Abstract
Protein modification by the small ubiquitin-related modifier (SUMO) was simultaneously discovered by several groups at the middle of the 1990s. Although distinct names were proposed including Sentrin, GMP1, PIC1, or SMT3, SUMO became the most popular. Early studies on the functions of SUMOylation focused on activities in the nucleus, including transcription activation, chromatin structure, and DNA repair. However, it is now recognized that SUMOylation affects a large diversity of cellular processes both in the nucleus and the cytoplasm and functions of SUMOylation appear to have undefined limits. SUMO-conjugating enzymes and specific proteases actively regulate the modification status of target proteins. The recent discoveries of ubiquitin-SUMO hybrid chains, multiple SUMO-interacting motifs, and macromolecular complexes regulated by SUMOylation underscore the high complexity of this dynamic reversible system. New conceptual frameworks suggested by these findings have motivated the development of new methodologies to study pre- and post-SUMOylation events in vitro and in vivo, using distinct model organisms. Here we summarize some of the new developments and methodologies in the field, particularly those that will be further elaborated on in the chapters integrating this book.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 A Brief History of the Discovery of Small Ubiquitin-Related Modifiers
In 1978, Avram Hershko and his graduate student, Aaron Ciechanover, discovered that proteins added to reticulocyte extracts become covalently conjugated to a protein called ubiquitin [1]. They subsequently demonstrated that these ubiquitylated proteins are degraded in an ATP-dependent process, thus establishing entirely new fields of cellular, molecular, and biochemical research [2]. These pioneering studies paved the way for subsequent work by a large number of laboratories that revealed regulatory roles for ubiquitylation in virtually every aspect of cell function [3], and ultimately earned Hershko, Ciechanover, and their collaborator Irwin Rose the 2004 Nobel Prize in Chemistry [4].
Despite the clear utility of the ubiquitylation pathway for regulating cellular processes through exquisite spatial and temporal control of posttranslational protein modification, the existence of related pathways remained uncertain for nearly 20 years following the initial discoveries of Hershko and Ciechanover. This abruptly changed in the mid-1990s with the near-concomitant discoveries by several laboratories of the family of proteins now known as small ubiquitin-related modifiers, or SUMOs. The newly emergent SUMO field benefited enormously from the groundwork and insights obtained through more than 20 years of ubiquitin research. Thus, it was established very quickly that SUMOylation proceeds through an enzyme cascade paralleling ubiquitylation and functions as a posttranslational modification with impact and consequences as broad and profound as ubiquitylation [5]. This review highlights the earliest studies and experimental evidence that identified either the genes coding for SUMOs or the SUMO proteins themselves, and thus pointed to the existence of parallel, ubiquitin-like posttranslational protein modification pathways. Notably, the discovery of SUMOs represented an important milestone, paving the way for the subsequent discovery and characterization of more than ten functionally distinct ubiquitin-like proteins and pathways (UBLs) [6].
1.1 First Discoveries of Genes and cDNAs Coding for SUMO Proteins
The earliest reported studies hinting at the existence of SUMO proteins came in 1995 through the work of Pamela Meluh and Douglas Koshland. Their discovery of SUMO resulted from work focused on analysis of Mif2 , a protein linked to accurate chromosome transmission in the budding yeast , Saccharomyces cerevisiae [7]. Through genetic and molecular analysis, they verified the importance of Mif2 in chromosome segregation and provided evidence that it is a centromere-associated protein with homology to human CENP-C . Importantly, they also identified several high-copy suppressors of a temperature-sensitive Mif2 mutant allele, including the Smt3 (suppressor of mif two, clone 3) gene that codes for what we now know to be the yeast ortholog of human SUMO1 . Although evidence that Smt3 is conjugated to Mif2 or other centromere-associated proteins was not presented in this early study, it is now recognized that multiple centromere and kinetochore-associated proteins are regulated through SUMOylation and that SUMOylation is essential for accurate chromosome segregation in organisms ranging from yeast to human [8]. In this regard, it is also interesting to note that the SUMO E2- conjugating enzyme, Ubc9 , was also first characterized in 1995 as an essential protein required for chromosome segregation and progression through mitosis in S. cerevisiae [9]. In this study, however, Ubc9 was misidentified as a ubiquitin E2-conjugating enzyme.
Following the identification of the yeast SUMO gene, three studies reporting the identification of cDNAs coding for human SUMO1 appeared in 1996. All three studies involved yeast two-hybrid screens using varying proteins as bait. In the first of these studies, Michael Boddy and colleagues found that SUMO1 interacts with the promyelocytic leukemia (PML) protein and it was therefore referred to as PIC1 (for PML-interacting clone 1) [10]. Again, this study did not present evidence to suggest that SUMO1 is covalently conjugated to PML or any other proteins. However, immunofluorescence microscopy was used to document for the first time what later emerged to be a very important association of SUMO1 with PML nuclear bodies. A large body of subsequent work from many groups has shown that PML is directly SUMOylated and moreover that SUMOylation is vital to the assembly and functions of PML nuclear bodies [11].
The second protein found to interact with human SUMO1 in yeast two-hybrid screens was the Fas/APO-1 receptor [12]. In this study by Edward Yeh and his colleagues, transient overexpression of SUMO1 was shown to protect cells from anti-Fas/APO1-mediated cell death, and thus the protein was named Sentrin (after sentry, because of its guardian effect against cell death). It is now clear that SUMOylation can act at multiple points to affect signal transduction pathways, and particularly through effects on gene expression [13]. The molecular basis for how SUMOylation affects anti-Fas/APO1-mediated cell death, however, remains unknown.
The third protein involved in the early identification of SUMO1 using yeast two-hybrid screens was the DNA recombinase, RAD51 [14]. This study is notable as being the first to suggest a functional link between SUMOylation and DNA damage repair. A large body of work has subsequently established that SUMOylation is intimately involved in nearly all facets of DNA damage repair and that cross talk between SUMOylation and ubiquitylation pathways is crucial for efficient and accurate maintenance of genome integrity [15]. Notably, several studies have specifically reported that non-covalent interactions between RAD51 and SUMO are important for RAD51 recruitment to DNA double-strand breaks, validating the functional significance of this early discovery [16, 17]. In Chapter 2, Wilson and Hochstrasser review in more detail the broad roles of SUMOylation in regulating chromatin structure and function [18].
1.2 Early Discoveries of SUMO Proteins as Posttranslational Protein Modifications
Although an important part of the early discovery of SUMO and the functions of SUMOylation, the above studies all fell short of providing evidence that SUMO proteins are in fact covalently conjugated to other proteins and thus function as posttranslational protein modifications. It did not take long however, for two independent studies, one published in late 1996 by Michael Matunis and colleagues [19] and the other in early 1997 by Frauke Melchior and colleagues [20], to establish for the first time that SUMO1 is covalently and reversibly conjugated to the Ran GTPase-activating protein, RanGAP1. In both studies, antibodies specific to RanGAP1 were found to detect two proteins differing in molecular mass by ~15 kDa. Peptide sequence analysis confirmed the identity of both proteins as forms of RanGAP1, but also revealed the presence of unique peptides specific to the larger protein. Both groups identified expressed sequence tagged (EST) clones in available cDNA databases that encoded the unique peptides and also a predicted 11.5 kDa protein with 18 % sequence identity to ubiquitin. Matunis and colleagues originally referred to the predicted protein as GAP-modifying protein 1 (GMP1) whereas Melchior and colleagues called the protein small ubiquitin-related modifier 1 (SUMO1).
Due to the absence of stop codons 5′ to the predicted methionine in available EST clones, both groups initially considered the possibility that alternative mRNA splicing could explain the two forms of RanGAP1. However, Melchior and colleagues were ultimately able to demonstrate an ATP-dependent interconversion of the higher and lower molecular mass forms of RanGAP1 using isolated cell-free cell extracts. Matunis and colleagues demonstrated that extraction of rat liver nuclear envelopes in the presence of DTT led to the conversion of the higher molecular mass form of RanGAP1 to the lower form, with the concomitant release of the 11.5 kDa SUMO1. This conversion could be inhibited by extraction in the presence of NEM, providing the first evidence for the association of cysteine-dependent SUMO isopeptidases with nuclear pore complexes. Both groups subsequently went on to demonstrate that SUMOylation functions to promote the association of RanGAP1 with Nup358/RanBP2 at the nuclear pore complex, a finding that was also supported by work from Hisato Saitoh, Mary Dasso, and their colleagues [21–23].
These early studies of RanGAP1 SUMOylation established a number of important paradigms that have proven useful for thinking about the functions and regulation of SUMOylation: (1) that SUMO is reversibly conjugated to proteins and affects protein fate through molecular mechanisms similar to ubiquitylation, and (2) that SUMOylation functions to affect protein-protein interactions and assembly of multi-protein complexes, without necessarily affecting protein degradation . At the same time, RanGAP1 has also proven to be a highly unusual SUMO substrate, and in some cases an exception to more universal paradigms. In this regard, RanGAP1 is unusual in that it is stably SUMO1 modified as a consequence of its tight association with Nup358/RanBP2 and consequent protection from deconjugation by isopeptidases [24]. It is now recognized that the majority of sumoylated proteins are modified only transiently, and at relatively low steady-state levels. In addition, RanGAP1 is modified at a single site by a single SUMO1 protein, whereas it is now recognized that proteins can also be modified at multiple sites and by polymeric SUMO chains. It has also become increasingly clear during the past several years that cross talk between the SUMOylation and ubiquitylation pathways includes roles for SUMO as a signal for protein degradation [25].
1.3 Initial Characterization of SUMO Paralogs
Although yeasts and invertebrate organisms express a single SUMO protein, vertebrates possess multiple genes encoding for unique SUMO paralogs. The presence of SUMO paralogs in mammalian cells was first suggested by sequence analysis of human cDNAs, whereby a family of up to three SUMO-related proteins (SUMO1, SUMO2 , and SUMO3 ) was originally identified [14, 26]. A gene coding for SUMO4 was subsequently identified through analysis of single-nucleotide polymorphisms associated with type 1 diabetes [27]. Experimental evidence that SUMO2 and SUMO3 function as posttranslational protein modifications similar to SUMO1 was first provided through transient transfection and overexpression studies that demonstrated modification of the PML protein [28]. Studies by Hisato Saitoh and Joseph Hinchey, however, were the first to report on the analysis of SUMOylation by endogenous SUMO2 and SUMO3 [29]. Notably, this study was also the first to suggest functionally distinct properties for SUMO1 in comparison with SUMO2 and SUMO3, based on their differential activations in response to environmental stresses.
SUMO2 and SUMO3 share ~95 % sequence identity (and are therefore often referred to as SUMO2/3 ) but are only ~45 % identical to SUMO1, further suggestive of possibly distinct signaling properties and functions. Among the sequence differences between SUMO1 and SUMO2/3, perhaps the most significant is the presence of a SUMOylation consensus sequence surrounding lysine 11 that is specific to SUMO2/3. This consensus sequence is efficiently recognized as a SUMO conjugation site, and SUMO2/3 therefore readily form polymeric chains both in vitro and in vivo, as first documented by Tatham and colleagues [30]. Polymeric chains can also form through other lysines in SUMO2/3, and whether chain linkages affect downstream signaling is an important question that remains to be fully evaluated. Polymeric chains have, however, been shown to be functionally distinct from monomeric SUMO due to enhanced affinity for proteins containing tandem SUMO-interacting motifs. Thus, proteins modified by polymeric SUMO2/3 chains are preferentially recognized and ubiquitylated by SUMO-targeted ubiquitin ligases (STUbLs) which contain tandem SUMO-interacting motifs (SIMs) and RING E3 ligase domains [25, 31–33]. Whereas monomeric SUMO1 modification may antagonize ubiquitylation and protein degradation [34], polymeric SUMO2/3 chains have the ability to direct ubiquitin-mediated protein degradation . Consistent with unique signaling properties and functions, multiple studies have also provided evidence for selective modification of proteins by SUMO1 and SUMO2/3 [24, 35–37]. Thus, it was somewhat surprising that gene knockout studies in mice revealed nonessential roles for SUMO1 and SUMO3 expression [38, 39]. Whether these mice have subtle growth defects or conditional phenotypes and whether SUMO paralogs have essential and unique functions in other organisms including humans are important questions for future studies.
2 The Complexity of Protein SUMOylation
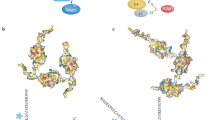
As other members of the ubiquitin family, SUMOs can be attached to target proteins as monomers but also as polymers. PolySUMO-2/3 chains have been identified by mass spectrometry under different stress conditions [40–42]. Furthermore, hybrid SUMO-ubiquitin chains were also reported by several groups [32, 33]. The status of these homologous or heterologous SUMO chains is actively regulated by modifying and de-modifying enzymes in a cellular compartment or time/stimuli-dependent response (Fig. 1). The way these heterologous chains are recognized and connected with distinct functions is still an open domain of investigation. Tandem SUMO-interacting motifs (SIMs) and ubiquitin-interacting motifs (UIMs) appear to play an important role in hybrid chain recognition [43–45]; however, it is not clear if other motifs are also relevant to recognize hybrid chain architectures.
Regulation of protein SUMOylation and chapters contributing to explore these molecular and cellular events. Multiple steps control the status of SUMOylated proteins and its connection with effector functions. The chapters included in this book contribute to explore some of these events using different biological models and systems. This conceptual and methodological framework should contribute to progress in our knowledge of protein SUMOylation and the development of translational research
2.1 SUMO Conjugation Machinery
SUMOs are conjugated to target proteins by an enzymatic cascade involving an activating enzyme (E1) , a conjugating enzyme (E2) , and a ligase (E3) (Fig. 1). The E1 is a heterodimer containing SAE1 and SAE2 subunits (known as Aos1 and Uba2 in yeast ) [46–49]. The E1 catalyzes the formation of SUMO-AMP and the subsequent transfer of SUMO to the E1 active-site cysteine sulfhydryl group. In the second step of the enzyme cascade, SUMO is transferred from the E1 to the active-site cysteine of the E2-conjugating enzyme, Ubc9 [50, 51]. Ubc9 has the ability to directly recognize substrate proteins and catalyze formation of an isopeptide bond between SUMO and the ε-amino group of a lysine in the substrate protein. Alternatively, SUMO E3 ligases may also bind Ubc9 and increase the rate of SUMOylation. Most SUMO E3s appear to target multiple proteins with recognizably similar features, although exact mechanisms of specificity in many cases are not well understood [52]. SUMOylation is highly dynamic, with the removal of SUMO from proteins (deSUMOylation ) being mediated by SUMO-specific proteases /SENtrin proteases (SUSPs/SENPs) that also contribute to the processing of the SUMO precursors [53]. The six members of the SENP family of proteases localize to unique subcellular compartments and the distribution of SUSPs/SENPs is therefore thought to play an important role in the spatial regulation of SUMO turnover and function [53–56].
2.2 SUMO Consensus and Interacting Sequences
Ubc9 recognizes a SUMO consensus motif , ψKxE (where ψ is a large hydrophobic residue and x is any residue, K a lysine, and E/D a glutamic or aspartic amino acid) [57, 58]. SUMOylation of a majority of substrates occurs within this consensus motif; nevertheless ~30 % of proteins are modified on lysine residues not conforming to this consensus sequence [59, 60]. Furthermore, not all proteins containing the ψKxE sequence are SUMOylated, indicating that other factors such as protein structure or localization may influence modification [61, 62]. Interestingly, SUMO-2/3 have functional SUMO consensus motifs used to form polymeric SUMO chains [63]. In addition, polymeric chains are also formed using other non-consensus lysine residues in both SUMO1 and SUMO-2/3 [60, 64]. To avoid the time-consuming approach of systematic mutation of lysine residues on target proteins, bioinformatic tools were developed to identify SUMO consensus sites, including SUMOplot (http://www.abgent.com/tools/sumoplot/) and SUMOsp (http://sumosp.biocuckoo.org/). Unfortunately, since these tools do not consider atypical sequences, structural, temporal, or cellular distribution requirements, predicted SUMOylation sites have not always been confirmed. The contribution of mass spectrometry (MS) approaches has been crucial to identify and/or confirm SUMOylation sites [60] and reviewed in [65]. The identification of conjugation sites has been particularly advanced by recently developed MS-based approaches that allow for enrichment and identification of peptides containing modified lysine residues [66, 67].
Proteins can also interact with SUMO in a non-covalent manner due to the presence of SUMO-interacting motifs (SIMs). The first evidence of SIMs was published by Minty and collaborators in 2000 [68]. Using a two-hybrid approach, the authors observed that some proteins interacted specifically with the SUMOylated version of p73, a member of the p53 family. Further analysis revealed that these interacting proteins contained a common SxS sequence, in which x is any amino acid surrounded by two serine residues, flanked by a hydrophobic core on one side and acidic amino acids on the other. Subsequent studies further confirmed the presence of a Val/Ile-x-Val/Ile-Val/Ile (V/I-x-V/I-V/I) consensus SIM in proteins of varying functions that could facilitate interactions with SUMO [69]. Several enzymes within the SUMO pathway, including the SUMO ligases PIASX and Ran-binding protein 2 (RanBP2 /Nup358) and multiple SENPs [55], contain consensus SIMs, suggesting that non-covalent interactions with SUMO facilitate modification and demodification of substrates. In support of this, the SIM in RanBP2/Nup358 is directly adjacent to the minimal IR1-IR2 domain that has E3 activity. However, although this SIM has been shown to bind SUMO, it does not appear to be essential for ligase activity in vitro [70]. The hydrophobic core of a SIM can bind to an interaction surface on SUMO in parallel or antiparallel orientations. Acidic residues, either upstream or downstream of the core, determine binding orientation and may also affect affinity and paralogue specificity [71]. From these initial SIM studies, a more complex type of SUMO recognition domain, named the SUMO-binding domain (SBDs), containing several hydrophobic cores of 3–4 residues often surrounded by a cluster of acidic amino acids was realized [72, 73]. Recent analysis performed by Hoffman revealed three different types of SIMs with the following PROSITE format: (SIMa) (PILVM)- (ILVM)-x-(ILVM)-(DES>) (3), (SIMb) (PILVM)-(ILVM)-D-L-T, and (SIMr) (DSE) (3)-(ILVM)-x-(ILVMF) (2) [74]. The identification and validation of these SIMs using site-directed mutagenesis has been an important approach to investigate the role of SUMO in the regulation of particular processes or pathways. The identification of new SIMs will be crucial for the integration of the many functions regulated by SUMOylation.
2.3 Diversity of SUMO E3s
The Siz proteins in S. cerevisiae were the first SUMO E3s identified. These ligases have domains that are homologous to the RING domains of ubiquitin E3 ligases [75]. Co-deletion of Siz1 and Siz2 genes in S. cerevisiae eliminates most SUMOylation and affects growth under a variety of conditions, underlining the importance of these SUMO E3 ligases [76, 77]. The protein inhibitors of activated STAT (PIAS) proteins are homologs of the Siz proteins in higher eukaryotes. However, PIAS proteins appear to play additional roles apart from being SUMO E3 ligases [78]. In humans the five PIAS proteins (PIAS1 , PIASxα , PIASxβ , PIASγ , and PIAS3 ) encoded by the genome contain the RING domain. Individual deletion of these genes results in distinct phenotypes. Using PIAS1−/− mice it was demonstrated that PIAS1 regulates interferon-inducible gene expression and is important in innate immunity [79]. Nevertheless, there was no detectable impact of PIAS1 deletion on total SUMOylation patterns as compared to WT mice. A similar situation was observed with the PIASγ−/− mice, where mild defects in transcriptional responses induced by interferon γ and Wnt agonists were also observed [80, 81]. Although PIAS E3 ligases show some redundancy in vitro and in transient overexpression studies, differences in substrate preferences and regulation likely exist under normal in vivo conditions.
The SUMO-specific ligases RanBP2 and polycomb group protein 2 (Pc2) are unrelated to known ubiquitin E3s. RanBP2/NUP358 is located at nuclear pore complexes and enhances Ubc9 -mediated conjugation of SUMO-1 and SUMO-2/3 to a variety of protein substrates. For instance, it enhances SUMOylation of Sp100 and histone deacetylase 4 (HDAC4) by SUMO-1 and preferentially modifies PML with SUMO-2 [30, 82, 83]. The domain of RanBP2 that contains SUMO E3 ligase activity includes the IR1, M1, and IR2 regions involved in binding Ubc9 [30, 84, 85]. The lack of interaction between the RanBP2 ligase domain and substrates indicates that RanBP2 alters the structure of the SUMO-Ubc9 thioester , thereby increasing the capacity to transfer SUMO to protein substrates. However, the mechanism used by RanBP2 to enhance SUMO1 or SUMO2/3 modification of substrates is distinct since M-IR2 binds SUMO1 but not SUMO2 .
The Pc2 component of the polycomb chromatin-modifying complex also possesses SUMO E3 ligase activity. One Pc2 substrate is the C-terminal-binding protein (CtBP) transcriptional co-repressor [86]. The N-terminal region of Pc2 alone binds Ubc9 and exhibits E3 ligase activity in vitro. However, the C-terminal region that binds CtBP is also required for activity in vivo. Although the mechanism of action of this SUMO E3 is not completely clear, it is likely that the C-terminal domain of Pc2 functions to recruit CtBP to PcG subnuclear domains where the active N-terminal domain recruits Ubc9 and drives SUMOylation of CtBP [86].
2.4 Regulation of SUMOylation by Specific Proteases
Protein SUMOylation is reversible and is removed from targets by specific cysteine proteases known as SUMO-specific proteases or SUMO isopeptidases. These enzymes remove SUMO from protein conjugates and depolymerize poly-SUMO chains. Some of these enzymes also function to process SUMO precursors by cleaving and releasing carboxy-terminal residues, thereby exposing the signature double glycine required for SUMO conjugation. SUMO-specific isopeptidases/proteases are classified into three families: the Ulp/SENP (ubiquitin-like protease/sentrin -specific protease) family, the Desi (deSUMOylating isopeptidase) family, and USPL1 (ubiquitin-specific peptidase-like protein 1) [53].
Ulp1 /Ulp2 , discovered in Saccharomyces cerevisiae , belong to the C48 family of thiol proteases [87, 88]. In higher eukaryotes, the family includes six enzymes called SENPs 1–3 and 5–7 [89]. SENP8 acts on the ubiquitin family member NEDD8, but not on SUMO paralogs [90, 91]. The catalytic domain of the Ulp/SENP family spans ~200 amino acids in the carboxy-terminal part of the enzyme. The catalytic domains of the human Ulp/SENP family members share 20–60 % sequence identity. The amino-terminal regions of all SENPs contain amino acids susceptible to phosphorylation or ubiquitylation , modifications that may affect their stability or interactions with substrates or adaptor proteins that determine their subcellular distribution [92–98]. The N-terminal domains of most SENPs also contain one or more SIMs which are believed to contribute to the recognition of SUMOylated substrates.
The deSUMOylating isopeptidases Desi-1 and Desi-2 belong to the C97 family of cysteine proteases [99]. No orthologs of Desi-1 and Desi-2 have been described in yeast . USPL1 is the only mammalian SUMO-specific protease of the C98 family [100]. Desi-1 and Desi-2 are small proteins characterized by PPPDE (permutated papain fold peptidases of the double-stranded RNA viruses and eukaryotes) domains of around 140 amino acids. Desi-1 and Desi-2 share about 20 % sequence identity within this region. The active site contains two conserved cysteine and histidine residues that form a catalytic dyad [99, 101]. The catalytic domain of USPL1 shows homology to the C19 family of ubiquitin-specific proteases. The catalytic domain of USPL1 contains a catalytic triad composed of Cys-His-Asp residues [100, 102].
SUMO isopeptidases show distinct subcellular distributions that limit their activity to specific sets of substrates. Ulp/SENP family members are mainly located in distinct sub-nuclear regions. SENP1 and SENP2 , however, also shuttle between the nucleus and the cytoplasm and are concentrated at the nuclear envelope through their interaction with components of the nuclear pore complex [98, 103–106]. SENP1 and SENP2 are excluded from the nucleolus, but can be detected in nuclear foci that show some overlap with PML nuclear bodies . SENP1 and SENP2 redistribute during mitosis from the nuclear envelope to the kinetochore [13]. SENP3 and SENP5 are located in sub-compartments of the nucleolus, where they act on proteins involved in the early steps of ribosome maturation [92, 107–109]. However, a small fraction of SENP3 and SENP5 also reside in the nucleoplasm and the cytoplasm. SENP5 translocates to the mitochondrial surface during the G2/M transition prior to nuclear envelope breakdown [110]. SENP6 and SENP7 mainly exhibit a nucleoplasmic distribution. Desi family members are primarily concentrated in the cytoplasm [110]. USPL1 is a predominantly nuclear protein and co-localizes with coilin in Cajal bodies [100, 102].
3 Methodologies to Study Protein SUMOylation
Due to the expanding interest in the study of protein SUMOylation in distinct fields, methodologies to improve our understanding of the functions regulated by this posttranslational modification are in constant innovation. Collected here are some recent methodologies that have been developed and used by well-recognized SUMO experts. Many of the reviewed techniques/approaches are versatile and can be adapted to different biological models or in cellulo or in vitro systems. The classification of included methods is somewhat practical and refers to one possible application. However, most techniques can be adapted according to the needs of specific projects and used in different in vitro, in cellulo, or in vivo models.
The first section of this book includes in vitro procedures to study protein SUMOylation. Considering the complexity of the SUMOylation analysis in vivo, in vitro procedures provide simplified systems that are more ideally suited to address mechanistic questions. The complete reconstitution of the recombinant RanBP2 SUMO E3 ligase complex proposed by Ritterhoff et al., in Chapter 3 [111], allows for quantitative SUMOylation of RanGAP1 but can be extrapolated to other RanBP2 substrates [112, 113]. A protocol to purify recombinant SUMOylated proteins from bacteria , as outlined by Brockly et al., in Chapter 4 [114], can be used to gain insights into biochemical aspects of specific SUMOylation targets [115–117]. Eisenhardt et al. present in Chapter 5 [118] a fluorescent -SUMO conjugation assay to evaluate E3-mediated chain formation activity in a paralog-specific manner. The application of fluorescent assays to study substrate modification in vitro provides fast procedures to investigate SUMO enzyme activities and mechanistic insights into SUMO chain formation [119]. Once SUMOylated, target proteins are recognized by effector proteins containing functional SIMs. SUMO-SIM interactions are far from being fully understood and deeper exploration is needed to better understand the molecular mechanisms regulating this connection. In Chapter 6, Husnjak et al. [120] describe two complementary approaches to identify SUMOylated proteins and characterize their interactions with SIMs. Their method has been validated and successfully applied to the identification of novel SUMO-binding proteins as well as the characterization of known SUMO-interacting modules [68, 72, 73, 121–123]. A complementary and quantitative method to characterize real-time SUMO-SIM interactions using surface plasmon resonance is outlined by Xolalpa et al. in Chapter 7 [124]. This method can be used to analyze the effect of SUMO or SIM point mutations, or regulatory proteins, on SUMO-SIM interactions. The analysis of SUMOylated proteins can also be carried out using chimeric SIMs arranged in tandem, also known as SUMO-traps [44, 125, 126]. In the approach proposed by Lang et al. in Chapter 8, [127] a biotinylated version of SUMO-traps is used to analyze SUMO substrates in vitro, but can also be used for in vivo studies. SUMO-SIM interactions are transient, in part due to the action of SUMO-specific proteases. To study the activity of SUMO proteases, Eckhoff and Dohmen have developed a method for fast and economic analysis [128]. The method reported in Chapter 9 [129] was developed for analysis of S. cerevisiae Ulp enzymes but can be expanded to SUMO-specific proteases from other species. Since the proteases are key regulatory molecules of protein SUMOylation, their inhibition represents not only a desirable approach to better characterize their functions, but also opens possibilities for clinical intervention.
Protocols for the analysis of SUMOylated proteins in cell cultures are grouped in the second part of this book. Diverse and imaginative approaches use chimeric proteins and other sophisticated strategies to identify SUMO targets and interacting cellular factors. A method proposed by Sahin et al. in Chapter 10 [130] allows detection of protein SUMOylation in situ by the now popular technique of proximity ligation assay (PLA) [131]. Yuasa and Saitoh present in Chapter 11 [132] an alternative technique to detect in situ protein SUMOylation and de-SUMOylation using fluorescence-based assays in permeabilized cells [133]. The analysis of total or individual SUMOylated proteins in cell lysates, but also in vivo, can be performed using tagged SUMO proteins. Some of the most popular tags are biotin [134, 135] and histidine [67, 136], as described by Pirone et al. in Chapter 12 [137] and Hendriks and Vertagaal in Chapter 13 [138]. The detailed protocols can be adapted to detect any SUMOylated target and can also be used for global MS analysis of SUMO conjugation signatures. One of the major bottlenecks to MS analysis of SUMOylated proteins is the enrichment of the SUMO-GG signature peptides, which are longer than those generated by tryptic digestion of ubiquitinylated proteins. Multiple approaches have been developed to address this issue, including the mutation of C-terminal amino acid residues in SUMO to generate shorter tryptic GG-peptide signatures. Alternatively, longer His10-tags have been developed to allow a single-step, high-yield purification of SUMOylated proteins which can then be digested and analyzed by high-resolution MS analysis, as described in Chapter 13 [138].
SIMs in downstream effector proteins function to integrate SUMOylation with specific cellular processes. For this reason, Aguilar-Martínez and Sharrocks used multimeric protein scaffolds to identify novel multi-SUMO-binding proteins, as outlined in Chapter 14 [139]. The isolation and identification of SUMOylated proteins associated specifically with chromatin represents an important challenge. Bawa-Khalfe report in Chapter 15 [140] a protocol for effectively purifying endogenous SUMOylated proteins from chromatin fractions prepared from cultured cells [141]. This approach has the potential to be used to evaluate chromatin-bound SUMO targets using varying cellular models and biological systems.
The final section of this book includes methods to study protein SUMOylation using distinct biological models. The SUMO pathway often targets protein groups that are functionally and physically connected [142, 143]. Psakhye and Jentsch present a method to identify SUMOylated protein groups in Saccharomyces cerevisiae in Chapter 16 [144]. This protocol can be easily adapted for studies of SUMOylation in mammalian cells. Caenorhabditis elegans represents another powerful genetic system to study protein SUMOylation, and Pelish and Hay expand in Chapter 17 [145] the existing set of tools to investigate the role of SUMOylation using this nematode [146]. These tools and reagents allow a combination of genetics, imaging, and biochemical approaches that will be useful to gain insights into the biological role of SUMOylation in the context of this multicellular organism [146]. Another attractive system to study protein SUMOylation is the model plant Arabidopsis thaliana . Based on the expression of modified SUMOs bearing epitope tags, Rytz and Vierstra combine standard and quantitative MS analysis methods to identify SUMOylated proteins, as detailed in Chapter 18 [147]. The role of protein SUMOylation during multiple human infections , including viral, bacterial, and parasitic infections, also has an increasing interest for microbiologist and immunologists. In Chapter 19 [148], Reiter and Matunis present methods to improve the functional analysis of protein SUMOylation in Plasmodium falciparum using antibodies specific for the parasite SUMO [149]. Considering that SUMOylation is essential, a more detailed understanding of its role during the parasite life cycle will be required for the further development of antimalarial drugs targeting SUMOylation. Preclinical studies often use rodent models to validate information collected from other experimental systems. In Chapter 20 [150], Tirad and Brose describe step-by-step methods to purify and analyze SUMO1-modified proteins from His6-HA-SUMO1 knock-in mouse brain based on an anti-HA immunopurification protocol. These methods are generally applicable and can be easily adapted to other cell types and tissues. This His6-HA-SUMO1 mouse line can be crossed into any disease model, thus providing the opportunity to study SUMO1 conjugation in a plethora of disease-relevant processes. These are clear advantages that make using His6-HA-SUMO1 knock-in mice a powerful model system for the analysis of SUMOylation [151].
4 Conclusions and Future Directions
After the first 20 years of study, it has become clear that SUMOylation is linked to virtually all cellular process, including intracellular transport, transcription, DNA replication and repair, chromatin assembly/accessibility and proteolysis , among others. All these processes can be fine-tuned through the action of SUMO-conjugating/de-conjugating enzymes that are activated during physiological and pathological events. How SUMO moieties are integrated into hybrid chains containing ubiquitin or other ubiquitin-like proteins (e.g., NEDD8) is still under intense investigation. One of the major bottlenecks to analysis of hybrid chains is that the available MS technologies do not reveal the order in which individual moieties are integrated into the chains, as these technologies read a single-branched peptide at a time. Thus, new tools and technological implementations will continue to be key for progress in the SUMO and other ubiquitin-related protein fields. In sum, since its original discoveries, the roles for protein SUMOylation have expanded in extraordinary and unanticipated ways. It will surely be fascinating to see how the field develops during the next two decades.
References
Ciechanover A, Hod Y, Hershko A (1978) A heat-stable polypeptide component of an ATP-dependent proteolytic system from reticulocytes. Biochem Biophys Res Commun 81:1100–1105
Ciechanover A, Heller H, Elias S et al (1980) ATP-dependent conjugation of reticulocyte proteins with the polypeptide required for protein degradation. Proc Natl Acad Sci U S A 77:1365–1368
Ciechanover A (2012) Intracellular protein degradation: from a vague idea through the lysosome and the ubiquitin-proteasome system and onto human diseases and drug targeting. Neurodegener Dis 10:7–22. doi:10.1159/000334283
Vogel G (2004) Nobel prizes. Gold medal from cellular trash. Science 306:400–401. doi:10.1126/science.306.5695.400b
Geiss-Friedlander R, Melchior F (2007) Concepts in sumoylation: a decade on. Nat Rev Mol Cell Biol 8:947–956. doi:10.1038/nrm2293
van der Veen AG, Ploegh HL (2012) Ubiquitin-like proteins. Annu Rev Biochem 81:323–357. doi:10.1146/annurev-biochem-093010-153308
Meluh PB, Koshland D (1995) Evidence that the MIF2 gene of Saccharomyces cerevisiae encodes a centromere protein with homology to the mammalian centromere protein CENP-C. Mol Biol Cell 6:793–807
Eifler K, Vertegaal AC (2015) SUMOylation-mediated regulation of cell cycle progression and cancer. Trends Biochem Sci 40:779–793. doi:10.1016/j.tibs.2015.09.006
Seufert W, Futcher B, Jentsch S (1995) Role of a ubiquitin-conjugating enzyme in degradation of S- and M-phase cyclins. Nature 373:78–81. doi:10.1038/373078a0
Boddy MN, Howe K, Etkin LD et al (1996) PIC 1, a novel ubiquitin-like protein which interacts with the PML component of a multiprotein complex that is disrupted in acute promyelocytic leukaemia. Oncogene 13:971–982
Sahin U, de The H, Lallemand-Breitenbach V (2014) PML nuclear bodies: assembly and oxidative stress-sensitive sumoylation. Nucleus 5:499–507. doi:10.4161/19491034.2014.970104
Okura T, Gong L, Kamitani T et al (1996) Protection against Fas/APO-1- and tumor necrosis factor-mediated cell death by a novel protein, sentrin. J Immunol 157:4277–4281
Cubenas-Potts C, Matunis MJ (2013) SUMO: a multifaceted modifier of chromatin structure and function. Dev Cell 24:1–12. doi:10.1016/j.devcel.2012.11.020
Shen Z, Pardington-Purtymun PE, Comeaux JC et al (1996) UBL1, a human ubiquitin-like protein associating with human RAD51/RAD52 proteins. Genomics 36:271–279
Jackson SP, Durocher D (2013) Regulation of DNA damage responses by ubiquitin and SUMO. Mol Cell 49:795–807. doi:10.1016/j.molcel.2013.01.017
Shima H, Suzuki H, Sun J et al (2013) Activation of the SUMO modification system is required for the accumulation of RAD51 at sites of DNA damage. J Cell Sci 126:5284–5292. doi:10.1242/jcs.133744
Ouyang KJ, Woo LL, Zhu J et al (2009) SUMO modification regulates BLM and RAD51 interaction at damaged replication forks. PLoS Biol 7:e1000252. doi:10.1371/journal.pbio.1000252
Wilson N, Hochstrasser M (2016) The regu- lation of chromatin by dynamic SUMO modifications. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_2
Matunis MJ, Coutavas E, Blobel G (1996) A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J Cell Biol 135:1457–1470
Mahajan R, Delphin C, Guan T et al (1997) A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88:97–107
Saitoh H, Pu R, Cavenagh M, Dasso M (1997) RanBP2 associates with Ubc9p and a modified form of RanGAP1. Proc Natl Acad Sci U S A 94:3736–3741
Matunis MJ, Wu J, Blobel G (1998) SUMO-1 modification and its role in targeting the Ran GTPase-activating protein, RanGAP1, to the nuclear pore complex. J Cell Biol 140:499–509
Mahajan R, Gerace L, Melchior F (1998) Molecular characterization of the SUMO-1 modification of RanGAP1 and its role in nuclear envelope association. J Cell Biol 140:259–270
Zhu S, Goeres J, Sixt KM et al (2009) Protection from isopeptidase-mediated deconjugation regulates paralog-selective sumoylation of RanGAP1. Mol Cell 33:570–580. doi:10.1016/j.molcel.2009.02.008
Sriramachandran AM, Dohmen RJ (2014) SUMO-targeted ubiquitin ligases. Biochim Biophys Acta 1843:75–85. doi:10.1016/j.bbamcr.2013.08.022
Lapenta V, Chiurazzi P, van der Spek P et al (1997) SMT3A, a human homologue of theS. cerevisiae SMT3gene, maps to chromosome 21qter and defines a novel gene family. Genomics 40:362–366
Guo D, Li M, Zhang Y et al (2004) A functional variant of SUMO4, a new IkBα modifier, is associated with type 1 diabetes. Nat Genet 36:837–841
Kamitani T, Nguyen HP, Kito K et al (1998) Covalent modification of PML by the sentrin family of ubiquitin-like proteins. J Biol Chem 273:3117–3120
Saitoh H, Hinchey J (2000) Functional heterogeneity of small ubiquitin-related protein modifiers SUMO-1 versus SUMO-2/3. J Biol Chem 275:6252–6258
Tatham MH, Kim S, Jaffray E et al (2005) Unique binding interactions among Ubc9, SUMO and RanBP2 reveal a mechanism for SUMO paralog selection. Nat Struct Mol Biol 12:67–74
Sun H, Leverson JD, Hunter T (2007) Conserved function of RNF4 family proteins in eukaryotes: targeting a ubiquitin ligase to SUMOylated proteins. EMBO J 26:4102–4112
Tatham MH, Geoffroy M-C, Shen L et al (2008) RNF4 is a poly-SUMO-specific E3 ubiquitin ligase required for arsenic-induced PML degradation. Nat Cell Biol 10:538–546
Lallemand-Breitenbach V, Jeanne M, Benhenda S et al (2008) Arsenic degrades PML or PML–RARα through a SUMO-triggered RNF4/ubiquitin-mediated pathway. Nat Cell Biol 10:547–555
Desterro JM, Rodriguez MS, Hay RT (1998) SUMO-1 modification of IkappaBalpha inhibits NF-kappaB activation. Mol Cell 2:233–239
Ayaydin F, Dasso M (2004) Distinct in vivo dynamics of vertebrate SUMO paralogues. Mol Biol Cell 15:5208–5218
Vertegaal AC, Andersen JS, Ogg SC et al (2006) Distinct and overlapping sets of SUMO-1 and SUMO-2 target proteins revealed by quantitative proteomics. Mol Cell Proteomics 5:2298–2310
Zhang X-D, Goeres J, Zhang H et al (2008) SUMO-2/3 modification and binding regulate the association of CENP-E with kinetochores and progression through mitosis. Mol Cell 29:729–741
Zhang F-P, Mikkonen L, Toppari J et al (2008) Sumo-1 function is dispensable in normal mouse development. Mol Cell Biol 28:5381–5390
Wang L, Wansleeben C, Zhao S et al (2014) SUMO2 is essential while SUMO3 is dispensable for mouse embryonic development. EMBO Rep 15(8):878–885
Vertegaal AC, Ogg SC, Jaffray E et al (2004) A proteomic study of SUMO-2 target proteins. J Biol Chem 279(32):33791–33798
Seifert A, Schofield P, Barton GJ, Hay RT (2015) Proteotoxic stress reprograms the chromatin landscape of SUMO modification. Sci Signal 8:rs7
Bursomanno S, Beli P, Khan AM et al (2015) Proteome-wide analysis of SUMO2 targets in response to pathological DNA replication stress in human cells. DNA Repair 25:84–96
Bruderer R, Tatham MH, Plechanovova A et al (2011) Purification and identification of endogenous polySUMO conjugates. EMBO Rep 12:142–148
Da Silva-Ferrada E, Xolalpa W, Lang V et al (2013) Analysis of SUMOylated proteins using SUMO-traps. Sci Rep. doi:10.1038/srep01690
Guzzo CM, Matunis MJ (2013) Expanding SUMO and ubiquitin-mediated signaling through hybrid SUMO-ubiquitin chains and their receptors. Cell Cycle 12:1015–1017
Desterro JM, Rodriguez MS, Kemp GD, Hay RT (1999) Identification of the enzyme required for activation of the small ubiquitin-like protein SUMO-1. J Biol Chem 274:10618–10624
Gong L, Li B, Millas S, Yeh ET (1999) Molecular cloning and characterization of human AOS1 and UBA2, components of the sentrin-activating enzyme complex. FEBS Lett 448:185–189
Johnson ES, Schwienhorst I, Dohmen RJ, Blobel G (1997) The ubiquitin-like protein Smt3p is activated for conjugation to other proteins by an Aos1p/Uba2p heterodimer. EMBO J 16:5509–5519
Okuma T, Honda R, Ichikawa G et al (1999) In vitroSUMO-1 modification requires two enzymatic steps, E1 and E2. Biochem Biophys Res Commun 254:693–698
Desterro JM, Thomson J, Hay RT (1997) Ubch9 conjugates SUMO but not ubiquitin. FEBS Lett 417:297–300
Johnson ES, Blobel G (1997) Ubc9p is the conjugating enzyme for the ubiquitin-like protein Smt3p. J Biol Chem 272:26799–26802
Melchior F, Schergaut M, Pichler A (2003) SUMO: ligases, isopeptidases and nuclear pores. Trends Biochem Sci 28:612–618
Nayak A, Müller S (2014) SUMO-specific proteases/isopeptidases: SENPs and beyond. Genome Biol 15:422
Hay RT (2007) SUMO-specific proteases: a twist in the tail. Trends Cell Biol 17:370–376
Mukhopadhyay D, Dasso M (2007) Modification in reverse: the SUMO proteases. Trends Biochem Sci 32:286–295
Cubeñas-Potts C, Goeres JD, Matunis MJ (2013) SENP1 and SENP2 affect spatial and temporal control of sumoylation in mitosis. Mol Biol Cell 24:3483–3495
Rodriguez MS, Dargemont C, Hay RT (2001) SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J Biol Chem 276:12654–12659
Sampson DA, Wang M, Matunis MJ (2001) The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J Biol Chem 276:21664–21669
Tammsalu T, Matic I, Jaffray EG et al (2014) Proteome-wide identification of SUMO2 modification sites. Sci Signal 7:rs2
Matic I, Schimmel J, Hendriks IA et al (2010) Site-specific identification of SUMO-2 targets in cells reveals an inverted SUMOylation motif and a hydrophobic cluster SUMOylation motif. Mol Cell 39:641–652
Lin D, Tatham MH, Yu B et al (2002) Identification of a substrate recognition site on Ubc9. J Biol Chem 277:21740–21748
Tatham MH, Kim S, Yu B et al (2003) Role of an N-terminal site of Ubc9 in SUMO-1,-2, and-3 binding and conjugation. Biochemistry (Mosc) 42:9959–9969
Tatham MH, Jaffray E, Vaughan OA et al (2001) Polymeric chains of SUMO-2 and SUMO-3 are conjugated to protein substrates by SAE1/SAE2 and Ubc9. J Biol Chem 276:35368–35374
Golebiowski F, Matic I, Tatham MH et al (2009) System-wide changes to SUMO modifications in response to heat shock. Sci Signal 2:24
Da Silva-Ferrada E, Lopitz-Otsoa F, Lang V et al (2012) Strategies to identify recognition signals and targets of SUMOylation. Biochem Res Int 2012:875148
Tammsalu T, Matic I, Jaffray EG et al (2015) Proteome-wide identification of SUMO modification sites by mass spectrometry. Nat Protoc 10:1374–1388
Hendriks IA, D’Souza RC, Chang J-G et al (2015) System-wide identification of wild-type SUMO-2 conjugation sites. Nat Commun 6:7289
Minty A, Dumont X, Kaghad M, Caput D (2000) Covalent Modification of p73α by SUMO-1 two-hybrid screening with p73 identifies novel SUMO-1-interacting proteins and a SUMO-1 interaction motif. J Biol Chem 275:36316–36323
Song J, Durrin LK, Wilkinson TA et al (2004) Identification of a SUMO-binding motif that recognizes SUMO-modified proteins. Proc Natl Acad Sci U S A 101:14373–14378
Merrill JC, Melhuish TA, Kagey MH et al (2010) A role for non-covalent SUMO interaction motifs in Pc2/CBX4 E3 activity. PLoS One 5:e8794
Song J, Zhang Z, Hu W, Chen Y (2005) Small Ubiquitin-like Modifier (SUMO) Recognition of a SUMO Binding Motif a reversal of the bound orientation. J Biol Chem 280:40122–40129
Hecker C-M, Rabiller M, Haglund K et al (2006) Specification of SUMO1-and SUMO2-interacting motifs. J Biol Chem 281:16117–16127
Hannich JT, Lewis A, Kroetz MB et al (2005) Defining the SUMO-modified proteome by multiple approaches in Saccharomyces cerevisiae. J Biol Chem 280:4102–4110
Vogt B, Hofmann K (2012) Bioinformatical detection of recognition factors for ubiquitin and SUMO. In: Ubiquitin Fam. Modif. Proteasome. Springer. pp 249–261
Hochstrasser M (2001) SP-RING for SUMO: new functions bloom for a ubiquitin-like protein. Cell 107:5–8
Johnson ES, Gupta AA (2001) An E3-like factor that promotes SUMO conjugation to the yeast septins. Cell 106:735–744
Takahashi Y, Kahyo T, Toh-e A et al (2001) Yeast Ull1/Siz1 is a novel SUMO1/Smt3 ligase for septin components and functions as an adaptor between conjugating enzyme and substrates. J Biol Chem 276:48973–48977
Rytinki MM, Kaikkonen S, Pehkonen P et al (2009) PIAS proteins: pleiotropic interactors associated with SUMO. Cell Mol Life Sci 66:3029–3041
Liu B, Mink S, Wong KA et al (2004) PIAS1 selectively inhibits interferon-inducible genes and is important in innate immunity. Nat Immunol 5:891–898
Roth W, Sustmann C, Kieslinger M et al (2004) PIASy-deficient mice display modest defects in IFN and Wnt signaling. J Immunol 173:6189–6199
Wong KA, Kim R, Christofk H et al (2004) Protein inhibitor of activated STAT Y (PIASy) and a splice variant lacking exon 6 enhance sumoylation but are not essential for embryogenesis and adult life. Mol Cell Biol 24:5577–5586
Kirsh O, Seeler J-S, Pichler A et al (2002) The SUMO E3 ligase RanBP2 promotes modification of the HDAC4 deacetylase. EMBO J 21:2682–2691
Pichler A, Melchior F (2002) Ubiquitin-related modifier SUMO1 and nucleocytoplasmic transport. Traffic 3:381–387
Saitoh H, Pizzi MD, Wang J (2002) Perturbation of SUMOlation enzyme Ubc9 by distinct domain within nucleoporin RanBP2/Nup358. J Biol Chem 277:4755–4763
Pichler A, Knipscheer P, Saitoh H et al (2004) The RanBP2 SUMO E3 ligase is neither HECT-nor RING-type. Nat Struct Mol Biol 11:984–991
Kagey MH, Melhuish TA, Powers SE, Wotton D (2005) Multiple activities contribute to Pc2 E3 function. EMBO J 24:108–119
Li S-J, Hochstrasser M (1999) A new protease required for cell-cycle progression in yeast. Nature 398:246–251
Li S-J, Hochstrasser M (2000) The yeast ULP2 (SMT4) gene encodes a novel protease specific for the ubiquitin-like Smt3 protein. Mol Cell Biol 20:2367–2377
Yeh ET, Gong L, Kamitani T (2000) Ubiquitin-like proteins: new wines in new bottles. Gene 248:1–14
Gan-Erdene T, Nagamalleswari K, Yin L et al (2003) Identification and characterization of DEN1, a deneddylase of the ULP family. J Biol Chem 278:28892–28900
Mendoza HM, Shen L, Botting C et al (2003) NEDP1, a highly conserved cysteine protease that deNEDDylates Cullins. J Biol Chem 278:25637–25643
Nishida T, Tanaka H, Yasuda H (2000) A novel mammalian Smt3-specific isopeptidase 1 (SMT3IP1) localized in the nucleolus at interphase. Eur J Biochem 267:6423–6427
Garvin AJ, Densham RM, Blair-Reid SA et al (2013) The deSUMOylase SENP7 promotes chromatin relaxation for homologous recombination DNA repair. EMBO Rep 14:975–983
Guo C, Hildick KL, Luo J et al (2013) SENP3-mediated deSUMOylation of dynamin-related protein 1 promotes cell death following ischaemia. EMBO J 32:1514–1528
Klein UR, Haindl M, Nigg EA, Muller S (2009) RanBP2 and SENP3 function in a mitotic SUMO2/3 conjugation-deconjugation cycle on Borealin. Mol Biol Cell 20:410–418
Kuo M-L, den Besten W, Thomas MC, Sherr CJ (2008) Arf-induced turnover of the nucleolar nucleophosmin-associated SUMO-2/3 protease Senp3. Cell Cycle 7:3378–3387
Wang Y, Yang J, Yang K et al (2012) The biphasic redox sensing of SENP3 accounts for the HIF-1 transcriptional activity shift by oxidative stress. Acta Pharmacol Sin 33:953–963
Hang J, Dasso M (2002) Association of the human SUMO-1 protease SENP2 with the nuclear pore. J Biol Chem 277:19961–19966
Shin EJ, Shin HM, Nam E et al (2012) DeSUMOylating isopeptidase: a second class of SUMO protease. EMBO Rep 13:339–346
Schulz S, Chachami G, Kozaczkiewicz L et al (2012) Ubiquitin-specific protease-like 1 (USPL1) is a SUMO isopeptidase with essential, non-catalytic functions. EMBO Rep 13:930–938
Suh H-Y, Kim J-H, Woo J-S et al (2012) Crystal structure of DeSI-1, a novel deSUMOylase belonging to a putative isopeptidase superfamily. Proteins 80:2099–2104
Hutten S, Chachami G, Winter U et al (2014) A role for the Cajal-body-associated SUMO isopeptidase USPL1 in snRNA transcription mediated by RNA polymerase II. J Cell Sci 127:1065–1078
Goeres J, Chan P-K, Mukhopadhyay D et al (2011) The SUMO-specific isopeptidase SENP2 associates dynamically with nuclear pore complexes through interactions with karyopherins and the Nup107-160 nucleoporin subcomplex. Mol Biol Cell 22:4868–4882
Takahashi Y, Mizoi J, Toh-e A, Kikuchi Y (2000) Yeast Ulpl, an Smt3-specific protease, associates with nucleoporins. J Biochem (Tokyo) 128:723–725
Zhang H, Saitoh H, Matunis MJ (2002) Enzymes of the SUMO modification pathway localize to filaments of the nuclear pore complex. Mol Cell Biol 22:6498–6508
Itahana Y, Yeh ET, Zhang Y (2006) Nucleocytoplasmic shuttling modulates activity and ubiquitination-dependent turnover of SUMO-specific protease 2. Mol Cell Biol 26:4675–4689
Gong L, Yeh ET (2006) Characterization of a family of nucleolar SUMO-specific proteases with preference for SUMO-2 or SUMO-3. J Biol Chem 281:15869–15877
Haindl M, Harasim T, Eick D, Muller S (2008) The nucleolar SUMO-specific protease SENP3 reverses SUMO modification of nucleophosmin and is required for rRNA processing. EMBO Rep 9:273–279
Yun C, Wang Y, Mukhopadhyay D et al (2008) Nucleolar protein B23/nucleophosmin regulates the vertebrate SUMO pathway through SENP3 and SENP5 proteases. J Cell Biol 183:589–595
Zunino R, Braschi E, Xu L, McBride HM (2009) Translocation of SenP5 from the nucleoli to the mitochondria modulates DRP1-dependent fission during mitosis. J Biol Chem 284:17783–17795
Ritterhoff T, Das H, Hao Y, et al (2016) Reconstitution of the recombinant RanBP1 SUMO E3 ligase complex. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_3
Werner A, Flotho A, Melchior F (2012) The RanBP2/RanGAP1*SUMO1/Ubc9 complex is a multisubunit SUMO E3 ligase. Mol Cell 46(3):287–98. doi:10.1016/j.molcel.2012.02.017
Sakin V1, Richter SM1, Hsiao HH2, Urlaub H3, Melchior F4 (2015) Sumoylation of the GTPase Ran by the RanBP2 SUMO E3 Ligase Complex. J Biol Chem 290(39):23589–602. doi:10.1074/jbc.M115.660118
Brockly F, Piechaczyk M, Bossis G (2016) Production and purification of recombinant SUMOylated proteins using engineered bacteria. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_4
Uchimura Y, Nakamura M, Sugasawa K et al (2004) Overproduction of eukaryotic SUMO-1-and SUMO-2-conjugated proteins in Escherichia coli. Anal Biochem 331:204–206
Lens Z, Dewitte F, Van Lint C et al (2011) Purification of SUMO-1 modified IkBα and complex formation with NF-kB. Protein Expr Purif 80:211–216
Weber AR, Schuermann D, Schär P (2014) Versatile recombinant SUMOylation system for the production of SUMO-modified protein. PLoS One 9(7):e102157
Eisenhardt N, Chaugule VK, Pichler A (2016) A fluorescent in vitro assay to investigate paralog specific SUMO conjugation. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_5
Eisenhardt N, Chaugule VK, Koidl S et al (2015) A new vertebrate SUMO enzyme family reveals insights into SUMO-chain assembly. Nat Struct Mol Biol 22:959–967
Husnjak K, Keiten-Schmitz J, Muller S (2016) Identification and characterization of SUMO- SIM interactions. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_6
Stehmeier P, Muller S (2009) Phospho-regulated SUMO interaction modules connect the SUMO system to CK2 signaling. Mol Cell 33:400–409
Kroetz M, Hochstrasser M (2009) Identification of SUMO-interacting proteins by yeast two-hybrid analysis. In: Ulrich H (ed) SUMO Protoc. Humana, Totowa, pp 107–120
Pfander B, Moldovan G-L, Sacher M, et al. (2005) SUMO-modified PCNA recruits Srs2 to prevent recombination during S phase. nature 436:428–433.
Xolalpa W, Rodriguez MS, England P (2016) Real-time surface plasmon resonance (SPR) for the analysis of interactions between SUMO traps and mono or poly SUMO moieties. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_7
Lang V, Aillet F, Da Silva-Ferrada E et al (2015) Analysis of PTEN ubiquitylation and SUMOylation using molecular traps. Methods 77:112–118
Ureña E, Pirone L, Chafino S et al (2015) Evolution of SUMO function and chain formation in insects. Mol Biol Evol. doi:10.1093/molbev/msv242
Lang V, Da Silva-Ferrada E, Barrio R, et al (2016) Using biotinylated SUMO-traps to analyse SUMOylated proteins. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi: 10.1007/978-1-4939-6358-4_8
Eckhoff J, Dohmen RJ (2015) In vitro studies reveal a sequential mode of chain processing by the yeast SUMO (small ubiquitin-related modifier)-specific protease Ulp2. J Biol Chem 290:12268–12281
Eckhoff J, Dohmen RJ (2016) In vitro char- acterization of chain depolymerization activities of SUMO-specific proteases. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_9
Sahin U, Lallemand-Breitenbach V, de Thé H (2016) Detection of protein sumoylation in situ by proximity ligation assays. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_10
Fredriksson S, Gullberg M, Jarvius J et al (2002) Protein detection using proximity-dependent DNA ligation assays. Nat Biotechnol 20:473–477
Yuasa E, Saitoh H (2016) In situ SUMOylation and deSUMOylation assays: fluorescent methods to visualize SUMOylation and deSUMOylation in permeabilized cells. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_11
Obata S, Yuasa E, Seki D et al (2013) Molecular cloning and bacterial expression of the catalytic domain of the SENP1 gene of Oryzias latipes. Biosci Biotechnol Biochem 77:1788–1791
Lectez B, Migotti R, Lee SY et al (2014) Ubiquitin profiling in liver using a transgenic mouse with biotinylated ubiquitin. J Proteome Res 13:3016–3026
Franco M, Seyfried NT, Brand AH et al (2011) A novel strategy to isolate ubiquitin conjugates reveals wide role for ubiquitination during neural development. Mol Cell Proteomics 10:M110–M002188
Hendriks IA, D’Souza RC, Yang B et al (2014) Uncovering global SUMOylation signaling networks in a site-specific manner. Nat Struct Mol Biol 21:927–936
Pirone L, Xolalpa W, Mayor U, et al (2016) Analysis of SUMOylated proteins in cells and in vivo using bioSUMO molecules. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_12
Hendriks IA, Vertegaal ACO (2016) Label-free identification and quantification of SUMO target proteins. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_13
Aguilar-Martinez E, Sharrocks AD (2016) The use of multimeric protein scaffolds for identifying multi-SUMO binding proteins. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_14
Bawa-Khalfe T (2016) Isolation of in vivo SUMOylated chromatin-bound proteins. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_15
Bawa-Khalfe T, Lu L-S, Zuo Y et al (2012) Differential expression of SUMO-specific protease 7 variants regulates epithelial–mesenchymal transition. Proc Natl Acad Sci 109:17466–17471
Psakhye I, Jentsch S (2012) Protein group modification and synergy in the SUMO pathway as exemplified in DNA repair. Cell 151:807–820
Jentsch S, Psakhye I (2013) Control of nuclear activities by substrate-selective and protein-group SUMOylation. Annu Rev Genet 47:167–186
Psakhye I, Jentsch S (2016) Identification of substrates of protein-group SUMOylation. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi: 10.1007/978-1-4939-6358-4_16
Pelisch F, Hay RT (2016) Tools to study SUMO conjugation in Caenorhabditis elegans. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi:10.1007/978-1-4939-6358-4_17
Pelisch F, Sonneville R, Pourkarimi E et al (2014) Dynamic SUMO modification regulates mitotic chromosome assembly and cell cycle progression in Caenorhabditis elegans. Nat Commun 5:5485
Rytz T, Miller MJ, Vierstra RD (2016) Purification of SUMO conjugates from Arabidopsis for mass spectrometry analysis. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi: 10.1007/978-1-4939-6358-4_18
Reiter K, Matunis MJ (2016) Detection of SUMOylation in Plasmodium falciparum. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, doi: 10.1007/978-1-4939-6358-4_19
Reiter K, Mukhopadhyay D, Zhang H et al (2013) Identification of biochemically distinct properties of the small ubiquitin-related modifier (SUMO) conjugation pathway in Plasmodium falciparum. J Biol Chem 288:27724–27736
Tirard M, Brose N (2016) Systematic localization and identification of SUMOylation substrates in knock-in mice expressing affinity tagged SUMO1. SUMO: Methods and Protocols, Methods in Molecular Biology, vol. 1475, DOI 10.1007/978-1-4939-6358-4_20
Tirard M, Hsiao H-H, Nikolov M et al (2012) In vivo localization and identification of SUMOylated proteins in the brain of His6-HA-SUMO1 knock-in mice. Proc Natl Acad Sci U S A 109:21122–21127
Acknowledgments
This work was supported by the National Institutes of Health (GM060980 to M.J.M.). M.S.R. would like to acknowledge the Proteostasis COST action (BM1307).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Matunis, M.J., Rodriguez, M.S. (2016). Concepts and Methodologies to Study Protein SUMOylation: An Overview. In: Rodriguez, M. (eds) SUMO. Methods in Molecular Biology, vol 1475. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6358-4_1
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6358-4_1
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6356-0
Online ISBN: 978-1-4939-6358-4
eBook Packages: Springer Protocols