Abstract
Despite the well-described importance of bacteriophages to bacterial pathogens, little is known about their influence on the many bacterial species that form beneficial symbioses with eukaryotes. Most insect species, for example, are infected with one or more maternally inherited symbionts, which provide nutritional and defensive services in exchange for housing. The pea aphid, Acyrthosiphon pisum, harbors at least seven common heritable symbionts that mediate a range of ecological interactions and has emerged as a model for studies of beneficial symbionts. One common pea aphid defensive symbiont, Hamiltonella defensa, protects against parasitic wasps, which are important natural enemies. The bacterium is itself infected by temperate bacteriophages, called APSEs (A cyrthosiphon p isum secondary endosymbionts), which are necessary for H. defensa-mediated protection. This represents the first known instance of a bacteria-insect mutualism requiring a viral partner. APSEs play other key roles in the regulation and maintenance of H. defensa: APSE loss results in high titers of the bacterial symbiont, which is correlated with severe fitness costs to the aphid host. These costs to the aphid incurred by phage loss likely lead to phage-free H. defensa-infected aphids being rapidly removed from the population, thus limiting the invasion potential of H. defensa. Below we review the roles of APSEs in the H. defensa-aphid defensive symbiosis and suggest a framework for future studies. We then make comparisons with another well-studied phage-symbiont interaction (Wolbachia-WO) and consider the roles of bacteriophages in the evolution of heritable symbioses and how they may influence other insect-associated bacteria, such as the gut microbiota.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
1.1 Mutualistic Viruses
Viruses are the most abundant and genetically diverse segment of the biosphere (Weinbauer 2004; Hatfull 2008). Due to their obligate reproductive parasitism, viruses have historically been associated with pathogenicity, yet they are now known to participate in mutualisms with most major organismal groups, including bacteria, fungi, plants, and animals. While the provision of pathogenicity loci to their bacterial hosts is a well-known attribute of phage infection, viruses may also be conditional or even obligate mutualists of eukaryotes. In some eukaryotic hosts, for example, infection by nonpathogenic viruses interferes with the establishment of pathogens via a number of mechanisms, including resource competition and immune system priming (Barton et al. 2007; Strive et al. 2010; Tillmann et al. 2001). Viruses may also mediate key ecological interactions for their hosts: persistent viruses confer cold and drought tolerance to plants, while insect-associated viruses improve food-plant quality for herbivores (reviewed in: Roossinck 2011). In a highly specialized obligate mutualism, many parasitic wasps employ viruses to overcome the defenses of the insects they attack. In this interaction, the Polydnavirus is stably integrated into the wasp genome and replicates only in specialized cells in the female wasp’s reproductive tract. The encapsidated form of the virus is injected into the wasp’s host along with the wasp egg and suppresses the host’s immune response to allow for the successful development of the wasp offspring (reviewed in: Strand 2010). Viruses thus engage in diverse beneficial interactions, which vary in complexity and specialization, and most are involved with providing hosts with a range of nutritional and defensive services.
Virus-mediated symbioses also exist at one remove: bacteriophages can infect insect-associated bacteria with important consequences for the insect host. Insects are easily the most diverse and abundant group of animals, and they have varied and dynamic interactions with microbes, especially bacteria. Insect guts, for example, often contain microbial communities (including bacteria, archaea, fungi, and protists) that assist in breaking down difficult to digest materials, such as cellulose (reviewed in: Engel and Moran 2013). These animal-microbe holobionts are therefore major players in nutrient recycling and other ecosystem services. Gut microbiota, which persist extracellularly and are often environmentally transmitted, have also been shown to detoxify plant allelochemicals, changing the range of permissible herbivore diets, and to modulate insect innate immunity and confer protection against pathogens and parasites (Dillon and Dillon 2004; Engel and Moran 2013). The majority of insect species are also likely infected with maternally transmitted symbionts, a phenomenon that is uncommon in mammals (Moran et al. 2008; Zug and Hammerstein 2012). Maternally transmitted symbionts contribute an additional source of heritable genetic variation that can be acted on by natural selection, so many invade and persist in natural populations by conferring net benefits relative to uninfected individuals (Oliver et al. 2010; Dillon and Dillon 2004; Hedges et al. 2008; Xie et al. 2010). Though bacteriophages are the most abundant organisms on earth, and are present as prophage in the majority of sequenced bacterial genomes (Bordenstein and Wernegreen 2004), little is known about their associations with the diverse and ecologically important array of insect symbionts described above. Recent findings, however, indicate the potential for a wide assortment of ecologically and evolutionarily important roles for these proxy mutualists (Table 7.1).
1.2 Heritable Symbionts Profoundly Influence Insect Ecology and Evolution
The acquisition of heritable microbial infections has allowed insects to exploit and specialize on nutrient poor diets, including plant sap and vertebrate blood (Gibson and Hunter 2010; Douglas 1998). Insects, like other animals, cannot synthesize essential amino acids and must acquire them from their diet or other sources. In aphids, which are small, soft-bodied herbivores, an ancient infection with the gamma-proteobacterium Buchnera aphidicola allowed its host to subsist on only plant phloem: a sugar-rich but nitrogen-poor diet. This ancient symbiosis (160–280 mya) allowed aphids to first exploit a previously unavailable resource and subsequently specialize and diversify (ca. 4400 spp.) with flowering plants (Moran et al. 1993). More generally, at least 10 % (>100,000 spp.) of extant insect species are hosts to one or more obligate nutrient-provisioning bacterial symbionts, demonstrating their importance in the evolutionary success of insects (Wernegreen 2002). While most obligate symbionts characterized perform a nutritional role, and those not explicitly examined are expected to, given that they occur in hosts specialized on restricted diets (Akman et al. 2002; Moran and Wernegreen 2000; Baumann 2005), obligate symbionts may possibly provide other, or additional, mutualistic benefits to their insect hosts, such as defense (Nakabachi et al. 2013).
Obligate nutritional symbionts cannot survive or reproduce outside of their host insects, and the insects themselves cannot reproduce, and generally do not survive, after antibiotic removal of nutritional symbionts (Douglas 1989). This inextricable mutual dependence results in a number of key characteristics shared among obligate symbionts. Insects control obligate symbionts by cordoning them off in specialized cells known as bacteriocytes, which are often grouped into organs called bacteriomes. Their strict maternal inheritance renders the bacteria’s fitness directly dependent on the host’s reproductive success (Vautrin et al. 2008), and insect hosts and obligate symbionts typically show patterns of cospeciation (Thao et al. 2000; Sauer et al. 2000; Clark et al. 2000). These domesticated bacteria also exhibit many of the smallest, most A-T-rich genomes yet sequenced and have lost entire functional gene groups (including those involved in DNA repair) yet retain pathways critical for essential nutrient provisioning (Moran and Wernegreen 2000; McCutcheon and Moran 2012; Gil et al. 2002, 2003).
The facultative symbionts of insects are even more common, likely infecting most species, and they exert far more varied effects on hosts (Feldhaar 2011). Facultative symbionts exhibit intermediate genome sizes between obligate symbionts and free-living bacteria, yet often contain a higher percentage of MGEs than either (Burke and Moran 2011; Degnan et al. 2009, 2010; Newton and Bordenstein 2011; Belda et al. 2010). Some facultative symbionts confer ecologically important traits valuable to the host organism, including heat protection, defense against natural enemies, and host-plant specification (Ferrari et al. 2007; Koga et al. 2003; Montllor et al. 2002; Scarborough et al. 2005; Tsuchida et al. 2004). Others are reproductive parasites, manipulating the host’s reproduction to favor infected female hosts at the expense of males or uninfected females to ensure their own propagation (Werren et al. 2008). The line between parasite and mutualist is often blurred: some bacterial clades are manipulators or beneficial depending on the host, and others are both at once. For example, Wolbachia spp., which may infect 40 % of arthropod species (Zug and Hammerstein 2012), are generally considered reproductive manipulators, but many strains also provide protection against a range of pathogens (reviewed in: Hamilton and Perlman 2013). In addition, many facultative symbionts confer benefits to infected individuals under some conditions but costs under others (Oliver et al. 2008). Tissue tropism of facultative symbionts is variable, depending on symbiont and host species, but they are not typically isolated in the bacteriocytes and hence are more likely, relative to obligate symbionts, to interact with other bacteria and their mobile elements. However, the limited diversity of bacterial taxa found in tissues generally frequented by facultative bacteria constrains their bacteriophage exposure relative to free-living bacteria (Gottlieb et al. 2008).
While the prevalence of bacteriophages in insect facultative symbionts is not known, many widespread symbionts, including Wolbachia, Arsenophonus, Spiroplasma, and Hamiltonella, are associated with active bacteriophages, and there are remnants of inactivated prophage in other species, including Regiella insecticola and Sodalis glossinidius (Bordenstein et al. 2006; Darby et al. 2010; Belda et al. 2010). One of the best studied phage-facultative symbiont interactions, and the only currently known instance of mutualism by proxy, involves the pea aphid, Acyrthosiphon pisum, its bacterial symbiont Hamiltonella defensa, and its temperate phage APSE (A cyrthosiphon p isum secondary endosymbiont). Below we will review the diverse roles attributed to APSE in the maintenance and regulation of a defensive symbiosis, in which infected aphids are protected against a common natural enemy, the parasitic wasp Aphidius ervi (Oliver et al. 2009). We will also draw comparisons to the other well-studied tripartite interaction between insects, Wolbachia and the temperate phage WO.
1.3 Introduction to Pea Aphids: Their Symbionts and Their Natural Enemies
The pea aphid, Acyrthosiphon pisum, is a cosmopolitan pest of cultivated legumes, including clover and alfalfa. In addition to the obligate nutritional symbiont Buchnera aphidicola (Sect. 7.1.2), individual pea aphids are usually infected with one or more facultative symbionts (Russell et al. 2013). This aphid has emerged as a model organism for identifying the phenotypic effects of infection with heritable symbionts, due to its cyclically parthenogenetic reproduction and the relative ease with which facultative symbionts can be manipulated among clonal lines. At present, seven facultative symbionts are known to frequently infect pea aphids, and all seven facultative symbionts are known or suspected to mediate important ecological interactions including heat shock protection, host-plant specification, and protection from natural enemies (Oliver et al. 2010; Lukasik et al. 2013b).
One of the most prevalent natural enemies of the pea aphid in North America is the hymenopteran parasitic wasp Aphidius ervi (Braconidae). A female wasp injects an egg into the aphid hemocoel and the resulting wasp larva feeds and develops within a still living aphid. When larval development is complete, the wasp kills and consumes the remaining contents of the aphid and pupates within the desiccated and hardened cuticle, called a “mummy” (Fig. 7.1) (Oliver et al. 2005). The parasitoid wasp manipulates the aphid to support the wasp larva’s stage-specific nutritional needs (Caccia et al. 2005); as part of these manipulations, Ap. ervi injects a venom that degrades the aphid’s reproductive system, which, depending on timing of attack, can completely castrate the aphid (Digilio et al. 2000). Aphid embryos that survive venom castration may then be degraded by teratocytes, large polyploid cells that dissociate from the developing wasp’s extraembryonic membrane (Falabella et al. 2009; Digilio et al. 2000). Parasitoids may also specifically manipulate the nutritional symbiosis. In susceptible aphids uninfected by facultative symbionts, parasitism results in an increased abundance of Buchnera, as well as the number and mass of bacteriocytes relative to unparasitized controls (Cloutier and Douglas 2003). Parasitism influences the type and quantity of amino acid provisioning performed by Buchnera, presumably to the parasitoid’s benefit (Rahbe et al. 2002); wasps fail to thrive in aposymbiotic (Buchnera-free) aphids (Falabella et al. 2000).
Developmental trajectories of pea aphids (Acyrthosiphon pisum) parasitized by the wasp Aphidius ervi in the presence and absence of facultative endosymbiont Hamiltonella defensa and its associated phage, APSE (A cyrthosiphon p isum secondary endosymbiont). Development times assume a 16L:8D light cycle and constant 20C temperature. (a) H. defensa-free aphid: Wasp injects a single egg into a preadult aphid and embryonic development occurs from 0 to 3 days after oviposition. Wasp larval development continues from 3 to 9 days post-parasitism. The wasp larva pupates at approximately 9 days, killing and “mummifying” the aphid. Single adult wasp emerges from aphid “mummy” 14–15 days post-parasitism (He 2008). (b) H. defensa-infected aphid without APSE: wasp development proceeds as in “1a.” (c) H. defensa-infected aphid with phage APSE: wasp mortality at either egg or larval stage, depending on APSE strain (brackets indicate that larval development stage may or may not occur) (Martinez et al. 2014b). Aphids survive to reproduce (Oliver et al. 2009)
1.4 Bacteria-Mediated Defense Against Wasp Aphidius ervi
An early study indicated that pea aphid clonal lines varied tremendously in susceptibility to the wasp Ap. ervi, with some lines entirely resistant and others highly susceptible (Henter and Via 1995). Originally, it was assumed that the basis for this variation was encoded in the aphid genome, although this was surprising given that aphids, including Ac. pisum, lack a strong encapsulation response, i.e., the typical insect innate cellular immune response to internal parasites (Carver and Sullivan 1988; Laughton et al. 2011). Shortly after this, several facultative symbionts were characterized in this aphid (Chen and Purcell 1997; Fukatsu et al. 2000; Sandstrom et al. 2001; Darby et al. 2001), and two of these symbionts, Hamiltonella defensa and Serratia symbiotica, were found to contribute to variation in susceptibility to parasitism (Oliver et al. 2003). Subsequent studies confirmed that resistance to wasps was largely due to infection with H. defensa, and in the absence of facultative symbionts, most Ac. pisum are highly susceptible to Ap. ervi (Oliver et al. 2005, 2009), although some aphid-genotype-based variation in susceptibility also occurs (Martinez et al. 2014a). Furthermore, it was shown that symbiont-mediated resistance was due to intra-aphid factors: wasps readily attack infected and uninfected aphids but are far less likely to complete development in infected aphids (Oliver et al. 2003). H. defensa also rescues the reproductive capacity of parasitized aphids, which are normally castrated by parasitoid venoms: the fact that resistant aphids not only survive but reproduce demonstrates a direct, heritable, benefit to H. defensa infection in the presence of parasitoid wasps (Oliver et al. 2008).
H. defensa is estimated to infect 14 % of aphid species and occurs in other hemipteran insects (Oliver et al. 2010). In addition to Ac. pisum, an anti-parasitoid role for H. defensa has only been demonstrated in the black bean aphid, Aphis fabae, and the cowpea aphid, Aphis craccivora (Schmid et al. 2012; Asplen et al. 2014). In the grain aphid, Sitobion avenae, and the blackberry-cereal aphid, Sitobion fragariae, H. defensa provides no or very low protection from either Aphidius ervi or another parasitoid wasp, Ephedrus plagiator (Lukasik et al. 2013a; Lukasik et al. 2015). In all three aphid species where protection is conferred, H. defensa’s protection is specific to parasitoid species identity (Asplen et al. 2014; Cayetano and Vorburger 2015; McLean and Godfray 2015; Martinez et al. 2016).
2 The Role of Phage in an Aphid Defensive Mutualism
Given the clear benefits of protection owing to infection with H. defensa, the next question emerged: what mechanisms underlie symbiont-based protection? Unfortunately, like most heritable bacterial symbionts, H. defensa is uncultivable and hence not amenable to standard microbiological assays. In order to identify potential mechanisms underlying symbiont-based protection, Moran et al. produced a preliminary sequence of the H. defensa genome (2005). This effort reported a number of putative toxins and pathogenicity loci but also confirmed the presence of an intact prophage called APSE, which has proven to be highly influential in the beneficial heritable symbiosis between aphid Ac. pisum and H. defensa: APSE is, in fact, the first active phage known to be required by an insect mutualism. As in heritable bacteria, the lack of horizontal escape routes in isolated animal hosts renders phage fitness dependent on the vertical transmission of their host bacterial line to the next generation of insect hosts (Vautrin and Vavre 2009). Phages in mutualistic symbioses may therefore encode factors critical to protective or nutritional services, which could act alone or in conjunction with those of bacterial origin (Moran et al. 2005). On the other hand, phage effects on symbiont abundance may reduce the effectiveness of conferred benefits and limit symbiont and phage invasion into insect host populations. Finally, increasing the number of required players may generally lead to instability as each will have their own biotic and abiotic optima, such as overlapping but distinct ideal temperature ranges, resulting in a mutualism that functions well only under restricted conditions.
2.1 Phage Can Provision Functions Important to Bacteria-Insect Mutualisms
APSE was first reported in a European line of Ac. pisum (van der Wilk et al. 1999), and it was subsequently determined that its host was H. defensa (Sandstrom et al. 2001). APSEs are dsDNA viruses with capsid morphology similar to the Podoviridae and share sequence similarity with P22 (Degnan and Moran 2008b; van der Wilk et al. 1999). Synteny and nucleotide identity are conserved among APSE haplotypes with the exception of a virulence cassette region (VCR), which varies in length and contains the toxic effector molecules hypothesized to provide or support the defensive symbiosis (Moran et al. 2005; Degnan and Moran 2008a, b). Seven APSE variants (APSE1-7) have been designated on the basis of the assortment of putative eukaryotic toxins and bacterial lysis genes; their genomes vary in length from 36 to 39 kbp (van der Wilk et al. 1999; Degnan and Moran 2008a). The presence of toxin-encoding APSEs in a defensive symbiont strongly suggested its involvement in the protective phenotype (Moran et al. 2005).
APSE variants are each associated with one of three different eukaryotic toxins: cytolethal-distending toxin subunit B homolog (CdtB; APSEs 2, 6, and 7), a Shiga-like toxin (Stx; APSEs 1, 4, and 5), and a putative toxin containing a YD repeat—a motif characterized by a highly conserved tyrosyl-aspartate dipeptide (Ydp; APSE3) (Degnan and Moran 2008a; Moran et al. 2005; van der Wilk et al. 1999). In North American pea aphids, where only APSE2 and APSE3 have been reported, aphids infected with H. defensa containing either APSE variant receive protection against parasitoids, and the phage variant generally correlates with the intensity of the defensive phenotype: APSE2s are associated with moderate protection, while APSE3s are associated with high to complete protection from the wasp Ap. ervi (Oliver et al. 2005). It was next found that some descendants of a pea aphid clonal lab line infected with H. defensa and APSE3 spontaneously lost their bacteriophage (more on this below) while retaining the bacterial symbiont. This allowed the creation of experimental lines that shared the same pea aphid and H. defensa genotype, with or without APSE3. After it was confirmed that components other than APSE3 were not lost from the H. defensa chromosome, these experimental lines were used to determine the contribution of APSE to the protective phenotype. As previously reported, pea aphids with neither H. defensa nor APSE were susceptible to parasitism, while those with H. defensa and APSE were highly resistant; however, the line infected with H. defensa without APSE was just as susceptible to the wasp as the H. defensa-free line (Fig. 7.1) (Oliver et al. 2009). This same study confirmed that this result was not a quirk of either the particular H. defensa strain or aphid clonal genotype, as numerous additional lines that lost APSE also lost resistance to wasps. Together, the experimental and correlation-based evidence makes a strong case that infection with APSE3, and probably APSE2, is required for H. defensa to produce the protective phenotype.
Some APSE-associated toxins have been found in pathogenic bacteria where their functions have been characterized. The APSE2-associated CdtBs (also found in APSE6 from the aphid Chaitophorus and APSE7 from the whitefly Bemisia tabaci), for example, were first identified in E. coli (Johnson and Lior 1987). CdtB is the DNase I subunit of the cyclomodulin cytolethal-distending toxin, which disrupts actively dividing eukaryotic cells (Ohara et al. 2004). Homologs of the Shiga-like toxin found in APSE1 (Ac. pisum from the Netherlands), APSE4 (the cowpea aphid, Aphis craccivora), and APSE5 (aphid Uroleucon rudbeckia) are also found in other enteric pathogens, including E. coli, where they were shown to prevent protein synthesis by cleaving ribosomal RNA via N-glycosidase action (Endo et al. 1988). The Ydp encoded by APSE3 is associated with the H. defensa strains that confer the most protection, but they are understood least at the functional level: however the YD dipeptide motif in the reading frame (ORF) is associated with binding carbohydrates, and at least one Ydp is associated with eukaryotic toxicity (Degnan and Moran 2008a). It is important to remember that the inability to culture H. defensa and APSEs limits the tools available for functional studies, so whether phage toxins, acting alone or in concert with H. defensa or aphid-encoded factors, cause harm to wasps is not yet known.
The potential contribution of other APSE variants to the protective phenotype has received little attention. APSE4 (Stx) wielding H. defensa protect their native cowpea aphid host against some, but not all, parasitoids (Asplen et al. 2014), but an APSE4/H. defensa strain in the grain aphid Sitobion avenae provided no protection from either of the two parasitoid species assayed, although it may confer protection against other wasps attacking this aphid (Lukasik et al. 2013, 2015). Some H. defensa strains may be maintained in host populations by conferring thermal protection or other benefits rather than defense against parasitoids (Russell and Moran 2006).
APSE also commonly infects the widespread insect symbiont Arsenophonus (Enterobacteriaceae), though less is known about the phage’s roles in these hosts. That Arsenophonus-associated APSEs exhibit much greater genetic diversity, and H. defensa-infecting APSEs form a distinct branch within the larger Arsenophonus-APSE phylogenetic tree, suggests that Arsenophonus was APSE’s original host (Duron 2014). While Arsenophonus has not been found in pea aphids, it has been reported in other aphids and is widespread in some Ap. craccivora populations (Nováková et al. 2009; Brady and White 2013; Duron 2014), where Arsenophonus is known to influence dietary breadth (Wagner et al. 2015). Eukaryotic toxins homologous to an open reading frame (called ORF D) in H. defensa-APSE’s variable cassette region have been found in wasp-associated Arsenophonus-APSEs, where they may contribute to a male-killing reproductively manipulative phenotype, but whether any potential eukaryotic toxins are found in hemipteran-infecting Arsenophonus-APSEs is unknown (Wilkes et al. 2010).
2.2 Role of Phage in Horizontal Transmission of Ecologically Important Traits
Phages are well-known vectors of lateral gene transmission among bacterial lineages, contributing functional pathways that profoundly affect host ecology and evolution (Ochman et al. 2000). In heritable symbionts, phage may move traits that influence not only the bacterial host but also the animal host (Moran et al. 2005). Phylogenetic studies reveal that APSEs move among H. defensa strains and hence likely move traits important in aphid protection (Degnan and Moran 2008b). The high G + C content of the virulence cassette region, where the putative toxins are found, suggests that it is foreign to both APSE and H. defensa, further highlighting the potential for lateral exchange of genes relevant to defensive symbioses among phages (Degnan and Moran 2008a). Phylogenetic evidence and transfection experiments show that facultative symbionts, including H. defensa, also move horizontally within and among arthropod species, transferring ecologically significant bacterial or phage-encoded traits in the process (Oliver et al. 2010). In the lab, an experimental transfer of APSE4-infected H. defensa from cowpea aphids (Ap. craccivora) to pea aphids (Ac. pisum) resulted in the instant acquisition of increased resistance to parasitism (Oliver et al. 2005). The observation that phages move traits among heritable symbionts, and symbionts among arthropods, has led to speculation of a reservoir of ecologically important traits that are shared among communities of interacting species (Henry et al. 2013; Jaenike 2012; Moran 2007). However, aphid species which share host plants (e.g., pea aphids and cowpea aphids) are associated with distinct H. defensa strains and APSE variants, which suggests that there may be factors limiting the spread of both bacteria and phage between insect species (Degnan and Moran 2008a; Dykstra et al. 2014; Weldon 2015). While the historical evidence for lateral phage movement between H. defensa strains is strong, experimental movement of APSEs in the laboratory has not been reported.
2.3 Phage Roles in the Regulation and Maintenance of Heritable Symbiosis
Intra-host bacterial symbiont density can be important for the maintenance and performance of heritable symbionts and their insect hosts. Within-host bacterial abundance can affect conferred phenotypes (e.g., Noda et al. 2001; Ikeda et al. 2003), rates of horizontal transfer, the establishment of novel infections (Chafee et al. 2010), and the maintenance of tripartite symbioses (Jaenike 2009; Bordenstein and Bordenstein 2011). All stable beneficial heritable symbiont infections must be coordinated between host and symbiont to strike a balance between a sufficient titer to produce the beneficial phenotype and ensure vertical transmission to progeny and an excessive titer that might deplete host resources and negatively affect both host reproduction and the bacterium’s chances of reaching the next generation of insects (Jaenike 2009).
Virulent phages are predacious on their hosts and hence capable of influencing bacterial abundance. Temperate phages capable of integrating into the host genome can also influence bacterial abundance, either through excision and lysis or by influencing the ecology of the bacterial host, such as by modifying reproductive rates relative to uninfected cells (Lin et al. 1977). Phage-bacteria interactions are often only considered in terms of integrated prophages and lytic behavior, but a wide range of complex interactions exists, including persistent infections distinguished by, for example, episomal prophage or phage shedding or budding (Weinbauer 2004; Calendar and Abedon 2005). As these different phage lifestyles will differentially influence bacterial abundance, it will be important to understand the basic biology of these interactions.
2.4 APSEs Are Temperate, with Variable Lifecycles and No Known Lytic Triggers
Within-aphid H. defensa populations are heterogeneous for APSE prophage integration (Degnan and Moran 2008a), and APSEs tend to outnumber their bacterial hosts 10- to 100-fold within the pea aphid hemolymph, suggesting that most are not maintained as lysogens (Moran et al. 2005; Martinez et al. 2014b; Oliver et al. 2009). The lytic triggers or density-dependent factors that control APSE’s interactions with its host-limited bacterial population are unknown, though there is some evidence wasp parasitism may trigger APSE-induced lysis of H. defensa cells (Degnan and Moran 2008a; Martinez et al. 2014b). Lysis of a subset of H. defensa may be a prerequisite to the release of phage-encoded toxins into the aphid hemolymph to disrupt wasp development (Moran et al. 2005; Martinez et al. 2014b); this has not, however, been examined mechanistically, and H. defensa retains potential transporters for phage-encoded toxins that would render lysis superfluous (Degnan et al. 2009). Lytic activity may not be a specialized response to wasp parasitism, but rather APSE’s general reaction to stressors acting on the host insect. Several APSE genes have been proposed as orthologs to lysogenic cycle control genes found in other lambdoid phages: ORF1 (or P1) is homologous to lambdoid phage HB19’s C1 protein, based both off of sequence similarity and genomic placement; ORF2 (or P2) has been identified as an ortholog to the cro-like gene in lambdoid phage 434 (van der Wilk et al. 1999). Sequencing results suggest that some APSE variants may now be inactivated prophage, though none of these variants are present in the pea aphid defense mutualism and their biology is largely unknown (Degnan and Moran 2008a). In laboratory-reared pea aphid clones, H. defensa is vertically transmitted at rates approaching 100 % (Oliver et al. 2010). The highly protective APSE3 phage infections are also transmitted with very high fidelity but can be spontaneously lost at very low rates as described above (Oliver et al. 2009). Moderately protective APSE2s, however, have never been lost from lab-held H. defensa lines (Weldon et al. 2013). The underlying basis for the differential persistence of APSE2 and APSE3 across aphid generations is currently unclear but may be influenced by differential rates of phage integration between variants. Further, if not all H. defensa cells within an individual aphid carry APSEs, then intraspecific competition between APSE-infected and APSE-uninfected H. defensa could reduce the proportion of infected bacteria, making it more likely that only uninfected symbionts will pass through the tight trans-generational bottleneck.
2.5 APSE3 Loss Is Associated with Consistent and Immediate Increases in H. defensa Titer
One study has examined the effects of APSE on within-host H. defensa abundance and reports results consistent with phage roles in the regulation of the protective symbiont (Weldon et al. 2013). That APSE3s are often lost from H. defensa allows for crossline comparisons between APSE-infected and APSE-uninfected lines, and in all cases, APSE-free aphid lines carried significantly higher H. defensa titers than aphid lines with related H. defensa strains that maintained APSE3 infections. Similarly, APSE losses make it possible, although difficult due to the rarity of the event, to establish experimental lines comprised of the same aphid host clonal lineage and identical H. defensa genotypes but varying with respect to APSE infection status, to experimentally show the effects of phage loss on symbiont abundance. In this case, estimates of symbiont abundance revealed that phage-free pea aphids contained significantly more H. defensa than aphids with APSE3 at all examined ages in aphid development. The increase in H. defensa abundance following APSE loss was dramatic, up to ninefold at its maximum. This, however, was not the gradual result of H. defensa equilibrium divergence over the course of many generations reared apart. Some phage-harboring aphid mothers produced a mix of phage-infected and phage-free offspring, and comparisons among their offspring found that siblings without phage had significantly higher titers of H. defensa than their phage-harboring clonal sisters. While the other common North American pea aphid-associated phage variant, APSE2, has proven too stably infective to naturally produce phage-free identical H. defensa strains, the same study reported an inverse correlation between APSE2 numbers and H. defensa titers consistent with phage effects on bacterial abundance (Weldon et al. 2013).
2.6 APSE3 Loss Is Severely Deleterious to Pea Aphid Fitness
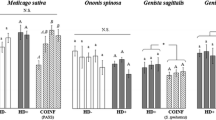
The dramatically increased H. defensa titers associated with the loss of APSE3 suggested potentially harmful effects for the aphid host. In component fitness assays, costs to infection with H. defensa + APSE have generally been small or difficult to show (Oliver et al. 2006, 2008; Russell and Moran 2005). H. defensa, however, relies on B. aphidicola for some of the same nutrients it provides its aphid host (Degnan et al. 2009), and at higher H. defensa densities, competition with aphid tissues for B. aphidicola-provisioned resources may increase. To assess the effects of phage loss on aphid performance, three component fitness parameters (fecundity, development time, and fresh weight at adulthood) were examined in phage-free and phage-harboring pea aphids (Weldon et al. 2013). In each instance, it was found that the absence of APSE3 significantly increased fitness costs to the aphid host (Fig. 7.2). On average, aphids lacking APSE3 reproduced 18 h later, weighed 20 % less at adulthood, and produced roughly half as many total offspring over the course of their lifespans compared to clonal aphids with APSE3 (Weldon et al. 2013). Thus, APSE loss resulted in a huge fitness deficit for genetically identical aphids. The negative effects associated with high H. defensa abundance, combined with the loss of protection from parasitoids (Sect. 7.2.1), essentially convert a heritable mutualist, H. defensa, into a heritable pathogen.
Aphids infected with APSE-free H. defensa weigh ~20 % less at adulthood, first reproduce on average 18 h later, and produce half as many total offspring as their uninfected clonemates. Aphids infected with H. defensa + APSE show no significant component fitness assay detriments relative to uninfected clonemates (Weldon et al. 2013; Oliver et al. 2006)
2.7 APSEs May Influence the Maintenance of H. defensa Diversity in Natural Populations
As noted above, vertically transmitted symbionts, such as H. defensa, provide heritable variation that can be acted on by host-level selection. While more work is needed in field populations, laboratory-based population cage studies show that aphids infected with H. defensa and APSE3 increase rapidly in the population when parasitism pressure is present but decrease in the absence of natural enemies (Oliver et al. 2008). Thus, benefits in the presence and costs in the absence of parasitism likely partially explain why field populations exhibit varying infection frequencies that never reach fixation (Russell et al. 2013). The recurrent loss of APSE3 furthermore produces aphids that suffer doubly from the loss of anti-parasitoid protection and the introduction of additional and quite severe costs to H. defensa infection (Weldon et al. 2013). Thus negative (i.e., purifying) selection at the host level likely rapidly removes these phage-free, H. defensa-infected aphids from the population. This would explain the infrequency of phage-free H. defensa in field surveys (Oliver et al. 2009; Weldon et al. 2015). It may also explain the maintenance of APSE2 H. defensa strains, which are more prevalent in field populations despite being inferior defenders (Weldon et al. 2013, 2015). Hence, there may be a trade-off between defensive ability (APSE3 > APSE2) and stability of the interaction (APSE2 > APSE3), such that APSE3 H. defensa strains are only maintained in populations with heavy parasitism pressure. Temporal variation in parasitism pressure may lead to cyclical replacement of highly protective strains with highly stable strains or vice versa.
3 WO Phage of Wolbachia spp.
3.1 Wolbachia Are Associated with a Group of Bacteriophages Known as WO
Roles for viruses in heritable symbiosis have been substantially investigated in one other tripartite interaction: the Rickettsiales symbiont Wolbachia, its temperate phage WO, and their diverse insect hosts. The Wolbachia group comprises both facultative strains with the capacity for host-swapping (primarily infecting arthropods) and obligate mutualist strains (mostly infecting filarial nematodes) which undergo far stricter vertical inheritance (Bordenstein and Reznikoff 2005; Kent and Bordenstein 2010; Foster et al. 2005; Masui et al. 2000). Facultative Wolbachia strains are associated with both beneficial defensive phenotypes, such as improved resistance to microbial pathogens (reviewed in: Hamilton and Perlman 2013), and reproductive parasitism (reviewed in: Stouthamer et al. 1999; Werren et al. 2008). Most facultative strains are infected with one or more active temperate phages, called WO, which can move laterally among Wolbachia strains occupying different phylogenetic supergroups (Bordenstein and Wernegreen 2004; Baldo et al. 2006; Chafee et al. 2010; Masui et al. 2000; Tanaka et al. 2009). Given that WO has primarily been studied in parasitic rather than beneficial strains, and has been reviewed recently (Kent and Bordenstein 2010; Metcalf and Bordenstein 2012), we will limit our coverage to summarizing major findings and making comparisons with APSE.
3.2 WO and Wolbachia Density Interactions Affect Host Phenotype and Are in Turn Affected by Host Genotype and Abiotic Factors
Wolbachia has been described as the “master manipulator” of arthropod reproduction (Werren et al. 2008). It is the only bacterial species known to employ all four strategies of reproductive parasitism, allowing invasion and maintenance in host populations by enhancing the fitness of infected female insects at the expense of males and uninfected females. These strategies are cytoplasmic incompatibility (CI) between infected males and uninfected females, feminization of male offspring, male killing (MK), and parthenogenesis induction (PI) (Stouthamer et al. 1999). Microscopy and inverse correlations between Wolbachia and WO abundance indicate that WO likely lyses bacterial cells and thereby influences bacterial titers (Bordenstein et al. 2006). As Wolbachia density is the best predictor of CI levels in several divergent insect taxa (Noda et al. 2001; Ikeda et al. 2003), WO may routinely affect the strength of CI, and other density-dependent phenotypes (Hurst et al. 2000), through its effects on symbiont abundance. By reducing the strength of CI, phage WO may limit the invasion potential of both phage and symbiont to new hosts. This may be unavoidable, however, if routine lysis is required for within-host maintenance of WO infections. Of course, if pathogen defense or other Wolbachia-mediated beneficial phenotypes are also density dependent, then WO activity may similarly affect these traits. In vitro experiments suggest that Wolbachia-mediated viral resistance, at least, is directly dependent on per-cell Wolbachia density (Frentiu et al. 2010). While there is no correlation between the strength of APSE-H. defensa-mediated protection and the within-host densities of either partner (Martinez et al. 2014b), APSE’s role in limiting H. defensa’s fitness costs in aphids indicates that phage effects on symbiont density in both systems are important determinants of reproductive success for the insect hosts and their heritable bacteria.
Phage-bacteria density interactions are dependent on a number of biotic and abiotic factors. Temperature, for example, decreases the abundance of Wolbachia but increases the abundance of WO in the wasp Nasonia (Bordenstein and Bordenstein 2011). As heat shock is a common trigger for lytic activity in lysogenic phages, WO may often mediate temperature effects on Wolbachia-conferred phenotypes. The protective phenotype conferred by H. defensa is lost at higher temperatures (Bensadia et al. 2006; Guay et al. 2009), though this has not been associated with changes in either APSE or H. defensa abundance. Establishment of equilibria in phage-bacteria density interactions may also be specific to host genotype: introgression of Nasonia giraulti’s genome into the cytoplasm of N. vitripennis infected with Wolbachia led to a dramatic increase in Wolbachia densities and an equally precipitous decline in WO densities (Chafee et al. 2011). Thus, novel environments, or genotype x genotype x genotype interactions, may routinely result in varying effects of phage on symbiont abundance and thereby influence the stability of the interaction, the strength of conferred phenotypes, and the likelihood of successful establishment of a horizontally acquired symbiont.
3.3 Phage WO May Provide Effector Molecules Necessary for the Induction of Reproductive Parasitism
While no association between WO and Wolbachia-induced defensive phenotypes has been identified, there is speculation that WO may contribute to manipulative Wolbachia phenotypes through more than density interactions. This is unlikely to be universally the case, however, as some WO-free strains of Wolbachia are effective manipulators (Gavotte et al. 2007). A proliferation of ankyrin-repeat proteins is encoded in Wolbachia genomes and is often located on, or near, WOs. Given that ankyrin-repeat motifs are involved in a range of cell functions in eukaryotes, such as protein-protein interactions, and are otherwise rare in bacteria, they have been hypothesized to contribute to Wolbachia’s facility for reproductive manipulation (reviewed in: Metcalf and Bordenstein 2012; Kent and Bordenstein 2010).
Alternately, WO-encoded factors may limit the detrimental effects of CI. CI occurs when a male insect (a dead end for Wolbachia) harbors a strain of Wolbachia not present in his mate, decreasing the fitness of uninfected females relative to infected females, which can mate with infected or uninfected males (reviewed in: Werren et al. 2008). Often though, multiple strains of Wolbachia infect both individuals in a mating pair, resulting in complex compatibility dynamics. It has been hypothesized that WO associated with some Wolbachia strains encode “rescue” factors capable of negotiating among strain incompatibilities (Saridaki et al. 2011). In brief, WO prophages encode a DNA methyltransferase, Met2, found in all CI-rescuing strains of “A” group Wolbachia but absent from strains not capable of rescuing CI. Phage-encoded factors could therefore render these CI-rescuing strains of Wolbachia circumstantial reproductive mutualists in particular coinfections. Note that Met2 is not present in CI-rescuing Wolbachia from other phylogenetic groups, indicating that this mechanism, if valid, is not universal. The importance of methylation to CI induction, however, is unclear: while methylation patterns have been found to underlie some Wolbachia-induced feminizations, there is no similar evidence in CI induction (Metcalf and Bordenstein 2012). Unlike the APSE system, no direct comparisons of phenotypes induced by phage-harboring and phage-free bacterial populations have been possible, so only genomic and correlation-based evidence currently inform this aspect of the association.
4 Bacteriophage Roles in the Evolution of Heritable Symbiont Genomes
4.1 Bacteriophages Contribute to the Content and Architecture of Symbiont Genomes
High rates of nucleotide change are associated with obligate, bacteriocyte-associated symbionts due to relaxations in selection pressure, loss of repair mechanisms, and genetic bottlenecks, but a lack of mobile genetic elements provides them with fairly stable architectures (e.g., Tamas et al. 2002; Degnan et al. 2005; van Ham et al. 2003). In contrast, many facultative symbionts, such as Wolbachia, exhibit low rates of sequence divergence but high levels of rearrangement due to the relatively high percentage of mobile genetic elements in their genomes (Wu et al. 2004; Degnan et al. 2010).
Though bacteriophages are absent in obligate bacterial mutualists, most of those are ancient, and phages may have proliferated early in the development of the association. Any evidence of MGEs, however, has been lost due to genetic drift and a bias toward deletions (Moran and Plague 2004; Kuo et al. 2009). In species which have recently transitioned to a symbiotic lifestyle, such as Sodalis glossinidius and Serratia symbiotica, there is a proliferation of all classes of mobile genetic elements (MGEs), including phages (Belda et al. 2010; Burke and Moran 2011). Phages can transport other MGEs into the genome, as in the case of the insertion sequence (IS)-carrying WO phage in Wolbachia, indicating that MGE proliferation can occur as a self-reinforcing treadmill (Tanaka et al. 2009). Phages provided nearly 18 % of the coding sequence in the S. glossinidius genome, although more than half were pseudogenized, a major process in the transition to a host-dependent lifestyle (Moya et al. 2008; Belda et al. 2010; Burke and Moran 2011). MGEs, including phages, can inactivate genes; however, in S. glossinidius, the role of MGEs in pseudogenization is dwarfed by the role of frameshift and nonsense mutations (Belda et al. 2010). Inactivated genes may also proliferate due to the lack of purifying selection (Burke and Moran 2011). Prior phage infections may also give rise to modern extrachromosomal DNA features: S. glossinidius’s plasmid pSOG3 may be producing virions (occasionally spotted in culture and the ORFs for assembly are present) and appears to be a coalescence of two ancestral phages, one P22-like and the other epsilon 15-like (Clark et al. 2007).
4.2 Bacteriophages Allow for Recombination Between Coinfecting Strains and More Divergent Taxa
Superinfections by multiple heritable symbionts (and different strains of the same symbiont species) are common in many insect groups, including aphids, although heritable symbiont “communities” are usually less complex than gut communities, and coinfecting mutualists may even occupy separate tissues within the host (Engel and Moran 2013; Oliver et al. 2013). Superinfections provide opportunities for the exchange of phage and other MGEs, though there may be substantial barriers. For example, the sister symbionts H. defensa and R. insecticola are quite distinct, with high nucleotide divergence and only 55 % of genes shared in common. This is despite frequent host and tissue sharing and high percentages of their genomes dedicated to MGEs, two factors that would seem to facilitate genetic exchange (Degnan et al. 2010). On the other hand, comparative Wolbachia genomics suggests that significant recombination and mixing occurs between coinfecting Wolbachia strains, even those in different supergroups (Klasson et al. 2009). This includes whole-phage transfer and recombination of phage-encoded regions, not only among Wolbachia strains but also into other Rickettsiaceae. More dramatically, WO-associated regions have moved into the genomes of some of Wolbachia’s eukaryotic hosts (reviewed in: Metcalf and Bordenstein 2012).
5 Future Directions in Insect-Associated Phage Research
Given that only two heritable phage-symbiont systems have received much attention, it is unsurprising that there remain many basic questions unanswered about both APSE and WO and phage-symbiont interactions more broadly. Further investigation is needed both to determine the prevalence of active bacteriophages in insect symbionts and to understand how their interactions may be similar to or distinct from the behavior of phages in other bacterial communities.
5.1 Questions Remaining in the APSE-H. defensa System
As described in Sect. 7.2.1, the mere presence of the variable toxin cassette region in APSE led to speculation that APSE-encoded toxin homologs caused wasp mortality (Moran et al. 2005), yet no functional assays confirm this. APSE-associated toxins could be moved into expression vectors and the purified products assayed in vitro for toxicity to cultivable wasp tissues (eggs, larvae, and teratocytes) (e.g., Lawrence 1990; Okuda and Kadonookuda 1995; Vinson et al. 1994; Grbic and Strand 1998). Furthermore, it is unknown whether these putative toxins are capable of functioning in isolation or whether they require other factors located on the phage or H. defensa chromosomes. The CdtB subunit, for example, is integrated into a holotoxin in other organisms, including E. coli and Campylobacter spp., comprised of three subunits (Ohara et al. 2004); the CdtA and CdtC subunits bind to the target cell via an unknown receptor and allow the B subunit to enter the cell (Asakura et al. 2007). The three APSE variants that carry CdtB-homologs, however, do not include any ORFs homologous to the A and C subunits, indicating that they use a different delivery system (Degnan and Moran 2008a). Some CdtB-APSEs do contain uncharacterized ORFs in close association with the putative toxin, and these possibilities perform analogous roles (Degnan and Moran 2008a). Despite their lack of strong homology to known proteins, the genomic placement of these ORFs has led to speculation that they interact with APSEs’ toxins. Uncharacterized ORFs are also associated with other APSE toxins, including ydp (APSE3) and stx (APSE1, APSE4, and APSE5). In general, mechanisms of toxin delivery remain unclear across APSE variants and H. defensa strains. H. defensa has an intact type 3 secretion system that may allow for APSE toxin secretion, but toxin release into the aphid hemocoel may also be accomplished via lysis of a subset of H. defensa cells (Moran et al. 2005; Martinez et al. 2014b).
Facultative symbionts are generally uncultivable in cell-free media, and not easily amenable to transformation or reverse genetics, but some have been cultured in association with insect cells (Pontes and Dale 2006). H. defensa has been successfully grown and maintained in culture with a number of dipteran and lepidopteran cell lines (Darby et al. 2005), opening the possibility of in vitro studies aimed at understanding the functional mechanisms underlying host defense as well as key questions about phage lifestyle. As described in Sect. 7.2, APSE loss is a major determinant of both parasitoid protection and aphid fecundity. However, nothing is known about how or why APSEs are lost, in part because very little is known about how it is normally maintained; the ability to work with APSEs in culture may provide answers.
The range of effects attributable to APSEs will be, at least in part, a function of their diversity, which is poorly characterized. Three of the seven known APSE variants were identified from H. defensa strains infecting a single aphid species—the pea aphid Ac. pisum—but this likely reflects only depth of sampling (Degnan and Moran 2008a). Furthermore, the pea aphid has diversified into numerous genetically distinct host races that specialize on cultivated herbaceous legumes (e.g., Ferrari et al. 2007; Frantz et al. 2006; Peccoud et al. 2009; Via et al. 2000), and APSE sampling to date has concentrated on pea aphids found on alfalfa. Historical contingencies, natural enemy identity, and transmission opportunities relating to environmental factors may have each contributed to the development and maintenance of current APSE associations, and we need a better appreciation of APSE diversity to adequately investigate these components.
5.2 Determining the Prevalence and Roles of Active Phage in Other Heritable Bacteria
Mobile genetic elements, particularly prophage-like regions, are quite common in heritable facultative symbionts (as discussed in Sect. 7.4.1). In H. defensa, for example, mobile DNA, including a 59 kbp plasmid, one active prophage (APSE), and 22 phage-like regions, comprises 21 % of the genome (Degnan et al. 2009). However, this says little about the proportion of insect symbionts containing active phages. More genomic surveys, combined with microscopy and bioassays (such as those done for APSEs), are needed before we can assess whether the unique selective pressures on maternally inherited bacteria alter the likelihood of phage acquisition and maintenance.
When phage are present in heritable insect symbionts, then we expect that the general roles—mutualism factor provisioning, horizontal gene transfer, and symbiont population control—identified in the APSE and WO systems will occur, although each system is likely to exhibit its own particular features and dynamics. There are several potential advantages to phage-encoded effector molecules, such as increased genomic copy number of key mutualism factors, allowing for higher transcription rates. This potential benefit has been ascribed to plasmids encoding products for amino acid production, though in the polyploid obligate aphid symbiont B. aphidicola, this shift may instead decrease transcription (Plague et al. 2003). It is also possible that encoding factors critical for cooperative functions on mobile elements prevent nonproductive cheating (reviewed in: Rankin et al. 2011). Bacteria often secrete public goods, such as toxins enabling invasion of eukaryotic tissues, which benefit not only the secretor but also other bacteria in the population. Many components of the “cooperative” portion of the bacterial secretome are encoded by mobile genetic elements, including phages. This may simply be a coincidence of secreted goods’ inclusion in the general class of secondary metabolites: horizontal transmission of core functions is lethally dangerous, so the only molecules that are consistently exchanged are those that the bacterium could survive without. An alternative and intriguing explanation is that infectious public goods provide cooperators with a neat solution to the control of cheaters: noncooperators can be transformed into cooperators via infection.
The spread of phage-encoded public goods may be limited by the acquisition of phage resistance by noncooperators; however, resistance in itself may be as costly as the averted lysogenic conversion and may also be insufficient: some WO strains, for example, are capable of not only infecting cheaters but also of killing phage-resistant defectors via a toxin-antitoxin addiction system (Kent and Bordenstein 2010; Engelberg-Kulka and Glaser 1999). Of course, populations of infectious public-goods-carrying particles would themselves be susceptible to invasion by incompatible noncooperators. Heavy purifying selection, enhanced by trans-generational bottlenecks that increase within-host relatedness, may also limit cheaters among heritable symbiotic bacteria, decreasing the utility of correction by infection (Herre et al. 1999). Furthermore, phage in heritable bacterial systems may be more likely to lose their capacity for horizontal transmission relative to those found in free-living bacteria, due to a lack of access to uninfected hosts. Selection may also favor the transfer of phage-encoded mutualism factors to the bacterial genome to avert the costs of infection, and phages may be lost through vertical transmission failure more readily than bacterial symbionts from processes such as competition between infected and uninfected heritable bacterial symbionts.
Phages associated with heritable symbionts may be less prone to lateral exchange than their free-living relatives. A large majority of prokaryotic viruses are dsDNA-tailed bacteriophages in the order Caudovirales; functional and genetic diversity in this group is driven primarily by nonhomologous recombination and the promiscuous lateral exchange of whole functional modules (Krupovic et al. 2011). This modularity renders dsDNA phages highly effective carriers of secondary bacterial metabolites. However, while Caudovirales may generally evolve according to the modular theory (Botstein 1980), the extreme host-limited lifestyle of symbiont-associated phages may isolate them from common phage gene pools to the extent that modular exchange takes a backseat to other processes, such as non-modular recombination and mutation (reviewed in: Metcalf and Bordenstein 2012).
Finally, the role of phage in density interactions with host bacteria is particularly difficult to predict due to our limited understanding of phage reproductive behavior outside of readily manipulated model systems. Phage-bacteria interactions are dependent on transmission rate, length of survival outside the host, and timing of self-replication, all of which may be limited by phage, bacterial, and environmental factors (Abedon 2008). Phages occurring in closed systems, including insect-restricted bacteria, are at risk of running through their limited host population. This may be prevented by processes such as latent-time mutability (Heilmann et al. 2010), superinfection exclusion or immunity (Susskind 1980; Susskind et al. 1974; McGrath et al. 2002), and mechanical shielding by dead bacteria, where the number of dead reach a density where phages are likely to irreversibly inject DNA into dead cells rather than the remaining uninfected live cells (Rabinovitch et al. 2003). Nutritional limitations can also decrease bacterial receptivity to phage infection (Ptashne 2004) or phage reproductive rate (Ptashne 2004; Lenski 1988). Environmental hostility to free phage particles, such as pH extremes, can increase the rate of lysogenization, and rates of decay by insect host hemolymph factors may be a major unexplored determinant of bacteriophage lifestyle (Pantastico-Caldas et al. 1992). Of course, environmental stressors severe enough to damage DNA may lead to prophage induction, and abiotic influences may predictably and seasonally influence phage-bacteria-insect relations, as seen in other viral communities: marine phages, for example, produce fewer lysogens in summer months (McDaniel et al. 2002).
5.3 Technical Challenges of Phage Discovery
While detecting and characterizing the diversity of uncultivable insect-associated bacteria has become routine through the development of universal 16S rRNA techniques, no simple universal tool for identifying bacteriophages in symbionts is available. Instead they are often found by chance through electron microscopy or bacterial genome sequencing. High-throughput sequencing has allowed for the production of genomic data on total organismal and tissue-specific viral samples. Sequence data alone, however, is often uninformative due to the high genic richness and sequence divergence associated with viruses: phage metagenomic studies consistently report high proportions of totally novel sequences with no known function (Hatfull 2008). A study of mosquito viromes, for example, was unable to match nearly half of the sequences to any known viral genome (Ng et al. 2011). Furthermore, overall virome samples may miss phage diversity or fail to pick up on phage presence altogether, particularly if eukaryotic viruses are present. For instance, a metavirome sample taken from worker honeybees suffering from an unknown source of malaise was dominated by a small handful of eukaryotic viruses, and only 0.7 % of those reads were from known phages (Granberg et al. 2013).
The host ranges of even well-studied phages are seldom known (Flores et al. 2011), which can limit the interpretation of virome results. One possible way to estimate host ranges is to analyze the legacy of bacteriophage infections. Stern has suggested that clustered regularly interspaced short palindromic repeat (or CRISPR) spacers in microbiome samples be viewed as a “database of fragments from phage and plasmid genomes” (Stern et al. 2012). This could allow researchers to associate phages with at least a subset of their potential hosts even in the absence of direct experimental evidence or integrated prophage. Interestingly, there appears to be variation in the presence of CRISPR systems in two aphid heritable symbionts: H. defensa has CRISPR sequences, while its sister species, R. insecticola, does not (Degnan et al. 2009; Hansen et al. 2012), but the basis of this variation is unknown. More generally, it is unknown why CRISPR/cas systems are absent from 60 % of bacterial genomes despite their near universality in archaea and the ubiquity of bacteriophage predation (Horvath and Barrangou 2010).
5.4 Phage Roles in the More Complex Insect-Associated Bacterial Communities
Recently, much attention has been paid to the broader microbiome and to the nutritional and immune services it provides to its animal host. Studies in humans suggest that a characteristic, somewhat taxonomically restricted bacterial community may itself host a characteristic, somewhat taxonomically restricted bacteriophage community (Stern et al. 2012) and that microbiomes and viromes covary between individuals (Minot et al. 2011). Though functional studies of insect gut microbiota lag behind those in mammals, it is known that these consortia play significant parts in host development, nutrition, and immune function (Dillon and Dillon 2004; Engel and Moran 2013). The effects of phages in these more complex insect-associated bacterial communities are largely unknown but may differ from those found in heritable symbionts. Phages may influence community diversity through species-specific predation and community structure by lysing the most abundant species or strains (reviewed in: Weinbauer 2004). Such density-dependent “kill-the-winner” dynamics potentially reduce functions associated with mutualistic bacteria, limit population sizes, and may also drive bacterial strain diversification (Middelboe et al. 2009). Heterogeneity in bacterial groups tends to reduce group productivity, so bacterial diversity per se may lower the efficiency of mutualistic functions provisioned by the microbiota (Mendes-Soares et al. 2014). However, phage-driven diversification can increase a bacterial population’s adaptability to changing environments (Williams 2013), decreasing the likelihood that environmental perturbations will catastrophically disrupt the microbiome.
Bacteriophage-mediated horizontal gene transfer may also contribute to the development of functional redundancy and allow newly acquired bacteria to rapidly adapt to a shared host. This has been proposed in sponge-associated bacterial communities, with “forcible” adaptation spreading mutualistically important functions across taxa (Fan et al. 2012), which may limit the risk of function loss via density-dependent strain predation. Microbiome and virome diversity therefore reciprocally influence one another, and phage may contribute to the maintenance of diversity with or without disrupting the mutualistic function of the insect-associated bacterial community. However, the metabolic loads or receptor modifications associated with phage resistance could theoretically decrease the efficiency of a mutualism. Flyg and colleagues (1980) identified a strain of Serratia marcescens (an insect pathogen with symbiont relatives) that was more resistant to phage but more vulnerable to the cercropid immune response and less virulent in drosophilids.
6 Concluding Remarks
While bacteriophages are the most abundant biological entities on earth, we currently know little about their prevalence in insect-associated bacteria. When they are present, they can profoundly influence not only the ecology and evolution of their bacterial hosts but also that of the insect in which they both reside. While tripartite mutualisms in insects remain understudied, there is growing interest in their roles, as recent findings indicate that phages can provide key services required for symbiont function and dramatically alter the within-insect population dynamics of symbiotic microorganisms. These within-host effects can in turn be important in the spread and maintenance of heritable symbionts in host populations. Phages also likely play key roles in the horizontal transfer of ecologically relevant traits and in the transition to a host-associated lifestyle, potentially resulting in more dynamic and responsive bacterial symbionts. Even less is known about tripartite interactions operating under natural conditions. The additional complexity of tripartite interactions potentially renders them less stable when faced with environmental perturbations. Thus, phage infection in heritable symbiosis likely provides advantages and disadvantages for all interacting players.
References
Abedon ST (2008) Bacteriophage ecology: population growth, evolution, and impact of bacterial viruses. Cambridge University Press, Cambridge
Akman L, Yamashita A, Watanabe H, Oshima K, Shiba T, Hattori M, Aksoy S (2002) Genome sequence of the endocellular obligate symbiont of tsetse flies, Wigglesworthia glossinidia. Nat Genet 32(3):402–407. doi:10.1038/ng986
Asakura M, Hinenoya A, Alam MS, Shima K, Zahid SH, Shi L, Sugimoto N, Ghosh AN, Ramamurthy T, Faruque SM, Nair GB, Yamasaki S (2007) An inducible lambdoid prophage encoding cytolethal distending toxin (Cdt-I) and a type III effector protein in enteropathogenic Escherichia coli. Proc Natl Acad Sci USA 104(36):14483–14488. doi:10.1073/pnas.0706695104
Asplen MK, Bano N, Brady CM, Desneux N, Hopper KR, Malouines C, Oliver KM, White JA, Heimpel GE (2014) Specialisation of bacterial endosymbionts that protect aphids from parasitoids. Ecol Entomol 39(6):736–739
Baldo L, Bordenstein S, Wernegreen JJ, Werren JH (2006) Widespread recombination throughout Wolbachia genomes. Mol Biol Evol 23(2):437–449. doi:10.1093/molbev/msj049
Barton ES, White DW, Cathelyn JS, Brett-McClellan KA, Engle M, Diamond MS, Miller VL, Virgin HW (2007) Herpesvirus latency confers symbiotic protection from bacterial infection. Nature 447 (7142):326–U327. doi:10.1038/Nature05762
Baumann P (2005) Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu Rev Microbiol 59:155–189. doi:10.1146/annurev.micro.59.030804.121041
Belda E, Moya A, Bentley S, Silva FJ (2010) Mobile genetic element proliferation and gene inactivation impact over the genome structure and metabolic capabilities of Sodalis glossinidius, the secondary endosymbiont of tsetse flies. BMC Genomics 11(1):449
Bensadia F, Boudreault S, Guay JF, Michaud D, Cloutier C (2006) Aphid clonal resistance to a parasitoid fails under heat stress. J Insect Physiol 52(2):146–157. doi:10.1016/j.jinsphys.2005.09.011
Bordenstein SR, Bordenstein SR (2011) Temperature affects the tripartite interactions between bacteriophage WO, Wolbachia, and cytoplasmic incompatibility. PLoS One 6(12), e29106
Bordenstein SR, Reznikoff WS (2005) Mobile DNA in obligate intracellular bacteria. Nat Rev Microbiol 3(9):688–699. doi:10.1038/nrmicro1233
Bordenstein SR, Wernegreen JJ (2004) Bacteriophage flux in endosymbionts (Wolbachia): infection frequency, lateral transfer, and recombination rates. Mol Biol Evol 21(10):1981–1991. doi:10.1093/molbev/msh211
Bordenstein SR, Marshall ML, Fry AJ, Kim U, Wernegreen JJ (2006) The tripartite associations between bacteriophage, Wolbachia, and arthropods. PLoS Pathog 2(10):1024. doi:10.1371/journal.ppat.0020106
Botstein D (1980) A theory for the modular evolution of bacteriophages. Ann NY Acad Sci 354:484–490. doi:10.1111/j.1749-6632.1980.tb27987.x
Brady CM, White JA (2013) Cowpea aphid (Aphis craccivora) associated with different host plants has different facultative endosymbionts. Ecol Entomol 38(4):433–437
Burke GR, Moran NA (2011) Massive genomic decay in Serratia symbiotica, a recently evolved symbiont of aphids. Genome Biol Evol 3:195–208. doi:10.1093/Gbe/Evr002
Caccia S, Leonardi MG, Casartelli M, Grimaldi A, de Eguileor M, Pennacchio F, Giordana B (2005) Nutrient absorption by Aphidius ervi larvae. J Insect Physiol 51(11):1183–1192. doi:10.1016/j.jinsphys.2005.06.010
Calendar R, Abedon ST (2005) The bacteriophages, 2nd edn. Oxford University Press, Oxford
Carver M, Sullivan D (1988) Encapsulative defence reactions of aphids (Hemiptera: Aphididae) to insect parasitoids (Hymenoptera: Aphidiidae and Aphelinidae). Ecology and Effectiveness of Aphidophaga: 299–303
Cayetano L, Vorburger C (2015) Symbiont‐conferred protection against Hymenopteran parasitoids in aphids: how general is it? Ecol Entomol 40(1):85–93. doi:10.1111/een.12161
Chafee ME, Funk DJ, Harrison RG, Bordenstein SR (2010) Lateral phage transfer in obligate intracellular bacteria (Wolbachia): verification from natural populations. Mol Biol Evol 27(3):501–505. doi:10.1093/molbev/msp275
Chafee ME, Zecher CN, Gourley ML, Schmidt VT, Chen JH, Bordenstein SR, Clark ME (2011) Decoupling of host-symbiont-phage coadaptations following transfer between insect species. Genetics 187(1):203–215. doi:10.1534/genetics.110.120675
Chen DQ, Purcell AH (1997) Occurrence and transmission of facultative endosymbionts in aphids. Curr Microbiol 34(4):220–225
Clark MA, Moran NA, Baumann P, Wernegreen JJ (2000) Cospeciation between bacterial endosymbionts (Buchnera) and a recent radiation of aphids (Uroleucon) and pitfalls of testing for phylogenetic congruence. Evolution 54(2):517–525. doi:10.1554/0014-3820(2000)054[0517:Cbbeba]2.0.Co;2
Clark AJ, Pontes M, Jones T, Dale C (2007) A possible heterodimeric prophage-like element in the genome of the insect endosymbiont Sodalis glossinidius. J Bacteriol 189(7):2949–2951. doi:10.1128/Jb.00913-06
Cloutier C, Douglas AE (2003) Impact of a parasitoid on the bacterial symbiosis of its aphid host. Entomol Exp Appl 109(1):13–19
Darby AC, Birkle LM, Turner SL, Douglas AE (2001) An aphid-borne bacterium allied to the secondary symbionts of whitefly. FEMS Microbiol Ecol 36(1):43–50
Darby AC, Chandler SM, Welburn SC, Douglas AE (2005) Aphid-symbiotic bacteria cultured in insect cell lines. Appl Environ Microbiol 71(8):4833–4839. doi:10.1128/Aem.71.8.4833-4839.2005
Darby AC, Choi JH, Wilkes T, Hughes MA, Werren JH, Hurst GDD, Colbourne JK (2010) Characteristics of the genome of Arsenophonus nasoniae, son-killer bacterium of the wasp Nasonia. Insect Mol Biol 19:75–89. doi:10.1111/j.1365-2583.2009.00950.x
Degnan PH, Moran NA (2008a) Diverse phage-encoded toxins in a protective insect endosymbiont. Appl Environ Microbiol 74(21):6782–6791. doi:10.1128/Aem.01285-08
Degnan PH, Moran NA (2008b) Evolutionary genetics of a defensive facultative symbiont of insects: exchange of toxin-encoding bacteriophage. Mol Ecol 17(3):916–929
Degnan PH, Lazarus AB, Wernegreen JJ (2005) Genome sequence of Blochmannia pennsylvanicus indicates parallel evolutionary trends among bacterial mutualists of insects. Genome Res 15(8):1023–1033
Degnan PH, Yu Y, Sisneros N, Wing RA, Moran NA (2009) Hamiltonella defensa, genome evolution of protective bacterial endosymbiont from pathogenic ancestors. Proc Natl Acad Sci USA 106(22):9063–9068. doi:10.1073/pnas.0900194106
Degnan PH, Leonardo TE, Cass BN, Hurwitz B, Stern D, Gibbs RA, Richards S, Moran NA (2010) Dynamics of genome evolution in facultative symbionts of aphids. Environ Microbiol 12(8):2060–2069. doi:10.1111/j.1462-2920.2009.02085.x
Digilio MC, Isidoro N, Tremblay E, Pennacchio F (2000) Host castration by Aphidius ervi venom proteins. J Insect Physiol 46(6):1041–1050
Dillon RJ, Dillon VM (2004) The gut bacteria of insects: nonpathogenic interactions. Annu Rev Entomol 49:71–92. doi:10.1146/annurev.ento.49.061802.123416
Douglas AE (1989) Mycetocyte symbiosis in insects. Biol Rev 64(4):409–434. doi:10.1111/j.1469-185X.1989.tb00682.x
Douglas AE (1998) Nutritional interactions in insect-microbial symbioses: Aphids and their symbiotic bacteria Buchnera. Annu Rev Entomol 43:17–37
Duron O (2014) Arsenophonus insect symbionts are commonly infected with APSE, a bacteriophage involved in protective symbiosis. FEMS Microbiol Ecol 90(1):184–194
Dykstra HR, Weldon SR, Martinez AJ, White JA, Hopper KR, Heimpel GE, Asplen MK, Oliver KM (2014) Factors limiting the spread of the protective symbiont Hamiltonella defensa in Aphis craccivora aphids. Appl Environ Microbiol 80(18):5818–5827
Endo Y, Tsurugi K, Yutsudo T, Takeda Y, Ogasawara T, Igarashi K (1988) Site of action of a vero toxin (Vt2) from Escherichia coli O157-H7 and of shiga toxin on eukaryotic ribosomes - RNA N-glycosidase activity of the toxins. Eur J Biochem 171(1–2):45–50
Engel P, Moran NA (2013) The gut microbiota of insects - diversity in structure and function. FEMS Microbiol Rev 37(5):699–735. doi:10.1111/1574-6976.12025
Engelberg-Kulka H, Glaser G (1999) Addiction modules and programmed cell death and antideath in bacterial cultures. Annu Rev Microbiol 53:43–70. doi:10.1146/annurev.micro.53.1.43
Falabella P, Tremblay E, Pennacchio F (2000) Host regulation by the aphid parasitoid Aphidius ervi: the role of teratocytes. Entomol Exp Appl 97(1):1–9
Falabella P, Riviello L, De Stradis ML, Stigliano C, Varricchio P, Grimaldi A, de Eguileor M, Graziani F, Gigliotti S, Pennacchio F (2009) Aphidius ervi teratocytes release an extracellular enolase. Insect Biochem Mol 39(11):801–813. doi:10.1016/j.ibmb.2009.09.005
Fan L, Reynolds D, Liu M, Stark M, Kjelleberg S, Webster NS, Thomas T (2012) Functional equivalence and evolutionary convergence in complex communities of microbial sponge symbionts. Proc Natl Acad Sci USA 109(27):E1878–E1887. doi:10.1073/pnas.1203287109
Feldhaar H (2011) Bacterial symbionts as mediators of ecologically important traits of insect hosts. Ecol Entomol 36(5):533–543. doi:10.1111/j.1365-2311.2011.01318.x
Ferrari J, Scarborough CL, Godfray HCJ (2007) Genetic variation in the effect of a facultative symbiont on host-plant use by pea aphids. Oecologia 153(2):323–329. doi:10.1007/s00442-007-0730-2
Flores CO, Meyer JR, Valverde S, Farr L, Weitz JS (2011) Statistical structure of host-phage interactions. Proc Natl Acad Sci USA 108(28):E288–E297. doi:10.1073/pnas.1101595108
Flyg C, Kenne K, Boman HG (1980) Insect pathogenic properties of Serratia marcescens – phage-resistant mutants with a decreased resistance to Cecropia immunity and a decreased virulence to Drosophila. J Gen Microbiol 120(1):173–181
Foster J, Ganatra M, Kamal I, Ware J, Makarova K, Ivanova N, Bhattacharyya A, Kapatral V, Kumar S, Posfai J, Vincze T, Ingram J, Moran L, Lapidus A, Omelchenko M, Kyrpides N, Ghedin E, Wang S, Goltsman E, Joukov V, Ostrovskaya O, Tsukerman K, Mazur M, Comb D, Koonin E, Slatko B (2005) The Wolbachia genome of Brugia malayi: Endosymbiont evolution within a human pathogenic nematode. PLos Biol 3(4):599–614. ARTN e121
Frantz A, Plantegenest M, Mieuzet L, Simon JC (2006) Ecological specialization correlates with genotypic differentiation in sympatric host-populations of the pea aphid. J Evol Biol 19(2):392–401. doi:10.1111/j.1420-9101.2005.01025.x
Frentiu FD, Robinson J, Young PR, McGraw EA, O’Neill SL (2010) Wolbachia-mediated resistance to dengue virus infection and death at the cellular level. PLoS One 5(10):e13398. doi:10.1371/journal.pone.0013398
Fukatsu T, Nikoh N, Kawai R, Koga R (2000) The secondary endosymbiotic bacterium of the pea aphid Acyrthosiphon pisum (Insecta : Homoptera). Appl Environ Microbiol 66(7):2748–2758
Gavotte L, Henri H, Stouthamer R, Charif D, Charlat S, Boulétreau M, Vavre F (2007) A survey of the bacteriophage WO in the endosymbiotic bacteria Wolbachia. Mol Biol Evol 24(2):427–435
Gibson CM, Hunter MS (2010) Extraordinarily widespread and fantastically complex: comparative biology of endosymbiotic bacterial and fungal mutualists of insects. Ecol Lett 13(2):223–234. doi:10.1111/j.1461-0248.2009.01416.x
Gil R, Sabater-Munoz B, Latorre A, Silva FJ, Moya A (2002) Extreme genome reduction in Buchnera spp.: toward the minimal genome needed for symbiotic life. Proc Natl Acad Sci USA 99(7):4454–4458. doi:10.1073/pnas.062067299
Gil R, Silva FJ, Zientz E, Delmotte F, Gonzalez-Candelas F, Latorre A, Rausell C, Kamerbeek J, Gadau J, Holldobler B, van Ham RCHJ, Gross R, Moya A (2003) The genome sequence of Blochmannia floridanus: comparative analysis of reduced genomes. Proc Natl Acad Sci USA 100(16):9388–9393. doi:10.1073/pnas.1533499100
Gottlieb Y, Ghanim M, Gueguen G, Kontsedalov S, Vavre F, Fleury F, Zchori-Fein E (2008) Inherited intracellular ecosystem: symbiotic bacteria share bacteriocytes in whiteflies. FASEB J 22(7):2591–2599. doi:10.1096/fj.07-101162
Granberg F, Vicente-Rubiano M, Rubio-Guerri C, Karlsson OE, Kukielka D, Belak S, Sanchez-Vizcaino JM (2013) Metagenomic detection of viral pathogens in Spanish honeybees: co-infection by aphid lethal paralysis, Israel acute paralysis and Lake Sinai viruses. PLoS One 8(2), e57459
Grbic M, Strand MR (1998) Shifts in the life history of parasitic wasps correlate with pronounced alterations in early development. Proc Natl Acad Sci USA 95(3):1097–1101
Guay JF, Boudreault S, Michaud D, Cloutier C (2009) Impact of environmental stress on aphid clonal resistance to parasitoids: role of Hamiltonella defensa bacterial symbiosis in association with a new facultative symbiont of the pea aphid. J Insect Physiol 55(10):919–926. doi:10.1016/j.jinsphys.2009.06.006
Hamilton PT, Perlman SJ (2013) Host defense via symbiosis in Drosophila. Plos Pathog 9(12). doi:10.1371/journal.ppat.1003808
Hansen AK, Vorburger C, Moran NA (2012) Genomic basis of endosymbiont-conferred protection against an insect parasitoid. Genome Res 22(1):106–114
Hatfull GF (2008) Bacteriophage genomics. Curr Opin Microbiol 11(5):447–453. doi:10.1016/j.mib.2008.09.004
He XZ (2008) Reproductive behavior of Aphidius ervi Haliday (Hymenoptera: Aphidiidae). Massey University Dissertation, Palmerston North, New Zealand
Hedges LM, Brownlie JC, O’Neill SL, Johnson KN (2008) Wolbachia and virus protection in insects. Science 322(5902):702. doi:10.1126/science.1162418
Heilmann S, Sneppen K, Krishna S (2010) Sustainability of virulence in a phage-bacterial ecosystem. J Virol 84(6):3016–3022. doi:10.1128/Jvi.02326-09
Henry LM, Peccoud J, Simon JC, Hadfield JD, Maiden MJC, Ferrari J, Godfray HCJ (2013) Horizontally transmitted symbionts and host colonization of ecological niches. Curr Biol 23(17):1713–1717
Henter HJ, Via S (1995) The potential for coevolution in a host-parasitoid system. 1. Genetic-variation within an aphid population in susceptibility to a parasitic wasp. Evolution 49(3):427–438. doi:10.2307/2410267
Herre EA, Knowlton N, Mueller UG, Rehner SA (1999) The evolution of mutualisms: exploring the paths between conflict and cooperation. Trends Ecol Evol 14(2):49–53. doi:10.1016/S0169-5347(98)01529-8
Horvath P, Barrangou R (2010) CRISPR/Cas, the immune system of bacteria and archaea. Science 327(5962):167–170. doi:10.1126/science.1179555
Hurst GDD, Johnson AP, von der Schulenburg JHG, Fuyama Y (2000) Male-killing Wolbachia in Drosophila: a temperature-sensitive trait with a threshold bacterial density. Genetics 156(2):699–709
Ikeda T, Ishikawa H, Sasaki T (2003) Infection density of Wolbachia and level of cytoplasmic incompatibility in the Mediterranean flour moth, Ephestia kuehniella. J Invertebr Pathol 84(1):1–5. doi:10.1016/S0022-2011(03)00106-X
Jaenike J (2009) Coupled population dynamics of endosymbionts within and between hosts. Oikos 118(3):353–362. doi:10.1111/j.1600-0706.2008.17110.x
Jaenike J (2012) Population genetics of beneficial heritable symbionts. Trends Ecol Evol 27(4):226–232. doi:10.1016/j.tree.2011.10.005
Johnson WM, Lior H (1987) Response of Chinese-hamster ovary cells to a cytolethal distending toxin (Cdt) of Escherichia coli and possible misinterpretation as heat-labile (Lt) enterotoxin. FEMS Microbiol Lett 43(1):19–23
Kent BN, Bordenstein SR (2010) Phage WO of Wolbachia: lambda of the endosymbiont world. Trends Microbiol 18(4):173–181. doi:10.1016/j.tim.2009.12.011
Klasson L, Westberg J, Sapountzis P, Nasiund K, Lutnaes Y, Darby AC, Veneti Z, Chen LM, Braig HR, Garrett R, Bourtzis K, Andersson SGE (2009) The mosaic genome structure of the Wolbachia wRi strain infecting Drosophila simulans. Proc Natl Acad Sci USA 106(14):5725–5730. doi:10.1073/pnas.0810753106
Koga R, Tsuchida T, Fukatsu T (2003) Changing partners in an obligate symbiosis: a facultative endosymbiont can compensate for loss of the essential endosymbiont Buchnera in an aphid. Proc R Soc Lond B Biol 270(1533):2543–2550. doi:10.1098/rspb.2003.2537
Krupovic M, Prangishvili D, Hendrix RW, Bamford DH (2011) Genomics of bacterial and archaeal viruses: dynamics within the prokaryotic virosphere. Microbiol Mol Biol Rev 75(4):610. doi:10.1128/Mmbr.00011-11
Kuo CH, Moran NA, Ochman H (2009) The consequences of genetic drift for bacterial genome complexity. Genome Res 19(8):1450–1454. doi:10.1101/gr.091785.109
Laughton AM, Garcia JR, Altincicek B, Strand MR, Gerardo NM (2011) Characterisation of immune responses in the pea aphid, Acyrthosiphon pisum. J Insect Physiol 57(6):830–839. doi:10.1016/j.jinsphys.2011.03.015
Lawrence PO (1990) Serosal cells of Biosteres longicaudatus (Hymenoptera, Braconidae) – ultrastructure and release of polypeptides. Arch Insect Biochem 13(3–4):199–216
Lenski RE (1988) Dynamics of interactions between bacteria and virulent bacteriophage. Adv Microb Ecol 10:1–44
Lin L, Bitner R, Edlin G (1977) Increased reproductive fitness of Escherichia coli Lambda-lysogens. J Virol 21(2):554–559
Lukasik P, Dawid MA, Ferrari J, Godfray HC (2013a) The diversity and fitness effects of infection with facultative endosymbionts in the grain aphid, Sitobion avenae. Oecologia. doi:10.1007/s00442-013-2660-5
Lukasik P, van Asch M, Guo HF, Ferrari J, Godfray HCJ (2013b) Unrelated facultative endosymbionts protect aphids against a fungal pathogen. Ecol Lett 16(2):214–218. doi:10.1111/Ele.12031
Lukasik P, Weldon SR, van Asch M, Patel V, Dennis A, Husnik F, Vorburger C, Ferrari J, Godfray HCJ, Oliver KM, Russell JA (2016) Strain diversity of parasitoid-protective endosymbionts of aphids: correlating phenotype with genotype (in prep)
Martinez AJ, Ritter SG, Doremus MR, Russell JA, Oliver KM (2014a) Aphid-encoded variability in susceptibility to a parasitoid. BMC Evol Biol 14(1):127. doi:10.1186/1471-2148-14-127
Martinez AJ, Weldon SR, Oliver KM (2014b) Effects of parasitism on aphid nutritional and protective symbioses. Mol Ecol 23(6):1594–1607. doi:10.1111/mec.12550
Martinez AJ, Kim KL, Harmon JP, Oliver KM (2016) Specificity of multi-modal aphid defenses against two rival parasitoids. PLoS One
Masui S, Kamoda S, Sasaki T, Ishikawa H (2000) Distribution and evolution of bacteriophage WO in Wolbachia, the endosymbiont causing sexual alterations in arthropods. J Mol Evol 51(5):491–497
McCutcheon JP, Moran NA (2012) Extreme genome reduction in symbiotic bacteria. Nat Rev Microbiol 10(1):13–26. doi:10.1038/Nrmicro2670
McDaniel L, Houchin LA, Williamson SJ, Paul JH (2002) Plankton blooms – Lysogeny in marine Synechococcus. Nature 415(6871):496. doi:10.1038/415496a
McGrath S, Fitzgerald GF, van Sinderen D (2002) Identification and characterization of phage-resistance genes in temperate lactococcal bacteriophages. Mol Microbiol 43(2):509–520. doi:10.1046/j.1365-2958.2002.02763.x
McLean AH, Godfray HCJ (2015) Evidence for specificity in symbiont-conferred protection against parasitoids. Proc R Soc B Biol Sci 282(1811):20150977. doi:10.1098/rspb.2015.0977
Mendes-Soares H, Chen I-CK, Fitzpatrick K, Velicer GJ (2014) Chimaeric load among sympatric social bacteria increases with genotype richness. Proc R Soc B Biol Sci 281(1787). doi:10.1098/rspb.2014.0285
Metcalf JA, Bordenstein SR (2012) The complexity of virus systems: the case of endosymbionts. Curr Opin Microbiol 15(4):546–552. doi:10.1016/j.mib.2012.04.010
Middelboe M, Holmfeldt K, Riemann L, Nybroe O, Haaber J (2009) Bacteriophages drive strain diversification in a marine Flavobacterium: implications for phage resistance and physiological properties. Environ Microbiol 11(8):1971–1982. doi:10.1111/j.1462-2920.2009.01920.x
Minot S, Sinha R, Chen J, Li HZ, Keilbaugh SA, Wu GD, Lewis JD, Bushman FD (2011) The human gut virome: inter-individual variation and dynamic response to diet. Genome Res 21(10):1616–1625. doi:10.1101/gr.122705.111
Montllor CB, Maxmen A, Purcell AH (2002) Facultative bacterial endosymbionts benefit pea aphids Acyrthosiphon pisum under heat stress. Ecol Entomol 27(2):189–195
Moran NA (2007) Symbiosis as an adaptive process and source of phenotypic complexity. Proc Natl Acad Sci USA 104:8627–8633. doi:10.1073/pnas.0611659104
Moran NA, Plague GR (2004) Genornic changes following host restriction in bacteria. Curr Opin Genet Dev 14(6):627–633. doi:10.1016/j.gde.2004.09.003
Moran NA, Wernegreen JJ (2000) Lifestyle evolution in symbiotic bacteria: insights from genomics. Trends Ecol Evol 15(8):321–326
Moran NA, Munson MA, Baumann P, Ishikawa H (1993) A molecular clock in endosymbiotic bacteria is calibrated using the insect hosts. Proc R Soc Lond B Biol 253(1337):167–171
Moran NA, Degnan PH, Santos SR, Dunbar HE, Ochman H (2005) The players in a mutualistic symbiosis: insects, bacteria, viruses, and virulence genes. Proc Natl Acad Sci USA 102(47):16919–16926. doi:10.1073/pnas.0507029102
Moran NA, McCutcheon JP, Nakabachi A (2008) Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet 42:165–190. doi:10.1146/annurev.genet.41.110306.130119
Moya A, Pereto J, Gil R, Latorre A (2008) Learning how to live together: genomic insights into prokaryote-animal symbioses. Nat Rev Genet 9(3):218–229. doi:10.1038/Nrg2319
Nakabachi A, Ueoka R, Oshima K, Teta R, Mangoni A, Gurgui M, Oldham NJ, van Echten-Deckert G, Okamura K, Yamamoto K, Inoue H, Ohkuma M, Hongoh Y, Miyagishima S, Hattori M, Piel J, Fukatsu T (2013) Defensive bacteriome symbiont with a drastically reduced genome. Curr Biol 23(15):1478–1484. doi:10.1016/j.cub.2013.06.027
Newton ILG, Bordenstein SR (2011) Correlations between bacterial ecology and mobile DNA. Curr Microbiol 62(1):198–208. doi:10.1007/s00284-010-9693-3
Ng TFF, Willner DL, Lim YW, Schmieder R, Chau B, Nilsson C, Anthony S, Ruan YJ, Rohwer F, Breitbart M (2011) Broad surveys of DNA viral diversity obtained through viral metagenomics of mosquitoes. PLoS One 6(6), e20579
Noda H, Koizumi Y, Zhang Q, Deng KJ (2001) Infection density of Wolbachia and incompatibility level in two planthopper species, Laodelphax striatellus and Sogatella furcifera. Insect Biochem Mol 31(6–7):727–737
Nováková E, Hypša V, Moran NA (2009) Arsenophonus, an emerging clade of intracellular symbionts with a broad host distribution. BMC Microbiol 9(1):143
Ochman H, Lawrence JG, Groisman EA (2000) Lateral gene transfer and the nature of bacterial innovation. Nature 405(6784):299–304. doi:10.1038/35012500
Ohara M, Oswald E, Sugai M (2004) Cytolethal distending toxin: a bacterial bullet targeted to nucleus. J Biochem 136(4):409–413. doi:10.1093/jb/mvh154
Okuda T, Kadonookuda K (1995) Perilitus coccinellae teratocyte olypeptide – evidence for production of a teratocyte-specific 540 kda protein. J Insect Physiol 41(9):819–825
Oliver KM, Russell JA, Moran NA, Hunter MS (2003) Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. Proc Natl Acad Sci USA 100(4):1803–1807. doi:10.1073/pnas.0335320100
Oliver KM, Moran NA, Hunter MS (2005) Variation in resistance to parasitism in aphids is due to symbionts not host genotype. Proc Natl Acad Sci USA 102(36):12795–12800. doi:10.1073/pnas.0506131102
Oliver KM, Moran NA, Hunter MS (2006) Costs and benefits of a superinfection of facultative symbionts in aphids. Proc R Soc B Biol Sci 273(1591):1273–1280. doi:10.1098/rspb.2005.3436
Oliver KM, Campos J, Moran NA, Hunter MS (2008) Population dynamics of defensive symbionts in aphids. Proc R Soc B Biol Sci 275(1632):293–299. doi:10.1098/rspb.2007.1192
Oliver KM, Degnan PH, Hunter MS, Moran NA (2009) Bacteriophages encode factors required for protection in a symbiotic mutualism. Science 325(5943):992–994. doi:10.1126/science.1174463
Oliver KM, Degnan PH, Burke GR, Moran NA (2010) Facultative symbionts in aphids and the horizontal transfer of ecologically important traits. Annu Rev Entomol 55:247–266. doi:10.1146/annurev-ento-112408-085305
Oliver KM, Smith AH, Russell JA (2013) Defensive symbiosis in the real world–advancing ecological studies of heritable, protective bacteria in aphids and beyond. Funct Ecol 28(2):341–355. doi:10.1111/1365-2435.12133
Pantastico-Caldas M, Duncan KE, Istock CA, Bell JA (1992) Population-dynamics of bacteriophage and Bacillus subtilis in soil. Ecology 73(5):1888–1902. doi:10.2307/1940040
Peccoud J, Ollivier A, Plantegenest M, Simon JC (2009) A continuum of genetic divergence from sympatric host races to species in the pea aphid complex. Proc Natl Acad Sci USA 106(18):7495–7500. doi:10.1073/pnas.0811117106
Plague GR, Dale C, Moran NA (2003) Low and homogeneous copy number of plasmid-borne symbiont genes affecting host nutrition in Buchnera aphidicola of the aphid Uroleucon ambrosiae. Mol Ecol 12(4):1095–1100. doi:10.1046/j.1365-294X.2003.01782.x
Pontes MH, Dale C (2006) Culture and manipulation of insect facultative symbionts. Trends Microbiol 14(9):406–412. doi:10.1016/j.tim.2006.07.004
Ptashne M (2004) A genetic switch: phage lambda revisited, vol 3. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Rabinovitch A, Aviram I, Zaritsky A (2003) Bacterial debris – an ecological mechanism for coexistence of bacteria and their viruses. J Theor Biol 224(3):377–383. doi:10.1016/S0022-5193(03)00174-7
Rahbe Y, Digilio MC, Febvay G, Guillaud J, Fanti P, Pennacchio F (2002) Metabolic and symbiotic interactions in amino acid pools of the pea aphid, Acyrthosiphon pisum, parasitized by the braconid Aphidius ervi. J Insect Physiol 48(5):507–516
Rankin DJ, Rocha EPC, Brown SP (2011) What traits are carried on mobile genetic elements, and why? Heredity 106(1):1–10. doi:10.1038/Hdy.2010.24
Roossinck MJ (2011) The good viruses: viral mutualistic symbioses. Nat Rev Microbiol 9(2):99–108. doi:10.1038/Nrmicro2491
Russell JA, Moran NA (2005) Horizontal transfer of bacterial symbionts: heritability and fitness effects in a novel aphid host. Appl Environ Microbiol 71(12):7987–7994. doi:10.1128/Aem.71.12.7987-7994.2005
Russell JA, Moran NA (2006) Costs and benefits of symbiont infection in aphids: variation among symbionts and across temperatures. Proc R Soc B Biol Sci 273(1586):603–610. doi:10.1098/rspb.2005.3348
Russell JA, Weldon S, Smith AH, Kim KL, Hu Y, Łukasik P, Doll S, Anastopoulos I, Novin M, Oliver KM (2013) Uncovering symbiont-driven genetic diversity across North American pea aphids. Mol Ecol 22(7):2045–2059. doi:10.1111/mec.12211
Sandstrom JP, Russell JA, White JP, Moran NA (2001) Independent origins and horizontal transfer of bacterial symbionts of aphids. Mol Ecol 10(1):217–228
Saridaki A, Sapountzis P, Harris HL, Batista PD, Biliske JA, Pavlikaki H, Oehler S, Savakis C, Braig HR, Bourtzis K (2011) Wolbachia prophage DNA adenine methyltransferase genes in different Drosophila-Wolbachia associations. PLoS One 6(5), e19708
Sauer C, Stackebrandt E, Gadau J, Holldobler B, Gross R (2000) Systematic relationships and cospeciation of bacterial endosymbionts and their carpenter ant host species: proposal of the new taxon Candidatus Blochmannia gen. nov. Int J Syst Evol Microbiol 50:1877–1886
Scarborough CL, Ferrari J, Godfray HCJ (2005) Aphid protected from pathogen by endosymbiont. Science 310(5755):1781. doi:10.1126/science.1120180
Schmid M, Sieber R, Zimmermann YS, Vorburger C (2012) Development, specificity and sublethal effects of symbiont-conferred resistance to parasitoids in aphids. Funct Ecol 26(1):207–215. doi:10.1111/j.1365-2435.2011.01904.x
Stern A, Mick E, Tirosh I, Sagy O, Sorek R (2012) CRISPR targeting reveals a reservoir of common phages associated with the human gut microbiome. Genome Res 22(10):1985–1994. doi:10.1101/gr.138297.112
Stouthamer R, Breeuwer JAJ, Hurst GDD (1999) Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu Rev Microbiol 53:71–102. doi:10.1146/annurev.micro.53.1.71
Strand MR (2010) Polydnaviruses. In: Asagari S, Johnson K (eds) Insect virology. Caister Scientific Press, Norfolk, pp 171–197
Strive T, Wright J, Kovaliski J, Botti G, Capucci L (2010) The non-pathogenic Australian lagovirus RCV-A1 causes a prolonged infection and elicits partial cross-protection to rabbit haemorrhagic disease virus. Virology 398(1):125–134. doi:10.1016/j.virol.2009.11.045
Susskind MM (1980) A new gene of bacteriophage-P22 which regulates synthesis of anti-repressor. J Mol Biol 138(4):685–713. doi:10.1016/0022-2836(80)90060-1
Susskind MM, Botstein D, Wright A (1974) Superinfection exclusion by P22 prophage in lysogens of Salmonella typhimurium. 3. Failure of superinfecting phage DNA to enter siea + lysogens. Virology 62(2):350–366. doi:10.1016/0042-6822(74)90398-5
Tamas I, Klasson L, Canback B, Naslund AK, Eriksson AS, Wernegreen JJ, Sandstrom JP, Moran NA, Andersson SGE (2002) 50 million years of genomic stasis in endosymbiotic bacteria. Science 296(5577):2376–2379
Tanaka K, Furukawa S, Nikoh N, Sasaki T, Fukatsu T (2009) Complete WO phage sequences reveal their dynamic evolutionary trajectories and putative functional elements required for integration into the Wolbachia genome. Appl Environ Microbiol 75(17):5676–5686. doi:10.1128/Aem.01172-09
Thao ML, Moran NA, Abbot P, Brennan EB, Burckhardt DH, Baumann P (2000) Cospeciation of psyllids and their primary prokaryotic endosymbionts. Appl Environ Microbiol 66(7):2898–2905. doi:10.1128/Aem.66.7.2898-2905.2000
Tillmann HL, Heiken H, Knapik-Botor A, Heringlake S, Ockenga J, Wilber JC, Goergen B, Detmer J, McMorrow M, Stoll M, Schmidt RE, Manns MP (2001) Infection with GB virus C and reduced mortality among HIV-infected patients. N Engl J Med 345(10):715–724. doi:10.1056/NEJMoa010398
Tsuchida T, Koga R, Fukatsu T (2004) Host plant specialization governed by facultative symbiont. Science 303(5666):1989–1989
van der Wilk F, Dullemans AM, Verbeek M, van den Heuvel JFJM (1999) Isolation and characterization of APSE-1, a bacteriophage infecting the secondary endosymbiont of Acyrthosiphon pisum. Virology 262(1):104–113
van Ham RCHJ, Kamerbeek J, Palacios C, Rausell C, Abascal F, Bastolla U, Fernandez JM, Jimenez L, Postigo M, Silva FJ, Tamames J, Viguera E, Latorre A, Valencia A, Moran F, Moya A (2003) Reductive genome evolution in Buchnera aphidicola. Proc Natl Acad Sci USA 100(2):581–586. doi:10.1073/pnas.0235981100
Vautrin E, Vavre F (2009) Interactions between vertically transmitted symbionts: cooperation or conflict? Trends Microbiol 17(3):95–99. doi:10.1016/j.tim.2008.12.002
Vautrin E, Genieys S, Charles S, Vavre F (2008) Do vertically transmitted symbionts co-existing in a single host compete or cooperate? A modelling approach. J Evol Biol 21(1):145–161. doi:10.1111/j.1420-9101.2007.01460.x
Via S, Bouck AC, Skillman S (2000) Reproductive isolation between divergent races of pea aphids on two hosts. II. Selection against migrants and hybrids in the parental environments. Evolution 54(5):1626–1637
Vinson SB, Mourad AK, Sebesta DK (1994) Sources of possible host regulatory factors in Cardiochiles nigriceps (Hymenoptera, Braconidae). Arch Insect Biochem 26(2–3):197–209
Wagner SM, Martinez AJ, Ruan YM, Kim KL, Lenhart PA, Dehnel AC, Oliver KM, White JA (2015) Facultative endosymbionts mediate dietary breadth in a polyphagous herbivore. Funct Ecol. doi:10.1111/1365-2435.12459
Weinbauer MG (2004) Ecology of prokaryotic viruses. FEMS Microbiol Rev 28(2):127–181. doi:10.1016/j.femsre.2003.08.001
Weldon S (2015) Matryoshka mutualisms: developing the bacteriophage APSE-Hamiltonella defensa-Acyrthosiphon pisum system as a model for tripartite symbioses. University of Georgia Dissertation, Athens, GA
Weldon SR, Strand MR, Oliver KM (2013) Phage loss and the breakdown of a defensive symbiosis in aphids. Proc R Soc B Biol Sci 280(1751):20122103. doi:10.1098/rspb.2012.2103
Wernegreen JJ (2002) Genome evolution in bacterial endosymbionts of insects. Nat Rev Genet 3(11):850–861. doi:10.1038/nrg931
Werren JH, Baldo L, Clark ME (2008) Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol 6(10):741–751. doi:10.1038/nrmicro1969
Wilkes TE, Darby AC, Choi JH, Colbourne JK, Werren JH, Hurst GDD (2010) The draft genome sequence of Arsenophonus nasoniae, son-killer bacterium of Nasonia vitripennis, reveals genes associated with virulence and symbiosis. Insect Mol Biol 19(1):59–73
Williams HTP (2013) Phage-induced diversification improves host evolvability. BMC Evol Biol 13(1):17
Wu M, Sun LV, Vamathevan J, Riegler M, Deboy R, Brownlie JC, McGraw EA, Martin W, Esser C, Ahmadinejad N, Wiegand C, Madupu R, Beanan MJ, Brinkac LM, Daugherty SC, Durkin AS, Kolonay JF, Nelson WC, Mohamoud Y, Lee P, Berry K, Young MB, Utterback T, Weidman J, Nierman WC, Paulsen IT, Nelson KE, Tettelin H, O’Neill SL, Eisen JA (2004) Phylogenomics of the reproductive parasite Wolbachia pipientis wMel: a streamlined genome overrun by mobile genetic elements. PLoS Biol 2(3):327–341. doi:10.1371/journal.pbio.0020069
Xie JL, Vilchez I, Mateos M (2010) Spiroplasma bacteria enhance survival of Drosophila hydei attacked by the parasitic wasp Leptopilina heterotoma. PLoS One 5(8):e12149. doi:10.1371/journal.pone.0012149
Zug R, Hammerstein P (2012) Still a host of hosts for Wolbachia: analysis of recent data suggests that 40% of terrestrial arthropod species are infected. Plos One 7(6). Doi:10.1371/journal.pone.0038544
Acknowledgements
The authors thank Adam Martinez for commenting on an earlier version of this manuscript. Funding support was provided by the National Science Foundation award 1256794.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Weldon, S.R., Oliver, K.M. (2016). Diverse Bacteriophage Roles in an Aphid-Bacterial Defensive Mutualism. In: Hurst, C. (eds) The Mechanistic Benefits of Microbial Symbionts. Advances in Environmental Microbiology, vol 2. Springer, Cham. https://doi.org/10.1007/978-3-319-28068-4_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-28068-4_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-28066-0
Online ISBN: 978-3-319-28068-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)