Abstract
Reversible post-translational modification is a rapid and efficient system to control the activity of pre-existing proteins. Modifiers range from small chemical moieties, such as phosphate groups, to proteins themselves as the modifier. The patriarch of the protein modifiers is ubiquitin which plays a central role in protein degradation and protein targeting. Over the last 20 years, the ubiquitin family has expanded to include a variety of ubiquitin-related small modifier proteins that are all covalently attached to a lysine residue on target proteins via series of enzymatic reactions. Of these more recently discovered ubiquitin-like proteins, the SUMO family has gained prominence as a major regulatory component that impacts numerous aspects of cell growth, differentiation, and response to stress. Unlike ubiquitinylation which often leads to proteins turn over, sumoylation performs a variety of function such as altering protein stability, modulating protein trafficking, directing protein-protein interactions, and regulating protein activity. This chapter will introduce the basic properties of SUMO proteins and the general tenets of sumoylation.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1.1 The Sumo Proteins
Over two decades ago, a small cellular protein of 12 kDa, with 18% homology to the well-known ubiquitin protein, was co-discovered and termed Small Ubiquitin-like MOdifier or SUMO. SUMO was independently identified by four groups in 1996: Freemont’s group found it as a small ubiquitin -like protein associated with PML in an interacting complex and called it PIC1 (Boddy et al. 1996), Chen’s group identified it in a two-yeast hybrid screen of proteins associated with cellular DNA repair proteins (Shen et al. 1996), Yeh’s group identified it as a small modifier associated with Fas which they called sentrin (Okura et al. 1996), and Blobel’s group discovered that RanGAP was modified by a small ubiquitin-like protein which they designated GMP1 (Matunis et al. 1996). These modifiers were all the same protein that is now commonly referred to as SUMO .
SUMO is conserved from yeast to mammalian cells though the number of SUMO genes varies greatly (Chen et al. 1998). The budding yeast, Saccharomyces cerevisiae , possesses only one SUMO gene, Smt 3, whose protein product shares 48% identity and 75% similarity with the mammalian SUMO1 (Huang et al. 2004). Likewise, both Drosophila melanogaster (Lehembre et al. 2000) and Caenorhabditis elegans (Jones et al. 2002) each have a single SUMO gene. In contrast, plants express 8 SUMOs (Kurepa et al. 2003) and vertebrates have 4 SUMOs. There are four different genes in the human genome coding for the different SUMO modifiers, SUMO1, 2, 3, and 4. SUMO2 and 3 share about 92% identity but they only related to SUMO1 at 48% identity (Kamitani et al. 1998a). While SUMO1, 2, and 3 are expressed in all tissues tested, SUMO4 transcription is restricted primarily to the kidneys , lymph nodes and spleen (Bohren et al. 2004). SUMO4 has been less studied than the others, but seems to play a role in diabetes (see Chap. 18) and stress response (Wei et al. 2008). SUMO1 is a 12 kDa protein of 101 amino acids that is related in structure and in sequence to the 9 kDa ubiquitin protein, as both modifiers share ~18% primary structure identity to each other and have 48% similarity in their three-dimensional structure (Bayer et al. 1998). Ubiquitin is only a 76 amino acid polypeptide, and the difference between those two modifiers mainly resides in the extended N-terminal structure of SUMO as this extension is absent in ubiquitin.
At the tertiary level, the basic structures have been solved for SUMO1 (Bayer et al. 1998), SUMO2 (Huang et al. 2004), and SUMO 3 (Ding et al. 2005). All three SUMOs share a central compact, globular domain with the characteristic ββαββαβ ubiquitin fold. The SUMOs also each have both N- and C-terminal extensions, with the N-terminal extension being much longer than for ubiquitin. Within this extension in SUMO2/3 is a lysine at position 11 that can itself be conjugated with SUMO to yield SUMO2/3 chains (Tatham et al. 2001). SUMO1 lacks a suitable lysine for conjugation and does not appear to form chains in vivo, though in vitro chain formation has been observed (Yang et al. 2006a). The biological role and function(s) of the N-terminal extension are not well understood, but the C-terminal extension is important for direct contact with the SUMO activating enzyme, SAE1/2 (Lois and Lima 2005).
One of the ongoing questions about the SUMOs is the functional difference between the SUMO1 and SUMO2/3 families. Certain biological variations have already been identified, including different responses to environmental conditions (Saitoh and Hinchey 2000; Manza et al. 2004; Deyrieux et al. 2007), different susceptibilities to various SUMO proteases (Gong and Yeh 2006; Mikolajczyk et al. 2007), and differences in subcellular localization and abundance (Saitoh and Hinchey 2000; Manza et al. 2004; Ayaydin and Dasso 2004). The substrate pool for these two SUMO groups is also different with some substrates capable of being modified by either SUMO1 or SUMO2/3, and other substrates showing a clear preference for one or the other SUMO type (Saitoh and Hinchey 2000; Rosas-Acosta et al. 2005; Vertegaal et al. 2006; Citro and Chiocca 2013). While SUMO preference differences exist for individual substrates, in general for both the SUMO1 and SUMO2/3 modified proteins, the substrates are predominantly nuclear and are often involved in regulation of nucleic acid structure and function. Just how biologically important this demarcation in the substrate preference is remains unclear as SUMO1 knockout mouse studies have suggested that SUMO2/3 can compensate for absent SUMO1 (Evdokimov et al. 2008; Zhang et al. 2008), suggesting considerable redundancy between the SUMO paralogs. However, more recently it was shown that SUMO2 is essential during mouse embryonic development, while SUMO3 was dispensable (Wang et al. 2014), indicating that there are suitable functional differences even between these nearly identical paralogs. Interestingly, it was previously shown that SUMO3 can be phosphorylated at serine 2, while SUMO2 cannot be phosphorylated since it has an alanine at this position (Matic et al. 2008). This observation suggests one basis for functional, regulatory, or substrate preference differences between the highly identical SUMO2 and SUMO3 proteins could be related to differences in their own post-translational modification. Much additional work is needed to clarify the common and distinct roles of the various SUMO proteins.
1.2 The Enzymology of Sumoylation
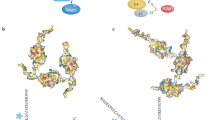
Sumoylation is the enzymatic activity which results in the covalent attachment of SUMO to a large number of proteins, including cellular and viral proteins. This multi-step enzymatic process (Fig. 1.1) includes a heterodimeric activating enzyme, SAE1/2 , a monomeric conjugating enzyme, Ubc9 , and multiple ligases and isopeptidases (Wilson 2004). SUMOs are translated as precursor forms which are initially processed by specific isopeptidases (SENPs ) to remove C-terminal residues and generate a mature SUMO , terminating with a C-terminal diglycine (Johnson et al. 1997). Interestingly, SUMO4 has a proline residue at position 90 that prevents this processing by the SENPs (Owerbach et al. 2005) and instead it is processed only under stress conditions by a stress-induced hydrolase (Wei et al. 2008). The mature forms of SUMOs then interact with the SUMO E1 E1 activating enzyme, SAE1/2 . SAE1 is a 346 amino acid polypeptide while SAE2 is 640 amino acids and contains the catalytic cysteine at residue 173; the SUMOs interact exclusively with the SAE2 subunit. The SAE2 subunit also contains a nuclear localization signal that may contribute to the enrichment of sumoylation components in the nucleus. Together, the SAE1 and 2 proteins form an U-shaped heterodimer complex with a large groove that has the ATP-binding motif at the base of the groove (Lois and Lima 2005). Binding of SUMOs to SAE2 positions the SUMO diglycine motif for adenylation, then the activated SUMO can be covalently attached to the catalytic cysteine via a thioester linkage.
Representation of the enzymatic cascades leading to the covalent attachment of SUMO to a substrate protein. The SUMO enzymes are the SENP isopeptidase, the SAE1/SAE2 activating enzyme, the Ubc9 conjugating enzyme, and the SUMO ligases. Attachment of SUMO to SAE2 and Ubc9 is via a thioester linkage to a cysteine residue in the enzymes. SUMO attached to the substrate is via a lysine residue to form a stable isopeptide bond
Subsequent to formation of the SAE1/2 -SUMO complex, the activating enzyme transfers SUMO to SUMO E2 conjugating enzyme, Ubc9 . Unlike the ubiquitin pathway that contains many E2 enzymes, Ubc9 is the sole conjugating enzyme for SUMO and functions with all 4 SUMOs. Once again, there is a conserved domain motif [αβββββ(ββ)ααα] common to all E2 enzymes known as the ubc superfold (Tong et al. 1997). Within this domain is the catalytic groove that contains the active site cysteine, amino acid 93. Binding of SAE1/2 to Ubc9 allows transfer of the SUMO C-terminus to cysteine 93, again through formation of a thioester linkage, and the structural contexts of the SAE2-Ubc9 interaction are highly conserved across species (Wang et al. 2010). Lastly, Ubc9 transfers SUMO to the substrate protein, where SUMO is covalently linked to a lysine residue through an isopeptide bond between the epsilon amino group of the lysine and the carboxyl group of the C-terminal glycine on SUMO.
The lysine residue utilized for sumoylation commonly falls in the ΨKxE/D motif, where Ψ is a hydrophobic residue (typically Val, Ile, Leu, Met, or Phe), K is the target lysine, x is any amino acid, and the fourth position is an acidic residue (Hay 2005). However, even early studies of sumoylated proteins found that not all were modified at lysines in sequence contexts that match the consensus motif, indicating that alternative sequence features could also specify a particular lysine for SUMO modification (Kamitani et al. 1998b; Rangasamy et al. 2000; Hoege et al. 2002). Subsequently, numerous proteomics approaches have identified hundreds of sumoylated proteins and characterized the SUMO addition sites in many of these substrates, revealing a site selection complexity much greater than the original consensus motif. Zhou et al. used a proteomics approach and found that five of the ten sumoylation sites determined for yeast proteins were in non-canonical sequences (Zhou et al. 2004). Similarly, Chung et al. examined SUMO2 conjugation sites for in vitro sumoylated proteins and found that half the identified sumoylation sites (three of six) where in sequences which did not conform to the ΨKxE/D motif (Chung et al. 2004). These and similar studies confirmed that while the ΨKxE/D motif is often associated with SUMO addition, only about half the identified SUMO substrates have the original consensus motif (Matic et al. 2010). In some cases sumoylation appears fairly promiscuous with many lysines in the substrate capable of serving a SUMO acceptors (Eladad et al. 2005; Chymkowitch et al. 2015; Gonzalez-Prieto et al. 2015), especially after mutation of the predominant SUMO target(s). In these cases, the substrate typically has a SUMO -interacting motif (SIM ; see below) that recruits the sumoylation machinery (Chang et al. 2011; Meulmeester et al. 2008). However, more commonly these other SUMO acceptor lysines fall within alternative SUMO conjugation motifs, including the inverted (E/DxKΨ) motif (Matic et al. 2010), the hydrophobic (ΨΨΨKxE) motif (Matic et al. 2010), the phosphorylation-dependent (PDSM ; ΨKxExxSPP) motif (Hietakangas et al. 2006), the negatively charged amino acid-dependent (ΨKxExxEEEE) motif (Yang et al. 2006b), the phosphorylated (ΨKxSPP) motif (Picard et al. 2012), and the extended phosphorylation (ΨKxSPPSPxxxSPP) motif (Picard et al. 2012). Collectively, this array of motifs helps explain the large number of lysines capable of being sumoylated and may contribute to paralog specific modification differences for individual substrates.
Unlike ubiquitinylation, which absolutely requires an E3 ubiquitin ligase for transfer of ubiquitin to the substrate, sumoylation occurs readily in vitro without a ligase requirement (Melchior 2000). Nonetheless, several SUMO ligases have now been identified, including SP-RING type ligases such as the PIAS family (Johnson and Gupta 2001) and MMS21 (Potts and Yu 2005). Members of this family share sequence homology with the RING domain of ubiquitin RING ligases. The SP-RING domain directly interacts with Ubc9 , inducing a conformational change that enhances transfer of SUMO from Ubc9 to the substrate (Rytinki et al. 2009). Additional identified SUMO ligases include RanBP2 (Pichler et al. 2002), Pc2 (Kagey et al. 2003), and TOPORS (Weger et al. 2005), as well as a few other proteins that appear to facilitate sumoylation but whose mechanisms are poorly defined. Given that there are roughly 600 ubiquitin ligase genes in the human genome (Deshaies and Joazeiro 2009), it is quite likely that many more SUMO ligases remain to be identified. Generally, all these SUMO ligases enhance sumoylation both in vitro and in vivo, and influence substrate selection (Gareau and Lima 2010). For instance, PIAS acts as a SUMO ligase, preferentially targeting the tumor suppressor p53 , c-Jun, STAT1, and the nuclear androgen receptor AR (Schmidt and Muller 2002; Ungureanu et al. 2003; Sachdev et al. 2001). RanBP2 stimulates sumoylation of the promyelocytic leukemia protein (PML) , the nuclear body SP100 protein, and the histone deacetylase HDAC4 (Pichler et al. 2002), while Pc2 is the unique E3 ligase for the transcriptional factor co-repressor CtBP (Kagey et al. 2003). In addition to enhancing the overall sumoylation reaction and substrate selection, these ligases likely also influence preferential utilization of the SUMO paralogs.
The SENPs , the SUMO isopeptidases , play a dual role; they are involved in the maturation of SUMO and in the de-conjugation of SUMO from its target proteins (Hang and Dasso 2002; Gong et al. 2000). There are 6 SENPs that function with SUMO, 1–3 and 5–7 (there is no SENP 4, and SENP 8 is a Nedd 8 protease). In mammalian cells these enzymes are differentially located, with SENP1 located at the PML bodies, SENP6 in the cytoplasm , SENP3 in the nucleolus, and SENP2 at the nuclear pore complexes (Gong and Yeh 2006). Therefore, it appears that de-sumoylation of conjugates is possible at different subcellular locations, and access of individual substrates to specific SENPs may provide an additional level of regulation. Additionally, specific functional differences have been observed among the 6 SENPs regarding their maturation and deconjugation activities. While SENP1 and SENP2 can generally process all the SUMOs 1–3 precursors (Nayak and Muller 2014), SENP5 preferentially processes the SUMO3 precursor (Di Bacco et al. 2006). With regard to deconjugation, SENP1 functions primarily with SUMO1 conjugates (Sharma et al. 2013), while the other SENPs strongly prefer SUMO2/3 substrates. Additionally, SENP6 and SENP7 are most adapt at disassembly of SUMO2/3 chains (Lima and Reverter 2008; Drag et al. 2008). Deletion of the SENP genes, like deletion of Ubc9 in yeast, stops cell cycle progression and further highlights that reversible sumoylation is an essential and critical function in the cell life cycle (Li and Hochstrasser 1999). Overall, the diversity and specificity of SENPs undoubtedly helps regulate the dynamic and reversible sumoylation process.
1.3 Sumoylation Functions
Functionally, sumoylation is a more diverse modifier than ubiquitin . Unlike ubiquitinylation, which has a major role of targeting proteins for proteasome degradation, addition of the SUMO moiety does not directly target proteins to the proteasome. Instead, there are examples of substrates where sumoylation blocks proteosomal degradation by competing with ubiquitinylation for a common lysine residue (Desterro et al. 1998; Klenk et al. 2006; Escobar-Ramirez et al. 2015). Since over 25% of the SUMO sites in human proteins are known ubiquitination sites (Hendriks et al. 2014), regulation of degradation through such competition may be more common than anticipated. Intriguingly, lysine residues are also targets for modification by acetylation and methylation, so sumoylation may also be competing with those events to regulate protein activity as has been shown for Iĸβα (Desterro et al. 1998), delta-lactoferrin (Escobar-Ramirez et al. 2015), and STAT5 (Van Nguyen et al. 2012).
Further cross-talk between the SUMO and ubiquitin systems is mediated by SUMO-Targeted ubiquitin ligases (STUbls ) (Xie et al. 2007; Prudden et al. 2007; Sun et al. 2007; Uzunova et al. 2007). This novel class of ubiquitin ligases functions by specifically interacting with SUMO moieties on sumoylated proteins, thereby causing ubiquitination and subsequent degradation (Perry et al. 2008). This interaction is depends on SUMO-interacting motifs (SIMs ) present on the STUbls . The canonical SIM is a hydrophobic motif with the consensus V/I-x-V/I-V/I (Song et al. 2004; Hecker et al. 2006), and the interaction between the SIM and the SUMO is through a β strand of the SIM and the β2 strand of SUMO (Sekiyama et al. 2008; Namanja et al. 2012). Both of the human STUbls, RNF 4 , RNF4 and RNF111, contain at least 3 SIM motifs, so they preferentially target proteins with poly-SUMO signals, either multiple SUMO moieties or SUMO chains (Tatham et al. 2008; Erker et al. 2013). Lastly, there is at least one example of a viral protein whose stability is indirectly tied to sumoylation levels (Wu et al. 2009). Through an undefined mechanism, the stability of the human papillomavirus E2 E2 protein is greatly enhanced when overall sumoylation levels increase, suggesting that further examples of cross-talk between the ubiquitin and SUMO pathways await discovery, and that these two systems may have an even richer interplay than currently imagined (see Chap. 6).
In contrast to its modest role in protein stability, it is now clear that SUMO has a major role in transcriptional regulation (see Chaps. 2 and 3), both through direct modification of individual transcription factors and co-factors (Verger et al. 2003; Garcia-Dominguez and Reyes 2009), and through chromatin remodeling (Cubenas-Potts and Matunis 2013). For most transcription factors, sumoylation reduces their transactivation capacity, though enhanced transcriptional activity has also been demonstrated for a few substrates, including heat shock factors (Goodson et al. 2001; Hong et al. 2001), Oct4 (Wei et al. 2007), and Smad4 on some promoters (Long et al. 2004). The negative transcriptional effects can be due to changes in transcription factor stability and/or subcellular localization, particularly the recruitment of sumoylated transcription factors into PML nuclear bodies as has been observed for HIPK2 (Kim et al. 1999), Sp3 (Ross et al. 2002), NACC1 (Tatemichi et al. 2015), and other proteins (Sahin et al. 2014). Alternatively, SUMO modification can have more global effects on transcription by affecting chromatin remodeling. Examples are plentiful of sumoylation facilitating the recruitment and/or modification of various remodeling enzymes including histone deacetylases (HDACs) (Wagner et al. 2015; de la Vega et al. 2012; Citro and Chiocca 2013; Girdwood et al. 2003; Yang et al. 2003), histone demethylases demethylase s (Huang et al. 2016; Bueno and Richard 2013), and methyltransferases (Spektor et al. 2011; Lee and Muller 2009; Riising et al. 2008), as well as directly modifying histones (Shiio and Eisenman 2003; Nathan et al. 2006; Zheng et al. 2015; Dhall et al. 2014). Clearly, all of these mechanisms could be reversed by desumoylation with SENPs , leading to dynamic and controllable effects on transcription of individual or groups of genes. Thus, sumoylation effects on transcriptional activity would reflect the overall dynamics of sumoylation/desumoylation that may vary with cell cycle , cell growth conditions, and disease state.
In addition to regulating transcriptional activity, sumoylation also has an important regulatory role for other nuclear functions, including RNA processing (see Chap. 2), genome maintenance and repair (see Chaps. 4 and 5), and nucleocytoplasmic transport (see Chap. 7). More recently, non-nuclear functions of sumoylation have been identified (Wasik and Filipek 2014), and Chaps. 8 and 9 will explore the role of SUMOs in regulating ion channel activity and metabolic pathways, respectively. Because of this pleiotropic ability to modify numerous proteins and affect transcriptional activity or cellular environment on a global scale, sumoylation in now recognized as a regulatory process involved in mitosis (Chap. 10), meiosis (Chap. 11), differentiation and development (Chap. 12), and senescence (Chap. 13). While much of the focus in the sumoylation field is on vertebrates, sumoylation is equally important for plants (Chap. 14) and invertebrates (Chap. 15). Much progress has been made in recent years in understanding the roles of sumoylation in these diverse areas of cell biology, particularly through global proteomics efforts (Tammsalu et al. 2015; Eifler and Vertegaal 2015; Hendriks et al. 2015a; Xiao et al. 2015; Yang and Paschen 2015), but much work remains, and for most of these processes there still are many more questions than answers.
One recently emerging theme that likely contributes to the ability of sumoylation to control the cellular processes mentioned above is the coordinate modification of functionally related groups of protein in response to specific stimuli (Jentsch and Psakhye 2013; Raman et al. 2013). Overall increases in cellular sumoylation levels have long been seen in response to various kinds of stress (the SUMO stress response, SSR) (Zhou et al. 2004; Manza et al. 2004; Tempe et al. 2008), and more recent studies are now revealing that much of this increased sumoylation is associated with networked proteins (Lewicki et al. 2015; Castro et al. 2012; Hendriks et al. 2015b; Xiao et al. 2015). For example, DNA damage has been shown to elicit the sumoylation of numerous proteins in the homologous recombination system (Psakhye and Jentsch 2012). Many of the proteins in these networks contain SIM motifs, so increased sumoylation would likely contribute to enhanced interactions and stability of these multi-protein complexes. Thus, by subtle interplay of sumoylation and desumoylation these protein complexes and functional pathway could be fine-tuned to produce rapid and appropriate levels of response to changing cellular conditions. Interestingly, at least in vertebrates, it is SUMO2/3 that are mostly involved in SSR, and the intracellular pools of free SUMO2/3 are rapidly lost after exposure to stress -inducing agents as SUMO2 and SUMO3 becomes largely conjugated to their substrates.
Lastly, given the breadth of SUMO modified targets and the critical pathways involved, it is not surprising that dysregulation of the SUMO system can contribute to disease states. Increasing evidence links over or under expression of various sumoylation components to diseases as diverse as neurodegeneration (Chap. 16), cancer (Chap. 17), diabetes (Chap. 18), craniofacial disorders (Chap. 19), and vascular disease (Chap. 20). It is also now apparent that utilization and/or modulation of the host sumoylation system are an important aspect of many infection diseases, both viral (Chap. 21) and bacterial (Chap. 22). This emerging recognition of a role for sumoylation in disease and infection is exciting as it may ultimately offer new insights for diagnosis, therapeutics, and prevention. The next several years should bring exciting new insight into the role of sumoylation, not only in fundamental cellular processes, but also in applications to understanding and managing disease states.
1.4 Conclusion
In the 20 plus years since its discovery, SUMO has gone from an obscure and functionally unknown protein to one that is recognized as a key regulator of multiple nuclear and cytoplasm ic events. The principal components of this modification system have been identified, their basic structures elucidated, and the general features of their enzymology understood. Thanks to the combination of individual targeted protein studies and more global proteomics approaches, hundreds of sumoylation targets are now known, providing a rich resource for subsequent functional studies. The sumoylation system has been shown to be an important player in many biological processes, such as cellular differentiation, transcriptional regulation, and cell growth (Deyrieux et al. 2007; Gill 2005; Ihara et al. 2007). Perturbing this biological system changes the cellular response to diverse signaling pathways (Sharrocks 2006) and likely leads to disease. In the chapters that follow, the role of sumoylation in a variety of cellular processes will be explored. The focus will range from effects on molecular targets through cell processes to the organismal level. While many exciting questions remain unanswered, by spanning from molecules to multicellular systems, the full impact and profound significance of the sumoylation system should become apparent. We hope that both newcomers to this field, as well as veterans, will find this comprehensive compilation of state-of-the-art reviews on current sumoylation topics useful and insightful.
References
Ayaydin F, Dasso M (2004) Distinct in vivo dynamics of vertebrate SUMO paralogues. Mol Biol Cell 15:5208–5218
Bayer P, Arndt A, Metzger S, Mahajan R, Melchior F, Jaenicke R, Becker J (1998) Structure determination of the small ubiquitin-related modifier SUMO-1. J Mol Biol 280:275–286
Boddy MN, Howe K, Etkin LD, Solomon E, Freemont PS (1996) PIC 1, a novel ubiquitin-like protein which interacts with the PML component of a multiprotein complex that is disrupted in acute promyelocytic leukaemia. Oncogene 13:971–982
Bohren KM, Nadkarni V, Song JH, Gabbay KH, Owerbach D (2004) A M55 V polymorphism in a novel SUMO gene (SUMO-4) differentially activates heat shock transcription factors and is associated with susceptibility to type I diabetes mellitus. J Biol Chem 279:27233–27238
Bueno MT, Richard S (2013) SUMOylation negatively modulates target gene occupancy of the KDM5B, a histone lysine demethylase. Epigenetics 8:1162–1175
Castro PH, Tavares RM, Bejarano ER, Azevedo H (2012) SUMO, a heavyweight player in plant abiotic stress responses. Cell Mol Life Sci 69:3269–3283
Chang CC, Naik MT, Huang YS, Jeng JC, Liao PH, Kuo HY, Ho CC, Hsieh YL, Lin CH, Huang NJ, Naik NM, Kung CCH, Lin SY, Chen RH, Chang KS, Huang TH, Shih HM (2011) Structural and functional roles of Daxx SIM phosphorylation in SUMO paralog-selective binding and apoptosis modulation. Mol Cell 42:62–74
Chen A, Mannen H, Li SS (1998) Characterization of mouse ubiquitin-like SMT3A and SMT3B cDNAs and gene/pseudogenes. Biochem Mol Biol Int 46:1161–1174
Chung TL, Hsiao HH, Yeh YY, Shia HL, Chen YL, Liang PH, Wang AHJ, Khoo KH, Li SSL (2004) In vitro modification of human centromere protein CENP-C fragments by small ubiquitin-like modifier (SUMO) protein – definitive identification of the modification sites by tandem mass spectrometry analysis of the isopeptides. J Biol Chem 279:39653–39662
Chymkowitch P, Nguea AP, Aanes H, Koehler CJ, Thiede B, Lorenz S, Meza-Zepeda LA, Klungland A, Enserink JM (2015) Sumoylation of Rap1 mediates the recruitment of TFIID to promote transcription of ribosomal protein genes. Genome Res 25:897–906
Citro S, Chiocca S (2013) SUMO paralogs: redundancy and divergencies. Front Biosci 5:544–553
Cubenas-Potts C, Matunis MJ (2013) SUMO: a multifaceted modifier of chromatin structure and function. Dev Cell 24:1–12
de la Vega L, Grishina I, Moreno R, Kruger M, Braun T, Schmitz ML (2012) A redox-regulated SUMO/acetylation switch of HIPK2 controls the survival threshold to oxidative stress. Mol Cell 46:472–483
Deshaies RJ, Joazeiro CA (2009) RING domain E3 ubiquitin ligases. Annu Rev Biochem 78:399–434
Desterro JM, Rodriguez MS, Hay RT (1998) SUMO-1 modification of IkappaBalpha inhibits NF-kappaB activation. Mol Cell 2:233–239
Deyrieux AF, Rosas-Acosta G, Ozbun MA, Wilson VG (2007) Sumoylation dynamics during keratinocyte differentiation. J Cell Sci 120:125–136
Dhall A, Wei S, Fierz B, Woodcock CL, Lee TH, Chatterjee C (2014) Sumoylated human histone H4 prevents chromatin compaction by inhibiting long-range internucleosomal interactions. J Biol Chem 289:33827–33837
Di Bacco A, Ouyang J, Lee HY, Catic A, Ploegh H, Gill G (2006) The SUMO-specific protease SENP5 is required for cell division. Mol Cell Biol 26:4489–4498
Ding HS, Xu YQ, Chen Q, Dai HM, Tang YJ, Wu JH, Shi YY (2005) Solution structure of human SUMO-3 C47S and its binding surface for Ubc9. Biochemist 44:2790–2799
Drag M, Mikolajczyk J, Krishnakumar IM, Huang ZW, Salvesen GS (2008) Activity profiling of human deSUMOylating enzymes (SENPs) with synthetic substrates suggests an unexpected specificity of two newly characterized members of the family. Biochem J 409:461–469
Eifler K, Vertegaal AC (2015) Mapping the SUMOylated landscape. FEBS J 282:3669–3680
Eladad S, Ye TZ, Hu P, Leversha M, Beresten S, Matunis MJ, Ellis NA (2005) Intra-nuclear trafficking of the BLM helicase to DNA damage-induced foci is regulated by SUMO modification. Hum Mol Genet 14:1351–1365
Erker Y, Neyret-Kahn H, Seeler JS, Dejean A, Atfi A, Levy L (2013) Arkadia, a novel SUMO-targeted ubiquitin ligase involved in PML degradation. Mol Cell Biol 33:2163–2177
Escobar-Ramirez A, Vercoutter-Edouart AS, Mortuaire M, Huvent I, Hardiville S, Hoedt E, Lefebvre T, Pierce A (2015) Modification by SUMOylation controls both the transcriptional activity and the stability of delta-lactoferrin. PLoS One 10:e0129965
Evdokimov E, Sharma P, Lockett SJ, Lualdi M, Kuehn MR (2008) Loss of SUMO1 in mice affects RanGAP1 localization and formation of PML nuclear bodies, but is not lethal as it can be compensated by SUMO2 or SUMO3. J Cell Sci 121:4106–4113
Garcia-Dominguez M, Reyes JC (2009) SUMO association with repressor complexes, emerging routes for transcriptional control. Biochim Biophys Acta 1789:451–459
Gareau JR, Lima CD (2010) The SUMO pathway: emerging mechanisms that shape specificity, conjugation and recognition. Nat Rev Mol Cell Biol 11:861–871
Gill G (2005) Something about SUMO inhibits transcription. Curr Opin Genet Dev 15:536–541
Girdwood D, Bumpass D, Vaughan OA, Thain A, Anderson LA, Snowden AW, Garcia-Wilson E, Perkins ND, Hay RT (2003) p300 transcriptional repression is mediated by SUMO modification. Mol Cell 11:1043–1054
Gong L, Yeh ETH (2006) Characterization of a family of nucleolar SUMO-specific proteases with preference for SUMO-2 or SUMO-3. J Biol Chem 281:15869–15877
Gong LM, Millas S, Maul GG, Yeh ETH (2000) Differential regulation of sentrinized proteins by a novel sentrin-specific protease. J Biol Chem 275:3355–3359
Gonzalez-Prieto R, Cuijpers SA, Kumar R, Hendriks IA, Vertegaal AC (2015) c-Myc is targeted to the proteasome for degradation in a SUMOylation-dependent manner, regulated by PIAS1, SENP7 and RNF4. Cell Cycle 14:1859–1872
Goodson ML, Hong Y, Rogers R, Matunis MJ, Park-Sarge OK, Sarge KD (2001) SUMO-1 modification regulates the DNA binding activity of heat shock transcription factor 2, a promyelocytic leukemia nuclear body associated transcription factor. J Biol Chem 276:18513–18518
Hang J, Dasso M (2002) Association of the human SUMO-1 protease SENP2 with the nuclear pore. J Biol Chem 277:19961–19966
Hay RT (2005) SUMO: a history of modification. Mol Cell 18:1–12
Hecker CM, Rabiller M, Haglund K, Bayer P, Dikic I (2006) Specification of SUMO1- and SUMO2-interacting motifs. J Biol Chem 281:16117–16127
Hendriks IA, D’Souza RC, Yang B, Verlaan-de Vries M, Mann M, Vertegaal AC (2014) Uncovering global SUMOylation signaling networks in a site-specific manner. Nat Struct Mol Biol 21:927–936
Hendriks IA, D’Souza RC, Chang JG, Mann M, Vertegaal AC (2015a) System-wide identification of wild-type SUMO-2 conjugation sites. Nat Commun 6:7289
Hendriks IA, Treffers LW, Verlaan-de Vries M, Olsen JV, Vertegaal AC (2015b) SUMO-2 orchestrates chromatin modifiers in response to DNA damage. Cell Rep 10:1778–1791
Hietakangas V, Anckar J, Blomster HA, Fujimoto M, Palvimo JJ, Nakai A, Sistonen L (2006) PDSM, a motif for phosphorylation-dependent SUMO modification. Proc Natl Acad Sci USA 103:45–50
Hoege C, Pfander B, Moldovan GL, Pyrowolakis G, Jentsch S (2002) RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419:135–141
Hong YL, Rogers R, Matunis MJ, Mayhew CN, Goodson M, Park-Sarge OK, Sarge KD (2001) Regulation of heat shock transcription factor 1 by stress-induced SUMO-1 modification. J Biol Chem 276:40263–40267
Huang WC, Ko TP, Li SSL, Wang AHJ (2004) Crystal structures of the human SUMO-2 protein at 1.6 angstrom and 1.2 angstrom resolution – implication on the functional differences of SUMO proteins. Eur J Biochem 271:4114–4122
Huang C, Cheng J, Bawa-Khalfe T, Yao X, Chin YE, Yeh ET (2016) SUMOylated ORC2 recruits a histone demethylase to regulate centromeric histone modification and genomic stability. Cell Rep 15:147–157
Ihara M, Koyama H, Uchimura Y, Saitoh H, Kikuchi A (2007) Noncovalent binding of small ubiquitin-related modifier (SUMO) protease to SUMO is necessary for enzymatic activities and cell growth. J Biol Chem 282:16465–16475
Jentsch S, Psakhye I (2013) Control of nuclear activities by substrate-selective and protein-group SUMOylation. Annu Rev Genet 47:167–186
Johnson ES, Gupta AA (2001) An E3-like factor that promotes SUMO conjugation to the yeast septins. Cell 106:735–744
Johnson ES, Schwienhorst I, Dohmen RJ, Blobel G (1997) The ubiquitin-like protein Smt3p is activated for conjugation to other proteins by an Aos1p/Uba2p heterodimer. EMBO J 16:5509–5519
Jones D, Crowe E, Stevens TA, Candido EP (2002) Functional and phylogenetic analysis of the ubiquitylation system in Caenorhabditis elegans: ubiquitin-conjugating enzymes, ubiquitin-activating enzymes, and ubiquitin-like proteins. Genome Biol 3:0002.1–0002.15
Kagey MH, Melhuish TA, Wotton D (2003) The polycomb protein Pc2 is a SUMO E3. Cell 113:127–137
Kamitani T, Kito K, Nguyen HP, Fukuda-Kamitani T, Yeh ET (1998a) Characterization of a second member of the sentrin family of ubiquitin-like proteins. J Biol Chem 273:11349–11353
Kamitani T, Kito K, Nguyen HP, Wada H, Fukuda-Kamitani T, Yeh ET (1998b) Identification of three major sentrinization sites in PML. J Biol Chem 273:26675–26682
Kim YH, Choi CY, Kim Y (1999) Covalent modification of the homeodomain-interacting protein kinase 2 (HIPK2) by the ubiquitin-like protein SUMO-1. Proc Natl Acad Sci U S A 96:12350–12355
Klenk C, Humrich J, Quitterer U, Lohse MJ (2006) SUMO-1 controls the protein stability and the biological function of phosducin. J Biol Chem 281:8357–8364
Kurepa J, Walker JM, Smalle J, Gosink MM, Davis SJ, Durham TL, Sung DY, Vierstra RD (2003) The small ubiquitin-like modifier (SUMO) protein modification system in Arabidopsis – accumulation of SUMO1 and −2 conjugates is increased by stress. J Biol Chem 278:6862–6872
Lee B, Muller MT (2009) SUMOylation enhances DNA methyltransferase 1 activity. Biochem J 421:449–461
Lehembre F, Badenhorst P, Muller S, Travers A, Schweisguth F, Dejean A (2000) Covalent modification of the transcriptional repressor tramtrack by the ubiquitin-related protein Smt3 in Drosophila flies. Mol Cell Biol 20:1072–1082
Lewicki MC, Srikumar T, Johnson E, Raught B (2015) The S. cerevisiae SUMO stress response is a conjugation-deconjugation cycle that targets the transcription machinery. J Proteomics 118:39–48
Li SJ, Hochstrasser M (1999) A new protease required for cell-cycle progression in yeast. Nature 398:246–251
Lima CD, Reverter D (2008) Structure of the human SENP7 catalytic domain and poly-SUMO deconjugation activities for SENP6 and SENP7. J Biol Chem 283:32045–32055
Lois LM, Lima CD (2005) Structures of the SUMO E1 provide mechanistic insights into SUMO activation and E2 recruitment to E1. EMBO J 24:439–451
Long JY, Wang GN, He DM, Liu F (2004) Repression of Smad4 transcriptional activity by SUMO modification. Biochem J 379:23–29
Manza LL, Codreanu SG, Stamer SL, Smith DL, Wells KS, Roberts RL, Liebler DC (2004) Global shifts in protein sumoylation in response to electrophile and oxidative stress. Chem Res Toxicol 17:1706–1715
Matic I, Macek B, Hilger M, Walther TC, Mann M (2008) Phosphorylation of SUMO-1 occurs in vivo and is conserved through evolution. J Proteome Res 7:4050–4057
Matic I, Schimmel J, Hendriks IA, van Santen MA, van de Rijke F, van Dam H, Gnad F, Mann M, Vertegaal ACO (2010) Site-specific identification of SUMO-2 targets in cells reveals an inverted SUMOylation motif and a hydrophobic cluster SUMOylation motif. Mol Cell 39:641–652
Matunis MJ, Coutavas E, Blobel G (1996) A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J Cell Biol 135:1457–1470
Melchior F (2000) SUMO – nonclassical ubiquitin. Annu Rev Cell Dev Biol 16:591–626
Meulmeester E, Kunze M, Hsiao HH, Urlaub H, Melchior F (2008) Mechanism and consequences for paralog-specific sumoylation of ubiquitin-specific protease 25. Mol Cell 30:610–619
Mikolajczyk J, Drag M, Bekes M, Cao JT, Ronai Z, Salvesen GS (2007) Small ubiquitin-related modifier (SUMO)-specific proteases – profiling the specificities and activities of human SENPs. J Biol Chem 282:26217–26224
Namanja AT, Li YJ, Su Y, Wong S, Lu J, Colson LT, Wu C, Li SS, Chen Y (2012) Insights into high affinity small ubiquitin-like modifier (SUMO) recognition by SUMO-interacting motifs (SIMs) revealed by a combination of NMR and peptide array analysis. J Biol Chem 287:3231–3240
Nathan D, Ingvarsdottir K, Sterner DE, Bylebyl GR, Dokmanovic M, Dorsey LA, Whelan KA, Krsmanovic M, Lane WS, Meluh PB, Johnson ES, Berger SL (2006) Histone sumoylation is a negative regulator in Saccharomyces cerevisiae and shows dynamic interplay with positive-acting histone modifications. Genes Dev 20:966–976
Nayak A, Muller S (2014) SUMO-specific proteases/isopeptidases: SENPs and beyond. Genome Biol 15:422
Okura T, Gong L, Kamitani T, Wada T, Okura I, Wei CF, Chang HM, Yeh ET (1996) Protection against Fas/APO-1- and tumor necrosis factor-mediated cell death by a novel protein, sentrin. J Immunol 157:4277–4281
Owerbach D, McKay EM, Yeh ET, Gabbay KH, Bohren KM (2005) A proline-90 residue unique to SUMO-4 prevents maturation and sumoylation. Biochem Biophys Res Commun 337:517–520
Perry JJP, Tainer JA, Boddy MN (2008) A SIM-ultaneous role for SUMO and ubiquitin. Trends Biochem Sci 33:201–208
Picard N, Caron V, Bilodeau S, Sanchez M, Mascle X, Aubry M, Tremblay A (2012) Identification of estrogen receptor beta as a SUMO-1 target reveals a novel phosphorylated sumoylation motif and regulation by glycogen synthase kinase 3beta. Mol Cell Biol 32:2709–2721
Pichler A, Gast A, Seeler JS, Dejean A, Melchior F (2002) The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell 108:109–120
Potts PR, Yu HT (2005) Human MMS21/NSE2 is a SUMO ligase required for DNA repair. Mol Cell Biol 25:7021–7032
Prudden J, Pebernard S, Raffa G, Slavin DA, Perry JJP, Tainer JA, McGowan CH, Boddy MN (2007) SUMO-targeted ubiquitin ligases in genome stability. EMBO J 26:4089–4101
Psakhye I, Jentsch S (2012) Protein group modification and synergy in the SUMO pathway as exemplified in DNA repair. Cell 151:807–820
Raman N, Nayak A, Muller S (2013) The SUMO system: a master organizer of nuclear protein assemblies. Chromosoma 122:475–485
Rangasamy D, Woytek K, Khan SA, Wilson VG (2000) SUMO-1 modification of bovine papillomavirus E1 protein is required for intranuclear accumulation. J Biol Chem 275:37999–38004
Riising EM, Boggio R, Chiocca S, Helin K, Pasini D (2008) The polycomb repressive complex 2 is a potential target of SUMO modifications. PLoS One 3:e2704
Rosas-Acosta G, Russell WK, Deyrieux A, Russell DH, Wilson VG (2005) A universal strategy for proteomic studies of SUMO and other ubiquitin-like modifiers. Mol Cell Proteomics 4:56–72
Ross S, Best JL, Zon LI, Gill G (2002) SUMO-1 modification represses Sp3 transcriptional activation and modulates its subnuclear localization. Mol Cell 10:831–842
Rytinki MM, Kaikkonen S, Pehkonen P, Jaaskelainen T, Palvimo JJ (2009) PIAS proteins: pleiotropic interactors associated with SUMO. Cell Mol Life Sci 66:3029–3041
Sachdev S, Bruhn L, Sieber H, Pichler A, Melchior F, Grosschedl R (2001) PIASy, a nuclear matrix-associated SUMO E3 ligase, represses LEF1 activity by sequestration into nuclear bodies. Genes Dev 15:3088–3103
Sahin U, de The H, Lallemand-Breitenbach V (2014) PML nuclear bodies: assembly and oxidative stress-sensitive sumoylation. Nucleus 5:499–507
Saitoh H, Hinchey J (2000) Functional heterogeneity of small ubiquitin-related protein modifiers SUMO-1 versus SUMO-2/3. J Biol Chem 275:6252–6258
Schmidt D, Muller S (2002) Members of the PIAS family act as SUMO ligases for c-Jun and p53 and repress p53 activity. Proc Natl Acad Sci U S A 99:2872–2877
Sekiyama N, Ikegami T, Yamane T, Ikeguchi M, Uchimura Y, Baba D, Ariyoshi M, Tochio H, Saitoh H, Shirakawa M (2008) Structure of the small ubiquitin-like modifier (SUMO)-interacting motif of MBD1-containing chromatin-associated factor 1 bound to SUMO-3. J Biol Chem 283:35966–35975
Sharma P, Yamada S, Lualdi M, Dasso M, Kuehn MR (2013) SENP1 is essential for desumoylating SUMO1-modified proteins but dispensable for SUMO2 and SUMO3 deconjugation in the mouse embryo. Cell Rep 3:1640–1650
Sharrocks AD (2006) PIAS proteins and transcriptional regulation – more than just SUMO E3 ligases? Genes Dev 20:754–758
Shen ZY, Pardingtonpurtymun PE, Comeaux JC, Moyzis RK, Chen DJ (1996) Ubl1, a human ubiquitin-like protein associating with human rad51/rad52 proteins. Genomics 36:271–279
Shiio Y, Eisenman RN (2003) Histone sumoylation is associated with transcriptional repression. Proc Natl Acad Sci U S A 100:13225–13230
Song J, Durrin LK, Wilkinson TA, Krontiris TG, Chen YA (2004) Identification of a SUMO-binding motif that recognizes SUMO-modified proteins. Proc Natl Acad Sci U S A 101:14373–14378
Spektor TM, Congdon LM, Veerappan CS, Rice JC (2011) The UBC9 E2 SUMO conjugating enzyme binds the PR-Set7 histone methyltransferase to facilitate target gene repression. PLoS One 6:e22785
Sun H, Leverson JD, Hunter T (2007) Conserved function of RNF4 family proteins in eukaryotes: targeting a ubiquitin ligase to SUMOylated proteins. EMBO J 26:4102–4112
Tammsalu T, Matic I, Jaffray EG, Ibrahim AF, Tatham MH, Hay RT (2015) Proteome-wide identification of SUMO modification sites by mass spectrometry. Nat Protoc 10:1374–1388
Tatemichi Y, Shibazaki M, Yasuhira S, Kasai S, Tada H, Oikawa H, Suzuki Y, Takikawa Y, Masuda T, Maesawa C (2015) Nucleus accumbens associated 1 is recruited within the promyelocytic leukemia nuclear body through SUMO modification. Cancer Sci 106:848–856
Tatham MH, Jaffray E, Vaughan OA, Desterro JMP, Botting CH, Naismith JH, Hay RT (2001) Polymeric chains of SUMO-2 and SUMO-3 are conjugated to protein substrates by SAE1/SAE2 and Ubc9. J Biol Chem 276:35368–35374
Tatham MH, Geoffroy MC, Shen L, Plechanovova A, Hattersley N, Jaffray EG, Palvimo JJ, Hay RT (2008) RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol 10:538–546
Tempe D, Piechaczyk M, Bossis G (2008) SUMO under stress. Biochem Soc Trans 36:874–878
Tong H, Hateboer G, Perrakis A, Bernards R, Sixma TK (1997) Crystal structure of murine/human Ubc9 provides insight into the variability of the ubiquitin-conjugating system. J Biol Chem 272:21381–21387
Ungureanu D, Vanhatupa S, Kotaja N, Yang J, Aittomaki S, Janne OA, Palvimo JJ, Silvennoinen O (2003) PIAS proteins promote SUMO-1 conjugation to STAT1. Blood 102:3311–3313
Uzunova K, Gottsche K, Miteva M, Weisshaar SR, Glanemann C, Schnellhardt M, Niessen M, Scheel H, Hofmann K, Johnson ES, Praefcke GJ, Dohmen RJ (2007) Ubiquitin-dependent proteolytic control of SUMO conjugates. J Biol Chem 282:34167–34175
Van Nguyen T, Angkasekwinai P, Dou H, Lin FM, Lu LS, Cheng J, Chin YE, Dong C, Yeh ET (2012) SUMO-specific protease 1 is critical for early lymphoid development through regulation of STAT5 activation. Mol Cell 45:210–221
Verger A, Perdomo J, Crossley M (2003) Modification with SUMO – a role in transcriptional regulation. EMBO Rep 4:137–142
Vertegaal ACO, Andersen JS, Ogg SC, Hay RT, Mann M, Lamond AI (2006) Distinct and overlapping sets of SUMO-1 and SUMO-2 target proteins revealed by quantitative proteomics. Mol Cell Proteomics 5:2298–2310
Wagner T, Kiweler N, Wolff K, Knauer SK, Brandl A, Hemmerich P, Dannenberg JH, Heinzel T, Schneider G, Kramer OH (2015) Sumoylation of HDAC2 promotes NF-kappaB-dependent gene expression. Oncotarget 6:7123–7135
Wang J, Taherbhoy AM, Hunt HW, Seyedin SN, Miller DW, Miller DJ, Huang DT, Schulman BA (2010) Crystal Structure of UBA2(ufd)-Ubc9: insights into E1-E2 interactions in SUMO pathways. PLoS One 5:e15805
Wang L, Wansleeben C, Zhao S, Miao P, Paschen W, Yang W (2014) SUMO2 is essential while SUMO3 is dispensable for mouse embryonic development. EMBO Rep 15:878–885
Wasik U, Filipek A (2014) Non-nuclear function of sumoylated proteins. Biochim Biophys Acta 1843:2878–2885
Weger S, Hammer E, Heilbronn R (2005) Topors acts as a SUMO-1 E3 ligase for p53 in vitro and in vivo. FEBS Lett 579:5007–5012
Wei F, Scholer HR, Atchison ML (2007) Sumoylation of Oct4 enhances its stability, DNA binding, and transactivation. J Biol Chem 282:21551–21560
Wei WZ, Yang P, Pang JF, Zhang S, Wang Y, Wang MH, Dong Z, She JX, Wang CY (2008) A stress-dependent SUMO4 sumoylation of its substrate proteins. Biochem Biophys Res Commun 375:454–459
Wilson VG (2004) Sumoylation: molecular biology and biochemistry. Horizon Biosciences, Norfolk
Wu YC, Bian XL, Heaton PH, Deyrieux AF, Wilson VG (2009) Host cell sumoylation level influences papillomavirus E2 protein stability. Virology 387:176–183
Xiao Z, Chang JG, Hendriks IA, Sigurethsson JO, Olsen JV, Vertegaal AC (2015) System-wide analysis of SUMOylation dynamics in response to replication stress reveals novel small ubiquitin-like modified target proteins and acceptor lysines relevant for genome stability. Mol Cell Proteomics 14:1419–1434
Xie Y, Kerscher O, Kroetz MB, McConchie HF, Sung P, Hochstrasser M (2007) The yeast HEX3-SLX8 heterodimer is a ubiquitin ligase stimulated by substrate sumoylation. J Biol Chem 282:34176–34184
Yang W, Paschen W (2015) SUMO proteomics to decipher the SUMO-modified proteome regulated by various diseases. Proteomics 15:1181–1191
Yang SH, Jaffray E, Hay RT, Sharrocks AD (2003) Dynamic interplay of the SUMO and ERK pathways in regulating Elk-1 transcriptional activity. Mol Cell 12:63–74
Yang ML, Hsu CT, Ting CY, Liu LF, Hwang JL (2006a) Assembly of a polymeric chain of SUMO1 on human topoisomerase I in vitro. J Biol Chem 281:8264–8274
Yang SH, Galanis A, Witty J, Sharrocks AD (2006b) An extended consensus motif enhances the specificity of substrate modification by SUMO. EMBO J 25:5083–5093
Zhang FP, Mikkonen L, Toppari J, Palvimo JJ, Thesleff I, Janne OA (2008) SUMO-1 function is dispensable in normal mouse development. Mol Cell Biol 28:5381–5390
Zheng J, Liu L, Wang S, Huang X (2015) SUMO-1 promotes Ishikawa cell proliferation and apoptosis in endometrial cancer by increasing Sumoylation of histone H4. Int J Gynecol Cancer 25:1364–1368
Zhou W, Ryan JJ, Zhou H (2004) Global analyses of sumoylated proteins in Saccharomyces cerevisiae – induction of protein sumoylation by cellular stresses. J Biol Chem 279:32262–32268
Acknowledgments
We wish to thank other current and former members of the Wilson lab for discussions that helped form much of the work presented here.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Wilson, V.G. (2017). Introduction to Sumoylation. In: Wilson, V. (eds) SUMO Regulation of Cellular Processes. Advances in Experimental Medicine and Biology, vol 963. Springer, Cham. https://doi.org/10.1007/978-3-319-50044-7_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-50044-7_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-50043-0
Online ISBN: 978-3-319-50044-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)