Abstract
RNA-binding proteins affect cellular metabolic programs through development and in response to cellular stimuli. Though much work has been done to elucidate the roles of a handful of RNA-binding proteins and their effect on RNA metabolism, the progress of studies to understand the effects of post-translational modifications of this class of proteins is far from complete. This chapter summarizes the work that has been done to identify the consequence of post-translational modifications to some RNA-binding proteins. The effects of these modifications have been shown to increase the panoply of functions that a given RNA-binding protein can assume. We will survey the experimental methods that are used to identify the presence of several protein modifications and methods that attempt to discern the consequence of these modifications.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
RNA-binding proteins (RBPs) regulate RNAs at every stage of their existence. This includes processes that govern RNA metabolism from capping and polynucleotide extension, RNA splicing, subcellular RNA localization, cellular export, translation (initiation elongation and extension), to RNA destruction. This class of ~1200–1600 proteins has important roles in the etiology of disease and therefore advances in the understanding of these proteins hold the promise to be directly applicable to the treatment of human neurodegenerative diseases, cancers and developmental disorders [1–5]. It has been known for some time that mRNA splicing is coupled to signal transduction and posttranslational modifications (PTMs) [6]. A full understanding of this intertwined network of processes has been complicated by the realization that RNA-binding proteins are a diverse class of regulators which themselves undergo extensive regulation via splicing, alternative 5′ and 3′ ends and various post-translational modifications.
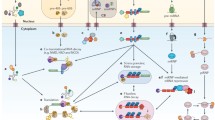
Post-translational modifications follow from various signaling pathways to cause activation of enzymes that add or remove PTM moieties (Fig. 12.1a). The set of post-translational modifications known to affect RNA-binding protein function includes at least: the reversible addition/removal of phosphate groups (PO3) by kinases/phosphatases, of methyl groups (CH3) by methylases/demethylases, of acetyl groups (C2H30) by acetylases/deacetylases, of the small protein ubiquitin (~8.5 kDa protein) by ubiquitin ligases/deubiquitinating enzymes, of SUMO (~12 kDa proteins) by ubiquitin ligases/SUMO proteases, and of glycans (polysaccharides) by glycosyltransferases/exoglycosidases or proteolytic cleavage by proteases. Although classically ubiquitination is associated with proteassomal degradation, some studies point to other functional roles for ubiquitin conjugation including localization and regulating protein interaction partners. Further, there have been observations of functional differences in the activity of polyubiquitin chains, depending on which lysine position links the ubiquitin monomers [7].
Schematic of the effects of PTM on RNA-binding proteins. (a) Signal integration. Various signals from inside a cell and from external sources activate signaling cascades that converge on the regulators of PTM state. (b) PTM may activate or deactivate the functions of an RNA-binding protein, including altering RNA targets, protein partners, mediating protein degradation or intrinsic enzymatic activities. (c) The altered functions of RNA-binding proteins lead to overall differences in the metabolism of RNAs at every stage of their existence, from transcription through destruction
RBPs are affected by these PTMs in diverse and currently unpredictable ways. RBP subcellular localization, affinity for RNA, enzymatic activity and association with cofactors have all been shown to be directed by PTM state (Fig. 12.1b). Since these consequences on protein function on RNA fate are often indistinguishable without close inspection, the exact stage where a PTM effects change is sometimes unclear. We have organized this chapter following the life-cycle of an mRNA from transcription through splicing, then nuclear export, subcellular targeting, translation and ultimately destruction. We highlight research at each stage of the RNA life-cycle that shows the indispensability of PTMs for fidelity of RNA regulation (Fig. 12.1c). Where possible we point to studies that discuss the exact effect of PTMs and describe experiments that can reveal this information. Overall, this chapter aims to be a review of the work done to bridge the gap between proteomics and transcriptomics and answer vital questions about the diversity of ways PTMs alter RBP function.
As mass spectroscopy methodologies improve and become more accessible, protein modifications are mapped with high accuracy and with little cost [8]. Cataloguing modifications is the first step in order to understand how they participate in RBP function, and how RBP function is regulated by a certain pathway. Therefore, our intention in this chapter is far from reporting all the identified PTMs in translation control of RBPs. Instead, we will focus on some of the most studied cases hoping that they will serve as examples and predictions of what we may expected from the other members of this protein family.
There are several proteins or families of proteins discussed below that may have diverse roles that are not fully explored. As these are involved in several stages of RNA maturation, it is appropriate to first introduce them (Fig. 12.2):
Large families of RBPs and their modifications. Proteins are shown as rectangles along rows. Functional domains of the protein are labelled and the proteins are sorted to group proteins with similar domains. PTMs listed in the Uniprot database are depicted as square shapes on each gene. Each vertical line is 50 amino acid residues. (Note: The QR code in the bottom-right links to an interactive version of this figure where references can also be visualized. Web link: here: https://rawgit.com/mlovci/12365bcafbef4a32d35a/raw/f781d50b3fd96ef83d019baf9e7984374420fdc6/Figure%25202.html)
HNRNPs are a diverse family of RNA-binding proteins for which post-translational modifications were discovered in the first descriptions of the proteins [9]. PTMs regulate the ability for HNRNP members to effect splicing changes and control the localization of RNA, as well as regulate translation, which will be discussed later in this chapter.
SR proteins were originally discovered for their role as splicing activators, but this view has been complicated by the nuances of effects due to PTMs [10, 11]. An abundance of literature points to the importance of PTMs in the activity of this family of proteins. Classically, SR proteins are modified by SR-protein kinases (SRPK1 and SRPK2) [12] and CDC2-like kinases (CLK1-4) [13–15]. The activity of these kinases is regulated through cell cycle, through development, and in response to cellular stresses like heat shock [10, 15, 16]. For several of the “classical” SR-proteins, including SRSF1 (aka Asf/Sf2) and SRSF2 (aka Sc35), phosphorylation induces changes in the intranuclear distribution of these phosphoproteins, causing release from nuclear speckles [17].
ELAV (aka Hu) proteins are exemplified by a well-studied member of the family, ELAVL1. ELAVL1 (aka HuR, human antigen R) has 2 N-terminal RRM domains followed by a hinge region that contains a nucleo/cytoplasmic shuttling domain (HNS), and another C-terminal RRM domain. It recognizes and binds to Adenylate-Urydinilate rich elements (AREs) present in 3′UTR and/or 5′-UTR of transcripts in the nucleus and regulates their splicing, processing, nuclear export, localization, half-life and translation [18, 19]. Several features of its behavior are partially explained by PTMs. ELAVL1 has a dual effect on translation, being capable of activating translation of certain mRNAs (hypoxia-inducible factor (HIF)-1α, p53, prothymosin-α, MKP-1, cytochrome c, heme oxygenase-1, and cationic amino acid transporter 1 [20–25]) and inhibiting others (IGFR, p27, Wnt5a e c-Myc, [25–30] for instance) with altered RNA affinity dependent upon phosphorylation. Phosphorylation on the hinge domain affects ELAVL1 localization [31], as do several phosphorylation sites on the RRM domains. There is evidence that RRM3 domain phosphorylation modulates dimerization inducing higher substrate affinity and altered protein localization.
2 PTM-Mediated Regulation of Pre-mRNA Processing
2.1 Transcription
The integration of signaling cascades in the control of RNA begins with RNA polymerase. Synthesis, 5′ capping and 3′ polyadenylation of almost all protein-coding transcripts is orchestrated by phosphorylation of the C-terminal domain (CTD) of DNA-directed RNA-polymerase 2 (POLR2A and homologs B-M) [32]. CTD hyperphosphorylation by carboxy-terminal kinases induces the recruitment of capping enzymes [33, 34] and 3′ end cleavage and polyadenylation factors [35]. Phosphorylation of the CTD of POLR1* also regulates transcriptional activity of ribosomal RNAs [36]. In fact, there is an extensive body of literature devoted to the consequence of RNA-polymerase CTD phosphorylation and interested readers should read [32, 37] for a more comprehensive review of the effectors and effects of this specific PTM target. Further, acetylation by p300, indicated by a p300 dose-dependent shift in POLR2 CTD molecular weight, seems to be involved in transcription initiation or early transcription elongation of growth-factor induced genes [38]. The production of specific species of RNA that bind FUS nucleates the formation of nuclear FUS aggregates [39]. POLR2 CTD phosphorylation is reinforced by these FUS aggregates [40]. Finally, FUS N-terminal phosphorylation by DNA protein-kinases (DNA-PK) removes FUS granules from the nucleus causing them to lose the potential to directly regulate POLR2 [41]. Thus, complex signaling and feedback control the activity of POLR2 through regulation of RBP PTM state.
Some HNRNPK PTMs serve to gate cell division checkpoints in response to DNA damage sensing. The activity of p53 tumor suppressor targets is tied to DNA damage-induced PTMs on the RNA-binding protein HNRNPK, which alter p53-HNRNPK protein-protein affinities. Methylation , phosphorylation and sumoylation of HNRNPK all regulate the p53-dependent cell cycle checkpoint [42–44]. Several signalling pathways converge to alter the function and stability of HNRNPK in response to DNA damage, including reduced expression of the E3 ligase MDM2 that targets HNRNPK to proteasomal destruction [45].
2.2 Splicing
Splicing is the RNA-catalyzed concatenation of exons that requires several protein scaffolds for which PTM state can control outcomes. Splicing in the nucleus is controlled by upstream signalling for DNA damage and cell cycle [46]. Indeed, it is closely tied to transcription and PTM state of histone proteins. For splicing components to mature, SMN complexes interact with U snRNAs and sm proteins to form snRNPs. This spliceosome formation occurs at Cajal bodies and requires the SMN complex. SMN components are localized, in part, by phosphorylation of the GEMIN proteins and deficiencies in this are linked to serious defects in intron recognition [47–49]. During spliceosome assembly, the targeted PTM of specific residues of snRNP must be required as both kinases and phosphatases are required for spliceosome assembly [50, 51].
S20 phosphorylation of SRSF1 initiates spliceosome assembly at intronic splice sites and is required for pre-mRNA processing fidelity [52]. This was reported to be regulated by the KIS kinase and important for bridging SRSF1 and U2AF2 in ternary SRSF1/U2AF2/RNA complexes [53]. Recent work shows with X-ray crystallography exactly the conformational shifts involved with phosphorylation of SRSF1 and reveals that only the phosphorylated version of SRSF1 can interact with U2AF65 [54]. SRSF10 (aka SRp38), is normally an unphosphorylated splicing repressor, but switches to a sequence-specific splicing activator when it is phosphorylated [17, 55, 56], presumably by inducing formation of spliceosomal complex A along with S100 [57].
2.3 Alternative Splicing
Alternative splicing (AS) is the regulated process of selective inclusion of specific exons into processed mRNA transcripts at specific stages of development or in response to external stimulation. Alternative splicing results from inefficient recognition by the spliceosome or competition among 3′ splice-sites for ligation to 5′ splice-sites. Splicing factors are regulated through signalling cascades to either activate or repress splicing in certain environmental conditions. These include pathways that recognize extracellular signals like EGF, Wnt, insulin, cytokines and heat stress [16, 55, 58, 59].
Beside sub-nuclear localization and interactions with snRNPs, phosphorylation of SR-proteins has been shown to cause shuttling between the nucleus to the cytoplasm, usually resulting in the loss of inclusion of their splicing targets [60–62]. Proline-directed SRSF1 phosphorylation causes conformational shifts that affect enzymatic activity of the protein [63]. Lines of evidence implicating non-phosphorylation PTMs like ubiquitination and acetylation are less common but do exist, and are associated with regulation of protein turnover [64, 65]. In the case of acetylation, lysine-acetylated SRSF2 proteins by KAT5 (aka Tip60) are more likely to be subject to degradation and HDAC6-mediated deacetylation causes SRSF2 accumulation. However, KAT5-mediated PTM is accompanied by concomitant acetylation of SRPK1 and SRPK2 that causes these kinases to be excluded from the nucleus, thus the accumulated SR proteins are not actively regulating splicing in these cells [64]. Ubiquitination of SRSF1 was shown to be increased in activated T-cells, causing proteasomal destruction of the protein, but the E3 ligase that mediates this PTM is not yet known [65]. Development of small-molecule inhibitors of these SR-related PTMs has been the focus of recent research with potential applications in treatment of cancers like metastatic melanoma [66, 67].
HNRNPL S52 phosphorylation mediates signal integration via the PI3K/AKT pathway. Phosphorylated HNRNPL, but not non-phosphorylated HNRNPL out-competes HNRNPU for binding at a cis-element. RNA-binding assays with an antibody specific for S52-phosphorylated HNRNPL shows that when HNRNPL is phosphorylated, it is associates with RNA while HNRNPU binding is diminished leading to exclusion of a pro-apoptotic caspase-9 exon; HNRNPU phosphorylation alone did not account for this change [68]. Similarly, upon neuron depolarization CAMK4 kinase activation causes phosphorylation of HNRNPL. This S513 phosphorylation increases HNRNPL affinity for CaM-kinase responsive RNA elements, out-competing assembling spliceosomal components and inhibiting exon inclusion [69]. Data obtained with a methylation-sensitive antibody indicates that PRMT1 causes constitutive methylation on HNRNPU, but the authors did not observe methylation-dependent localization shifts and could not discern a regulated function for the methylation of this protein [70].
Splicing of the stress-induced isoform of the TRA2B transcript by ELAVL1 is accomplished only when nuclear-localized ELAVL1 is phosphorylated downstream of Chk2 and p38-MAPK at positions S88 and T118 [71]. These phosphorylated residues increase ELAVL1’s affinity for an intronic binding site near an exon that causes an in-frame stop-codon, in turn causing higher levels of exon inclusion and subsequent nonsense-mediated decay of the TRA2B transcript.
KHDRBS1 (aka Sam68), a member of the STAR family of RNA-binding proteins, stands out in that it has reports of multiple classes of PTMs modify its RNA regulatory activity. Phosphorylation or acetylation increases its affinity for RNA and splicing regulatory activity as shown with point-mutants and chemical small-molecule inhibitors of phosphorylation [59, 72–75] while methylation decreases affinity for poly-(U) targets [76]. Mutational studies showed that S58, T71 and T84 phosphorylation were required for splicing activation and authors note ATP-gS, a phosphotase-resistant ATP analog, was necessary to observe this effect in in vitro splicing assays [59]. In addition, tyrosine phosphorylation may influence the ability of KHDRBS1 to effectively form dimers, which are required for splicing-regulatory activity [77]. SUMOylation has also been reported on KHDRBS1 but not with an RNA-regulatory effect [78].
These are just a few of the hundreds of RNA-binding protein PTMs have clear roles in regulating downstream splicing. For example, RBFOX1 (by WNK3) and RBFOX2 (by PRKCA/B) are shown to be shuttled out of the nucleus and degraded, respectively, by phosphorylation; thus, excluding these proteins from regulating their target exons [79, 80]. TRA2B has a reduced affinity for the mRNA that encodes the TRA2B protein when it is phosphorylated by CLK2 [81]. CELF1 phosphorylation downstream of Akt signaling causes changes in subcellular CELF1 distributions, affecting splicing and translational control (reviewed in detail in [82]). Hyperphosphorylation of CELF1 by PKCA/B/C was shown to be downstream of accumulation of toxic DM1 repetitive RNA in myotonic dystrophy and important for proper splicing regulation [83].
2.4 mRNA 5′ G-Capping and Decapping
RNA 5′ 7-methyl guanosine capping by RNA guanlyltransferase , which protects RNAs from 5′ exonucleases, promotes translation and nuclear export, is tied to the phosphorylation state of RNA polymerase II CTD and this function is evolutionarily conserved to yeast [33, 35]. Decapping conversely is the first step of RNA decay and inhibits translation initiation. Decapping enzymes 1 and 2 in mammals are subject to rapid decay by ubiquitination and subsequent proteasomal degradation; thus, leading to longer RNA half-lives in general, in this case shown for a selection of targets that are subject to AU-rich element-mediated decay [84].
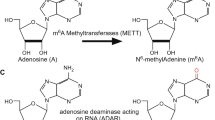
2.5 RNA Editing
ADAR protein levels, and consequently the extent of adenosine-to-inosine editing, are linked to the PTM state of these proteins. SUMO modification of ADAR1 at a lysine residue causes reduced editing efficacy in vitro and in vivo [85]. ADAR2 levels are decreased when phosphorylated by c-Jun kinase, resulting in reduced ADAR2-mediated A-to-I editing in pancreas [86].
3 PTM Regulation of Subcellular Localization
RBP PTMs commonly affect the ability for RBPs to move among cellular compartments. Thus, by virtue of their binding to RNA, RBPs regulate RNA localization based on their PTM state. In general, phosphorylation that affects RBP location also affects the set of bound RNAs, as may be expected since the availability of particular RNA species is not uniform across cells.
3.1 Nuclear/Cytoplasmic Shuttling
A few HNRNP proteins are sorted into the nucleus based on their PTM state. Nichols and colleagues showed with tritiated S-Adenosyl methionine then immunostaining after PRMT1 knockdown and GST-PRMT1 pull-down that HNRNPA2 arginine methylation in the RGG domain by PRMT1 is responsible for nuclear localization which is required for its regulation of alternative splicing [87]. HNRNPA1 localization is also regulated by PTM, with phosphorylation causing nuclear exclusion in a process that is activated by cellular stressors [88]. HNRNPQ has roles in splicing and mRNA stability and its localization is controlled by PRMT1-mediated methylation [89]; this may be important for controlling stability of RNA targets.
ELAVL1 is modified by an ubiquitin-like protein called NEDD8 on K313, K326 by MDM2 [90]. NEDD8 has 60 % homology with ubiquitin and its classical substrates are the cullin subunits of SCF ubiquitin E3 ligases [91]. Recently, it has been shown that MDM2 can associate with Ubc12 (the NEDD8 conjugating enzyme) and act as a NEDD8 ligase for p53 [92]. In the case of ELAVL1, neddylation promotes nuclear localization and inhibition of degradation [90].
Several kinases have been shown to be able to phosphorylate ELAVL1 and modulate its subcellular localization. For instance in the RRM domains: T118 phosphorylation by Chk2 or p38-MAPK [93, 94]; S158 phosphorylation by PRKCA and S318 phosphorylation by PRKCD [95, 96]. The hinge region (residues 186–244) is a hotspot for phosphorylation. Modifications on the hinge region affect ELAVL1 nucleo/cytoplasmic localization. Phosphorylation at S202 by CDK1 or CDK5, phosphorylation at S221 by PKC family members (PRKCA, PRKCD) and S242 phosphorylation by an unknown kinase all promote nuclear retention of the protein [31].
3.2 RNA Granules, P-Bodies and Nuclear Speckles
RNA-granules , processing bodies and nuclear speckles are functionally different aggregations of proteins and RNA that have modified activity and membership due to regulated changes in PTM state.
ELAVL1 phosphorylation on the hinge region outside of the HNS on Y200 by JAK3 inhibits ELAV1’s localization to stress granules upon arsenite stress, leading to accelerated degradation of some of its mRNA targets (e.g. SIRT1 and VHL), but it is unclear whether mRNAs are bound to ELAVL1 during the transition to stress granules [97].
TARDBP (aka TDP-43) is acetylated at K145, K192 by CREBBP (Creb-binding protein). Based on crystal structure mapping of acetylated side-chains, the conformation of TARDBP RRM may shift and alter its ability to bind to RNA. Using glutamine to mimic acetylated lysine and forced acetylation by CREBBP, Cohen and colleagues show that acetylated lysine on TARDBP reduces RNA-binding and results in aggregation of TARDBP into cytoplasmic inclusions. When not bound to RNAs, TARDBP exits the nucleus, joins RNA granules and is phosphorylated at S410 [98], perhaps by CSK1 (casein kinase 1) [99] or TTBK1/2 (Tau tubulin kinases 1 & 2 ) [100]. This may represent coordinated handoff between post-translational modifiers to place TARDBP in granules. While it is possible to prevent neurodegeneration by blocking TARDBP phosphorylation at S409/S410 [101] or by preventing acetylation, it is not clear how these mechanisms interact to cause RNA granules. Further, there are other modifications that will certainly need to be considered, including ubiquitination , which likely follows aggregation and precedes proteassomal degradation [102]. These processes are of biomedical importance because phosphorylated TARDBP is a hallmark of cytoplasmic inclusions in Amyotrophic Lateral Sclerosis (ALS) [103] and mutations that are predicted to increase TARDBP phosphorylation are linked to ALS [104].
3.3 Exosome
Exosome loading of RNAs including microRNAs (miRNAs) and longer classes of noncoding RNA are controlled with PTMs of a few proteins. RBM7 phosphorylation downstream of the MAPK-mediated stress response sorts ncRNA in the nucleus to exosomes [105]. Exosome targeting for certain unprocessed miRNAs is similarly HNRNPA2B1 sumoylation-dependent [106]. KHSRP is an RBP phosphorylated through ATM and PI3K/ATM kinases in response to DNA damage that guides RNAs to the exosome, where they are targeted for destruction [107, 108]. Indeed exosome destruction of several RNAs is coordinated through signal integration/kinase activation of several proteins including RBM7, KHSRP, TTP and others [109].
4 PTM Regulation of Translation
Some cytoplasmic RBPs modulate protein output by either contributing to initiation, elongation or termination of translation of their mRNA substrates, thus having a large effect via downstream cellular processes coded by targeted mRNAs. Below, we describe a few translation-regulatory RBPs, which are modified by phosphorylation, methylation, ubiquitination and oxidation.
Besides modulating subcellular localization and splicing, phosphorylation of the ELAVL1, as discussed above, PTM in ELAVL1 RRM domains can also modulate substrate affinity. For instance S38 and S100 phosphorylation by Chk2 can modulate mRNA substrate recognition. Interestingly, S100 phosphorylation seems to define the selectivity in ELAVL1 targeting. Thus, while phosphorylation induces release of Sirt1 mRNA and subsequent destabilization of Sirt1 mRNA, [93], the opposite is observed where S100 phosphorylation has been reported to increase affinity of ELAVL1 for Occludin mRNA (increasing its translation efficiency, [110]). This dual effect suggests the interesting possibility that phosphorylation may be used as a way for ELAVL1 to discriminate its different substrates and integrate dynamic signaling cues to translation decisions. It is unclear to what extent ELAVL1’s promiscuity in target selection is subject to this S100 phosphorylation event or other PTMs.
At least methylation and phosphorylation modulate FMR1 protein (aka FMRP) function. Absence of FMR1 expression in neurons leads to developmental abnormalities, such as immature, thin and highly branched dendritic spines. FMR1 has two Agenet domains followed by a NLS (nuclear localization signal) sequence close to its N-terminal, two KH domains (hnRNPK-homology domain) followed by a NES (nuclear export signal) signal in its middle and a RGG domain (arginine-glycine-glycine domain). The KH domain and RGG domains bind RNA. Phosphorylated FMR1 is associated with stalled polyribosome complexes. FMR1 forms a translation-inhibitory complex with the target mRNA and Cytoplasmic FMR1 Interacting Protein (CYFIP1) . This complex binds translation protein EIF4E, thereby inhibiting its interaction with EIF4G. Phosphorylation is in fact necessary for FMR1 to carry out its roles in developmental timing [111]. FMR1 may also bind directly to the ribosome in polysomes to inhibit translation elongation [112]. Indeed phosphorylation status of FMR1 has also been proposed to regulate its association with translating ribosomes and stalled ribosomes. Arginine methylation of the FMR1 RGG domain by PRMT1 has being proposed to inhibit its ability to recognize target mRNAs and its assembly in translation initiation inhibitory complexes [113–115].
CPEB1 contains a PEST sequence , two conserved RRM domains and a c-terminal znf-domain [116]. This RBP modulates translation of target mRNAs which contain a Cytoplasmic polyadenylation element (CPE) in their 3′UTR [117]. CEPB1 binds to CPE mRNA substrates, keeping them in a translation inhibited state. When CEBP1 is phosphorylated on S174 (outside of the RRM domains) by Aurora A kinase [118–120], it recruits CPSF [119, 121] and induces polyadenylation of the mRNA, greatly inducing their expression. CEBP1 can also be phosphorylated sequentially by Cdc2 on T125 and multiple Serine residues, which recruits Plx1 that phosphorylate S191 on the PEST sequence. Once PEST is phosphorylated, the hyperphosphorylated CPEB1 is recognized by the SCFb-TrCP E3 ubiquitin ligase complex, and polyubiquitinated leading to its proteassomal degradation [122]. CPEB1 plays important roles in the development of Xenopus oocyte and synapse formation/long-term potentiation [123, 124].
5 PTM Regulation of RNA Stability and Destruction
5.1 miRNA Related Repression
miRNAs are produced through a two-stage double-stranded RNA cleavage and processing. The miRNA pathway can either affect mRNA stability or translational output through miRNA:mRNA base-pairing mediated in a RNA-protein complex called RNA-induced silencing complex (RISC) . Canonical miRNAs originate from long primary mRNA transcripts, which are initially processed to miRNA precursors by the nuclear microprocessor complex in animals. The ribonuclease DROSHA and its RBP partner DGCR8 (DiGeorge syndrome critical region gene 8) are two key components of this complex (reviewed in [125]). PTM regulates this step in miRNA maturation, evidenced by co-immunoprecipitation of DROSHA and DGCR8 with components of the class I of Histone deacetylases (HDAC) [126]. In addition, the overexpression of HDAC1 in HEK293 cells results in increased affinity for primary miRNAs and higher mature miRNA availability without increasing the in corresponding primary miRNA’s expression levels [126]. This was attributed to deacetylation of the DGCR8 RNA-binding domain and increased affinity to primary miRNAs [126]. Other studies have also proposed that Drosha and DGCR8 can be stabilized by phosphorylation, and it was shown that anti-MAPK/CDK substrate antibodies recognized immunopurified DGCR8 [127].
Microprocessor components are not the only proteins targeted by PTM for regulation of nuclear miRNA processing. Certain factors that regulate miRNA biogenesis are altered by PTMs. In early differentiation, LIN28 is expressed and binds to the let-7 primary transcript, but acetylation by PCAF and ubiquitination by TRIM71 causes destruction of LIN28 leading to de-repressed let-7 processing and allowing cells to progress through differentiation [128, 129]. The E3 ligase TRIM65 represses RISC assembly by targeting TNRC6 (aka GW182) for destruction [130].
After miRNA precursors are loaded into the RISC and exported from the nucleus, DICER1, an RNAseIII, is responsible for recognizing the hairpin precursor sequences and processes them to mature miRNAs. FMR1 has been shown to interact with DICER1, argonaute 2 (AGO2) and specific miRNAs. Phosphorylation holds FMR1 in association with its targets and prevents translation, perhaps by AGO2 interactions with targets. While at the same time, phosphorylation of FMRP inhibits association with DICER and reduces DICER activity [131, 132].
Argonaute proteins facilitate the interactions between the 22nt long microRNAs and target mRNAs. Several signaling pathways converge on these AGO proteins to control their activity in various cellular contexts [133, 134]. AGO protein is phosphorylated by p38-MAPK under cellular stress treatments like sodium arsenite, causing it to localize to processing bodies [135]. Certain AGO family members are preferentially subject to hydroxylation, stabilizing these proteins and potentiating the effect of miRNAs in hypoxia [134, 136].
One study suggests that phosphorylation at S499 (in the RGG domain) of FMR1 modulates translation of its target mRNA and via AGO2 [132]. Activation of mGluR pathway leads to dephosphorylation of FMR1, followed by disassembly of its associated translation inhibitory complex and induction of PSD-95 translation/protein expression. It has being proposed that RPS6KB1 (aka S6K1) is the kinase that phosphorylates FMR1 and PP2A is the phosphatase that dephosphorylates it in the mGluR pathway [137, 138]. A separate study showed that HOXB8 mRNA is subject to the same phospho-FMR1/AGO2/miR-196a inhibitory complex [139]. This mechanism may be a way to induce gene expression of several FMR1 targets during long-term synaptic depression as depolarization leads to PP2A activation [138].
5.2 RNA Decay
QKI, a STAR-family RBP, is an essential regulator of myelination in oligodendrocytes. At early stages of development, QKI binds and stabilizes MBP RNA. C-terminal phosphorylation by Src-Protein Tyrosine Kinases (PTK) at Y285, Y288, Y290, Y292 and Y303 decreases QKI’s affinity for MBP (myelin basic protein) mRNA [140]. As src-PTK activity is reduced in early myelin development, mRNA can associate with QKI and accumulate. Indeed QKI sits downstream of several developmentally and disease-relevant pathways and understanding how PTMs affect its function will be an important goal for future studies ([140], reviewed in [141]).
ELAVL1 ubiquitination is related to the stability of its targets. ELAVL1 K48-linked ubiquitination on K182 by an unknown ubiquitin ligase promotes its proteassomal degradation [142]. However, K29-linked ubiquitination on ELAVL1 K313/K326 is reported to be a signal for protein-RNA complex disassembly. These modifications induce release of some ELAVL1 substrates (p21, MKP-1, and SIRT1 mRNAs) from ELAVL1, through recruitment of the p97–UBXD8 complex, leading to their destabilization [143]. Localization is also changed upon methylation of the hinge region by CARM1 (co-activator-associated arginine methyltransferase 1 ) at R217 [144]. Although the functional consequence of this modification is not completely understood, it has been shown to enhance ELAVL1’s ability to regulate turnover of some of its substrate mRNAs (TNF-alpha, cyclin A, cyclin B1, c-fos, SIRT1, and p16) [145].
6 Conclusion
As the effects of PTMs are broad and unpredictable, careful follow-up on the dynamic changes in protein function are necessary. With regard to RBP function, the essential question is whether a particular PTM will affect many of the RBPs in ways we have listed above, including:
-
1.
RNA-binding ability (i.e. QKI)
-
2.
Protein complex formation (i.e. SRSF1/U2AF65)
-
3.
Subcellular localization (i.e. CELF1, KDHRBS1)
-
(a)
Are RNAs bound during the transit?
-
(b)
What mechanisms drive RBP motility?
-
(a)
-
4.
Enzymatic activity of the RBP (i.e. ADAR)
-
5.
Initiation of RBP for destruction (i.e. LIN28)
Although the downstream consequences of these processes are varied and will be intensely studied, these are the basic features of RBP functions affected by PTMs. Answers to these simple questions for the library of RBP PTMs will be critical for accurate modeling of the effect of context and signal integration into the mRNP code. No doubt, advances in methods for probing protein structure will glean insight into the potential roles for PTMs on RBP function and provide a guide to prioritize the search for PTMs that have an impact on RNA maturation.
It should be noted again that this brief chapter is in no way a complete summary of the catalog of PTMs on RBPs. In the interest of space we had to restrict our discussions to the most well characterized examples and have left out many examples that may be relevant for basic biology or disease. These include rare post-translational modifications like nitration, which has evidence for affecting HNRNPA2B1 proteins [146], prolyl isomerization of POL2R [147], myristoylation which affects the axonal distribution of FXR2 [148] and PARylation which can globally repress the miRNA pathway in stress [149]. Several reviews cited herein have approached the relevance of PTMs in a particular pathway, family of genes, biological process or disease. Interested readers should follow this text with a thorough examination of these and the associated primary literature. There is a great deal still unknown about the cumulative and cross-regulatory effects that each PTM on each RBP holds. If the history of DNA-binding proteins and histone modifications is any indicator, this will be an area ripe for discovery and will advance basic biology and drug development.
References
Castello A, Fischer B, Hentze MW, Preiss T (2013) RNA-binding proteins in Mendelian disease. Trends Genet 29:318–327
Gerstberger S, Hafner M, Tuschl T (2014) A census of human RNA-binding proteins. Nat Rev Genet 15:829–845
Kapeli K, Yeo GW (2012) Genome-wide approaches to dissect the roles of RNA-binding proteins in translational control: implications for neurological diseases. Front Neurosci 6:144
Lukong KE, Chang KW, Khandjian EW, Richard S (2008) RNA-binding proteins in human genetic disease. Trends Genet 24:416–425
Nussbacher JK, Batra R, Lagier-Tourenne C, Yeo GW (2015) RNA-binding proteins in neurodegeneration: Seq and you shall receive. Trends Neurosci 38:226–236
Konig H, Ponta H, Herrlich P (1998) Coupling of signal transduction to alternative pre-mRNA splicing by a composite splice regulator. EMBO J 17:2904–2913
Michel MA, Elliott PR, Swatek KN, Simicek M, Pruneda JN, Wagstaff JL, Freund SM, Komander D (2015) Assembly and specific recognition of k29- and k33-linked polyubiquitin. Mol Cell 58:95–109
Dephoure N, Zhou C, Villen J, Beausoleil SA, Bakalarski CE, Elledge SJ, Gygi SP (2008) A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci U S A 105:10762–10767
Soulard M, Della Valle V, Siomi MC, Pinol-Roma S, Codogno P, Bauvy C, Bellini M, Lacroix JC, Monod G, Dreyfuss G et al (1993) hnRNP G: sequence and characterization of a glycosylated RNA-binding protein. Nucleic Acids Res 21:4210–4217
Sanford JR, Bruzik JP (1999) Developmental regulation of SR protein phosphorylation and activity. Genes Dev 13:1513–1518
Xiang-Yang Z, Pingping W, Joonhee H, Michael GR, Xiang-Dong F (2009) SR proteins in vertical integration of gene expression from transcription to RNA processing to translation. Mol Cell 35(1):1–10
Wang HY, Lin W, Dyck JA, Yeakley JM, Songyang Z, Cantley LC, Fu XD (1998) SRPK2: a differentially expressed SR protein-specific kinase involved in mediating the interaction and localization of pre-mRNA splicing factors in mammalian cells. J Cell Biol 140:737–750
Aubol B, Plocinik R, Hagopian J, Ma C, McGlone M, Bandyopadhyay R, Fu X, Adams J (2013) Partitioning RS domain phosphorylation in an SR protein through the CLK and SRPK protein kinases. J Mol Biol 425:2894–2909
Aubol B, Plocinik R, Keshwani M, McGlone M, Hagopian J, Ghosh G, Fu X, Adams J (2014) N-terminus of the protein kinase CLK1 induces SR protein hyperphosphorylation. Biochem J 462:143–152
Ninomiya K, Kataoka N, Hagiwara M (2011) Stress-responsive maturation of Clk1/4 pre-mRNAs promotes phosphorylation of SR splicing factor. J Cell Biol 195:27–40
Shin C, Manley JL (2002) The SR protein SRp38 represses splicing in M phase cells. Cell 111:407–417
Colwill K, Pawson T, Andrews B, Prasad J, Manley JL, Bell JC, Duncan PI (1996) The Clk/Sty protein kinase phosphorylates SR splicing factors and regulates their intranuclear distribution. EMBO J 15:265–275
Hinman MN, Lou H (2008) Diverse molecular functions of Hu proteins. Cell Mol Life Sci 65:3168–3181
López de Silanes I, Zhan M, Lal A, Yang X, Gorospe M (2004) Identification of a target RNA motif for RNA-binding protein HuR. Proc Natl Acad Sci U S A 101:2987–2992
Bhattacharyya SN, Habermacher R, Martine U, Closs EI, Filipowicz W (2006) Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 125:1111–1124
Kawai T, Lal A, Yang X, Galban S, Mazan-Mamczarz K, Gorospe M (2006) Translational control of cytochrome c by RNA-binding proteins TIA-1 and HuR. Mol Cell Biol 26:3295–3307
Kuwano Y, Kim HH, Abdelmohsen K, Pullmann R, Martindale JL, Yang X, Gorospe M (2008) MKP-1 mRNA stabilization and translational control by RNA-binding proteins HuR and NF90. Mol Cell Biol 28:4562–4575
Kuwano Y, Rabinovic A, Srikantan S, Gorospe M, Demple B (2009) Analysis of nitric oxide-stabilized mRNAs in human fibroblasts reveals HuR-dependent heme oxygenase 1 upregulation. Mol Cell Biol 29:2622–2635
Lal A, Kawai T, Yang X, Mazan-Mamczarz K, Gorospe M (2005) Antiapoptotic function of RNA-binding protein HuR effected through prothymosin alpha. EMBO J 24:1852–1862
Mazan-Mamczarz K, Galbán S, López de Silanes I, Martindale JL, Atasoy U, Keene JD, Gorospe M (2003) RNA-binding protein HuR enhances p53 translation in response to ultraviolet light irradiation. Proc Natl Acad Sci U S A 100:8354–8359
Abdelmohsen K, Kuwano Y, Kim HH, Gorospe M (2008) Posttranscriptional gene regulation by RNA-binding proteins during oxidative stress: implications for cellular senescence. Biol Chem 389:243–255
Kim HH, Kuwano Y, Srikantan S, Lee EK, Martindale JL, Gorospe M (2009) HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev 23:1743–1748
Kullmann M, Göpfert U, Siewe B, Hengst L (2002) ELAV/Hu proteins inhibit p27 translation via an IRES element in the p27 5′UTR. Genes Dev 16:3087–3099
Leandersson K, Riesbeck K, Andersson T (2006) Wnt-5a mRNA translation is suppressed by the Elav-like protein HuR in human breast epithelial cells. Nucleic Acids Res 34:3988–3999
Meng Z, King PH, Nabors LB, Jackson NL, Chen C-Y, Emanuel PD, Blume SW (2005) The ELAV RNA-stability factor HuR binds the 5′-untranslated region of the human IGF-IR transcript and differentially represses cap-dependent and IRES-mediated translation. Nucleic Acids Res 33:2962–2979
Kim HH, Yang X, Kuwano Y, Gorospe M (2008) Modification at HuR(S242) alters HuR localization and proliferative influence. Cell Cycle 7:3371–3377
Phatnani HP, Greenleaf AL (2006) Phosphorylation and functions of the RNA polymerase II CTD. Genes Dev 20:2922–2936
Cho EJ, Takagi T, Moore CR, Buratowski S (1997) mRNA capping enzyme is recruited to the transcription complex by phosphorylation of the RNA polymerase II carboxy-terminal domain. Genes Dev 11:3319–3326
McCracken S, Fong N, Rosonina E, Yankulov K, Brothers G, Siderovski D, Hessel A, Foster S, Shuman S, Bentley DL (1997) 5′-Capping enzymes are targeted to pre-mRNA by binding to the phosphorylated carboxy-terminal domain of RNA polymerase II. Genes Dev 11:3306–3318
McCracken S, Fong N, Yankulov K, Ballantyne S, Pan G, Greenblatt J, Patterson SD, Wickens M, Bentley DL (1997) The C-terminal domain of RNA polymerase II couples mRNA processing to transcription. Nature 385:357–361
Bouchoux C, Hautbergue G, Grenetier S, Carles C, Riva M, Goguel V (2003) CTD kinase I is involved in RNA polymerase I transcription. Nucleic Acids Res 32:5851–5860
Hsin J-PP, Xiang K, Manley JL (2014) Function and control of RNA polymerase II C-terminal domain phosphorylation in vertebrate transcription and RNA processing. Mol Cell Biol 34:2488–2498
Schröder S, Herker E, Itzen F, He D, Thomas S, Gilchrist DA, Kaehlcke K, Cho S, Pollard KS, Capra JA et al (2013) Acetylation of RNA polymerase II regulates growth-factor-induced gene transcription in mammalian cells. Mol Cell 52:314–324
Schwartz JC, Wang X, Podell ER, Cech TR (2013) RNA seeds higher-order assembly of FUS protein. Cell Rep 5:918–925
Kwon I, Kato M, Xiang S, Wu L, Theodoropoulos P, Mirzaei H, Han T, Xie S, Corden JL, McKnight SL (2013) Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155:1049–1060
Deng Q, Holler CJ, Taylor G, Hudson KF, Watkins W, Gearing M, Ito D, Murray ME, Dickson DW, Seyfried NT et al (2014) FUS is phosphorylated by DNA-PK and accumulates in the cytoplasm after DNA damage. J Neurosci 34:7802–7813
Lee SW, Lee MH, Park JH, Kang SH, Yoo HM, Ka SH, Oh YM, Jeon YJ, Chung CH (2012) SUMOylation of hnRNP-K is required for p53-mediated cell-cycle arrest in response to DNA damage. EMBO J 31:4441–4452
Pelisch F, Pozzi B, Risso G, Muñoz MJ, Srebrow A (2012) DNA damage-induced heterogeneous nuclear ribonucleoprotein K sumoylation regulates p53 transcriptional activation. J Biol Chem 287:30789–30799
Yang JH, Chiou YY, Fu SL, Shih IY, Weng TH, Lin WJ, Lin CH (2014) Arginine methylation of hnRNPK negatively modulates apoptosis upon DNA damage through local regulation of phosphorylation. Nucleic Acids Res 42:9908–9924
Moumen A, Masterson P, O’Connor MJ, Jackson SP (2005) hnRNP K: an HDM2 target and transcriptional coactivator of p53 in response to DNA damage. Cell 123:1065–1078
Moore MJ, Wang Q, Kennedy CJ, Silver PA (2010) An alternative splicing network links cell-cycle control to apoptosis. Cell 142:625–636
Husedzinovic A, Oppermann F, Draeger-Meurer S, Chari A, Fischer U, Daub H, Gruss OJ (2014) Phosphoregulation of the human SMN complex. Eur J Cell Biol 93:106–117
Husedzinovic A, Neumann B, Reymann J, Draeger-Meurer S, Chari A, Erfle H, Fischer U, Gruss OJ (2015) The catalytically inactive tyrosine phosphatase HD-PTP/PTPN23 is a novel regulator of SMN complex localization. Mol Biol Cell 26:161–171
Renvoisé B, Quérol G, Verrier ER, Burlet P, Lefebvre S (2012) A role for protein phosphatase PP1γ in SMN complex formation and subnuclear localization to Cajal bodies. J Cell Sci 125:2862–2874
Kamoun M, Filali M, Murray MV, Awasthi S, Wadzinski BE (2013) Protein phosphatase 2A family members (PP2A and PP6) associate with U1 snRNP and the spliceosome during pre-mRNA splicing. Biochem Biophys Res Commun 440:306–311
Mathew R, Hartmuth K, Mohlmann S, Urlaub H, Ficner R, Luhrmann R (2008) Phosphorylation of human PRP28 by SRPK2 is required for integration of the U4/U6-U5 tri-snRNP into the spliceosome. Nat Struct Mol Biol 15:435–443
Wang X, Bruderer S, Rafi Z, Xue J, Milburn PJ, Krämer A, Robinson PJ (1999) Phosphorylation of splicing factor SF1 on Ser20 by cGMP-dependent protein kinase regulates spliceosome assembly. EMBO J 18:4549–4559
Manceau V, Swenson M, Le Caer J-PP, Sobel A, Kielkopf CL, Maucuer A (2006) Major phosphorylation of SF1 on adjacent Ser-Pro motifs enhances interaction with U2AF65. FEBS J 273:577–587
Wang W, Maucuer A, Gupta A, Manceau V, Thickman KR, Bauer WJ, Kennedy SD, Wedekind JE, Green MR, Kielkopf CL (2013) Structure of phosphorylated SF1 bound to U2AF(6)(5) in an essential splicing factor complex. Structure 21:197–208
Shin C, Feng Y, Manley JL (2004) Dephosphorylated SRp38 acts as a splicing repressor in response to heat shock. Nature 427:553–558
Shin C, Kleiman FE, Manley JL (2005) Multiple properties of the splicing repressor SRp38 distinguish it from typical SR proteins. Mol Cell Biol 25:8334–8343
Feng Y, Chen M, Manley JL (2008) Phosphorylation switches the general splicing repressor SRp38 to a sequence-specific activator. Nat Struct Mol Biol 15:1040–1048
Fu XD, Ares M Jr (2014) Context-dependent control of alternative splicing by RNA-binding proteins. Nat Rev Genet 15:689–701
Matter N, Herrlich P, Konig H (2002) Signal-dependent regulation of splicing via phosphorylation of Sam68. Nature 420:691–695
Caceres J, Screaton G, Krainer A (1998) A specific subset of SR proteins shuttles continuously between the nucleus and the cytoplasm. Genes Dev 12:55–66
Lai MC, Tarn WY (2004) Hypophosphorylated ASF/SF2 binds TAP and is present in messenger ribonucleoproteins. J Biol Chem 279:31745–31749
Lai MC, Lin RI, Huang SY, Tsai CW, Tarn WY (2000) A human importin-beta family protein, transportin-SR2, interacts with the phosphorylated RS domain of SR proteins. J Biol Chem 275:7950–7957
Keshwani MM, Aubol BE, Fattet L, Ma CT, Qiu J, Jennings PA, Fu XD, Adams JA (2015) Conserved proline-directed phosphorylation regulates SR protein conformation and splicing function. Biochem J 466:311–322
Edmond V, Moysan E, Khochbin S, Matthias P, Brambilla C, Brambilla E, Gazzeri S, Eymin B (2011) Acetylation and phosphorylation of SRSF2 control cell fate decision in response to cisplatin. EMBO J 30:510–523
Moulton VR, Gillooly AR, Tsokos GC (2014) Ubiquitination regulates expression of the serine/arginine-rich splicing factor 1 (SRSF1) in normal and systemic lupus erythematosus (SLE) T cells. J Biol Chem 289:4126–4134
Gammons MV, Lucas R, Dean R, Coupland SE, Oltean S, Bates DO (2014) Targeting SRPK1 to control VEGF-mediated tumour angiogenesis in metastatic melanoma. Br J Cancer 111:477–485
Ohe K, Hagiwara M (2015) Modulation of alternative splicing with chemical compounds in new therapeutics for human diseases. ACS Chem Biol 10:914–924
Vu NT, Park MA, Shultz JC, Goehe RW, Hoeferlin LA, Shultz MD, Smith SA, Lynch KW, Chalfant CE (2013) hnRNP U enhances caspase-9 splicing and is modulated by AKT-dependent phosphorylation of hnRNP L. J Biol Chem 288:8575–8584
Liu G, Razanau A, Hai Y, Yu J, Sohail M, Lobo VG, Chu J, Kung SK, Xie J (2012) A conserved serine of heterogeneous nuclear ribonucleoprotein L (hnRNP L) mediates depolarization-regulated alternative splicing of potassium channels. J Biol Chem 287:22709–22716
Herrmann F, Bossert M, Schwander A, Akgün E, Fackelmayer FO (2004) Arginine methylation of scaffold attachment factor A by heterogeneous nuclear ribonucleoprotein particle-associated PRMT1. J Biol Chem 279:48774–48779
Akaike Y, Masuda K, Kuwano Y, Nishida K, Kajita K, Kurokawa K, Satake Y, Shoda K, Imoto I, Rokutan K (2014) HuR regulates alternative splicing of the TRA2β gene in human colon cancer cells under oxidative stress. Mol Cell Biol 34:2857–2873
Babic I, Jakymiw A, Fujita D (2004) The RNA-binding protein Sam68 is acetylated in tumor cell lines, and its acetylation correlates with enhanced RNA-binding activity. Oncogene 23:3781–3789
Paronetto MP, Achsel T, Massiello A, Chalfant CE, Sette C (2007) The RNA-binding protein Sam68 modulates the alternative splicing of Bcl-x. J Cell Biol 176:929–939
Taylor SJ, Shalloway D (1994) An RNA-binding protein associated with Src through its SH2 and SH3 domains in mitosis. Nature 368:867–871
Wang L, Richard S, Shaw A (1995) P62 association with RNA is regulated by tyrosine phosphorylation. J Biol Chem 270(5):2010–2013
Rho J, Choi S, Jung C-RR, Im D-SS (2007) Arginine methylation of Sam68 and SLM proteins negatively regulates their poly(U) RNA-binding activity. Arch Biochem Biophys 466:49–57
Meyer NH, Tripsianes K, Vincendeau M, Madl T, Kateb F, Brack-Werner R, Sattler M (2010) Structural basis for homodimerization of the Src-associated during mitosis, 68-kDa protein (Sam68) Qua1 domain. J Biol Chem 285:28893–28901
Babic I, Cherry E, Fujita DJ (2006) SUMO modification of Sam68 enhances its ability to repress cyclin D1 expression and inhibits its ability to induce apoptosis. Oncogene 25:4955–4964
Lee AY, Chen W, Stippec S, Self J, Yang F, Ding X, Chen S, Juang YC, Cobb MH (2012) Protein kinase WNK3 regulates the neuronal splicing factor Fox-1. Proc Natl Acad Sci U S A 109:16841–16846
Verma SK, Deshmukh V, Liu P, Nutter CA, Espejo R, Hung M-LL, Wang G-SS, Yeo GW, Kuyumcu-Martinez MN (2013) Reactivation of fetal splicing programs in diabetic hearts is mediated by protein kinase C signaling. J Biol Chem 288:35372–35386
Stoilov P, Daoud R, Nayler O, Stamm S (2004) Human tra2-beta1 autoregulates its protein concentration by influencing alternative splicing of its pre-mRNA. Hum Mol Genet 13:509–524
Giudice J, Cooper T (2014) RNA-binding proteins in heart development. Adv Exp Med Biol 825:389–429
Kuyumcu-Martinez NM, Wang G-SS, Cooper TA (2007) Increased steady-state levels of CUGBP1 in myotonic dystrophy 1 are due to PKC-mediated hyperphosphorylation. Mol Cell 28:68–78
Erickson SL, Corpuz EO, Maloy JP, Fillman C, Webb K, Bennett EJ, Lykke-Andersen J (2015) Competition between decapping complex formation and ubiquitin-mediated proteasomal degradation controls human Dcp2 decapping activity. Mol Cell Biol 35:2144–2153
Desterro JM, Keegan LP, Jaffray E, Hay RT, O’Connell MA, Carmo-Fonseca M (2005) SUMO-1 modification alters ADAR1 editing activity. Mol Biol Cell 16:5115–5126
Yang L, Huang P, Li F, Zhao L, Zhang Y, Li S, Gan Z, Lin A, Li W, Liu Y (2012) c-Jun amino-terminal kinase-1 mediates glucose-responsive upregulation of the RNA editing enzyme ADAR2 in pancreatic beta-cells. PLoS One 7(11):e48611
Nichols RC, Wang XW, Tang J, Hamilton BJ, High FA, Herschman HR, Rigby WF (2000) The RGG domain in hnRNP A2 affects subcellular localization. Exp Cell Res 256:522–532
van der Houven van Oordt W, Diaz-Meco MT, Lozano J, Krainer AR, Moscat J, Cáceres JF (2000) The MKK(3/6)-p38-signaling cascade alters the subcellular distribution of hnRNP A1 and modulates alternative splicing regulation. J Cell Biol 149:307–316
Passos DO, Quaresma AJ, Kobarg J (2006) The methylation of the C-terminal region of hnRNPQ (NSAP1) is important for its nuclear localization. Biochem Biophys Res Commun 346:517–525
Embade N, Fernández-Ramos D, Varela-Rey M, Beraza N, Sini M, Gutiérrez de Juan V, Woodhoo A, Martínez-López N, Rodríguez-Iruretagoyena B, Bustamante FJ et al (2012) Murine double minute 2 regulates Hu antigen R stability in human liver and colon cancer through NEDDylation. Hepatology 55:1237–1248
Enchev RI, Schulman BA, Peter M (2015) Protein neddylation: beyond cullin-RING ligases. Nat Rev Mol Cell Biol 16:30–44
Batuello CN, Hauck PM, Gendron JM, Lehman JA, Mayo LD (2015) Src phosphorylation converts Mdm2 from a ubiquitinating to a neddylating E3 ligase. Proc Natl Acad Sci U S A 112:1749–1754
Abdelmohsen K, Pullmann R, Lal A, Kim HH, Galban S, Yang X, Blethrow JD, Walker M, Shubert J, Gillespie DA et al (2007) Phosphorylation of HuR by Chk2 regulates SIRT1 expression. Mol Cell 25:543–557
Lafarga V, Cuadrado A, Lopez de Silanes I, Bengoechea R, Fernandez-Capetillo O, Nebreda AR (2009) p38 Mitogen-activated protein kinase- and HuR-dependent stabilization of p21(Cip1) mRNA mediates the G(1)/S checkpoint. Mol Cell Biol 29:4341–4351
Doller A, Huwiler A, Müller R, Radeke HH, Pfeilschifter J, Eberhardt W (2007) Protein kinase C alpha-dependent phosphorylation of the mRNA-stabilizing factor HuR: implications for posttranscriptional regulation of cyclooxygenase-2. Mol Biol Cell 18:2137–2148
Doller A, Akool E-S, Huwiler A, Müller R, Radeke HH, Pfeilschifter J, Eberhardt W (2008) Posttranslational modification of the AU-rich element binding protein HuR by protein kinase Cdelta elicits angiotensin II-induced stabilization and nuclear export of cyclooxygenase 2 mRNA. Mol Cell Biol 28:2608–2625
Yoon J-H, Abdelmohsen K, Srikantan S, Guo R, Yang X, Martindale JL, Gorospe M (2014) Tyrosine phosphorylation of HuR by JAK3 triggers dissociation and degradation of HuR target mRNAs. Nucleic Acids Res 42:1196–1208
Cohen T, Hwang A, Restrepo C, Yuan C, Trojanowski J, Lee V (2015) An acetylation switch controls TDP-43 function and aggregation propensity. Nat Commun 6:5845
Kametani F, Nonaka T, Suzuki T, Arai T, Dohmae N, Akiyama H, Hasegawa M (2009) Identification of casein kinase-1 phosphorylation sites on TDP-43. Biochem Biophys Res Commun 382:405–409
Liachko NF, McMillan PJ, Strovas TJ, Loomis E, Greenup L, Murrell JR, Ghetti B, Raskind MA, Montine TJ, Bird TD et al (2014) The tau tubulin kinases TTBK1/2 promote accumulation of pathological TDP-43. PLoS Genet 10:e1004803
Liachko N, McMillan P, Guthrie C, Bird T, Leverenz J, Kraemer B (2013) CDC7 inhibition blocks pathological TDP-43 phosphorylation and neurodegeneration. Ann Neurol 74:39–52
Hans F, Fiesel FC, Strong JC, Jackel S, Rasse TM, Geisler S, Springer W, Schulz JB, Voigt A, Kahle PJ (2014) UBE2E ubiquitin-conjugating enzymes and ubiquitin isopeptidase Y regulate TDP-43 protein ubiquitination. J Biol Chem 289:19164–19179
Arai T, Hasegawa M, Nonoka T, Kametani F, Yamashita M, Hosokawa M, Niizato K, Tsuchiya K, Kobayashi Z, Ikeda K et al (2010) Phosphorylated and cleaved TDP-43 in ALS, FTLD and other neurodegenerative disorders and in cellular models of TDP-43 proteinopathy. Neuropathology 30:170–181
Kühnlein P, Sperfeld A-DD, Vanmassenhove B, Van Deerlin V, Lee VM, Trojanowski JQ, Kretzschmar HA, Ludolph AC, Neumann M (2008) Two German kindreds with familial amyotrophic lateral sclerosis due to TARDBP mutations. Arch Neurol 65:1185–1189
Tiedje C, Lubas M, Tehrani M, Menon MB, Ronkina N, Rousseau S, Cohen P, Kotlyarov A, Gaestel M (2015) p38MAPK/MK2-mediated phosphorylation of RBM7 regulates the human nuclear exosome targeting complex. RNA 21:262–278
Villarroya-Beltri C, Gutierrez-Vazquez C, Sanchez-Cabo F, Perez-Hernandez D, Vazquez J, Martin-Cofreces N, Martinez-Herrera DJ, Pascual-Montano A, Mittelbrunn M, Sanchez-Madrid F (2013) Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat Commun 4:2980
Gherzi R, Trabucchi M, Ponassi M, Ruggiero T, Corte G, Moroni C, Chen CY, Khabar KS, Andersen JS, Briata P (2006) The RNA-binding protein KSRP promotes decay of beta-catenin mRNA and is inactivated by PI3K-AKT signaling. PLoS Biol 5:e5
Liu Y, Liu Q (2011) ATM signals miRNA biogenesis through KSRP. Mol Cell 41:367–368
Tiedje C, Holtmann H, Gaestel M (2014) The role of mammalian MAPK signaling in regulation of cytokine mRNA stability and translation. J Interferon Cytokine Res 34:220–232
Yu T-X, Wang P-Y, Rao JN, Zou T, Liu L, Xiao L, Gorospe M, Wang J-Y (2011) Chk2-dependent HuR phosphorylation regulates occludin mRNA translation and epithelial barrier function. Nucleic Acids Res 39:8472–8487
Say E, Tay H-GG, Zhao Z-S, Baskaran Y, Li R, Lim L, Manser E (2010) A functional requirement for PAK1 binding to the KH(2) domain of the fragile X protein-related FXR1. Mol Cell 38:236–249
Darnell JC, Van Driesche SJ, Zhang C, Hung KYS, Mele A, Fraser CE, Stone EF, Chen C, Fak JJ, Chi SW et al (2011) FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146:247–261
Dolzhanskaya N, Merz G, Aletta JM, Denman RB (2006) Methylation regulates the intracellular protein-protein and protein-RNA interactions of FMRP. J Cell Sci 119:1933–1946
Dolzhanskaya N, Merz G, Denman RB (2006) Alternative splicing modulates protein arginine methyltransferase-dependent methylation of fragile X syndrome mental retardation protein. Biochemistry 45:10385–10393
Stetler A, Winograd C, Sayegh J, Cheever A, Patton E, Zhang X, Clarke S, Ceman S (2006) Identification and characterization of the methyl arginines in the fragile X mental retardation protein Fmrp. Hum Mol Genet 15:87–96
Hake LE, Richter JD (1994) CPEB is a specificity factor that mediates cytoplasmic polyadenylation during Xenopus oocyte maturation. Cell 79:617–627
de Moor CH, Richter JD (1999) Cytoplasmic polyadenylation elements mediate masking and unmasking of cyclin B1 mRNA. EMBO J 18:2294–2303
Mendez R, Hake LE, Andresson T, Littlepage LE, Ruderman JV, Richter JD (2000) Phosphorylation of CPE binding factor by Eg2 regulates translation of c-mos mRNA. Nature 404:302–307
Mendez R, Murthy KG, Ryan K, Manley JL, Richter JD (2000) Phosphorylation of CPEB by Eg2 mediates the recruitment of CPSF into an active cytoplasmic polyadenylation complex. Mol Cell 6:1253–1259
Mendez R, Barnard D, Richter JD (2002) Differential mRNA translation and meiotic progression require Cdc2-mediated CPEB destruction. EMBO J 21:1833–1844
Dickson KS, Bilger A, Ballantyne S, Wickens MP (1999) The cleavage and polyadenylation specificity factor in Xenopus laevis oocytes is a cytoplasmic factor involved in regulated polyadenylation. Mol Cell Biol 19:5707–5717
Setoyama D, Yamashita M, Sagata N (2007) Mechanism of degradation of CPEB during Xenopus oocyte maturation. Proc Natl Acad Sci U S A 104:18001–18006
Di Nardo AA, Nedelec S, Trembleau A, Volovitch M, Prochiantz A, Montesinos ML (2007) Dendritic localization and activity-dependent translation of Engrailed1 transcription factor. Mol Cell Neurosci 35:230–236
Wu L, Wells D, Tay J, Mendis D, Abbott MA, Barnitt A, Quinlan E, Heynen A, Fallon JR, Richter JD (1998) CPEB-mediated cytoplasmic polyadenylation and the regulation of experience-dependent translation of alpha-CaMKII mRNA at synapses. Neuron 21:1129–1139
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524
Wada T, Kikuchi J, Furukawa Y (2012) Histone deacetylase 1 enhances microRNA processing via deacetylation of DGCR8. EMBO Rep 13:142–149
Herbert KM, Pimienta G, DeGregorio SJ, Alexandrov A, Steitz JA (2013) Phosphorylation of DGCR8 increases its intracellular stability and induces a progrowth miRNA profile. Cell Rep 5:1070–1081
Lee SH, Cho S, Kim MS, Choi K, Cho JY, Gwak HS, Kim YJ, Yoo H, Lee SH, Park JB et al (2014) The ubiquitin ligase human TRIM71 regulates let-7 microRNA biogenesis via modulation of Lin28B protein. Biochim Biophys Acta 1839:374–386
Wang L-X, Wang J, Qu T-T, Zhang Y, Shen Y-F (2014) Reversible acetylation of Lin28 mediated by PCAF and SIRT1. Biochim Biophys Acta 1843:1188–1195
Li S, Wang L, Fu B, Berman MA, Diallo A, Dorf ME (2014) TRIM65 regulates microRNA activity by ubiquitination of TNRC6. Proc Natl Acad Sci U S A 111:6970–6975
Cheever A, Ceman S (2009) Phosphorylation of FMRP inhibits association with Dicer. RNA 15:362–366
Muddashetty RS, Nalavadi VC, Gross C, Yao X, Xing L, Laur O, Warren ST, Bassell GJ (2011) Reversible inhibition of PSD-95 mRNA translation by miR-125a, FMRP phosphorylation, and mGluR signaling. Mol Cell 42:673–688
Jee D, Lai EC (2014) Alteration of miRNA activity via context-specific modifications of Argonaute proteins. Trends Cell Biol 24:546–553
Wu C, So J, Davis-Dusenbery BN, Qi HH, Bloch DB, Shi Y, Lagna G, Hata A (2011) Hypoxia potentiates microRNA-mediated gene silencing through posttranslational modification of Argonaute2. Mol Cell Biol 31:4760–4774
Zeng Y, Sankala H, Zhang X, Graves PR (2008) Phosphorylation of Argonaute 2 at serine-387 facilitates its localization to processing bodies. Biochem J 413:429–436
Qi HH, Ongusaha PP, Myllyharju J, Cheng D, Pakkanen O, Shi Y, Lee SW, Peng J, Shi Y (2008) Prolyl 4-hydroxylation regulates Argonaute 2 stability. Nature 455:421–424
Narayanan U, Nalavadi V, Nakamoto M, Thomas G, Ceman S, Bassell GJ, Warren ST (2008) S6K1 phosphorylates and regulates fragile X mental retardation protein (FMRP) with the neuronal protein synthesis-dependent mammalian target of rapamycin (mTOR) signaling cascade. J Biol Chem 283:18478–18482
Niere F, Wilkerson JR, Huber KM (2012) Evidence for a fragile X mental retardation protein-mediated translational switch in metabotropic glutamate receptor-triggered Arc translation and long-term depression. J Neurosci 32:5924–5936
Li Y, Tang W, Zhang L-R, Zhang C-Y (2014) FMRP regulates miR196a-mediated repression of HOXB8 via interaction with the AGO2 MID domain. Mol Biosyst 10:1757–1764
Zhang Y, Lu Z, Ku L, Chen Y, Wang H, Feng Y (2003) Tyrosine phosphorylation of QKI mediates developmental signals to regulate mRNA metabolism. EMBO J 22:1801–1810
Feng Y, Bankston A (2009) The star family member QKI and cell signaling. Adv Exp Med Biol 693:25–36
Abdelmohsen K, Srikantan S, Yang X, Lal A, Kim HH, Kuwano Y, Galban S, Becker KG, Kamara D, de Cabo R et al (2009) Ubiquitin-mediated proteolysis of HuR by heat shock. EMBO J 28:1271–1282
Zhou H-L, Geng C, Luo G, Lou H (2013) The p97-UBXD8 complex destabilizes mRNA by promoting release of ubiquitinated HuR from mRNP. Genes Dev 27:1046–1058
Li H, Park S, Kilburn B, Jelinek MA, Henschen-Edman A, Aswad DW, Stallcup MR, Laird-Offringa IA (2002) Lipopolysaccharide-induced methylation of HuR, an mRNA-stabilizing protein, by CARM1. Coactivator-associated arginine methyltransferase. J Biol Chem 277:44623–44630
Pang L, Tian H, Chang N, Yi J, Xue L, Jiang B, Gorospe M, Zhang X, Wang W (2013) Loss of CARM1 is linked to reduced HuR function in replicative senescence. BMC Mol Biol 14:15
Miyagi M, Sakaguchi H, Darrow RM, Yan L, West KA, Aulak KS, Stuehr DJ, Hollyfield JG, Organisciak DT, Crabb JW (2002) Evidence that light modulates protein nitration in rat retina. Mol Cell Proteomics 1:293–303
Yogesha SD, Mayfield JE, Zhang Y (2013) Cross-talk of phosphorylation and prolyl isomerization of the C-terminal domain of RNA Polymerase II. Molecules 19:1481–1511
Stackpole EE, Akins MR, Fallon JR (2014) N-myristoylation regulates the axonal distribution of the fragile X-related protein FXR2P. Mol Cell Neurosci 62:42–50
Leung AKL, Vyas S, Rood JE, Bhutkar A, Sharp PA, Chang P (2011) Poly (ADP-ribose) regulates stress responses and microRNA activity in the cytoplasm. Mol Cell 42:489–499
Acknowledgments
We would like to thank members of the Massirer and Bengston labs for their critical reading of this manuscript. We also would like to acknowledge the many works not included herein that have contributed to the understanding of the role of RNA-binding protein post-translational modifications.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Lovci, M.T., Bengtson, M.H., Massirer, K.B. (2016). Post-Translational Modifications and RNA-Binding Proteins. In: Yeo, G. (eds) RNA Processing. Advances in Experimental Medicine and Biology, vol 907. Springer, Cham. https://doi.org/10.1007/978-3-319-29073-7_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-29073-7_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-29071-3
Online ISBN: 978-3-319-29073-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)