylated residue; AGO2 argonaute-2; DICER endoribonuclease Dicer or helicase with RNase motif; Dnmt1 DNA (Cytosine-5-)-Methyltransferase 1; HDAC histone deacetylases; GW182 trinucleotide repeat-containing gene 6A (TNRC6A); Me methylated residue; miR microRNA; miRISC miRNA-mediated silencing complex; mRNA messenger RNA; pre-miR precursor microRNA; pri-miR primary microRNA; SNCA Synuclein, Alpha; TRBP transactivation-responsive binding protein; UTR untranslated region; XPO5 exportin 5
In 2013, a genome-wide epigenome study (EWAS) in frontal cortex and in blood of five PD patients and six controls was reported [70]. Less than 1 % of probes on the microarray presented significant differences in methylation levels in the cases vs controls, but more than 80 % of differentially methylated sites identified were hypomethylated in PD cases. Similar patterns were observed in brains and blood. Among the top hits of PD, hypermethylated genes was MAPT, which encodes protein tau, and is also one of the top PD GWAS hits [79]. Coupland et al. studied MAPT methylation in bisulfite-treated leukocytes (358 PD patients and 1084 controls) and brain tissues (3 brain regions from 28 PD patients and 12 controls) [15]. In tissue from healthy controls, they observed higher methylation in women compared to men and 1.5-fold higher levels of methylation in MAPT H1/H1 diplotype vs H2/H2; higher methylation correlated with lower expression levels of MAPT. In PD samples, lower methylation was detected in the putamen of PD patients compared to controls. Being female, having a later age at onset and carrying the MAPT H1 haplotype were all associated with higher methylation levels. It is not clear why the MAPT H1 risk haplotype is more methylated than the protective H2 haplotype, since previous studies have shown that H1 leads to increased MAPT expression [2, 61], but it is possible that other epigenetic mechanisms regulate MAPT gene expression.
Nalls et al. examined CpG methylation related to GWAS hits in cortex and cerebellar tissue samples from neurologically normal individuals [79], and identified 30 significant associations between GWAS variants and CpG methylation or mRNA expression across six loci. Interestingly, while no differential methylation pattern was observed in MAPT itself, four of the associations observed implicated genes at the chromosome 17q MAPT locus: long non-coding RNA MGC57346 and genes PLEKHM1, ARL17A, and KANSL1. These findings suggest that tag SNPs from GWAS hits can be related to non-coding functional variants located in regulatory regions and influencing gene expression. In addition, these findings suggest that functional studies are necessary complement of GWAS studies to adequately nominate causal genes within GWAS loci.
In the last few years, EWAS have become more and more popular to study methylation profiles in PD. Since the first reported study by Masliah et al. [70], sample size has increased exponentially. Moore and colleagues [77] were particularly interested in methylation patterns specific to anxiety in PD patients. Blood samples from 45 individuals (15 PD, 15 PD with anxiety, and 15 controls) were analyzed and led to the identification of more than 12,000 genes with differential methylation patterns between PD with and without anxiety. These genes are involved in brain-centric pathways, such as neuroactive ligand-receptor interaction and the neurotrophin signaling, among others. More than 9900 differentially methylated genes in PD vs controls were identified. A subset of the top hits was followed up in 219 PD subjects and 223 controls, and two genes, FANCC and TNKS2, which are, respectively, involved in neuronal apoptosis and post-translational signaling, showed significant differential methylation.
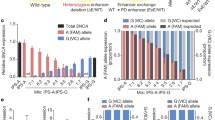
To determine whether methylation patterns in leukocytes reflect methylation patterns in the brain, the use of blood leukocytes as a proxy has been popular (because the tissues are more readily accessible) but controversial. Some studies have not been able to observe significant hypomethylation at the well-characterized SNCA intron 1 CpG island in blood leukocytes DNA from PD patients [93, 105], while others have [87, 107]. These differences might be attributable in part to sample size, but Fernandez-Santiago et al. have shown differential methylation profiles in different cell types from PD patients [25]. They derived dopaminergic neurons from induced pluripotent stem cells (iPSCs) of PD patients and healthy controls. They showed that dopaminergic neurons presented with differential methylation patterns in PD and controls, but these changes were not seen in fibroblasts or the parental iPSC. This suggests studies that aim to decipher how epigenetic regulation affects neurons should be performed in carefully chosen neuronal models or brain tissue.
Methylation has been proposed as a biomarker for cognitive impairment in PD by Obeil et al. [81]. Given S-adenosyl methionine (SAM) is a co-substrate involved in methyl group transfer and S-adenosyl homocysteine (SAH) is a product of the methylation reaction involving SAM, a higher SAM/SAH ratio is a sign of higher methylation potential. Obeid et al. discovered an association between plasma SAM/SAH ratio and cognitive function scores [81]. PD patients with no cognitive impairment had a significantly higher plasma SAM/SAH ratio than patients presenting with mild or severe cognitive dysfunction. Better cognitive functions were also related to higher levels of vitamin B6, a vitamin known to improve methylation status and protect against the production of amyloid-β [29, 34].
Horvath [39] created an epigenetic clock which predicts age based on the methylation status at 353 CpG sites in the human genome. The clock was developed using publicly available methylation array data from 7844 non-cancer samples from several different studies. Horvath named age acceleration the difference between chronological age and epigenetic age predicted by the epigenetic clock. By analyzing blood samples for PD patient and controls, Horvath et al. were able to show that there was an increased age acceleration in PD patients compared to controls [40]. They suggest that this accelerated aging of blood cells precede the onset of motor and non-motor symptoms, and could be used as a biomarker.
Methylation mechanisms affect not only DNA, but also mRNA [72, 117]. N6-methyladenosine (m6A) and 5-methylcytosine (m5C) are two mRNA methylation post-transcriptional modifications. Adenosine methylation occurs mainly in brain tissue, and it has been suggested that m6A methylation could play a role in the intracellular response to neuronal signaling by downregulating the expression of neuronal mRNAs [97]. One of the m6A demethylases is encoded by the FTO (fat mass and obesity-associated) gene. Mutations in FTO are involved in neurodegenerative disorders, such as AD [51, 92] and in impaired brain function [92]. In addition, the inactivation of FTO in a mouse model affects dopamine receptor type 2 (D2R) and dopamine receptor type 3 (D3R)-dependent control of dopaminergic neuron activity [37]. This dysfunction was caused by increased m6A levels of a specific subset of mRNAs (KCNJ6, GRIN1, and DRD3) directly involved in the dopaminergic signaling pathway, resulting in low protein expression [37].
In addition, a recent study showed that PD patients (n = 10) have a loss of m5C levels in the displacement loop (D-loop) of the substantia nigra mitochondrial DNA when compared with controls (n = 10) [4]. This could be attributable to dopaminergic neuronal loss in the substantia nigra, but methylation levels of mitochondrial NADH dehydrogenase 6 (ND6) in this region were similar between PD patients and controls, suggesting that the m5C methylation changes in the mitochondrial D-loop are not due to neuronal loss [4]. Interestingly, nuclear DNA is not the only type showing differential methylation patterns. Blanch et al. reported loss of mitochondrial DNA 5-methylation levels in the substantia nigra of PD patients [4]. These findings could support the hypothesis that mitochondrial dysfunction is a common molecular mechanism in PD pathogenesis.
DNA methylation is the most intensely studied epigenetic mechanism, based on the number of papers available in Pubmed, yet the role it plays in PD pathogenesis is just starting to be explored. Methylation correlates with aging of tissues and cognitive impairment and directly influences the expression of SNCA and other PD genes. Furthermore, the influence of methylation on the regulation of gene expression can only be understood by taking into account the other epigenetic mechanisms at play.
Chromatin remodeling
Chromatin remodeling, orchestrated by histone modifications, is a dynamic process by which important physiological functions, such as gene expression, are regulated. Post-translational modifications of histone proteins, such as methylation and acetylation, can modify chromatin structure and regulate accessibility of DNA for transcription. Histone acetylation is associated with transcriptionally active genes. In general, there are higher levels of histone acetylation in the midbrain of dopaminergic neurons from PD patients compared to controls [84]. Specific PD genes and molecules are regulated by histone modifications. The case of α-synuclein is interesting, because it is regulated by histone acetylation and it can also regulate histone acetylation, possibly due to a feedback loop. It was shown that α-synuclein interacts with histones in the nucleus and this accelerates α-synuclein fibrillation and toxicity [31]. Kontopoulos et al. confirmed these findings and further demonstrated that α-synuclein binds directly to histones and inhibits the acetylation of histone H3 through interaction with SIRT2, a deacetylase [57]. Furthermore, they showed that the α-synuclein disease-related mutants p.A30P and p.A53T present an increased nuclear localization compared with wild-type α-synuclein. Voutsinas et al. looked at SNCA gene expression in a PD patient heterozygous for the p.A53T mutation. They observed that the p.A53T allele was silenced and it could be reactivated through the use of inhibitors of histone deacetylases (HDACs), demonstrating that the silencing of the mutant SNCA allele is due to histone deacetylation [114]. Supporting these findings, a histone 3 lysine 27 acetylation (H3K27ac)-enriched enhancer sequence has been identified at the SNCA locus [113]. Interestingly, HDACs inhibitors have also been shown to be neuroprotective against α-synuclein -mediated toxicity [57, 76, 83].
Other PD-associated genes also seem to be regulated by histones modifications. MAPT haplotype H1 is preferentially associated with H3K4me3 histone modification, which normally indicates gene activation, whereas the H2 haplotype is associated with the repressive H3K27me3 histone modification [90]. The PINK1 protein can bind to HDAC3, a transcriptional repressor recruited to specific promoters, and upregulate its histone deacetylase activity through phosphorylation in neuronal cell lines [13]. This event leads to increased binding of phosphorylated HDAC3 to p53, which decreases p53 acetylation and stability, thus inhibiting p53-mediated neuronal apoptosis. The knockdown of HDAC3 abolishes the effect of PINK1 on p53 [13]. Known PD-associated variants in PINK1 do not promote phosphorylation of HDAC3 suggesting PINK1 mutations lead to a deregulation of HDAC3 activity and increased susceptibility to p53-dependent neuronal apoptosis and neurodegeneration [13].
RNA-based mechanisms of gene regulation
A number of RNA-based regulatory pathways based on non-coding RNAs and miRNAs influence gene expression. In this section, we will discuss specific examples of long non-coding RNAs (lncRNA) and microRNAs (miRs) which regulate the expression of PD genes. LncRNA are long (more than 200 nucleotides) non-protein coding transcripts, which are involved in gene expression regulation [118]. LncRNA RP11-115D19.1 binds to SNCA 3′-flanking region and can be stimulated prominently by YY1, a ubiquitous transcription factor involved in several biological pathways. This stimulation does not significantly modify SNCA expression levels in an SH-SY5Y cell model [75]; however, the knockdown of RP11-115D19.1 in the same cell model results in a significant increase of α-synuclein expression. This suggests that this lncRNA has a repressive effect on SNCA expression [75]. Another study using four different neuroblastoma cell lines revealed a lncRNA complementary to PINK1 3′-mRNA sequence, which was able to stabilize and increase PINK1 expression levels, which may prove beneficial [98].
MicroRNAs (miRs) are small non-coding nucleotide RNA molecules (21-25 nucleotides) that bind to imperfect complementary sequences within the 3′UTR region of their target messenger RNAs (mRNAs) [6, 43]. This binding inhibits the expression of proteins encoded by these mRNAs [111]. Several studies show that miRs are crucial in the regulation of PD-related gene expression (Fig. 1) and that they affect specific neuronal functions. MiR-133b is highly expressed in midbrain dopaminergic neurons of healthy subjects, but has lower levels in brains from patients with PD [55]. MiR-133b is normally expressed in mice midbrain and levels have been shown to be reduced in two different PD mice models: an adult Aphakia mouse model, which has a reduced expression of transcription factor Pitx3, and the classical 6-OHDA mouse model. Pitx3 promotes midbrain dopaminergic neuron gene expression [69, 102] and induces miR-133b transcription, while miR-133b overexpression reduces Pitx3 protein levels. These results suggest a negative-feedback loop in which Pitx3 induces miR-133b transcription and miR-133b overexpression downregulates Pitx3 activity [55].
It has been shown that miR-7 and miR-153, which bind specifically to SNCA, downregulate its expression and that their effect is additive (Fig. 2) [21]. The brain expression patterns of miR-7 and miR-153 mimic that of SNCA in adult brains as well as neuronal development, suggesting a tight balance in the regulatory mechanisms. Overexpression of either or both miRs reduces endogenous SNCA levels, whereas their inhibition promotes the translation of a chimeric luciferase mRNA bearing SNCA 3′UTR region in primary neurons [21]. MiR-7 is highly conserved across species, and it has been suggested to confer stability and robustness to certain regulatory networks against environmental fluctuation during development [38, 65], and to protect neurons against oxidative stress [50]. Moreover, a recent study suggests that miR-7 and miR-153 may protect neurons against neurotoxin 1-Methyl-4-Phenyl-Pyridinium (MPP+)-induced cell death by upregulating mTOR and SAPK/JNK signaling pathways [27]. Both pathways have been involved in apoptotic cell death mechanisms in PD [94]. In addition, miR-7 protects neurons against (MPP+) cell damage by targeting RelA [12], a structural component of inflammatory transcription factor NF-κB. These studies support miR-7 and miR-153 as potential therapeutic targets for PD and several other α-synucleinopathies.
A recent study showed that miR-155 was upregulated early in an in vivo PD model overexpressing α-synuclein [108]. A miR-155 knockout mouse model revealed that the absence of miR-155 reduced neuronal proinflammatory responses to α-synuclein and blocked α-synuclein-related neurodegeneration. The underlying mechanisms of this blockage involved the microglia’s response to α-synuclein aggregation. Primary microglia from miR-155 knockout mice presented with reduced inflammatory response to α-synuclein fibrils, including a diminished major histocompatibility complex class II and nitric oxide synthase expression. The treatment of this microglia with a synthetic miR-155 replicated their inflammatory response to α-synuclein fibrils [108]. These results suggest that miR-155 plays a key role in brain inflammatory response and in α-synuclein-dependent neurodegeneration and implies that miR-155 could be used as therapeutic target to modulate brain inflammatory responses to α-synuclein in patients with PD. In addition, it has been suggested that variant rs12720208 located in the miR-433 binding site of fibroblast growth factor 20 gene (FGF20), located in a GWAS locus, increases PD risk by upregulating the expression of α-synuclein [115], but this mechanism remains controversial [17, 119].
MiR-205 regulates the expression of LRRK2, a familial PD gene, and GWAS hit, where the most frequent causal mutation in sporadic and familial PD is located (LRRK2 p.G2019S). A study found that expression of the LRRK2 protein was increased in frontal cortex of sporadic PD patients when compared to controls, while mRNA levels were not significantly different between both groups, suggesting post-transcriptional modifications of LRRK2 protein expression [11]. The same study showed that miR-205, which is conserved across different vertebrate species, binds specifically to the 3′UTR LRRK2 mRNA region and that this binding suppressed LRRK2 protein expression. MiR-205 was also under-expressed in sporadic PD subjects with an increased LRRK2 protein expression compared to controls. These results were supported by experiments which showed that overexpression of miR-205 in cultured mouse hippocampal neurons expressing PD-causing LRRK2 p.R1441G mutation prevented neurite outgrowth defects suggesting miR-205 has a protective effect against neurodegeneration [11].
Proteins encoded by PARK2 (Parkin RBR E3 Ubiquitin Protein Ligase) and PARK7 (DJ1) genes are implicated in the disruption of the mitochondrial pathway in PD patients [1]. Their expression is tightly regulated by miR-34b and miR-34c. The expression of miR-34b and miR-34c is decreased in several brain areas (amygdala, frontal cortex, substantia nigra, and cerebellum) in PD patients with Braak stages 4 and 5 but also in PD patients in pre-motor stages (Braak stages 1–3) compared to controls, suggesting that this is an early event in PD pathogenesis. The decreased expression of miR-34b and miR-34c correlates with a reduced DJ1 and Parkin expression in human PD brains. However, overexpression of these miRs does not lead to any change in DJ1 or Parkin expression levels [74], suggesting that they are not direct targets of miR-34b and miR-34c, but that the miRs are involved in an intermediate regulatory step. Additional experiments in differentiated SH-SY5Y dopaminergic neuronal cells showed that a depletion of miR-34b or miR-34c resulted in a reduction in DJ1 and Parkin expression levels and cell viability along with mitochondrial dysfunction altered oxidative stress and a decrease of total cellular adenosine triphosphate [74]. Interestingly, miR-34b and miR-34c share a common non-coding precursor and they both play a key role in apoptosis pathways, as they are direct targets of the tumor suppressor gene P53 [36].
There is evidence that miRs can modulate tau expression. In primary hippocampal neurons extracted from an Alzheimer’s disease (AD) rat model, overexpression of miR-125b increased tau hyperphosphorylation and upregulated p35, cdk5, and p44/42-MAPK signaling [3]. Overexpression of miR-125b downregulated its direct targets, DUSP6 and PPP1CA phosphatases, and the anti-apoptotic factor Bcl-W. When overexpressed, PPP1CA and Ccl-W prevented miR-125-induced tau phosphorylation, suggesting that they intervene in the action of miR-125b on tau. Likewise, miR-125b suppression in neurons reduced tau phosphorylation and its kinase activity [3]. In addition, miR-125b hippocampal injection in mice leads to tau phosphorylation and produces learning impairment [3]. Similarly, miR-138 induces tau phosphorylation by targeting the retinoic acid receptor alpha (RARA)/glycogen synthase kinase-3β (GSK-3β) pathway [116]. A study by Wang et al. showed that miR-138 was increased in AD cell models [116]. MiR-138 overexpression activates GSK-3β and increases tau phosphorylation in HEK293/tau cells. It was also shown that 3′-UTR of RARA mRNA is a direct target of miR-138 and that overexpression of RARA attenuates GSK-3β activity and reduces tau phosphorylation induced by miR-138 [116].
Finally, a recent study analyzing variants in miRs or miR-binding sites on their target genes based on the largest PD GWAS [79], showed a significant association of two miR variants (rs897984 [chr16:30886643T>C] in miR-4519 and rs11651671 [chr17:40646803A>G] in miR-548at-5p) with PD [30]. In both cases, the variant’s alternative allele modified the secondary structure of the miR and decreased their expression level. However, only miR-4519 expression was detectable in brain tissue. An analysis of genetic variants in all putative miR-4519 target genes revealed a significant association of four genes with PD: NSF, TMEM163, CCNT2, and SH3GL2 [30]. Interestingly, the NSF gene lies in the PD-associated MAPT locus. In addition, 13 miR-binding sites located in ten known PD genes (RAB29 formerly known as RAB7L1, DGKQ, TMEM175, SNCA, STX1B, PLEKHM1, ARHGAP27, CRHR1, MAPT, and SPPL2B) and three other genes (CSTB, IGSF9B, and HSD3B7) showed a significant association with PD [30].
RNA editing
RNA-based epigenetics mechanisms can be disrupted by RNA editing, a process that is catalyzed by ADAR (adenosine deaminase acting on RNA) enzymes. These proteins convert adenosines (A) located in intronic regions, 5′ and 3′ UTR regions of mRNAs into an inosine (I) [101]. The cell machinery recognizes I as guanosine (G), because they have the same pairing properties. This process is similar to silent mutations at the mRNA level. While not affecting the nature of the mRNA, these changes can disrupt the binding of the miRs to the mRNA. Although the process is common and necessary for normal life and development [44, 112], in certain circumstances, it can disrupt the normal epigenetic regulation performed by non-coding RNA machinery. It has been described in several neurodegenerative diseases, such as amyotrophic lateral sclerosis [60], but so far, not in PD.
Taken together, these studies exemplify how small or long non-coding RNAs can be crucial to finely regulate gene and protein expression in both healthy and PD affected subjects. Moreover, these mechanisms hint to a new era in therapeutic tools development, where the focus is on managing protein aggregation and PD-causing dysfunctional pathways.
Neurotoxins and drugs interfere with epigenetic mechanisms in PD
It is widely accepted that certain environmental factors, such as pesticides, can induce parkinsonian symptoms. In fact, certain neurotoxins have been used to generate animal models that mimic parkinsonian symptoms of PD patients. The first animal models developed using 6-OHDA [110], methamphetamine [109], or rotenone [35] have certain features similar to the loss of dopaminergic neurons observed in PD patients, but they do not replicate all the pathogenic mechanisms present in PD. For that reason, the discovery of the selective nigral toxicity of 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) driven by its MPP+ metabolite was a breakthrough. The MPTP model [5] mimics several PD clinical and pathogenic mechanisms: oxidative stress, reactive oxygen species, energy failure, and inflammation. However, only a few studies using the MPTP model have been able to replicate the definitive pathological PD hallmark: the development of Lewy bodies [26, 58].
Several neurotoxins not only produce a PD phenotype, but also affect different epigenetic mechanisms, such as methylation, demethylation, hyperacetylation, and deacetylation of certain genomic regions (Table 1). In addition, some histone acetyltransferase inhibitors and histone deacetylases inhibitors may counteract the effects of toxin- or disease-related epigenetic changes and are potential therapeutic avenues to explore (Table 1).
Harnessing epigenetics mechanism for therapeutics
Epigenetic-based therapies are attractive strategies to treat neurodegenerative disorders. The key to their usefulness is specificity. Ideally, a desired outcome would be downregulation of a specific pathogenic gene-like SNCA or upregulation of gatekeeper genes or both. As mentioned previously, several molecules can affect DNA methylation, but their effects are highly unspecific which probably would lead to considerable adverse effects. Interestingly, l-dopa appears to increase SNCA methylation in vitro and in vivo [99]. However, its long-term use induces dyskinesia, which appears to be correlated with increased H4 deacetylation [80]. Similar to molecules involved in methylation, the effects of HDACs inhibitors are wide ranging. We highlight some of the outcomes and rescue mechanisms related to HDACs inhibitors, such as Vorinostat (SAHA) and sodium butyrate (Table 1). All the proposed HDACs related compounds are still in experimental stages in terms of their effect on PD.
The power to specifically downregulate or knockdown genes of interest makes RNA molecules particularly interesting therapeutic strategies. Small interfering RNAs (siRNA) are short (20–25 bp) double-stranded RNA molecules complementary to a specific mRNA region. Unstable in their native form, siRNA are often coupled to a vector for transfection into cells. Nonviral vectors specific for neuronal cells can cross the blood–brain barrier rapidly and deliver SNCA-specific siRNAs to neurons knocking down α-synuclein protein expression, thereby preventing PD-like symptoms in in vitro and in vivo experimental models [46, 63]. Pre-treatment of M17 cells with a vector/siRNA complex in in vitro models resulted in a greater survival rate when exposed to MPP+ toxin, than untreated M17 cells [46]. In addition, in MPTP mouse models, the release of these vector/siRNAs complexes improved the motor symptoms generated by the MPP+ toxin via downregulation of SNCA [46]. The inhibition SIRT2, a deacetylase, can also rescue α-synuclein toxicity in a cellular model of PD [83]. In fact, when transfecting human neuroglioma cells (H4) with SNCA and synthetic siRNA against SIRT2 or SIRT3, only those transfected with SIRT2 siRNA were rescued from α-synuclein toxicity [83].
Mirtrons are a specific class of miRNAs that are encoded in the introns of genes. They have been shown to silence certain genes in both in vitro and in vivo PD models [18]. Deng et al. created artificial mirtrons based on miR-1224 to target LRRK2 and SNCA. They achieved an 85 % LRRK2 gene expression reduction in HEK293 cells cotransfected with artificial mirtrons and exogenous LRRK2, but the reduction of endogenous LRRK2 was only 36 % in SH-SY5Y cells. A similar assay targeting SNCA could only achieve a 26 % reduction in its expression [18], suggesting that artificial mirtrons may have therapeutic uses but optimization is needed.
The off-target effects of a miR could seriously damage normal cell machinery. For RNA-based epigenetic treatments to be successful, specific neurons, as opposed to every cell in a tissue have to be targeted. For this purpose, the chosen delivery system is of utmost importance. MiRs have to: (1) cross the blood–brain barrier, (2) enter the targeted neurons, and (3) remain in the brain long enough to perform their actions.
Another aspect to be considered is the potential effects of therapeutic miRs on different isoforms that could lead to undesirable results. Additional experiments need to be carried out to identify which gene isoforms need to be downregulated to improve PD symptoms and delay pathological effects and which ones should be upregulated in neurons, so that they perform their expected role to maintain normal neuronal function. Mutations in miRs target regions of PD genes could impair binding of miRs to their target mRNA, thus pre-treatment sequencing of the targeted region in patients should be performed to avoid potential therapeutic failures.
While inherently more specific than HDAC inhibitors, RNA-based epigenetic mechanisms have several caveats that need to be considered before they can be harnessed into safe and generic treatments for PD patients. Epigenetic-based cancer treatments aim to kill cells that are proliferating without control. The situation is somewhat different for neurodegenerative diseases in that the goal is to prevent cell death. The ideal treatment for PD would be to overcome the selective vulnerability of dopaminergic neurons by administering treatment early in pre-clinical phases of disease.
Future perspectives
In genetic terminology, PD is a complex disease: one caused by the interplay of several genetic and environmental factors. As such, more often than not, predicting phenotype from DNA variation alone is difficult and this assessment might have led to the previous perception that PD is not a genetic disease. In truth, a diverse array of regulatory mechanisms is acting as mediator between genotype and phenotype and contributes to differential gene expression; some of these mechanisms are epigenetic-based. We have highlighted a number of these different levels of regulatory mechanisms involved in PD-related gene expression: methylation of promoter regions, histone modifications, and RNA-based mechanisms. Some of the factors involved in these regulatory processes are potential therapeutic targets or potential biomarkers for improved and earlier diagnosis. PD-related therapy using epigenetic processes is in its infancy but will most likely continue to develop and perhaps someday become part of routine treatment. It is becoming clear that epigenetic mechanisms play key roles in neurodegenerative diseases. Additional research efforts aimed at determining the complete epigenetic map of the human brain, both healthy and diseased, is needed to fully understand the etiology of PD and other complex diseases.
References
Abou-Sleiman PM, Muqit MM, Wood NW (2006) Expanding insights of mitochondrial dysfunction in Parkinson’s disease. Nat Rev Neurosci 7:207–219. doi:10.1038/nrn1868
Allen M, Kachadoorian M, Quicksall Z, Zou F, Chai HS, Younkin C, Crook JE, Pankratz VS, Carrasquillo MM, Krishnan S et al (2014) Association of MAPT haplotypes with Alzheimer’s disease risk and MAPT brain gene expression levels. Alzheimers Res Ther 6:39. doi:10.1186/alzrt268
Banzhaf-Strathmann J, Benito E, May S, Arzberger T, Tahirovic S, Kretzschmar H, Fischer A, Edbauer D (2014) MicroRNA-125b induces tau hyperphosphorylation and cognitive deficits in Alzheimer’s disease. EMBO J 33:1667–1680. doi:10.15252/embj.201387576
Blanch M, Mosquera JL, Ansoleaga B, Ferrer I, Barrachina M (2016) Altered mitochondrial DNA methylation pattern in Alzheimer disease-related pathology and in Parkinson disease. Am J Pathol 186:385–397. doi:10.1016/j.ajpath.2015.10.004
Burns RS, Chiueh CC, Markey SP, Ebert MH, Jacobowitz DM, Kopin IJ (1983) A primate model of parkinsonism: selective destruction of dopaminergic neurons in the pars compacta of the substantia nigra by N-methyl-4-phenyl-1,2,3,6-tetrahydropyridine. Proc Natl Acad Sci USA 80:4546–4550
Cannell IG, Kong YW, Bushell M (2008) How do microRNAs regulate gene expression? Biochem Soc Trans 36:1224–1231. doi:10.1042/bst0361224
Chartier-Harlin MC, Dachsel JC, Vilarino-Guell C, Lincoln SJ, Lepretre F, Hulihan MM, Kachergus J, Milnerwood AJ, Tapia L, Song MS et al (2011) Translation initiator EIF4G1 mutations in familial Parkinson disease. Am J Hum Genet 89:398–406. doi:10.1016/j.ajhg.2011.08.009
Chen PS, Peng GS, Li G, Yang S, Wu X, Wang CC, Wilson B, Lu RB, Gean PW, Chuang DM et al (2006) Valproate protects dopaminergic neurons in midbrain neuron/glia cultures by stimulating the release of neurotrophic factors from astrocytes. Mol Psychiatry 11:1116–1125. doi:10.1038/sj.mp.4001893
Chen PS, Wang CC, Bortner CD, Peng GS, Wu X, Pang H, Lu RB, Gean PW, Chuang DM, Hong JS (2007) Valproic acid and other histone deacetylase inhibitors induce microglial apoptosis and attenuate lipopolysaccharide-induced dopaminergic neurotoxicity. Neuroscience 149:203–212. doi:10.1016/j.neuroscience.2007.06.053
Chiu S, Terpstra KJ, Bureau Y, Hou J, Raheb H, Cernvosky Z, Badmeav V, Copen J, Husni M, Woodbury-Farina M (2013) Liposomal-formulated curcumin [Lipocurc] targeting HDAC (histone deacetylase) prevents apoptosis and improves motor deficits in Park 7 (DJ-1)-knockout rat model of Parkinson’s disease: implications for epigenetics-based nanotechnology-driven drug platform. J Complement Integr Med. doi:10.1515/jcim-2013-0020
Cho HJ, Liu G, Jin SM, Parisiadou L, Xie C, Yu J, Sun L, Ma B, Ding J, Vancraenenbroeck R et al (2013) MicroRNA-205 regulates the expression of Parkinson’s disease-related leucine-rich repeat kinase 2 protein. Hum Mol Genet 22:608–620. doi:10.1093/hmg/dds470
Choi DC, Chae YJ, Kabaria S, Chaudhuri AD, Jain MR, Li H, Mouradian MM, Junn E (2014) MicroRNA-7 protects against 1-methyl-4-phenylpyridinium-induced cell death by targeting RelA. J Neurosci 34:12725–12737. doi:10.1523/JNEUROSCI.0985-14.2014
Choi HK, Choi Y, Kang H, Lim EJ, Park SY, Lee HS, Park JM, Moon J, Kim YJ, Choi I et al (2015) PINK1 positively regulates HDAC3 to suppress dopaminergic neuronal cell death. Hum Mol Genet 24:1127–1141. doi:10.1093/hmg/ddu526
Choong CJ, Sasaki T, Hayakawa H, Yasuda T, Baba K, Hirata Y, Uesato S, Mochizuki H (2016) A novel histone deacetylase 1 and 2 isoform-specific inhibitor alleviates experimental Parkinson’s disease. Neurobiol Aging 37:103–116. doi:10.1016/j.neurobiolaging.2015.10.001
Coupland KG, Mellick GD, Silburn PA, Mather K, Armstrong NJ, Sachdev PS, Brodaty H, Huang Y, Halliday GM, Hallupp M et al (2014) DNA methylation of the MAPT gene in Parkinson’s disease cohorts and modulation by vitamin E in vitro. Mov Disord 29:1606–1614. doi:10.1002/mds.25784
Dauer W, Przedborski S (2003) Parkinson’s disease: mechanisms and models. Neuron 39:889–909
de Mena L, Cardo LF, Coto E, Miar A, Diaz M, Corao AI, Alonso B, Ribacoba R, Salvador C, Menendez M et al (2010) FGF20 rs12720208 SNP and microRNA-433 variation: no association with Parkinson’s disease in Spanish patients. Neurosci Lett 479:22–25. doi:10.1016/j.neulet.2010.05.019
Deng JH, Deng P, Lin SL, Ying SY (2015) Gene silencing in vitro and in vivo using intronic microRNAs. Methods Mol Biol 1218:321–340. doi:10.1007/978-1-4939-1538-5_20
Desplats P, Spencer B, Coffee E, Patel P, Michael S, Patrick C, Adame A, Rockenstein E, Masliah E (2011) Alpha-synuclein sequesters Dnmt1 from the nucleus: a novel mechanism for epigenetic alterations in Lewy body diseases. J Biol Chem 286:9031–9037. doi:10.1074/jbc.C110.212589
Dickson DW (2012) Parkinson’s disease and parkinsonism: neuropathology. Cold Spring Harb Perspect Med. doi:10.1101/cshperspect.a009258
Doxakis E (2010) Post-transcriptional regulation of alpha-synuclein expression by mir-7 and mir-153. J Biol Chem 285:12726–12734. doi:10.1074/jbc.M109.086827
Dumitriu A, Golji J, Labadorf AT, Gao B, Beach TG, Myers RH, Longo KA, Latourelle JC (2016) Integrative analyses of proteomics and RNA transcriptomics implicate mitochondrial processes, protein folding pathways and GWAS loci in Parkinson disease. BMC Med Genomics 9:5. doi:10.1186/s12920-016-0164-y
Feng Y, Liu T, Dong SY, Guo YJ, Jankovic J, Xu H, Wu YC (2015) Rotenone affects p53 transcriptional activity and apoptosis via targeting SIRT1 and H3K9 acetylation in SH-SY5Y cells. J Neurochem 134:668–676. doi:10.1111/jnc.13172
Fernandes S, Salta S, Summavielle T (2015) Methamphetamine promotes alpha-tubulin deacetylation in endothelial cells: the protective role of acetyl-l-carnitine. Toxicol Lett 234:131–138. doi:10.1016/j.toxlet.2015.02.011
Fernandez-Santiago R, Carballo-Carbajal I, Castellano G, Torrent R, Richaud Y, Sanchez-Danes A, Vilarrasa-Blasi R, Sanchez-Pla A, Mosquera JL, Soriano J et al (2015) Aberrant epigenome in iPSC-derived dopaminergic neurons from Parkinson’s disease patients. EMBO Mol Med 7:1529–1546. doi:10.15252/emmm.201505439
Forno LS, Langston JW, DeLanney LE, Irwin I, Ricaurte GA (1986) Locus ceruleus lesions and eosinophilic inclusions in MPTP-treated monkeys. Ann Neurol 20:449–455. doi:10.1002/ana.410200403
Fragkouli A, Doxakis E (2014) miR-7 and miR-153 protect neurons against MPP(+)-induced cell death via upregulation of mTOR pathway. Front Cell Neurosci 8:182. doi:10.3389/fncel.2014.00182
Fuchs J, Tichopad A, Golub Y, Munz M, Schweitzer KJ, Wolf B, Berg D, Mueller JC, Gasser T (2008) Genetic variability in the SNCA gene influences alpha-synuclein levels in the blood and brain. Faseb J 22:1327–1334. doi:10.1096/fj.07-9348com
Fuso A, Nicolia V, Cavallaro RA, Ricceri L, D’Anselmi F, Coluccia P, Calamandrei G, Scarpa S (2008) B-vitamin deprivation induces hyperhomocysteinemia and brain S-adenosylhomocysteine, depletes brain S-adenosylmethionine, and enhances PS1 and BACE expression and amyloid-beta deposition in mice. Mol Cell Neurosci 37:731–746. doi:10.1016/j.mcn.2007.12.018
Ghanbari M, Darweesh SK, de Looper HW, van Luijn MM, Hofman A, Ikram MA, Franco OH, Erkeland SJ, Dehghan A (2015) Genetic variants in MicroRNAs and their binding sites are associated with the risk of Parkinson disease. Hum Mutat. doi:10.1002/humu.22943
Goers J, Manning-Bog AB, McCormack AL, Millett IS, Doniach S, Di Monte DA, Uversky VN, Fink AL (2003) Nuclear localization of alpha-synuclein and its interaction with histones. Biochemistry 42:8465–8471. doi:10.1021/bi0341152
Goldberg AD, Allis CD, Bernstein E (2007) Epigenetics: a landscape takes shape. Cell 128:635–638. doi:10.1016/j.cell.2007.02.006
Hamza TH, Payami H (2010) The heritability of risk and age at onset of Parkinson’s disease after accounting for known genetic risk factors. J Hum Genet 55:241–243. doi:10.1038/jhg.2010.13
Hashim A, Wang L, Juneja K, Ye Y, Zhao Y, Ming LJ (2011) Vitamin B6s inhibit oxidative stress caused by Alzheimer’s disease-related Cu(II)-beta-amyloid complexes-cooperative action of phospho-moiety. Bioorg Med Chem Lett 21:6430–6432. doi:10.1016/j.bmcl.2011.08.123
Heikkila RE, Nicklas WJ, Vyas I, Duvoisin RC (1985) Dopaminergic toxicity of rotenone and the 1-methyl-4-phenylpyridinium ion after their stereotaxic administration to rats: implication for the mechanism of 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine toxicity. Neurosci Lett 62:389–394
Hermeking H (2010) The miR-34 family in cancer and apoptosis. Cell Death Differ 17:193–199. doi:10.1038/cdd.2009.56
Hess ME, Hess S, Meyer KD, Verhagen LA, Koch L, Bronneke HS, Dietrich MO, Jordan SD, Saletore Y, Elemento O et al (2013) The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry. Nat Neurosci 16:1042–1048. doi:10.1038/nn.3449
Horsham JL, Ganda C, Kalinowski FC, Brown RA, Epis MR, Leedman PJ (2015) MicroRNA-7: a miRNA with expanding roles in development and disease. Int J Biochem Cell Biol 69:215–224. doi:10.1016/j.biocel.2015.11.001
Horvath S (2013) DNA methylation age of human tissues and cell types. Genome Biol 14:R115. doi:10.1186/gb-2013-14-10-r115
Horvath S, Ritz BR (2015) Increased epigenetic age and granulocyte counts in the blood of Parkinson’s disease patients. Aging (Albany NY) 7:1130–1142
Hoss AG, Labadorf A, Beach TG, Latourelle JC, Myers RH (2016) microRNA profiles in Parkinson’s disease prefrontal cortex. Front Aging Neurosci 8:36. doi:10.3389/fnagi.2016.00036
Huang HY, Lin SZ, Chen WF, Li KW, Kuo JS, Wang MJ (2011) Urocortin modulates dopaminergic neuronal survival via inhibition of glycogen synthase kinase-3beta and histone deacetylase. Neurobiol Aging 32:1662–1677. doi:10.1016/j.neurobiolaging.2009.09.010
Jackson RJ, Standart N (2007) How do microRNAs regulate gene expression? Sci STKE. doi:10.1126/stke.3672007re1
Jacobs MM, Fogg RL, Emeson RB, Stanwood GD (2009) ADAR1 and ADAR2 expression and editing activity during forebrain development. Dev Neurosci 31:223–237. doi:10.1159/000210185
Jankovic J (2008) Parkinson’s disease: clinical features and diagnosis. J Neurol Neurosurg Psychiatry 79:368–376. doi:10.1136/jnnp.2007.131045
Javed H, Menon SA, Al-Mansoori KM, Al-Wandi A, Majbour NK, Ardah MT, Varghese S, Vaikath NN, Haque ME, Azzouz M et al (2015) Development of nonviral vectors targeting the brain as a therapeutic approach for Parkinson’s disease and other brain disorders. Mol Ther. doi:10.1038/mt.2015.232
Jiang W, Li J, Zhang Z, Wang H, Wang Z (2014) Epigenetic upregulation of alpha-synuclein in the rats exposed to methamphetamine. Eur J Pharmacol 745:243–248. doi:10.1016/j.ejphar.2014.10.043
Johnston TH, Huot P, Damude S, Fox SH, Jones SW, Rusche JR, Brotchie JM (2013) RGFP109, a histone deacetylase inhibitor attenuates L-DOPA-induced dyskinesia in the MPTP-lesioned marmoset: a proof-of-concept study. Parkinsonism Relat Disord 19:260–264. doi:10.1016/j.parkreldis.2012.07.001
Jowaed A, Schmitt I, Kaut O, Wullner U (2010) Methylation regulates alpha-synuclein expression and is decreased in Parkinson’s disease patients’ brains. J Neurosci 30:6355–6359. doi:10.1523/JNEUROSCI.6119-09.2010
Junn E, Lee KW, Jeong BS, Chan TW, Im JY, Mouradian MM (2009) Repression of alpha-synuclein expression and toxicity by microRNA-7. Proc Natl Acad Sci USA 106:13052–13057. doi:10.1073/pnas.0906277106
Keller L, Xu W, Wang HX, Winblad B, Fratiglioni L, Graff C (2011) The obesity related gene, FTO, interacts with APOE, and is associated with Alzheimer’s disease risk: a prospective cohort study. J Alzheimer’s Dis JAD 23:461–469. doi:10.3233/JAD-2010-101068
Keller MF, Saad M, Bras J, Bettella F, Nicolaou N, Simon-Sanchez J, Mittag F, Buchel F, Sharma M, Gibbs JR et al (2012) Using genome-wide complex trait analysis to quantify ‘missing heritability’ in Parkinson’s disease. Hum Mol Genet 21:4996–5009. doi:10.1093/hmg/dds335
Kidd SK, Schneider JS (2010) Protection of dopaminergic cells from MPP + -mediated toxicity by histone deacetylase inhibition. Brain Res 1354:172–178. doi:10.1016/j.brainres.2010.07.041
Kidd SK, Schneider JS (2011) Protective effects of valproic acid on the nigrostriatal dopamine system in a 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine mouse model of Parkinson’s disease. Neuroscience 194:189–194. doi:10.1016/j.neuroscience.2011.08.010
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E, Hannon G, Abeliovich A (2007) A MicroRNA feedback circuit in midbrain dopamine neurons. Science 317:1220–1224. doi:10.1126/science.1140481
Konno T, Ross OA, Puschmann A, Dickson DW, Wszolek ZK (2016) Autosomal dominant Parkinson’s disease caused by SNCA duplications. Parkinsonism Relat Disord 22(Suppl 1):S1–S6. doi:10.1016/j.parkreldis.2015.09.007
Kontopoulos E, Parvin JD, Feany MB (2006) Alpha-synuclein acts in the nucleus to inhibit histone acetylation and promote neurotoxicity. Hum Mol Genet 15:3012–3023. doi:10.1093/hmg/ddl243
Kowall NW, Hantraye P, Brouillet E, Beal MF, McKee AC, Ferrante RJ (2000) MPTP induces alpha-synuclein aggregation in the substantia nigra of baboons. NeuroReport 11:211–213
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. doi:10.1101/gr.092759.109
Kwak S, Kawahara Y (2005) Deficient RNA editing of GluR2 and neuronal death in amyotropic lateral sclerosis. J Mol Med (Berl) 83:110–120. doi:10.1007/s00109-004-0599-z
Kwok JB, Teber ET, Loy C, Hallupp M, Nicholson G, Mellick GD, Buchanan DD, Silburn PA, Schofield PR (2004) Tau haplotypes regulate transcription and are associated with Parkinson’s disease. Ann Neurol 55:329–334. doi:10.1002/ana.10826
Leng Y, Marinova Z, Reis-Fernandes MA, Nau H, Chuang DM (2010) Potent neuroprotective effects of novel structural derivatives of valproic acid: potential roles of HDAC inhibition and HSP70 induction. Neurosci Lett 476:127–132. doi:10.1016/j.neulet.2010.04.013
Lewis J, Melrose H, Bumcrot D, Hope A, Zehr C, Lincoln S, Braithwaite A, He Z, Ogholikhan S, Hinkle K et al (2008) In vivo silencing of alpha-synuclein using naked siRNA. Mol Neurodegener 3:19. doi:10.1186/1750-1326-3-19
Lewis PA, Cookson MR (2012) Gene expression in the Parkinson’s disease brain. Brain Res Bull 88:302–312. doi:10.1016/j.brainresbull.2011.11.016
Li X, Cassidy JJ, Reinke CA, Fischboeck S, Carthew RW (2009) A microRNA imparts robustness against environmental fluctuation during development. Cell 137:273–282. doi:10.1016/j.cell.2009.01.058
Maraganore DM, de Andrade M, Elbaz A, Farrer MJ, Ioannidis JP, Kruger R, Rocca WA, Schneider NK, Lesnick TG, Lincoln SJ et al (2006) Collaborative analysis of alpha-synuclein gene promoter variability and Parkinson disease. JAMA 296:661–670. doi:10.1001/jama.296.6.661
Marinova Z, Leng Y, Leeds P, Chuang DM (2011) Histone deacetylase inhibition alters histone methylation associated with heat shock protein 70 promoter modifications in astrocytes and neurons. Neuropharmacology 60:1109–1115. doi:10.1016/j.neuropharm.2010.09.022
Marinova Z, Ren M, Wendland JR, Leng Y, Liang MH, Yasuda S, Leeds P, Chuang DM (2009) Valproic acid induces functional heat-shock protein 70 via Class I histone deacetylase inhibition in cortical neurons: a potential role of Sp1 acetylation. J Neurochem 111:976–987. doi:10.1111/j.1471-4159.2009.06385.x
Martinat C, Bacci JJ, Leete T, Kim J, Vanti WB, Newman AH, Cha JH, Gether U, Wang H, Abeliovich A (2006) Cooperative transcription activation by Nurr1 and Pitx3 induces embryonic stem cell maturation to the midbrain dopamine neuron phenotype. Proc Natl Acad Sci USA 103:2874–2879. doi:10.1073/pnas.0511153103
Masliah E, Dumaop W, Galasko D, Desplats P (2013) Distinctive patterns of DNA methylation associated with Parkinson disease: identification of concordant epigenetic changes in brain and peripheral blood leukocytes. Epigenetics 8:1030–1038. doi:10.4161/epi.25865
Matsumoto L, Takuma H, Tamaoka A, Kurisaki H, Date H, Tsuji S, Iwata A (2010) CpG demethylation enhances alpha-synuclein expression and affects the pathogenesis of Parkinson’s disease. PLoS One 5:e15522. doi:10.1371/journal.pone.0015522
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149:1635–1646. doi:10.1016/j.cell.2012.05.003
Miller DW, Hague SM, Clarimon J, Baptista M, Gwinn-Hardy K, Cookson MR, Singleton AB (2004) Alpha-synuclein in blood and brain from familial Parkinson disease with SNCA locus triplication. Neurology 62:1835–1838
Minones-Moyano E, Porta S, Escaramis G, Rabionet R, Iraola S, Kagerbauer B, Espinosa-Parrilla Y, Ferrer I, Estivill X, Marti E (2011) MicroRNA profiling of Parkinson’s disease brains identifies early downregulation of miR-34b/c which modulate mitochondrial function. Hum Mol Genet 20:3067–3078. doi:10.1093/hmg/ddr210
Mizuta I, Takafuji K, Ando Y, Satake W, Kanagawa M, Kobayashi K, Nagamori S, Shinohara T, Ito C, Yamamoto M et al (2013) YY1 binds to alpha-synuclein 3′-flanking region SNP and stimulates antisense noncoding RNA expression. J Hum Genet 58:711–719. doi:10.1038/jhg.2013.90
Monti B, Gatta V, Piretti F, Raffaelli SS, Virgili M, Contestabile A (2010) Valproic acid is neuroprotective in the rotenone rat model of Parkinson’s disease: involvement of alpha-synuclein. Neurotox Res 17:130–141. doi:10.1007/s12640-009-9090-5
Moore K, McKnight AJ, Craig D, O’Neill F (2014) Epigenome-wide association study for Parkinson’s disease. Neuromolecular Med 16:845–855. doi:10.1007/s12017-014-8332-8
Mutez E, Lepretre F, Le Rhun E, Larvor L, Duflot A, Mouroux V, Kerckaert JP, Figeac M, Dujardin K, Destee A et al (2011) SNCA locus duplication carriers: from genetics to Parkinson disease phenotypes. Hum Mutat 32:E2079–E2090. doi:10.1002/humu.21459
Nalls MA, Pankratz N, Lill CM, Do CB, Hernandez DG, Saad M, DeStefano AL, Kara E, Bras J, Sharma M et al (2014) Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat Genet 46:989–993. doi:10.1038/ng.3043
Nicholas AP, Lubin FD, Hallett PJ, Vattem P, Ravenscroft P, Bezard E, Zhou S, Fox SH, Brotchie JM, Sweatt JD et al (2008) Striatal histone modifications in models of levodopa-induced dyskinesia. J Neurochem 106:486–494. doi:10.1111/j.1471-4159.2008.05417.x
Obeid R, Schadt A, Dillmann U, Kostopoulos P, Fassbender K, Herrmann W (2009) Methylation status and neurodegenerative markers in Parkinson disease. Clin Chem 55:1852–1860. doi:10.1373/clinchem.2009.125021
Omonijo O, Wongprayoon P, Ladenheim B, McCoy MT, Govitrapong P, Jayanthi S, Cadet JL (2014) Differential effects of binge methamphetamine injections on the mRNA expression of histone deacetylases (HDACs) in the rat striatum. Neurotoxicology 45:178–184. doi:10.1016/j.neuro.2014.10.008
Outeiro TF, Kontopoulos E, Altmann SM, Kufareva I, Strathearn KE, Amore AM, Volk CB, Maxwell MM, Rochet JC, McLean PJ et al (2007) Sirtuin 2 inhibitors rescue alpha-synuclein-mediated toxicity in models of Parkinson’s disease. Science 317:516–519. doi:10.1126/science.1143780
Park G, Tan J, Garcia G, Kang Y, Salvesen G, Zhang Z (2015) Regulation of histone acetylation by autophagy in Parkinson disease. J Biol Chem. doi:10.1074/jbc.M115.675488
Patel VP, Chu CT (2014) Decreased SIRT2 activity leads to altered microtubule dynamics in oxidatively-stressed neuronal cells: implications for Parkinson’s disease. Exp Neurol 257:170–181. doi:10.1016/j.expneurol.2014.04.024
Peng GS, Li G, Tzeng NS, Chen PS, Chuang DM, Hsu YD, Yang S, Hong JS (2005) Valproate pretreatment protects dopaminergic neurons from LPS-induced neurotoxicity in rat primary midbrain cultures: role of microglia. Brain Res Mol Brain Res 134:162–169. doi:10.1016/j.molbrainres.2004.10.021
Pihlstrom L, Berge V, Rengmark A, Toft M (2015) Parkinson’s disease correlates with promoter methylation in the alpha-synuclein gene. Mov Disord 30:577–580. doi:10.1002/mds.26073
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A, Pike B, Root H, Rubenstein J, Boyer R et al (1997) Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 276:2045–2047
Portela A, Esteller M (2010) Epigenetic modifications and human disease. Nat Biotechnol 28:1057–1068. doi:10.1038/nbt.1685
Prendergast JG, Tong P, Hay DC, Farrington SM, Semple CA (2012) A genome-wide screen in human embryonic stem cells reveals novel sites of allele-specific histone modification associated with known disease loci. Epigenet Chromatin 5:6. doi:10.1186/1756-8935-5-6
Rane P, Shields J, Heffernan M, Guo Y, Akbarian S, King JA (2012) The histone deacetylase inhibitor, sodium butyrate, alleviates cognitive deficits in pre-motor stage PD. Neuropharmacology 62:2409–2412. doi:10.1016/j.neuropharm.2012.01.026
Reitz C, Tosto G, Mayeux R, Luchsinger JA (2012) Genetic variants in the fat and obesity associated (FTO) gene and risk of Alzheimer’s disease. PLoS One 7:e50354. doi:10.1371/journal.pone.0050354
Richter J, Appenzeller S, Ammerpohl O, Deuschl G, Paschen S, Bruggemann N, Klein C, Kuhlenbaumer G (2012) No evidence for differential methylation of alpha-synuclein in leukocyte DNA of Parkinson’s disease patients. Mov Disord 27:590–591. doi:10.1002/mds.24907
Rodriguez-Blanco J, Martin V, Garcia-Santos G, Herrera F, Casado-Zapico S, Antolin I, Rodriguez C (2012) Cooperative action of JNK and AKT/mTOR in 1-methyl-4-phenylpyridinium-induced autophagy of neuronal PC12 cells. J Neurosci Res 90:1850–1860. doi:10.1002/jnr.23066
Ross OA, Braithwaite AT, Skipper LM, Kachergus J, Hulihan MM, Middleton FA, Nishioka K, Fuchs J, Gasser T, Maraganore DM et al (2008) Genomic investigation of alpha-synuclein multiplication and parkinsonism. Ann Neurol 63:743–750. doi:10.1002/ana.21380
Roy A, Ghosh A, Jana A, Liu X, Brahmachari S, Gendelman HE, Pahan K (2012) Sodium phenylbutyrate controls neuroinflammatory and antioxidant activities and protects dopaminergic neurons in mouse models of Parkinson’s disease. PLoS One 7:e38113. doi:10.1371/journal.pone.0038113
Satterlee JS, Basanta-Sanchez M, Blanco S, Li JB, Meyer K, Pollock J, Sadri-Vakili G, Rybak-Wolf A (2014) Novel RNA modifications in the nervous system: form and function. J Neurosci 34:15170–15177. doi:10.1523/JNEUROSCI.3236-14.2014
Scheele C, Petrovic N, Faghihi MA, Lassmann T, Fredriksson K, Rooyackers O, Wahlestedt C, Good L, Timmons JA (2007) The human PINK1 locus is regulated in vivo by a non-coding natural antisense RNA during modulation of mitochondrial function. BMC Genom 8:74. doi:10.1186/1471-2164-8-74
Schmitt I, Kaut O, Khazneh H, deBoni L, Ahmad A, Berg D, Klein C, Frohlich H, Wullner U (2015) L-dopa increases alpha-synuclein DNA methylation in Parkinson’s disease patients in vivo and in vitro. Mov Disord. doi:10.1002/mds.26319
Sharma S, Taliyan R, Singh S (2015) Beneficial effects of sodium butyrate in 6-OHDA induced neurotoxicity and behavioral abnormalities: modulation of histone deacetylase activity. Behav Brain Res 291:306–314. doi:10.1016/j.bbr.2015.05.052
Singh M (2012) Dysregulated A to I RNA editing and non-coding RNAs in neurodegeneration. Front Genet 3:326. doi:10.3389/fgene.2012.00326
Smidt MP, Smits SM, Burbach JP (2004) Homeobox gene Pitx3 and its role in the development of dopamine neurons of the substantia nigra. Cell Tissue Res 318:35–43. doi:10.1007/s00441-004-0943-1
Soldner F, Stelzer Y, Shivalila CS, Abraham BJ, Latourelle JC, Barrasa MI, Goldmann J, Myers RH, Young RA, Jaenisch R (2016) Parkinson-associated risk variant in distal enhancer of alpha-synuclein modulates target gene expression. Nature 533:95–99. doi:10.1038/nature17939
Song C, Kanthasamy A, Anantharam V, Sun F, Kanthasamy AG (2010) Environmental neurotoxic pesticide increases histone acetylation to promote apoptosis in dopaminergic neuronal cells: relevance to epigenetic mechanisms of neurodegeneration. Mol Pharmacol 77:621–632. doi:10.1124/mol.109.062174
Song Y, Ding H, Yang J, Lin Q, Xue J, Zhang Y, Chan P, Cai Y (2014) Pyrosequencing analysis of SNCA methylation levels in leukocytes from Parkinson’s disease patients. Neurosci Lett 569:85–88. doi:10.1016/j.neulet.2014.03.076
St Laurent R, O’Brien LM, Ahmad ST (2013) Sodium butyrate improves locomotor impairment and early mortality in a rotenone-induced Drosophila model of Parkinson’s disease. Neuroscience 246:382–390. doi:10.1016/j.neuroscience.2013.04.037
Tan YY, Wu L, Zhao ZB, Wang Y, Xiao Q, Liu J, Wang G, Ma JF, Chen SD (2014) Methylation of alpha-synuclein and leucine-rich repeat kinase 2 in leukocyte DNA of Parkinson’s disease patients. Parkinsonism Relat Disord 20:308–313. doi:10.1016/j.parkreldis.2013.12.002
Thome AD, Harms AS, Volpicelli-Daley LA, Standaert DG (2016) microRNA-155 regulates alpha-synuclein-induced inflammatory responses in models of parkinson disease. J Neurosci 36:2383–2390. doi:10.1523/JNEUROSCI.3900-15.2016
Trulson ME, Cannon MS, Faegg TS, Raese JD (1985) Effects of chronic methamphetamine on the nigral-striatal dopamine system in rat brain: tyrosine hydroxylase immunochemistry and quantitative light microscopic studies. Brain Res Bull 15:569–577
Ungerstedt U (1968) 6-Hydroxy-dopamine induced degeneration of central monoamine neurons. Eur J Pharmacol 5:107–110
Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20:515–524. doi:10.1101/gad.1399806
Valente L, Nishikura K (2005) ADAR gene family and A-to-I RNA editing: diverse roles in posttranscriptional gene regulation. Prog Nucleic Acid Res Mol Biol 79:299–338. doi:10.1016/S0079-6603(04)79006-6
Vermunt MW, Reinink P, Korving J, de Bruijn E, Creyghton PM, Basak O, Geeven G, Toonen PW, Lansu N, Meunier C et al (2014) Large-scale identification of coregulated enhancer networks in the adult human brain. Cell Rep 9:767–779. doi:10.1016/j.celrep.2014.09.023
Voutsinas GE, Stavrou EF, Karousos G, Dasoula A, Papachatzopoulou A, Syrrou M, Verkerk AJ, van der Spek P, Patrinos GP, Stoger R et al (2010) Allelic imbalance of expression and epigenetic regulation within the alpha-synuclein wild-type and p.Ala53Thr alleles in Parkinson disease. Hum Mutat 31:685–691. doi:10.1002/humu.21248
Wang G, van der Walt JM, Mayhew G, Li YJ, Zuchner S, Scott WK, Martin ER, Vance JM (2008) Variation in the miRNA-433 binding site of FGF20 confers risk for Parkinson disease by overexpression of alpha-synuclein. Am J Hum Genet 82:283–289. doi:10.1016/j.ajhg.2007.09.021
Wang X, Tan L, Lu Y, Peng J, Zhu Y, Zhang Y, Sun Z (2015) MicroRNA-138 promotes tau phosphorylation by targeting retinoic acid receptor alpha. FEBS Lett 589:726–729. doi:10.1016/j.febslet.2015.02.001
Wang Y, Li Y, Toth JI, Petroski MD, Zhang Z, Zhao JC (2014) N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat Cell Biol 16:191–198. doi:10.1038/ncb2902
Wapinski O, Chang HY (2011) Long noncoding RNAs and human disease. Trends Cell Biol 21:354–361. doi:10.1016/j.tcb.2011.04.001
Wider C, Dachsel JC, Soto AI, Heckman MG, Diehl NN, Yue M, Lincoln S, Aasly JO, Haugarvoll K, Trojanowski JQ et al (2009) FGF20 and Parkinson’s disease: no evidence of association or pathogenicity via alpha-synuclein expression. Mov Disord 24:455–459. doi:10.1002/mds.22442
Wirdefeldt K, Gatz M, Reynolds CA, Prescott CA, Pedersen NL (2011) Heritability of Parkinson disease in Swedish twins: a longitudinal study. Neurobiol Aging 32(1923):e1921–e1928. doi:10.1016/j.neurobiolaging.2011.02.017
Wu X, Chen PS, Dallas S, Wilson B, Block ML, Wang CC, Kinyamu H, Lu N, Gao X, Leng Y et al (2008) Histone deacetylase inhibitors up-regulate astrocyte GDNF and BDNF gene transcription and protect dopaminergic neurons. Int J Neuropsychopharmacol 11:1123–1134. doi:10.1017/s1461145708009024
Zhou W, Bercury K, Cummiskey J, Luong N, Lebin J, Freed CR (2011) Phenylbutyrate up-regulates the DJ-1 protein and protects neurons in cell culture and in animal models of Parkinson disease. J Biol Chem 286:14941–14951. doi:10.1074/jbc.M110.211029
Acknowledgments
The authors would like to thank all those who have contributed to our research, particularly the patients and families who donated DNA samples and brain tissue for this work. The Mayo Clinic is a Morris K. Udall Parkinson’s Disease Research Center of Excellence (NINDS P50 NS072187), an Alzheimer’s disease Research Center (NIA P50 AG16574) and is supported by The Little Family Foundation, the Mangurian Foundation for Lewy body research and the Mayo Clinic AD and related dementias genetics program. OAR is supported by NINDS R01 NS078086 and The Michael J. Fox Foundation. CL is the recipient of a FRSQ postdoctoral fellowship and is a 2015 Younkin Scholar supported by the Mayo Clinic Alzheimer’s Disease and Related Dementias Genetics program. The authors would like to thank Dr. Jungsu Kim for his careful review of the manuscript and Ms. Mariana Ruiz Villarreal for her design of the neuron used in Fig. 2.
Author information
Authors and Affiliations
Corresponding author
Additional information
C. Labbé and O. Lorenzo-Betancor contributed equally.
Appendix 1
Appendix 1
-
Epigenetics Study of the molecular changes that modify the final outcome of a locus or chromosome without changing the underlying DNA sequence.
-
Heritability Proportion of individual phenotypic differences in a population that is due to genetic variation within this population.
-
Methylation Process by which methyl groups are added to DNA, this occurs most often in promotor regions. Methylation generally represses gene expression.
-
Demethylation Process that removes a methyl group from DNA to make genes in these regions accessible for expression.
-
Acetylation Inclusion of an acetyl functional group which carries a negative charge into a free lysine of the N-terminal tail of the histone leading to transcriptional activation.
-
Deacetylation Removal of an acetyl group from the N-terminal tail of the histone repressing gene expression.
-
Long non-coding RNA (lncRNA) Non-protein coding transcripts longer than 200 nucleotides that can be involved in the regulation of gene expression.
-
Small RNA Non-protein coding transcript smaller than 200 nucleotides which are often involved in RNA silencing. This group includes MicroRNAs, Short interfering RNAs and Mirtrons.
-
MicroRNA (miR) Single hairpin RNA strand precursor of ~70 bp (black strand in A) located in exons or introns of RNAs that are processed sequentially by the Drosha ribonuclease and the Dicer enzyme into a ~22 mature nucleotide (red strand in A) with incomplete complementary sequence that can regulate the expression of multiple target mRNAs.

-
Short interfering RNA (siRNA) Double RNA strand with perfect complementary sequence which can regulate a limited number of target mRNAs. However, siRNAs can also have microRNA-like off-target effects (B).
-
Mirtron Special type of miR (A) located in the introns of pre-mature RNA coding genes. They are generated during the splicing process by bypassing the Drosha maturation step and act as mRNA expression regulators.
Rights and permissions
About this article
Cite this article
Labbé, C., Lorenzo-Betancor, O. & Ross, O.A. Epigenetic regulation in Parkinson’s disease. Acta Neuropathol 132, 515–530 (2016). https://doi.org/10.1007/s00401-016-1590-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-016-1590-9