Abstract
The cyclic di-GMP (c-di-GMP) second messenger represents a signaling system that regulates many bacterial behaviors and is of key importance for driving the lifestyle switch between motile loner cells and biofilm formers. This review provides an up-to-date summary of c-di-GMP pathways connected to biofilm formation by the opportunistic pathogen P. aeruginosa. Emphasis will be on the timing of c-di-GMP production over the course of biofilm formation, to highlight non-uniform and hierarchical increases in c-di-GMP levels, as well as biofilm growth conditions that do not conform with our current model of c-di-GMP.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Pseudomonas aeruginosa is a Gram-negative bacterium that is found in a diverse array of environments such as soil and water, and are found in plant, animal, or human hosts. This ubiquitous bacterium causes chronic, persistent infections in individuals with compromised immune systems, epithelial damage, or cystic fibrosis (CF), is the second leading cause of hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes, and is routinely the most common cause of fatalities in patients with nosocomial infections (CDC 2019, 2020; Costerton et al. 1999). Matrix-enclosed aggregates known as biofilms are associated with many nosocomial and chronic, persistent infections. According to the National Institutes of Health, biofilms are responsible for >80% of microbial infections and >60% of all nosocomial infections, with the Center for Disease Control (CDC) estimating biofilms to be the etiologic agent in 60% of all chronic infections. P. aeruginosa is the second leading cause of these infections.
Bacteria display to modes of growth, planktonically or as biofilms. In the planktonic mode of growth, bacteria appear as single and independent free-floating cells, while in the biofilm mode of growth, bacteria are organized in surface-attached sessile aggregates that are encased in a biofilm matrix composed of exopolysaccharides, extracellular DNA (eDNA) and proteins during biofilm formation (Flemming 2016; Flemming et al. 2007). The mode of growth is consequential to bacterial behavior. The biofilm mode of growth is beneficial for bacteria in that it allows cells to maintain close proximity to nutrients, promotes exchange of genetic material, and confers cells protection from a variety of chemical and environmental stresses (e.g., nutrient limitation, desiccation, and shear forces), as well as engulfment by protozoa in the environment or by phagocytes in a host (Davey and O’Toole 2000). Additionally, biofilm bacteria are protected from killing by antibiotics, with bacteria living in biofilms being up to 1000 times less susceptible to antimicrobial compounds when compared to their planktonic counterparts (Davies 2003). The reduced antibiotic susceptibility of biofilm bacteria contributes to the persistence of biofilm infections.
Biofilm formation by P. aeruginosa has been described as a sequential, highly regulated process, involving at least five phenotypically distinct stages, with each biofilm developmental stage corresponding to unique patterns of protein production and gene expression (Stoodley et al. 2002; Petrova and Sauer 2009; Petrova et al. 2017; O’Toole and Kolter 1998; Davey and O’Toole 2000; Sauer et al. 2002). The stages are referred to as reversible and irreversible attachment, biofilm maturation I and II involving cluster and microcolony formation, respectively, and dispersion (Stoodley et al. 2002; Sauer et al. 2002). Key features relating to the sessile mode of growth include loss of flagella gene expression, production of biofilm matrix components, induction of antibiotic tolerance mechanisms (including efflux-pumps even when biofilms were grown in the absence of antibiotics) and increased levels of virulence determinants (Heacock-Kang et al. 2017; Williamson et al. 2012; Liao et al. 2013). Dispersion is generally characterized as the terminal stage of biofilm development, as cells escape from the biofilm structure to return to the single cell, planktonic mode of growth (reviewed in Rumbaugh and Sauer 2020). Dispersion is assumed to lead to the translocation of bacteria to new sites for colonization. The phenotypic changes noted upon transition to the sessile mode of growth as well as the return to the planktonic mode of growth are driven by multiple signaling systems such as HptB, Wsp, and Pil-Chp systems capable of modulating the intracellular level of the second messenger molecule, cyclic di-GMP (c-di-GMP).
c-di-GMP has emerged as a key regulator of biofilm formation (Valentini and Filloux 2016; Römling et al. 2013; Jenal et al. 2017). C-di-GMP is a ubiquitous bacterial second messenger capable of regulating a myriad of cellular functions in bacteria, including group behavior such as surface-associated motility, surface adaptation, biofilm formation, and cell cycle progression, as well as virulence (Valentini and Filloux 2016; Römling et al. 2013; Jenal et al. 2017). The level of c-di-GMP in the cell is contingent on the activity of two groups of enzymes: diguanylate cyclase (DGC) and phosphodiesterase (PDE) (Simm et al. 2004; Ryjenkov et al. 2005; Chan et al. 2004). DGCs, with a canonical GG(D/E)EF motif, synthesize c-di-GMP from two molecules of GTP, while PDEs, with a canonical E(A/E/V)L motif, degrade c-di-GMP to pGpG (Hengge 2009; Römling and Amikam 2006) which is hydrolyzed into two GMP molecules by the oligoribonuclease Orn (Cohen et al. 2015; Orr et al. 2015). More recently, proteins with an HD-GYP domain were also shown to demonstrate c-di-GMP phosphodiesterase activity (Dow et al. 2006). It is now widely recognized that high c-di-GMP levels favor the biofilm mode of growth, while low levels favor the motile, planktonic mode of growth (Römling et al. 2013). Estimates suggest that P. aeruginosa planktonic cells harbor less than 30 pmol/mg c-di-GMP, whereas biofilms harbored close to 100 pmol/mg c-di-GMP (Gupta et al. 2014).
In many bacterial species, including P. aeruginosa, elevated c-di-GMP results in repression of flagellar-driven motility, by repressing the transcription of flagellar genes or acting post-transcriptionally to regulate flagellar reversals and/or speed, either by interacting with specific flagellar motor proteins or by altering the chemotactic signal transduction system (Baraquet and Harwood 2013; Boehm et al. 2010; Orr and Lee 2016). In contrast, elevated c-di-GMP promotes the expression of genes involved in producing a biofilm matrix (Römling et al. 2013). In P. aeruginosa, the biofilm matrix is composed of polysaccharides (Pel, Psl, alginate), extracellular (e)DNA, and matrix stabilizing proteins such as CdrA and LecB (Flemming 2016; Flemming et al. 2007; Flemming and Wingender 2010; Ryder et al. 2007; Rybtke et al. 2015; Borlee et al. 2010; Reichhardt et al. 2018; Ma et al. 2009). The cdrA, pel, and psl genes are all transcriptionally induced under conditions of high c-di-GMP (Starkey et al. 2009). While P. aeruginosa PAO2 produces both Pel and Psl polysaccharides, P. aeruginosa PA14 is only capable of producing the Pel polysaccharide (Friedman and Kolter 2004a, b).
The current thinking in the field is that once cells attach to a surface, they uniformly respond by producing c-di-GMP. This is supported by the finding that flooding the cell with c-di-GMP induces phenotypic changes associated with high c-di-GMP levels (e.g., aggregative behavior, matrix production, cessation of motility), and the transition to surface-associated lifestyle, while draining the cell induces transition to the motile mode of growth (Römling et al. 2013). However, given that biofilm formation progresses linearly through a number of distinct steps and coinciding with the formation of subpopulations in response to chemical gradients (reviewed in Serra and Hengge 2014), it is more likely that c-di-GMP levels are not produced in a uniform manner but increase in a hierarchical or stepwise manner. It is thus likely that our current model of c-di-GMP-dependent biofilm formation underestimates the complexity of the c-di-GMP signaling network. This review will focus on events linked to c-di-GMP production at different stages of biofilm development by the model organism P. aeruginosa. Emphasis will be on the timing of c-di-GMP production to highlight non-uniform and hierarchical increases in c-di-GMP levels, as well as biofilm growth conditions that do not conform with our current model of the role of c-di-GMP in biofilm development.
2 The First Contact with the Surface: Timing and Heterogeneity of Contact-Dependent Modulation of c-di-GMP Levels
The transition from a planktonic to the biofilm mode of growth is key to the survival of microbes in nature, and it is initiated by bacteria first sensing a surface. Surface sensing is contact dependent and involves transmitting the surface signal intracellularly, followed by responding to that signal to activate downstream pathways, ultimately leading to attachment and subsequent biofilm formation. A key regulator of this switch is the second messenger c-di-GMP (Römling et al. 2013; Jenal et al. 2017). In P. aeruginosa, at least two surface-sensing systems that produce c-di-GMP in response to surface contact have been reported. These are the Wsp and Pil-Chp systems (Fig. 3.1).
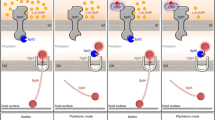
Surface behavior regulation in P. aeruginosa. (a) The Wsp chemosensory pathway. The Wsp system are composed of a methyl-accepting protein (WspA), a CheR-like methyltransferase (WspC), a CheB-like methylesterase (WspF), two CheW homologs (WspB and WspD), a CheA homolog (WspE), and a diguanylate cyclase (WspR). The association of surfaces leads to the activation of WspA, which promotes the autophosphorylation of WspE. WspE phosphorylates its two response regulators, WspR and WspF. Phosphorylated WspR catalyzes the synthesis of c-di-GMP via its GGDEF domain, while phosphorylated WspF resets the Wsp system by removing methyl groups from WspA. (b) The Chp/FimS/AlgR network. A surface-associated signal sensed by PilJ activates the Pil-Chp complex and induces the production of cAMP via activation of CyaB. The cAMP then binds to and activates the transcription factor Vfr, which stimulates pilY1 gene expression along with the response regulator AlgR. Upon complete assembly of TFP apparatus, PilY1 is secreted. The external PilY1 acts as a signal to induce SadC activity. The diguanylate cyclase SadC and the phosphodiesterase BifA inversely regulate c-di-GMP levels. P, protein phosphorylation or phosphotransfer reaction; OM, outer membrane; PG, peptidoglycan; IM, inner membrane
The Wsp system senses an unknown surface-related signal (recently proposed to be membrane perturbation (Chen et al. 2014)) through WspA, a membrane-bound protein homologous to methyl-accepting chemotaxis proteins (MCPs). Activation of this system stimulates phosphorylation of the diguanylate cyclase WspR, with phosphorylation of WspR increasing its activity, resulting in increased c-di-GMP levels that, in turn, induces the production of the biofilm matrix polysaccharide Pel and Psl (Hickman et al. 2005; Huangyutitham et al. 2013) (Fig. 3.1a). The Wsp system is localized laterally along the cell (O’Connor et al. 2012), suggesting that P. aeruginosa deploys a laterally localized systems to promote c-di-GMP synthesis in response to a surface.
The second surface-sensing mechanism involves the Pil-Chp system. This surface-sensing mechanism is more complex and involves a cascade of intracellular signaling molecules. The Pil-Chp system, comprised by the PilGHIJK-ChpABC complex functions as a chemotaxis-like system (Whitchurch et al. 2004; Darzins 1994; Sampedro et al. 2014) to detect surface contact on short timescales, using the mechanical activity of its type IV pili, a major surface adhesin (Persat et al. 2015) (Fig. 3.1b). Surface contact sensing requires attachment of type IV pili to a solid surface, followed by pilus retraction, which exerts tension on type IV pili. This modulates the interaction between the periplasmic domain of the methyl-accepting chemotaxis protein-like chemosensor PilJ with the major pilin subunit PilA (Persat et al. 2015), resulting in a signal transduction event to the cytoplasm, as follows: PilJ activates the CheA homolog ChpA complexed with the CheW homolog PilI, ultimately stimulating the adenylate cyclase CyaB to produce cyclic AMP (cAMP) (Persat et al. 2015; Luo et al. 2015) (Fig. 3.1b). cAMP then binds to the receptor protein Vfr, which in turn, together with the FimS-AlgR two-component system (TCS), upregulates not only the expression of numerous virulence factors, but also the fimU-pilVWXY1Y2E operon (Persat et al. 2015; Luo et al. 2015). Amongst the components being thus produced is the cell surface-associated protein PilY1 (Fig. 3.1b). Upon complete assembly of the type IV pili alignment complex (PilMNOP) at the cell surface, PilY1 is secreted. As the secretion of PilY1 is dependent on the type IV pili assembly system, PilY1 is not deployed until the pilus is assembled (Luo et al. 2015). Cell surface-associated PilY1, in turn, provides the signal to activate c-di-GMP production generated by the diguanylate cyclase SadC (Fig. 3.1b). Recent findings indicate that SadC’s diguanylate cyclase activity is dependent on the type IV pili alignment complex PilMNOP and more specifically, PilO (Webster et al. 2021). The findings suggested that the Pil-Chp and type IV pili alignment complexes work in concert to regulate cAMP and c-di-GMP production, thereby employing a hierarchical regulatory cascade of second messengers, cAMP and c-di-GMP, to coordinate its program of surface behaviors.
But why two surface contact systems? Initial attachment by P. aeruginosa has been reported to occur in two phases, reversible and irreversible attachment. The reversible attachment phase is characterized by cells first attaching to a surface by a single pole. Such cells either return to the bulk phase or commit to a more stable surface interaction by attaching via their longitudinal axis, referred to as irreversible attachment (Fig. 3.2). Once cells are irreversibly attached, P. aeruginosa will stop moving and ultimately multiply and form aggregates. The two points of attachment by P. aeruginosa are linked to the location of the contact sensory system. In P. aeruginosa, PilY1 is co-localized with type IV pili at the cell pole (Luo et al. 2015; Kuchma et al. 2010), while the Wsp system is localized laterally along the cell (O’Connor et al. 2012). Considering that P. aeruginosa first makes surface contact via its pole and then attaches longitudinal, the finding suggests that the Pil-Chp contact sensory system is the first of the two surface-sensing systems to be activated (Fig. 3.2). Recent findings also suggest that the Pil-Chp system likely contributes to a gradual buildup of c-di-GMP in surface-associated cells that are reversibly attached, likely through successive surface interactions and detachments (Armbruster and Parsek 2018; Lee et al. 2018). Repeated surface exposure thus enables cells to become surface adapted, with surface adaptation coinciding with a progressive increase in cellular cAMP levels and likely subsequent increase in c-di-GMP (Armbruster and Parsek 2018; Lee et al. 2018) (Fig. 3.2). The finding of progressive surface adaptation prior to cells becoming irreversibly attached may furthermore explain the time needed for cells to transition from the reversible to the irreversible attachment stage. Based on proteomic approaches and reporter gene studies, the transition to the irreversible attachment stage has been reported to occur within 6 h post initial attachment by P. aeruginosa when grown under flowing conditions (Chambers et al. 2014; Petrova and Sauer 2011). The timing is in agreement with findings by Lee et al. (2018) of the cAMP signal increase and its impact on type IV pili being offset by about 5 h. However, evidence suggests that attached cells may not uniformly elevate c-di-GMP levels upon attachment. Using a plasmid-based c-di-GMP reporter (pPcdrA::gfp) to monitor c-di-GMP levels in a clonal population of P. aeruginosa as they began to form biofilms, Armbruster et al. (2019) demonstrated heterogeneity in c-di-GMP levels by the attached population. In contrast to the general belief of attached cells uniformly producing c-di-GMP, the study revealed two physiologically distinct subpopulations of cells were noted, cells with high and low c-di-GMP levels. The two subpopulations were found to perform complementary and important tasks in the early stages of biofilm formation. The subpopulation with elevated c-di-GMP represented “biofilm founders” and produced biofilm matrix, whereas the low c-di-GMP cells represented “surface explorers” that engaged in surface motility, allowing for exploration of the surface (Armbruster et al. 2019). The heterogeneity was generated by the Wsp system (Armbruster et al. 2019).
Model of c-di-GMP modulating systems contributing to surface contact sensing and transition from the reversible to the irreversible attachment stage in P. aeruginosa. (a) Surface contact sensing requires the Pil-Chp and Wsp system, with activation leading to a hierarchical regulatory cascade of second messengers, cAMP and c-di-GMP, to coordinate the initial surface behaviors. cAMP levels are increased prior to c-di-GMP. (b) Psl/c-di-GMP feedback loop. Psl polysaccharide deposited by P. aeruginosa migrating across a surface acts as a signal to stimulate the two diguanylate cyclases, SiaD and SadC, to produce c-di-GMP. (c) Transition to the irreversible attachment stage requires c-di-GMP modulating systems SadC/BifA, HptB/Gac, FleQ, Lap, and SagS/NicD/PA3177. C-di-GMP levels likely increase in a hierarchical or stepwise manner upon first contact with the surface to ensure P. aeruginosa remains engaged with the surface. This increase may be linked to the sequential activation of c-di-GMP modulating systems, although little is known about the timing of activation relative to each other. As a consequence of increased c-di-GMP production, flagellar-driven motility ceases while matrix production is increased
3 Consequences of Surface Contact and Downstream Factors
Following the first contact with a surface and in response to elevated c-di-GMP, the phenotype of surface-associated P. aeruginosa cells switches. Several c-di-GMP effectors are responsible for the regulatory function of c-di-GMP through transcriptional, posttranscriptional, translational, and protein–protein interaction mechanisms. So far, five types of c-di-GMP receptors have been found: (1) effector proteins, such as PilZ domain proteins that bind to c-di-GMP to regulate other proteins or enzymes via domain–domain or protein–protein interactions; (2) degenerate GGDEF proteins that are no longer catalytically proficient; (3) proteins with enzymatic activity that demonstrated enhanced activity upon c-di-GMP binding; (4) c-di-GMP-responsive transcription factors or repressors, and (5) c-di-GMP riboswitches. c-di-GMP riboswitches are structured RNAs located in the 5′-untranslated regions (5′-UTRs) of mRNAs that regulate expression of downstream genes in response to changing concentrations of the second messenger c-di-GMP. While riboswitches have been identified in pathogens, such as Clostridium difficile, Vibrio cholerae, and Bacillus anthracis, P. aeruginosa lacks c-di-GMP riboswitches. The remaining c-di-GMP receptors, however, are present in P. aeruginosa. An overview of these receptors and their contribution to the phenotypic switch by P. aeruginosa upon transition to the surface-associated mode of growth is given below.
High c-di-GMP levels negatively affect swarming motility. In P. aeruginosa, the rotation of the polar flagellum is controlled by two distinct stator complexes, MotAB, which cannot support swarming motility, and MotCD, which promotes swarming motility (Toutain et al. 2007). At elevated c-di-GMP levels, swarming motility is repressed by the PilZ domain-containing protein FlgZ (PA3353), with the c-di-GMP-bound FlgZ impeding motility via its increased interaction with the MotCD stator at the cell pole (Baker et al. 2016).
Additional regulatory systems contribute to the inverse regulation of motility and matrix production. These include the HptB pathway, AmrZ(Jones et al. 2014) and FleQ. The HptB pathway regulates behaviors such as chemotaxis, twitching, swimming, swarming motility and biofilm formation by partially interfering with flagellar gene expression and by partially intersecting with the Gac/Rsm cascade (Hsu et al. 2008; Bordi et al. 2010; Bhuwan et al. 2012; Valentini et al. 2016) (Fig. 3.3). The pathway is composed of the histidine phosphotransfer protein HptB (PA3345), the response regulator HsbR (PA3346), the anti-anti sigma factor HsbA, and the diguanylate cyclase HsbD. Briefly, when HptB is phosphorylated, it activates the phosphatase domain of HsbR, which in turn dephosphorylates HsbA. Dephosphorylated form of HsbA sequesters the anti-sigma factor FlgM from the FliA sigma factor (σ28), with free FliA activating flagellar gene expression and hence swimming and swarming motility (Hsu et al. 2008; Bhuwan et al. 2012) (Fig. 3.3). In contrast, when HptB is inactive, HsbR is dephosphorylated and its Ser/Thr kinase domain is promoted. Under these conditions, phosphorylated HsbA interacts via the kinase activity of HsbR with the diguanylate cyclase HsbD (Valentini et al. 2016). The switch in HsbA interaction partners from FlgM to HsbD coincides with enhanced HsbD diguanylate cylcase activity, and thus, increased c-di-GMP levels (Fig. 3.3). Additionally, increased levels of the small regulatory RNA (sRNA) RsmY have been reported. The HptB pathway thus contributes to the repression of swimming motility while enhancing twitching motility and biofilm formation.
Modulation of the c-di-GMP pool by HptB and intersecting regulatory systems. Expression of RsmY/Z through the HptB system is GacA-dependent. Under planktonic conditions, SagS regulates small regulatory RNA (sRNA) levels in an HptB-dependent manner. Under biofilm growth conditions, SagS contributes via BfiSR to the suppression of the sRNA RsmZ level. Arrows and blunt-ended lines indicate stimulatory and inhibitory interactions, respectively. P, protein phosphorylation or phosphotransfer reaction. Proteins harboring diguanylate cyclase activity are shown in green
An additional c-di-GMP receptor affecting motility is FleQ. More specifically, FleQ is a c-di-GMP-responsive transcription factor that inversely regulates flagellar-driven motility and matrix production. At low c-di-GMP levels, FleQ positively regulates flagellar-driven motility the flhA gene to start the cascade of flagellar gene expression (Dasgupta et al. 2003). At high c-di-GMP levels, FleQ upregulates genes encoding the biofilm matrix components, Pel and Psl, and the adhesion CdrA. The activity of FleQ is modulated by at least two factors: the putative ATP/GTP binding protein, FleN, and c-di-GMP (Baraquet and Harwood 2013; Dasgupta and Ramphal 2001; Hickman and Harwood 2008; Baraquet et al. 2012). The binding constant (Kd) for c-di-GMP binding by FleQ has been reported to be 15–20 μM (Hickman and Harwood 2008; Baraquet et al. 2012). FleN inhibits FleQ ATPase activity by directly interaction. Moreover, binding by c-di-GMP to FleQ induces a conformational change of FleQ, which is furthermore linked to the inhibition of the FleQ ATPase activity, apparent by FleQ ATPase activity being more inhibited by c-di-GMP in the presence of FleN than in its absence. Therefore, FleN and c-di-GMP cooperate to inhibit FleQ activity, resulting in the repression of flagellar genes but the relieve of transcriptional repression of cdrA, pel, and psl genes necessary for biofilm formation (Borlee et al. 2010; Baraquet et al. 2012). The latter are components of the P. aeruginosa biofilm matrix (P. aeruginosa PA14 does not produce the Psl polysaccharide (Friedman and Kolter 2004a, b)). Given the high level of c-di-GMP required for FleQ to repress flagellar gene expression (or enable expression of genes linked to the biofilm matrix), it is likely that FleQ activity represents a second level of control, post-regulation by c-di-GMP binding proteins capable of domain–domain or protein–protein interactions such as FlgZ. In addition to FleQ, the Wsp system contribute to Pel and Psl polysaccharide production (Hickman et al. 2005; Huangyutitham et al. 2013). The Pel and Psl polysaccharide promote irreversible attachment, apparent by P. aeruginosa PA14 Δpel and P. aeruginosa PAO1 Δpsl mutants being both arrested at the monolayer stage of biofilm development, with concurrent reduction in accumulated biofilm biomass compared to their respective parental strains (Colvin et al. 2012). Moreover, the Psl polysaccharide itself enhances attachment. P. aeruginosa PAO1 migrating across a surface has been demonstrated to leave a trail of the Psl polysaccharide (Zhao et al. 2013) which acts as a signal to stimulate two diguanylate cyclases, SiaD and SadC, that in turn produce the intracellular secondary messenger molecule c-di-GMP (Irie et al. 2012) (Fig. 3.2). Elevated intracellular concentrations of c-di-GMP then lead to the increased production of Psl and other components required for attachment and biofilm formation. The positive feedback regulatory circuit of an extracellular polysaccharide promotes biofilm growth has been reported to be analogous to autocrine signaling in eukaryotes (Irie et al. 2012).
CdrA is an adhesin that reinforces the biofilm matrix by binding to and cross-linking Psl and Pel (Rybtke et al. 2015; Borlee et al. 2010; Reichhardt et al. 2020). At high c-di-GMP levels, CdrA is located at the outer membrane in a cell-associated form. However, at low intracellular c-di-GMP levels, the periplasmic cysteine protease LapG targets the membrane-anchor of CdrA, resulting in the release of CdrA from the outer membrane (Chatterjee et al. 2014). The c-di-GMP sensing component of the Lap system is LapD, a c-di-GMP receptor protein located in the inner membrane that senses c-di-GMP via its degenerate EAL domain (Rybtke et al. 2015; Cooley et al. 2016). Under condition of high c-di-GMP, c-di-GMP interacts with the EAL domain of LapD, which in turn sequesters LapG. When c-di-GMP levels are low, the protease LapG is released into periplasm, where it cleaves CdrA at a C-terminal TAAG site, resulting in CdrA being secreted. Secretion of CdrA promotes dispersion and the planktonic mode of growth, while tethered CdrA promotes biofilm stability and aggregative behavior (Reichhardt et al. 2020; Cherny and Sauer 2020). A homologous system regulating the transition from reversible to irreversible attachment is present in P. fluorescens and P. putida in which CdrA is substituted by LapA (Hinsa et al. 2003; Monds et al. 2007; Newell et al. 2009; Gjermansen et al. 2010; Newell et al. 2011).
Other regulatory systems that are activated during the attachment phase and either directly or indirectly contribute to the phenotypic switch, include the SadBC/BifA system (Figs. 3.1b and 3.2), GcbA, and SagS (Figs. 3.2 and 3.3). The SadBC/BifA system is composed of the DGCs SadB and SadC and PDE BifA, and contributes to the transition from the reversible to the irreversible attachment by regulating several surface-associated behaviors, including swarming, polysaccharide production, and modulation of flagellar reversal rates via the chemotaxis cluster IV in a c-di-GMP-dependent manner (Kuchma et al. 2007; Merritt et al. 2007). DGCs SadB and SadC and PDE BifA inversely modulate c-di-GMP levels (Kuchma et al. 2007; Merritt et al. 2007). The DGC GcbA likewise affects flagellum-driven motility by suppressing flagellar reversal rates in a manner independent of viscosity, surface hardness, and polysaccharide production (Petrova et al. 2014). SagS is a membrane-bound sensor that under planktonic conditions functions within the HptB pathway to inversely regulate motility and c-di-GMP levels produced by the DGC HsbD (Hsu et al. 2008; Bhuwan et al. 2012; Valentini et al. 2016) (Fig. 3.3). Upon P. aeruginosa transition to the surface, however, SagS has been reported to interact with and phosphorylate the two-component system BfiSR (Petrova and Sauer 2011; Petrova and Sauer 2010). The switch in interactions partners coincides with differential phosphorylation of SagS (Petrova and Sauer 2009; Petrova and Sauer 2011). Moreover, in the sessile mode, SagS (in)directly interacting with two DGCs, NicD and PA3177 (Park et al. 2021; Poudyal and Sauer 2018a) (Fig. 3.3). Inactivation of sagS correlated with biofilms being arrested at the irreversible attachment state, with biofilms formed by ΔsagS exhibited significantly reduced c-di-GMP levels relative to wild-type biofilms (Petrova et al. 2017; Gupta et al. 2014; Park et al. 2021). While wild-type P. aeruginosa biofilms harbored approximately 75–78 pmol/mg c-di-GMP, ΔsagS biofilms only displayed 33 ± 2 pmol/mg c-di-GMP. Nevertheless, ΔsagS biofilms harbor 2–3 times higher c-di-GMP levels compared to planktonic cells, suggesting the notion of stepwise increase of c-di-GMP.
Several c-di-GMP modulating systems are activated upon attachment by P. aeruginosa to the surface (Figs. 3.2 and 3.4). While little is known about the timing of activation relative to each other, it is furthermore apparent that each system adds to the increasing c-di-GMP pool present in attached cells as they progress through the initial attachment stages of biofilm formation (Fig. 3.2). Given the multitude of systems, it is unclear why so many seemingly redundant systems are required to generate c-di-GMP, considering that inactivation of either system arrests biofilms at an early attachment stage. As findings by Armbruster et al. (2019) indicated that attached cells do not uniformly produce c-di-GMP at the same level, it is likely that the systems are activated in a sequential manner to ensure the progressive increase in c-di-GMP levels. If this is the case, c-di-GMP levels likely increase in a hierarchical manner, starting with planktonic cells contacting the surface, followed by the transition to the irreversible attachment stage, and beyond (Figs. 3.2 and 3.4). Support for a stepwise increase in c-di-GMP levels stems from mutants arrested at the transition to the irreversible attachment stage, demonstrating intermediate c-di-GMP levels relative to planktonic and mature biofilm cells.
Model of c-di-GMP modulating systems contributing to biofilm development, including biofilm maturation and maintenance. The formation of biofilms is a cyclic process that occurs in a stage-specific and progressive manner. The process is initiated following surface contact by single planktonic cells. Several developmental steps are discernable as reversible attachment (step I), irreversible attachment (step II) and biofilm maturation (steps III and IV) (Petrova and Sauer 2009; Sauer et al. 2002). During reversible attachment, bacteria sense surface contact via the Pil-Chp and Wsp system, apparent by bacteria attaching to the substratum via the cell pole or via the flagellum (step I), followed by longitudinal attachment. Additional c-di-GMP modulating enzymes likely active at this stage include diguanylate cyclases GcbA, SiaD, and SadC. Transition to the irreversible attachment stage requires c-di-GMP modulating systems HptB/Gac, SadC/BifA, and SagS/NicD/PA3177 and coincides with a reduction in flagellar reversal rates, reduction in flagella gene expression and the production of biofilm matrix components (step II). This stage is also characterized by attached cells demonstrating drug tolerance, with drug tolerance having been linked to the action of diguanylate cyclase PA3177. Biofilm maturation stages are characterized by the appearance of cell clusters that are several cells thick and are embedded in the biofilm matrix (step III) which subsequently fully mature into microcolonies (step IV). Screens of c-di-GMP modulating enzymes revealed GGDEF domain only enzymes PA0169 (SiaD), PA0290, PA0338, PA3702 (WspR), PA1120 (YfiN), PA1107 (RoeA), PA2870, PA3177, PA4332 (SadC), PA4843 (GcbA), PA5487, HDGYP-domain only enzymes PA2572, PA4781, and EAL-GGDEF domain enzymes PA4367 (BifA), PA4602 (MorA), PA5017 (DipA) to contribute to the maturation of P. aeruginosa biofilms. Maintenance of the mature biofilm architecture, although not recognized as a distinct stage, has been shown to require the phosphodiesterases MorA and RmcA, likely by controlling polysaccharide production. Dispersion (stage V) coincides with the return to the planktonic mode of growth and low c-di-GMP levels, requiring the action of several c-di-GMP modulating enzymes, including GcbA (PA4843), NicD (PA4929), DipA (PA5017), RbdA (PA0861), and NbdA (PA3311). Dispersion has been reported to coincide with the decrease in and degradation of matrix components, with dispersed cells being motile and demonstrating increased drug susceptibility relative to biofilm cells. Biofilm matrix is shown in beige. Variations in the c-di-GMP level over the course of biofilm development are indicated by the thickness of the red arrows
4 Key Players Contributing to Biofilm Maturation
The genome of P. aeruginosa PA14 harbors 40 genes with predicted GGDEF, EAL, and HD-GYP domains either singly or in combination (Kulesekara et al. 2006). These c-di-GMP modulating proteins can be divided into four subclasses: 16 GGDEF-only, 5 EAL-only, 16 dual-domain GGDEF/EAL, and 3 HD-GYP-only proteins. The genome of P. aeruginosa PAO1 encodes a similar number of predicted genes, 41, including 17 GGDEF-only, 5 EAL-only, and 16 dual-domain GGDEF/EAL (Jacobs et al. 2003; Ryan et al. 2009). The two strains differ in that strain PA14 lacks one DGC- (PA2771) and one PDE-encoding gene (PA2818, arr) but harbors one additional gene, pvrR, encoding a PDE domain, relative to strain PAO1. Two studies explored the contribution of c-di-GMP metabolism (biosynthesis or degradation) on biofilm formation (Kulesekara et al. 2006; Ha et al. 2014). The studies overall indicated that inactivation of genes encoding diguanylate cyclases (DGC, harboring GGDEF-only) coincided with reduced biofilm formation, while inactivation of enzymes and factors contributing to the degradation of c-di-GMP enhanced biofilm formation. An overview is given in Table 3.1 and Fig. 3.4. These studies supported the notion that high intracellular c-di-GMP levels correlate with a sessile lifestyle, or a biofilm state, while low levels of this signal promote motility and/or planktonic growth. Interestingly, the studies also demonstrated that not every c-di-GMP-producing enzyme contributes to biofilm formation, as some DGCs had no effect on P. aeruginosa PA14 biofilm formation (Table 3.1). These included DGCs (GGDEF-only) encoded by PA14_20820, PA14_40570, PA14_53310, PA14_57140, and PA14_65090 formed wild-type like biofilms (Ha et al. 2014). The studies differed with respect to the role of genes encoding dual-domain GGDEF/EAL in biofilm formation. While Kulesakara et al. (2006) found insertions in genes encoding dual-domain GGDEF/EAL to negatively affect biofilm formation, Ha et al. (2014) found strains inactivated in the respective genes to be less predictable with respect to biofilm formation, with the effect being dependent on whether the GGDEF or EAL domain was functional or degenerate. For instance, dual-domain GGDEF/EAL genes PA14_21190, PA14_56790 (bifA), and PA14_66320 (dipA) have been reported to encode proteins with PDE activity (Kuchma et al. 2007; Kulesekara et al. 2006; Basu Roy et al. 2012), with inactivation of the respective genes coinciding with enhanced or hyperbiofilm formation (Ha et al. 2014). Several studies furthermore revealed strain differences with respect to dual-domain GGDEF/EAL proteins. While inactivation of PA14_53140 (rbdA, RbdA has PDE activity) in P. aeruginosa PA14 had no effect on biofilm formation (Kulesekara et al. 2006; Ha et al. 2014), inactivation of rbdA in P. aeruginosa PAO1 caused hyperbiofilm formation (An et al. 2010). Similar strain differences have been reported for MucR, a bifunctional enzyme capable of both c-di-GMP biosynthesis and degradation (Li et al. 2013). While P. aeruginosa PAO1 carrying a mucR mutation formed wild-type-like biofilms (Li et al. 2013; Hay et al. 2009), the PA14_42220 (mucR) mutant formed biofilms at 60% of wild-type levels (Ha et al. 2014).
The two screens also revealed differences in the contribution of no role of PDEs (EAL-only, HDGYP-only) and dual-domain GGDEF/EAL demonstrating PDE activity. For instance, while mutants in genes encoding dual-domain GGDEF/EAL proteins capable of degrading c-di-GMP appear to form hyperbiofilms, PDE mutants belonging to the EAL-only subclass showed no significant difference from the wild-type strain (Kulesekara et al. 2006; Ha et al. 2014). This is in contrast to PDE mutants belonging to the HD-GYP-only subclass. Of the three 3 HD-GYP-only proteins, only PA14_30830 and PA14_ 63210 contributed to biofilm formation, apparent by mutants forming significantly reduced biofilms relative to wild type (Ha et al. 2014). It is of interest to note that the respective mutants also demonstrated reduced swarming and twitching motility as well as reduced congo red binding indicative of reduced biofilm matrix production, compared to the wild type (Ha et al. 2014). The finding of c-di-GMP-degrading phosphodiesterases thus affecting biofilm formation is surprising and not in line with the currently accepted model wherein elevated c-di-GMP levels promote biofilm formation. The findings suggest that our current model is too simple or oversimplified while also raising several questions:
-
Does c-di-GMP turnover occur in mature biofilms to prevent the continued increase in c-di-GMP levels, and if so, is the turnover essential for the maintenance of the biofilm structure?
-
Is c-di-GMP production and turnover spatially controlled within the biofilm? And if so, where in the biofilm does the degradation of c-di-GMP-degrading occur?
-
Is this spatial control restricted to typically c-di-GMP-controlled biofilm functions such as matrix production?
-
And what is the role of PDEs in biofilm formation?
5 c-di-GMP Levels and Maintenance of the Mature Biofilm Structure
Although high c-di-GMP levels are a key to enable the formation of biofilms, evidence suggests that c-di-GMP levels in mature biofilms are not uniformly high. Using a plasmid-based c-di-GMP reporter (pPcdrA::gfp) to monitor c-di-GMP levels for which the fluorescence intensity is directly proportional to the concentration of intracellular c-di-GMP (Rybtke et al. 2012), a striking difference in c-di-GMP concentrations was seen across the P. aeruginosa biofilm structure (Nair et al. 2017). Relatively high amounts of c-di-GMP were detected at the outer boundary of large, mature biofilms and hence, lower levels at the center of microcolonies (Nair et al. 2017). In contrast, smaller and less developed biofilms showed a more uniform distribution of c-di-GMP (Nair et al. 2017), likely suggesting that gradients (oxygen, nutrients, waste) contribute to the spatial distribution of c-di-GMP in biofilms (Rumbaugh and Sauer 2020; Serra and Hengge 2014). Moreover, the finding suggested c-di-GMP levels to fluctuate or decrease as the biofilm matures. Similar observations were made for Escherichia coli macrocolony biofilms gown on agar using fluorescent-based detection of the expression of pdeH (formerly yhjH) encoding the master phosphodiesterase PdeH in E. coli (Klauck et al. 2018). PdeH maintains low cellular c-di-GMP level in E. coli and thus, can serve as a reporter for low c-di-GMP zones within the biofilm (Sarenko et al. 2017). The study revealed differences in c-di-GMP levels in vertically cryo-sectioned macrocolony biofilms, with differences coinciding with stratified matrix production along nutrient gradients (Klauck et al. 2018).
It is likely that the spatial distribution of c-di-GMP within the biofilm structure is not only influenced by microenvironmental conditions but also linked to the maintenance of the mature biofilm structure. Maintaining a steady state of mature biofilms may involve localized matrix production/degradation or dispersion events to enable restructuring and/or continued growth of the biofilm. In support of biofilm maintenance coinciding with lower c-di-GMP levels is the finding that cell lysis-dependent production of extracellular DNA (eDNA), which is necessary for the formation of biofilms by many species, including P. aeruginosa, is stimulated by low c-di-GMP levels (Ueda and Wood 2010). Likewise, the architecture of mature biofilms is affected by motile cells on the biofilm surface, with lower c-di-GMP levels contributing to the sessility-to-motility transition of biofilm cells (Römling et al. 2013). Additionally, biofilm dispersion, an integral part of long-term biofilm maintenance, requires the presence of specific PDEs in P. aeruginosa (Rumbaugh and Sauer 2020; Petrova and Sauer 2016). However, lowering c-di-GMP levels does not appear to be solely a function of localized matrix remodeling to maintain mature biofilms in a steady state. Instead, P. aeruginosa biofilms make use of specific PDEs to ensure survival in times under conditions of nutrient limitations. The PDEs required for the maintenance of the biofilm architecture were identified as RmcA (PA0575, PA14_07500) and MorA (PA4601, PA14_60870) (Katharios-Lanwermeyer et al. 2021) (Fig. 3.4). Strains inactivated in rmcA and morA were unable to maintain the biofilm structure when exposed to carbon-limited conditions, resulting in subsequent loss of P. aeruginosa PA14 biofilm viability. Moreover, RmcA and MorA were found to interact with the Pel biosynthesis machinery (Katharios-Lanwermeyer et al. 2021), allowing the PDEs to interfere or modulate matrix production. The biofilm maintenance deficiency phenotype observed for the PDE mutants was also found for the stringent response mutant ΔrelAΔspoT, suggesting that a regulatory intersection between c-di-GMP signaling, extracellular polysaccharide biosynthesis, and the nutrient limitation response is important for biofilm persistence. Taken together, the findings suggested that unregulated Pel biosynthesis contributes to cell death of established biofilms under nutrient-limiting conditions, suggesting that too high c-di-GMP levels can be deleterious to biofilm cells (Katharios-Lanwermeyer et al. 2021). It is of interest to note that the PDEs RmcA and MorA are unrelated to those required for establishing or dispersing a biofilm, suggesting that a wide variety of c-di-GMP metabolizing enzymes in organisms such as P. aeruginosa allows for discrete control over the formation, maintenance, or dispersion of biofilms.
6 Biofilm Dispersion: The Return to Low c-di-GMP Levels
Biofilm dispersion is an integral part of the biofilm developmental life cycle as well as a mechanism to maintain biofilms in a steady-state (Rumbaugh and Sauer 2020; Petrova and Sauer 2016; Kim and Lee 2016; Sauer 2020) (Fig. 3.4). As a biofilm grows in size, some cells will become increasingly separated from the bulk liquid interface and essential sources of energy or nutrients. Accumulation of waste products and toxins in the interior of biofilms poses additional challenges. Being trapped deep within a biofilm can, therefore, threaten cell survival. Thus, biofilm cells have evolved mechanisms which enables escaping the sessile mode of growth as a means of self-preservation, by liberating themselves from matrix-encased biofilms, and reverting back to the planktonic mode of growth. The transition to the planktonic mode of growth is referred to as dispersion (Davies 1999), and is characterized by single cells actively escaping from the biofilm, leaving behind eroded biofilms and microcolonies having central voids (Stoodley et al. 2002; Sauer et al. 2002; Basu Roy et al. 2012; Petrova and Sauer 2016; Davies and Marques 2009; Basu Roy and Sauer 2014; Morgan et al. 2006; Petrova and Sauer 2012a; Petrova et al. 2015; Davies 2011). Dispersion rarely involves the entire biofilm, with no more than 80% of the biofilm biomass being removed upon induction of dispersion (Davies and Marques 2009; Morgan et al. 2006; Sauer et al. 2004; Barraud et al. 2009a, b). Instead, selected microcolonies or areas within a biofilm will undergo a dispersion event at any particular time, in a manner often dependent on microcolony diameter (Purevdorj-Gage et al. 2005). In P. aeruginosa, native dispersion occurs in response to the fatty acid messenger molecule cis-2-decenoic acid (Davies and Marques 2009) as well as in response to a variety of environmental stimuli, including nutrient cues (glucose, succinate, glutamate), nitric oxide (NO), and others (iron, heavy metals, nutrient and oxygen limitation, etc.) (Gjermansen et al. 2010; Morgan et al. 2006; Barraud et al. 2009a).
Contrary to the formation of biofilms, dispersion is generally linked to low c-di-GMP levels. Intracellular c-di-GMP levels decreased up to 50% in biofilms in response to sensing of the dispersion cue nitric oxide (Barraud et al. 2009b), and >50% in response to glutamate (Basu Roy et al. 2012). Considering that dispersal is primarily initiated in the center of biofilm microcolonies and coincides without completely removing the entire biomass (Sauer et al. 2004), it is likely that dispersion contributes to the non-uniform distribution of c-di-GMP throughout the biofilm, especially the low c-di-GMP levels in the center of biofilms (Nair et al. 2017). Moreover, dispersed cells demonstrate significantly increased PDE activity relative to biofilms, and c-di-GMP concentrations that closely resemble those of planktonic cells (Basu Roy et al. 2012).
Dispersion in response to environmental cues, such as glutamate or NO, requires the presence of c-di-GMP-degrading enzymes. These have been identified as DipA, RbdA, NbdA, and MucR (Basu Roy et al. 2012; An et al. 2010; Li et al. 2013) (Fig. 3.5). Phosphodiesterase NbdA contributes to dispersion in a cue-specific manner (Li et al. 2013; Basu Roy and Sauer 2014), with NbdA also perceiving the dispersion cue nitric oxide (Li et al. 2013). In contrast, phosphodiesterases DipA and RbdA are central to the dispersion response regardless of the dispersion cue (Basu Roy et al. 2012; Morgan et al. 2006; Petrova and Sauer 2012a, b; Petrova et al. 2015; Barraud et al. 2009b; Li et al. 2014). In support of PDEs and low c-di-GMP levels being required for dispersion, draining the cell of c-di-GMP—by overexpression of native PDEs, BifA, and PA2133, or the E. coli PDE PdeH (YhjH)—lead to the dispersal of P. aeruginosa biofilms (Andersen et al. 2021a; Christensen et al. 2013). However, PDEs do not appear to be the only c-di-GMP modulating enzymes required for dispersion to occur. Additional factors include two diguanylate cyclases, GcbA and NicD, and the chemotaxis-like MCP homolog BdlA (Basu Roy and Sauer 2014; Morgan et al. 2006; Petrova et al. 2015; Petrova and Sauer 2012a, b) (Fig. 3.5). BdlA is central to the dispersion response. Sensing of dispersion cues results in the activation of BdlA via c-di-GMP-dependent phosphorylation, apparent by a sudden rise or burst in c-di-GMP (Basu Roy and Sauer 2014), followed by non-processive proteolysis of BdlA (Petrova and Sauer 2012a, b). The source of this burst in c-di-GMP has been reported to be generated by the DGCs GcbA (PA4843) and NicD (PA4929) (Fig. 3.5) (Basu Roy and Sauer 2014; Petrova and Sauer 2012b). The two DGCs have been linked to the c-di-GMP-dependent phosphorylation and activation of BdlA (Basu Roy and Sauer 2014; Petrova and Sauer 2012b). Once activated, BdlA then recruits PDE RbdA and activates PDE DipA, ultimately resulting in an overall reduction of c-di-GMP levels and subsequent dispersion events (Li et al. 2013; Basu Roy and Sauer 2014; Cutruzzolà and Frankenberg-Dinkel 2016) (Fig. 3.5).
Dispersion cue perception and relay resulting in the modulation of c-di-GMP levels. The membrane-associated MucR (EAL-GGDEF domain), NbdA (EAL domain only) and NicD (GGDEF domain only) are involved in perceiving and relaying dispersion cues to promote the modulation of c-di-GMP levels. The diguanylate cyclase NicD and the phosphodiesterase NbdA contribute to dispersion in a cue-specific manner, with NbdA sensing NO and NicD sensing nutrient cues. MucR has been reported to perceive both NO and nutrient cues. Receptors for other dispersion cues, such as heavy metals or the fatty acid cis-2-decenoic acid (cis-DA), have not yet been elucidated (indicated by the question mark). Relay of dispersion cues resulting in overall reduced c-di-GMP levels requires the chemotaxis-like MCP homolog, BdlA, and the c-di-GMP phosphodiesterases DipA and RbdA. BdlA, DipA, and RbdA are central to the dispersion response by NO, nutrients, and heavy metals. It is unclear, however, whether BdlA, DipA, and RbdA form a signaling cascade with NbdA or are involved in cis-DA induced dispersion (see question mark). BdlA is activated post dispersion cue sensing. Activation of BdlA involves phosphorylation and a burst of c-di-GMP, generated by diguanylate cyclases GcbA and NicD. Active BdlA recruits phosphodiesterase RbdA and enhances the activity of phosphodiesterase DipA, resulting in an overall reduction in c-di-GMP levels and subsequently, dispersion. No domain resolution is shown. For BdlA, domains are indicated only to indicate non-processive cleavage for BdlA activation. P, phosphorylation. Arrows indicate increased enzyme activity. IM, inner membrane; unknown
The phenotype of dispersed cells is distinct to biofilm cells, with dispersed cells demonstrating increased susceptibility to antimicrobial agents (Cherny and Sauer 2020; Cherny and Sauer 2019; Chambers et al. 2017) and flagellar-driven motility (Sauer et al. 2004). Recent findings suggest dispersion to coincide with the induction of several matrix-degrading enzymes, including the hydrolases PelA and PslG (Cherny and Sauer 2020) and the DNAses EndA and EddA (Cherny and Sauer 2019). Dispersion was furthermore enhanced by the release of the c-di-GMP responsive adhesin CdrA from the surface, with CdrA release likely contributing to the untethering of the matrix as a first step to enable Psl polysaccharide degradation (Cherny and Sauer 2020). The findings underscore that dispersion is likely a multi-step process to enable cells to revert back to a planktonic phenotype, a process that requires both high and low levels of c-di-GMP. Additionally, dispersion events are likely contributors to subpopulations within mature biofilms having low c-di-GMP levels.
7 Turnover and Modulation of the c-di-GMP Pool in Biofilms
Several small molecules have been reported to affect c-di-GMP pool. For instance, the intracellular pool of c-di-GMP is affected by 5′-phosphoguanylyl-(3′,5′)-guanosine (pGpG). pGpG is produced during the degradation of c-di-GMP, a two-step process involving EAL-dependent phosphodiesterases to linearize the c-di-GMP into pGpG, followed by hydrolysis by an oligoribonuclease into two GMPs. In P. aeruginosa, oligoribonuclease Orn (PA4951) has been identified as an oligoribonuclease required for c-di-GMP turnover and maintaining c-di-GMP homeostasis (Cohen et al. 2015; Orr et al. 2015). This was supported by loss of orn, resulting in increased intracellular levels of c-di-GMP and pGpG (Cohen et al. 2015). Moreover, loss of orn coincided with increased P. aeruginosa surface attachment and biofilm development, aggregation in liquid, and increased biofilm matrix production (Cohen et al. 2015). Moreover, high levels of pGpG have been shown to reduce the rate of c-di-GMP degradation in cell lysates, by inhibiting the activity of EAL-dependent PDEs (e.g., PA2133, PvrR, RocR) from P. aeruginosa. The findings suggested Orn to positively affect c-di-GMP turnover as well as PDE activity in general.
Another modulator of c-di-GMP levels is cAMP. While cAMP is a positive regulator of initial attachment by P. aeruginosa, leading to elevated levels of c-di-GMP (Luo et al. 2015; Lee et al. 2018) (Figs. 3.1 and 3.2), several reports indicate that in mature biofilms, high c-di-GMP levels coincide with low levels of cAMP (Almblad et al. 2019; Almblad et al. 2015). In turn, increased levels of cAMP, caused by either a lack of degradation or increased production, inhibit P. aeruginosa biofilm formation in a manner dependent on the virulence factor regulator Vfr and coinciding with reduced c-di-GMP levels and matrix Psl production (Lee et al. 2018). A subset of c-di-GMP-degrading phosphodiesterases involved in cAMP-Vfr-mediated biofilm inhibition in P. aeruginosa has been identified as DipA, RbdA, and BifA. It is of interest to note that the PDEs have previously been linked to the transitional episodes, namely dispersion and repression of attachment (Kuchma et al. 2007; Basu Roy et al. 2012; An et al. 2010).
Exposure to terrein, a fungal metabolite isolated from Aspergillus terreus with strong cytotoxic activity against cells with colorectal carcinoma, has been reported to have a negative impact on biofilm formation. The inhibitory effect has been linked to a decrease in c-di-GMP levels, with terrain inhibiting DGC activity (rather than increasing PDE activity) to affect the c-di-GMP pool (Kim et al. 2018). In contrast to terrein, other environmental cues have been linked to elevated c-di-GMP levels. Exposure of P. aeruginosa PAO1 to hypochlorite, a phagocyte-derived host defense compound that is also frequently used as a disinfectant, has been reported to induce significantly enhanced initial cell attachment, Pel and Psl matrix production, and c-di-GMP levels (Strempel et al. 2017). The hypochlorite-induced DGC contributing to the increased c-di-GMP pool in response to hypochlorite was identified as PA3177 (Strempel et al. 2017). Interestingly, the DGC PA3177 has also been linked to the tolerance of P. aeruginosa PAO1 biofilms to tobramycin and hydrogen peroxide (Poudyal and Sauer 2018a). Lack of drug tolerance by biofilms formed by ΔPA3177 coincided with reduced abundance of the BrlR (biofilm resistance locus regulator, PA4878), a c-di-GMP responsive transcriptional regulator. BrlR contributes to the high-level antimicrobial tolerance of P. aeruginosa biofilms by activating the expression of the multidrug efflux pump operons, mexAB-oprM and mexEF-oprN, and several ABC transport systems, including PA1874-77, as well as by repressing the expression of the oprH-phoPQ operon (Liao et al. 2013; Chambers et al. 2014; Chambers and Sauer 2013; Liao and Sauer 2012; Poudyal and Sauer 2018b).
Using the inverse relationship between biofilm formation (attaching to a surface) and swarming motility, Bernier et al. (2011) assessed the impact of amino acids, at concentrations relevant to those measured in CF sputum, on c-di-GMP levels by P. aeruginosa PA14. Most of the tested amino acids promoted both in vitro biofilm formation and swarming motility. The exception was arginine, which boosted biofilm formation but completely repressed swarming motility. Further analysis indicated that growth on arginine increased the intracellular levels of c-di-GMP, and this increase was dependent on the DGCs SadC and RoeA (Bernier et al. 2011). Arginine has been reported to likewise increase c-di-GMP levels in a DGC-dependent manner in biofilms formed P. putida (Barrientos-Moreno et al. 2020).
8 Blocking c-di-GMP for Biofilm Control
Numerous efforts are underway to develop innovative and effective strategies to control biofilms, ranging from (1) changing the properties of susceptible surfaces to prevent biofilm formation; (2) regulating signaling pathways to inhibit biofilm formation; (3) applying external forces to eradicate the biofilm, to (4) degrading the biofilm matrix to induce dispersion (Yin et al. 2021; Fleming and Rumbaugh 2017; Limqueco et al. 2020; Lu and Collins 2007; Han et al. 2019; Pestrak et al. 2019). Efforts have also focused on identifying strategies or compounds capable of reducing the intracellular level of c-di-GMP. This approach, if successful, is anticipated to disable several biofilm-related phenotypes at once, by suppressing biofilm formation and/or inducing dispersion, thus maintaining cells in the planktonic, antibiotic-susceptible mode of growth.
A reduction in the c-di-GMP content in bacteria can be achieved by inhibiting precursors (e.g., via the immunosuppressive drug azathioprine (Antoniani et al. 2013)), targeting the messenger (Hee et al. 2020), inhibiting DGCs, or activating PDEs. Several studies have focused on identifying compounds that inhibit DGC activity using c-di-GMP analogs (Zhou et al. 2013; Ching et al. 2010; Fernicola et al. 2015), identifying DGC inhibitors through in silico (Fernicola et al. 2015) or in vitro screening of compound libraries (Lieberman et al. 2014; Christen et al. 2019; Zheng et al. 2016), as well as in vivo screening of compound libraries in a number of different bacterial species (Sambanthamoorthy et al. 2012; Groizeleau et al. 2016; Bernardes et al. 2017). For instance, a screen targeting DGC activity by Sambanthamoorthy et al. (2012) resulted in the identification of seven small molecules that antagonize DGCs and inhibit biofilm formation by Vibrio cholerae. Two of these compounds significantly reduced the total concentration of c-di-GMP in V. cholerae, one of which also inhibited biofilm formation by P. aeruginosa in a continuous-flow system. The latter compound was identified as DI-3. Interestingly, although DI-3 was able to significantly reduce biofilm formation in flow cell assays, DI-3 did not induce dispersion nor inhibited biofilm formation under static growth conditions. Using a high-throughput screen, Andersen et al. (2021b) identified a small molecule that stimulated the activity of a specific PDE, BifA, in P. aeruginosa, resulting in efficient depletion of c-di-GMP, inhibition of biofilm formation, and dispersion. The small molecule was identified as 4-[(2-fluorophenyl) hydrazinylidene]pyrazole-3,5-diamine (H6-335-P1). In contrast, Hee et al. (2020) made use of a rationally designed peptide to sequester c-di-GMP. The c-di-GMP-sequestering peptide (CSP) was derived from a CheY-like c-di-GMP effector protein, based on the finding that a short arginine-rich region located at the C-termini of a novel family of CheY-like (Cle) proteins in Caulobacter crescentus binds c-di-GMP with nanomolar affinity (Nesper et al. 2017). The resulting CSP peptide, when endogenously expressed, was found to sequester c-di-GMP with sub-micromolar affinity, and inhibited P. aeruginosa biofilm formation (Hee et al. 2020).
None of these compounds appear to be in clinical use as of yet. However, another compound affecting c-di-GMP levels to induce biofilm dispersion, is currently undergoing clinical trials. This compound is nitric oxide (NO) and its derivatives, so-called NO donor prodrugs, that are designed to release NO at specific sites. These include sulfathiazole, cephalosporin-3′-diazeniumdiolates, with nitric oxide being released following cleavage of its β-lactam moiety by bacterial β-lactamases (Soren et al. 2020; Barraud et al. 2012), and spermine NONOate (N-[4-[1-(3-aminopropyl)-2-hydroxy-2-nitrosohydrazino]butyl]-1,3-propanediamine, S150) that spontaneously dissociates in a pH-dependent manner (Cai and Webb 2020; Marvasi et al. 2014). Nitric oxide and NO donor prodrugs were shown to significantly reduce c-di-GMP levels by targeting PDEs, and inducing significant dispersal in vitro of biofilms formed by P. aeruginosa, E. coli and Salmonella enterica (Cai and Webb 2020; Marvasi et al. 2014). Moreover, when co-administered with tobramycin, NO and NO prodrugs improved dispersal, and nearly completely eradicated biofilms when combined with colistin (Soren et al. 2020).
9 Conclusion
The goal of this review was to provide an up-to-date account of our current understanding of pathways and mechanisms leading to the modulation of c-di-GMP over the course of biofilm formation by P. aeruginosa. While these findings added to our understanding of the complexity of the c-di-GMP signaling network, they also raised several questions. For example, recent findings indicate that surface-attached cells do not uniformly produce c-di-GMP (Armbruster et al. 2019). And why would they? After all, other than the Psl-based positive feedback response (Irie et al. 2012), attached cells have no mechanism to gauge each other's intracellular c-di-GMP levels. Homogenous c-di-GMP levels would require attached cells to activate the relevant c-di-GMP signaling pathways in a synchronized manner which in turn would require attached cells to experience identical microenvironments. Additionally, why are there so many systems that become activated upon attachment that all contribute to the pool of c-di-GMP? Why the redundancy? It is possible that each system is part of a hierarchical regulatory cascade to gradually (or hierarchically) increase the c-di-GMP levels in the cell. However, it is also likely that each system provides a set of unique factors, enabling P. aeruginosa biofilm formation to progress. Given the requirement of each system for P. aeruginosa biofilms to mature, it is likely a combination, although there is little evidence as of yet to support this notion. Recent findings also suggest that c-di-GMP levels are not uniform. For one, mature biofilms do not uniformly produce c-di-GMP but instead demonstrate areas of low c-di-GMP in response to nutrient (or oxygen) availability. Likewise, low c-di-GMP levels in mature biofilms may also be the result of continued biofilm growth which requires modulation of the biofilm matrix, or to maintain the biofilm structure by preventing excess matrix production. Low c-di-GMP levels in mature biofilms have been demonstrated to be linked to increased PDE activity, conditions that have been previously linked only with dispersion. And while dispersion has been associated with an overall reduction in c-di-GMP, it is now apparent that nutrient-induced dispersion requires DGCs to produce a sudden rise or burst of c-di-GMP to subsequently activate PDEs and reduce c-di-GMP levels. It will be interesting to know whether other dispersion cues likewise require such a burst. It is exciting to think what else is yet to be uncovered.
References
Almblad H, Harrison JJ, Rybtke M, Groizeleau J, Givskov M, Parsek MR, Tolker-Nielsen T (2015) The cyclic AMP-Vfr signaling pathway in Pseudomonas aeruginosa is inhibited by cyclic di-GMP. J Bacteriol 197:2190–2200
Almblad H, Rybtke M, Hendiani S, Andersen JB, Givskov M, Tolker-Nielsen T (2019) High levels of cAMP inhibit Pseudomonas aeruginosa biofilm formation through reduction of the c-di-GMP content. Microbiology 165:324–333
An S, Je W, Zhang L-H (2010) Modulation of Pseudomonas aeruginosa biofilm dispersal by a cyclic-di-GMP phosphodiesterase with a putative hypoxia-sensing domain. Appl Environ Microbiol 76:8160–8173
Andersen JB, Kragh KN, Hultqvist LD, Rybtke M, Nilsson M, Jakobsen TH, Givskov M, Tolker-Nielsen T (2021a) Induction of native c-di-GMP phosphodiesterases leads to dispersal of Pseudomonas aeruginosa biofilms. Antimicrob Agents Chemother 65:e02431–e02420
Andersen JB, Hultqvist LD, Jansen CU, Jakobsen TH, Nilsson M, Rybtke M, Uhd J, Fritz BG, Seifert R, Berthelsen J (2021b) Identification of small molecules that interfere with c-di-GMP signaling and induce dispersal of Pseudomonas aeruginosa biofilms. NPJ Biofilms Microbiomes 7:1–13
Antoniani D, Rossi E, Rinaldo S, Bocci P, Lolicato M, Paiardini A, Raffaelli N, Cutruzzolà F, Landini P (2013) The immunosuppressive drug azathioprine inhibits biosynthesis of the bacterial signal molecule cyclic-di-GMP by interfering with intracellular nucleotide pool availability. Appl Microbiol Biotechnol 97:7325–7336
Armbruster CR, Parsek MR (2018) New insight into the early stages of biofilm formation. Proc Natl Acad Sci USA 115:4317–4319
Armbruster CR, Lee CK, Parker-Gilham J, de Anda J, Xia A, Zhao K, Murakami K, Tseng BS, Hoffman LR, Jin F, Harwood CS, Wong GCL, Parsek MR (2019) Heterogeneity in surface sensing suggests a division of labor in Pseudomonas aeruginosa populations. Elife 8:e45084
Baker AE, Diepold A, Kuchma SL, Scott JE, Ha DG, Orazi G, Armitage JP, O’Toole GA, Silhavy TJ (2016) PilZ domain protein FlgZ mediates cyclic di-GMP-dependent swarming motility control in Pseudomonas aeruginosa. J Bacteriol 198:1837–1846
Baraquet C, Harwood CS (2013) Cyclic diguanosine monophosphate represses bacterial flagella synthesis by interacting with the Walker A motif of the enhancer-binding protein FleQ. Proc Natl Acad Sci USA 110:18478–18483
Baraquet C, Murakami K, Parsek MR, Harwood CS (2012) The FleQ protein from Pseudomonas aeruginosa functions as both a repressor and an activator to control gene expression from the pel operon promoter in response to c-di-GMP. Nucleic Acids Res 40:7207–7218
Barraud N, Storey MV, Moore ZP, Webb JS, Rice SA, Kjelleberg S (2009a) Nitric oxide-mediated dispersal in single- and multi-species biofilms of clinically and industrially relevant microorganisms. J Microbial Biotechnol 2:370–378
Barraud N, Schleheck D, Klebensberger J, Webb JS, Hassett DJ, Rice SA, Kjelleberg S (2009b) Nitric oxide signaling in Pseudomonas aeruginosa biofilms mediates phosphodiesterase activity, decreased cyclic di-GMP levels, and enhanced dispersal. J Bacteriol 191:7333–7342
Barraud N, Kardak BG, Yepuri NR, Howlin RP, Webb JS, Faust SN, Kjelleberg S, Rice SA, Kelso MJ (2012) Cephalosporin-3′-diazeniumdiolates: targeted NO-Donor Prodrugs for Dispersing Bacterial Biofilms. Angew Chem Int Ed 51:9057–9060
Barrientos-Moreno L, Molina-Henares MA, Ramos-González MI, Espinosa-Urgel M (2020) Arginine as an environmental and metabolic cue for cyclic diguanylate signalling and biofilm formation in Pseudomonas putida. Sci Rep 10:1–15
Basu Roy A, Sauer K (2014) Diguanylate cyclase NicD-based signalling mechanism of nutrient-induced dispersion by Pseudomonas aeruginosa. Mol Microbiol 94:771–793
Basu Roy A, Petrova OE, Sauer K (2012) The phosphodiesterase DipA (PA5017) is essential for Pseudomonas aeruginosa biofilm dispersion. J Bacteriol 194:2904–2915
Bernardes ET, Charron-Mazenod L, Reading DJ, Reckseidler-Zenteno SL, Lewenza S (2017) Exopolysaccharide-repressing small molecules with antibiofilm and antivirulence activity against Pseudomonas aeruginosa. Antimicrob Agents Chemother 61:e01997–e01916
Bernier SP, Ha D-G, Khan W, Merritt JH, O’Toole GA (2011) Modulation of Pseudomonas aeruginosa surface-associated group behaviors by individual amino acids through c-di-GMP signaling. Res Microbiol 162:680–688
Bhuwan M, Lee HJ, Peng HL, Chang HY (2012) Histidine-containing phosphotransfer protein-B (HptB) regulates swarming motility through partner-switching system in Pseudomonas aeruginosa PAO1 strain. J Biol Chem 287:1903–1914
Boehm A, Kaiser M, Li H, Spangler C, Kasper CA, Ackermann M, Kaever V, Sourjik V, Roth V, Jenal U (2010) Second messenger-mediated adjustment of bacterial swimming velocity. Cell 141:107–116
Bordi C, Lamy M-C, Ventre I, Termine E, Hachani A, Fillet S, Roche B, Bleves S, Méjean V, Lazdunski A, Filloux A (2010) Regulatory RNAs and the HptB/RetS signalling pathways fine-tune Pseudomonas aeruginosa pathogenesis. Mol Microbiol 76:1427–1443
Borlee BR, Goldman AD, Murakami K, Samudrala R, Wozniak DJ, Parsek MR (2010) Pseudomonas aeruginosa uses a cyclic-di-GMP-regulated adhesin to reinforce the biofilm extracellular matrix. Mol Microbiol 75:827–842
Cai Y-m, Webb JS (2020) Optimization of nitric oxide donors for investigating biofilm dispersal response in Pseudomonas aeruginosa clinical isolates. Appl Microbiol Biotechnol 104:8859–8869
Cai Y-m, Hutchin A, Craddock J, Walsh MA, Webb JS, Tews I (2020) Differential impact on motility and biofilm dispersal of closely related phosphodiesterases in Pseudomonas aeruginosa. Sci Rep 10:6232
CDC (2019) Antibitic resistance threats in the United States. U.S. Department of Health and Human Services, CDC, Atlanta, GA, pp 1–150
CDC (2020) National action plan for combating antibiotic resistant bacteria, 2020-2025
Chambers JR, Sauer K (2013) The MerR-like regulator BrlR impairs Pseudomonas aeruginosa biofilm tolerance to colistin by repressing PhoPQ. J Bacteriol 195:4678–4688
Chambers JR, Liao J, Schurr MJ, Sauer K (2014) BrlR from Pseudomonas aeruginosa is a c-di-GMP-responsive transcription factor. Mol Microbiol 92:471–487
Chambers JR, Cherny KE, Sauer K (2017) Susceptibility of Pseudomonas aeruginosa dispersed cells to antimicrobial agents is dependent on the dispersion cue and class of the antimicrobial agent used. Antimicrob Agents Chemother 61:e00846–e00817
Chan C, Paul R, Samoray D, Amiot NC, Giese B, Jenal U, Schirmer T (2004) Structural basis of activity and allosteric control of diguanylate cyclase. Proc Natl Acad Sci USA 101:17084–17089
Chatterjee D, Cooley RB, Boyd CD, Mehl RA, O’Toole GA, Sondermann H (2014) Mechanistic insight into the conserved allosteric regulation of periplasmic proteolysis by the signaling molecule cyclic-di-GMP. Elife 3:e03650
Chen AI, Dolben EF, Okegbe C, Harty CE, Golub Y, Thao S, Ha DG, Willger SD, O’Toole GA, Harwood CS (2014) Candida albicans ethanol stimulates Pseudomonas aeruginosa WspR-controlled biofilm formation as part of a cyclic relationship involving phenazines. PLoS Pathog 10:e1004480
Cherny KE, Sauer K (2019) Pseudomonas aeruginosa requires the DNA-specific endonuclease EndA to degrade eDNA to disperse from the biofilm. J Bacteriol. https://doi.org/10.1128/JB.00059-19
Cherny KE, Sauer K (2020) Untethering and degradation of the polysaccharide matrix are essential steps in the dispersion response of Pseudomonas aeruginosa biofilms. J Bacteriol 202:e00575–e00519
Ching SM, Tan WJ, Chua KL, Lam Y (2010) Synthesis of cyclic di-nucleotidic acids as potential inhibitors targeting diguanylate cyclase. Bioorg Med Chem 18:6657–6665
Christen M, Kamischke C, Kulasekara HD, Olivas KC, Kulasekara BR, Christen B, Kline T, Miller SI (2019) Identification of small-molecule modulators of diguanylate cyclase by FRET-based high-throughput screening. ChemBioChem 20:394–407
Christensen LD, van Gennip M, Rybtke MT, Wu H, Chiang W-C, Alhede M, Høiby N, Nielsen TE, Givskov M, Tolker-Nielsen T (2013) Clearance of Pseudomonas aeruginosa foreign-body biofilm infections through reduction of the cyclic di-GMP level in the bacteria. Infect Immun 81:2705–2713
Cohen D, Mechold U, Nevenzal H, Yarmiyhu Y, Randall TE, Bay DC, Rich JD, Parsek MR, Kaever V, Harrison JJ, Banin E (2015) Oligoribonuclease is a central feature of cyclic diguanylate signaling in Pseudomonas aeruginosa. Proc Natl Acad Sci USA 112:11359–11364
Colvin KM, Irie Y, Tart CS, Urbano R, Whitney JC, Ryder C, Howell PL, Wozniak DJ, Parsek MR (2012) The Pel and Psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ Microbiol 14:1913–1928
Cooley RB, Smith TJ, Leung W, Tierney V, Borlee BR, O’Toole GA, Sondermann H (2016) Cyclic di-GMP-regulated periplasmic proteolysis of a Pseudomonas aeruginosa type Vb secretion system substrate. J Bacteriol 198:66–76
Costerton JW, Stewart PS, Greenberg EP (1999) Bacterial biofilms: a common cause of persistent infections. Science 284:1318–1322
Cutruzzolà F, Frankenberg-Dinkel N (2016) Origin and impact of nitric oxide in Pseudomonas aeruginosa biofilms. J Bacteriol 198:55–65
Darzins A (1994) Characterization of a Pseudomonas aeruginosa gene cluster involved in pilus biosynthesis and twitching motility: sequence similarity to the chemotaxis proteins of enterics and the gliding bacterium Myxococcus xanthus. Mol Microbiol 11:137–153
Dasgupta N, Ramphal R (2001) Interaction of the antiactivator FleN with the transcriptional activator FleQ regulates flagellar number in Pseudomonas aeruginosa. J Bacteriol 183:6636–6644
Dasgupta N, Wolfgang MC, Goodman AL, Arora SK, Jyot J, Lory S, Ramphal R (2003) A four-tiered transcriptional regulatory circuit controls flagellar biogenesis in Pseudomonas aeruginosa. Mol Microbiol 50:809–824
Davey ME, O’Toole GA (2000) Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64:847–867
Davies DG (1999) Regulation of matrix polymer in biofilm formation and dispersion. In: Wingender J, Neu TR, Flemming H-C (eds) Microbial extrapolymeric substances, characterization, structure and function. Springer, Berlin, pp 93–112
Davies D (2003) Understanding biofilm resistance to antibacterial agents. Nat Rev Drug Discov 2:114
Davies DG (2011) Biofilm dispersion. In: Biofilm highlights. Springer, Berlin, pp 1–28
Davies DG, Marques CNH (2009) A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J Bacteriol 191:1393–1403
Dow JM, Fouhy Y, Lucey JF, Ryan RP (2006) The HD-GYP domain, cyclic di-GMP signaling, and bacterial virulence to plants. Mol Plant Microbe Interact 19:1378–1384
Drenkard E, Ausubel FM (2002) Pseudomonas biofilm formation and antibiotic resistance are linked to phenotypic variation. Nature 416:740–743
Feng Q, Ahator SD, Zhou T, Liu Z, Lin Q, Liu Y, Huang J, Zhou J, Zhang L-H (2020) Regulation of exopolysaccharide production by ProE, a cyclic-di-GMP phosphodiesterase in Pseudomonas aeruginosa PAO1. Front Microbiol 11:1226
Fernicola S, Paiardini A, Giardina G, Rampioni G, Leoni L, Cutruzzolà F, Rinaldo S (2015) In silico discovery and in vitro validation of catechol-containing sulfonohydrazide compounds as potent inhibitors of the diguanylate cyclase PleD. J Bacteriol 198:147–156
Fleming D, Rumbaugh K (2017) Approaches to dispersing medical biofilms. Microorganisms 5:15
Flemming H-C (2016) EPS—then and now. Microorganisms 4:41
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633
Flemming H-C, Neu TR, Wozniak DJ (2007) The EPS matrix: the “house of biofilm cells”. J Bacteriol 189:7945–7947
Friedman L, Kolter R (2004a) Two genetic loci produce distinct carbohydrate-rich structural components of the Pseudomonas aeruginosa biofilm matrix. J Bacteriol 186:4457–4465
Friedman L, Kolter R (2004b) Genes involved in matrix formation in Pseudomonas aeruginosa PA14 biofilms. Mol Microbiol 51:675–690
Gjermansen M, Nilsson M, Yang L, Tolker-Nielsen T (2010) Characterization of starvation-induced dispersion in Pseudomonas putida biofilms: genetic elements and molecular mechanisms. Mol Microbiol 75:815–826
Groizeleau J, Rybtke M, Andersen JB, Berthelsen J, Liu Y, Yang L, Nielsen TE, Kaever V, Givskov M, Tolker-Nielsen T (2016) The anti-cancerous drug doxorubicin decreases the c-di-GMP content in Pseudomonas aeruginosa but promotes biofilm formation. Microbiology 162:1797–1807
Gupta K, Liao J, Petrova OE, Cherny KE, Sauer K (2014) Elevated levels of the second messenger c-di-GMP contribute to antimicrobial resistance of Pseudomonas aeruginosa. Mol Microbiol 92:488–506
Ha D-G, Richman ME, O’Toole GA (2014) Deletion mutant library for investigation of functional outputs of cyclic diguanylate metabolism in Pseudomonas aeruginosa PA14. Appl Environ Microbiol 80:3384–3393
Han C, Goodwine J, Romero N, Steck KS, Sauer K, Doiron A (2019) Enzyme-encapsulating polymeric nanoparticles: a potential adjunctive therapy in Pseudomonas aeruginosa biofilm-associated infection treatment. Colloids Surf B Biointerfaces 184:110512
Hay ID, Remminghorst U, Rehm BH (2009) MucR, a novel membrane-associated regulator of alginate biosynthesis in Pseudomonas aeruginosa. Appl Environ Microbiol 75:1110–1120
Heacock-Kang Y, Sun Z, Zarzycki-Siek J, McMillan IA, Norris MH, Bluhm AP, Cabanas D, Fogen D, Vo H, Donachie SP, Borlee BR, Sibley CD, Lewenza S, Schurr MJ, Schweizer HP, Hoang TT (2017) Spatial transcriptomes within the Pseudomonas aeruginosa biofilm architecture. Mol Microbiol 106:976–985
Hee C-S, Habazettl J, Schmutz C, Schirmer T, Jenal U, Grzesiek S (2020) Intercepting second-messenger signaling by rationally designed peptides sequestering c-di-GMP. Proc Natl Acad Sci USA 117:17211–17220
Hengge R (2009) Principles of c-di-GMP signalling in bacteria. Nat Rev Microbiol 7:263–273
Hickman JW, Harwood CS (2008) Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol Microbiol 69:376–389
Hickman JW, Tifrea DF, Harwood CS (2005) A chemosensory system that regulates biofilm formation through modulation of cyclic diguanylate levels. Proc Natl Acad Sci USA 102:14422–14427
Hinsa SM, Espinosa-Urgel M, Ramos JL, O’Toole GA (2003) Transition from reversible to irreversible attachment during biofilm formation by Pseudomonas fluorescens WCS365 requires an ABC transporter and a large secreted protein. Mol Microbiol 49:905–918
Hoffman LR, D’Argenio DA, MacCoss MJ, Zhang Z, Jones RA, Miller SI (2005) Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 436:1171–1175
Hsu JL, Chen HC, Peng HL, Chang HY (2008) Characterization of the histidine-containing phosphotransfer protein B-mediated multistep phosphorelay system in Pseudomonas aeruginosa PAO1. J Biol Chem 283:9933–9944
Huangyutitham V, Güvener ZT, Harwood CS (2013) Subcellular clustering of the phosphorylated WspR response regulator protein stimulates its diguanylate cyclase activity. mBio 4
Irie Y, Borlee BR, O’Connor JR, Hill PJ, Harwood CS, Wozniak DJ, Parsek MR (2012) Self-produced exopolysaccharide is a signal that stimulates biofilm formation in Pseudomonas aeruginosa. Proc Natl Acad Sci USA 109:20632–20636
Jacobs MA, Alwood A, Thaipisuttikul I, Spencer D, Haugen E, Ernst S, Will O, Kaul R, Raymond C, Levy R, Chun-Rong L, Guenthner D, Bovee D, Olson MV, Manoil C (2003) Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc Natl Acad Sci USA 100:14339–14344
Jenal U, Reinders A, Lori C (2017) Cyclic di-GMP: second messenger extraordinaire. Nat Rev Microbiol 15:271
Jones CJ, Newsom D, Kelly B, Irie Y, Jennings LK, Xu B, Limoli DH, Harrison JJ, Parsek MR, White P, Wozniak DJ (2014) ChIP-Seq and RNA-Seq reveal an AmrZ-mediated mechanism for cyclic di-GMP synthesis and biofilm development by Pseudomonas aeruginosa. PLoS Pathog 10:e1003984
Katharios-Lanwermeyer S, Whitfield GB, Howell PL, O’Toole G (2021) Pseudomonas aeruginosa uses c-di-GMP phosphodiesterases RmcA and MorA to regulate biofilm maintenance. MBio 12:e03384–e03320
Kazmierczak BI, Lebron MB, Murray TS (2006) Analysis of FimX, a phosphodiesterase that governs twitching motility in Pseudomonas aeruginosa. Mol Microbiol 60:1026–1043
Kim S-K, Lee J-H (2016) Biofilm dispersion in Pseudomonas aeruginosa. J Microbiol 54:71–85
Kim B, Park J-S, Choi H-Y, Yoon SS, Kim W-G (2018) Terrein is an inhibitor of quorum sensing and c-di-GMP in Pseudomonas aeruginosa: a connection between quorum sensing and c-di-GMP. Sci Rep 8:8617
Kimura S, Mori N, Kai T, Ishii Y, Yamaguchi K, Tateda K (2017) Azithromycin modulates 3′, 5′-cyclic diguanylic acid signaling in Pseudomonas aeruginosa. J Infect Chemother 23:550–555
Klauck G, Serra DO, Possling A, Hengge R (2018) Spatial organization of different sigma factor activities and c-di-GMP signalling within the three-dimensional landscape of a bacterial biofilm. Open Biol 8:180066
Kuchma SL, Brothers KM, Merritt JH, Liberati NT, Ausubel FM, O’Toole GA (2007) BifA, a c-di-GMP phosphodiesterase, inversely regulates biofilm formation and swarming motility by Pseudomonas aeruginosa PA14. J Bacteriol 189:8165–8178
Kuchma S, Ballok A, Merritt J, Hammond J, Lu W, Rabinowitz JD, O’Toole GA (2010) Cyclic-di-GMP-mediated repression of swarming motility by Pseudomonas aeruginosa: the pilY1 gene and its impact on surface-associated behaviors. J Bacteriol 192(12):2950–2964
Kulesekara H, Lee V, Brencic A, Liberati N, Urbach J, Miyata S, Lee DG, Neely AN, Hyodo M, Hayakawa Y, Ausubel FM, Lory S (2006) Analysis of Pseudomonas aeruginosa diguanylate cyclases and phosphodiesterases reveals a role for bis-(3′-5′)-cyclic-GMP in virulence. Proc Natl Acad Sci USA 103:2839–2844
Lee CK, de Anda J, Baker AE, Bennett RR, Luo Y, Lee EY, Keefe JA, Helali JS, Ma J, Zhao K, Golestanian R, O’Toole GA, Wong GCL (2018) Multigenerational memory and adaptive adhesion in early bacterial biofilm communities. Proc Natl Acad Sci USA 115:4471–4476
Li Y, Heine S, Entian M, Sauer K, Frankenberg-Dinkel N (2013) NO-induced biofilm dispersion in Pseudomonas aeruginosa is mediated by a MHYT-domain coupled phosphodiesterase. J Bacteriol 195:3531–3542
Li Y, Petrova OE, Su S, Lau GW, Panmanee W, Na R, Hassett DJ, Davies DG, Sauer K (2014) BdlA, DipA and induced dispersion contribute to acute virulence and chronic persistence of Pseudomonas aeruginosa. PLoS Pathog 10:e1004168
Liao J, Sauer K (2012) The MerR-like transcriptional regulator BrlR contributes to Pseudomonas aeruginosa biofilm tolerance. J Bacteriol 194:4823–4836
Liao J, Schurr MJ, Sauer K (2013) The MerR-like regulator BrlR confers biofilm tolerance by activating multidrug-efflux pumps in Pseudomonas aeruginosa biofilms. J Bacteriol 195:3352–3363
Lieberman OJ, Orr MW, Wang Y, Lee VT (2014) High-throughput screening using the differential radial capillary action of ligand assay identifies ebselen as an inhibitor of diguanylate cyclases. ACS Chem Biol 9:183–192
Limqueco E, Passos Da Silva D, Reichhardt C, Su F-Y, Das D, Chen J, Srinivasan S, Convertine A, Skerrett SJ, Parsek MR, Stayton PS, Ratner DM (2020) Mannose conjugated polymer targeting P. aeruginosa biofilms. ACS Infect Dis 6:2866–2871
Lu TK, Collins JJ (2007) Dispersing biofilms with engineered enzymatic bacteriophage. Proc Natl Acad Sci USA 104:11197–11202
Luo Y, Zhao K, Baker AE, Kuchma SL, Coggan KA, Wolfgang MC, Wong GC, O’Toole GA (2015) A hierarchical cascade of second messengers regulates Pseudomonas aeruginosa surface behaviors. MBio 6:e02456–e02414
Ma L, Conover M, Lu H, Parsek MR, Bayles K, Wozniak DJ (2009) Assembly and development of the Pseudomonas aeruginosa biofilm matrix. PLoS Pathog 5:e1000354
Malone JG, Jaeger T, Manfredi P, Dötsch A, Blanka A, Bos R, Cornelis GR, Häussler S, Jenal U (2012) The YfiBNR signal transduction mechanism reveals novel targets for the evolution of persistent Pseudomonas aeruginosa in cystic fibrosis airways. PLoS Pathog 8:e1002760
Marvasi M, Chen C, Carrazana M, Durie IA, Teplitski M (2014) Systematic analysis of the ability of nitric oxide donors to dislodge biofilms formed by Salmonella enterica and Escherichia coli O157: H7. AMB Express 4:1–11
Merritt JH, Brothers KM, Kuchma SL, O’Toole GA (2007) SadC reciprocally influences biofilm formation and swarming motility via modulation of exopolysaccharide production and flagellar function. J Bacteriol 189:8154–8164
Merritt JH, Ha D-G, Cowles KN, Lu W, Morales DK, Rabinowitz J, Gitai Z, O’Toole GA (2010) Specific control of Pseudomonas aeruginosa surface-associated behaviors by two c-di-GMP diguanylate cyclases. MBio 1:pii: e00183-00110
Monds RD, Newell PD, Gross RH, O’Toole GA (2007) Phosphate-dependent modulation of c-di-GMP levels regulates Pseudomonas fluorescens Pf0-1 biofilm formation by controlling secretion of the adhesin LapA. Mol Microbiol 63:656–679
Morgan R, Kohn S, Hwang S-H, Hassett DJ, Sauer K (2006) BdlA, a chemotaxis regulator essential for biofilm dispersion in Pseudomonas aeruginosa. J Bacteriol 188:7335–7343
Nair HA, Periasamy S, Yang L, Kjelleberg S, Rice SA (2017) Real time, spatial, and temporal mapping of the distribution of c-di-GMP during biofilm development. J Biol Chem 292:477–487
Navarro MV, De N, Bae N, Wang Q, Sondermann H (2009) Structural analysis of the GGDEF-EAL domain-containing c-di-GMP receptor FimX. Structure 17:1104–1116
Nesper J, Hug I, Kato S, Hee C-S, Habazettl JM, Manfredi P, Grzesiek S, Schirmer T, Emonet T, Jenal U (2017) Cyclic di-GMP differentially tunes a bacterial flagellar motor through a novel class of CheY-like regulators. Elife 6:e28842
Newell PD, Monds RD, O’Toole GA (2009) LapD is a bis-(3′,5′)-cyclic dimeric GMP-binding protein that regulates surface attachment by Pseudomonas fluorescens Pf0-1. Proc Natl Acad Sci USA 106:3461–3466
Newell PD, Boyd CD, Sondermann H, O’Toole GA (2011) A c-di-GMP effector system controls cell adhesion by inside-out signaling and surface protein cleavage. PLoS Biol 9:e1000587
O’Connor JR, Kuwada NJ, Huangyutitham V, Wiggins PA, Harwood CS (2012) Surface sensing and lateral subcellular localization of WspA, the receptor in a chemosensory-like system leading to c-di-GMP production. Mol Microbiol 86:720–729
Orr MW, Lee VT (2016) A PilZ domain protein for chemotaxis adds another layer to c-di-GMP-mediated regulation of flagellar motility. Sci Signal 9:fs16
Orr MW, Donaldson GP, Severin GB, Wang J, Sintim HO, Waters CM, Lee VT (2015) Oligoribonuclease is the primary degradative enzyme for pGpG in Pseudomonas aeruginosa that is required for cyclic-di-GMP turnover. Proc Natl Acad Sci USA 112:E5048–E5057
O’Toole GA, Kolter R (1998) Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: a genetic analysis. Mol Microbiol 28:449–461
Park S, Dingemans J, Gowett M, Sauer K (2021) Glucose-6-phosphate acts as an extracellular signal of SagS to modulate Pseudomonas aeruginosa c-di-GMP levels, attachment, and biofilm formation. mSphere 6:e01231–e01220
Persat A, Inclan YF, Engel JN, Stone HA, Gitai Z (2015) Type IV pili mechanochemically regulate virulence factors in Pseudomonas aeruginosa. Proc Natl Acad Sci USA 112:7563–7568
Pestrak MJ, Baker P, Dellos-Nolan S, Hill PJ, Passos da Silva D, Silver H, Lacdao I, Raju D, Parsek MR, Wozniak DJ, Howell PL (2019) Treatment with the Pseudomonas aeruginosa glycoside hydrolase PslG combats wound infection by improving antibiotic efficacy and host innate immune activity. Antimicrob Agents Chemother 63:e00234–e00219
Petrova OE, Sauer K (2009) A novel signaling network essential for regulating Pseudomonas aeruginosa biofilm development. PLoS Pathog 5:e1000668
Petrova OE, Sauer K (2010) The novel two-component regulatory system BfiSR regulates biofilm development by controlling the small RNA rsmZ through CafA. J Bacteriol 192:5275–5288
Petrova OE, Sauer K (2011) SagS contributes to the motile-sessile switch and acts in concert with BfiSR to enable Pseudomonas aeruginosa biofilm formation. J Bacteriol 193:6614–6628
Petrova OE, Sauer K (2012a) Dispersion by Pseudomonas aeruginosa requires an unusual posttranslational modification of BdlA. Proc Natl Acad Sci USA 109:16690–16695
Petrova OE, Sauer K (2012b) PAS domain residues and prosthetic group involved in BdlA-dependent dispersion response by Pseudomonas aeruginosa biofilms. J Bacteriol 194:5817–5828
Petrova OE, Sauer K (2016) Escaping the biofilm in more than one way: desorption, detachment or dispersion. Curr Opin Microbiol 30:67–78
Petrova OE, Cherny KE, Sauer K (2014) The P. aeruginosa diguanylate cyclase GcbA, a homolog of the P. fluorescens GcbA, promotes initial attachment to surfaces, but not biofilm formation, via regulation of motility. J Bacteriol 196:2827–2841
Petrova OE, Cherny KE, Sauer K (2015) The diguanylate cyclase GcbA facilitates Pseudomonas aeruginosa biofilm dispersion by activating BdlA. J Bacteriol 197(1):174–187
Petrova OE, Gupta K, Liao J, Goodwine JS, Sauer K (2017) Divide and conquer: the Pseudomonas aeruginosa two-component hybrid SagS enables biofilm formation and recalcitrance of biofilm cells to antimicrobial agents via distinct regulatory circuits. Environ Microbiol 19:2005–2024
Poudyal B, Sauer K (2018a) PA3177 encodes an active diguanylate cyclase that contributes to the biofilm antimicrobial tolerance but not biofilm formation by P. aeruginosa. Antimicrob Agents Chemother 62:e01049–e01018
Poudyal B, Sauer K (2018b) The ABC of biofilm drug tolerance: the MerR-like regulator BrlR is an activator of ABC transport systems, with PA1874-77 contributing to the tolerance of Pseudomonas aeruginosa biofilms to tobramycin. Antimicrob Agents Chemother 62:e01981–e01917
Purevdorj-Gage B, Costerton WJ, Stoodley P (2005) Phenotypic differentiation and seeding dispersal in non-mucoid and mucoid Pseudomonas aeruginosa biofilms. Microbiology 151:1569–1576
Reichhardt C, Wong C, Passos da Silva D, Wozniak DJ, Parsek MR (2018) CdrA interactions within the Pseudomonas aeruginosa biofilm matrix safeguard it from proteolysis and promote cellular packing. MBio 9:e01376-01318
Reichhardt C, Jacobs HM, Matwichuk M, Wong C, Wozniak DJ, Parsek MR (2020) The versatile Pseudomonas aeruginosa biofilm matrix protein CdrA promotes aggregation through different extracellular exopolysaccharide interactions. J Bacteriol 202:e00216–e00220
Römling U, Amikam D (2006) Cyclic di-GMP as a second messenger. Curr Opin Microbiol 9:218–228
Römling U, Galperin MY, Gomelsky M (2013) Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol Mol Biol Rev 77:1–52
Rumbaugh KP, Sauer K (2020) Biofilm dispersion. Nat Rev Microbiol 18:571–586
Ryan RP, Lucey J, O’Donovan K, McCarthy Y, Yang L, Tolker-Nielsen T, Dow JM (2009) HD-GYP domain proteins regulate biofilm formation and virulence in Pseudomonas aeruginosa. Environ Microbiol 11:1126–1136
Rybtke MT, Borlee BR, Murakami K, Irie Y, Hentzer M, Nielsen TE, Givskov M, Parsek MR, Tolker-Nielsen T (2012) Fluorescence-based reporter for gauging cyclic di-GMP levels in Pseudomonas aeruginosa. Appl Environ Microbiol 78:5060–5069
Rybtke M, Berthelsen J, Yang L, Høiby N, Givskov M, Tolker-Nielsen T (2015) The LapG protein plays a role in Pseudomonas aeruginosa biofilm formation by controlling the presence of the CdrA adhesin on the cell surface. MicrobiologyOpen 4(6):917–930
Ryder C, Byrd M, Wozniak DJ (2007) Role of polysaccharides in Pseudomonas aeruginosa biofilm development. Curr Opin Microbiol 10:644–648
Ryjenkov DA, Tarutina M, Moskvin OV, Gomelsky M (2005) Cyclic diguanylate is a ubiquitous signaling molecule in bacteria: insights into biochemistry of the GGDEF protein domain. J Bacteriol 187:1792–1798
Sambanthamoorthy K, Sloup RE, Parashar V, Smith JM, Kim EE, Semmelhack MF, Neiditch MB, Waters CM (2012) Identification of small molecules that antagonize diguanylate cyclase enzymes to inhibit biofilm formation. Antimicrob Agents Chemother 56:5202–5211
Sampedro I, Parales RE, Krell T, Hill JE (2014) Pseudomonas chemotaxis. FEMS Microbiol Rev 39:17–46
Sarenko O, Klauck G, Wilke F, Pfiffer V, Richter A, Herbst S, Kaever V, Hengge R (2017) More than enzymes that make and break c-di-GMP-the protein interaction network of GGDEF/EAL domain proteins of Escherichia coli. MBio 8:e01639-01617
Sauer K (2020) Cyclic di-GMP and the regulation of biofilm dispersion. In: Chou S-H, Guiliani N, Lee VT, Römling U (eds) Microbial cyclic di-nucleotide signaling. Springer, Cham, pp 545–560
Sauer K, Camper AK, Ehrlich GD, Costerton JW, Davies DG (2002) Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J Bacteriol 184:1140–1154
Sauer K, Cullen MC, Rickard AH, Zeef LAH, Davies DG, Gilbert P (2004) Characterization of nutrient-induced dispersion in Pseudomonas aeruginosa PAO1 biofilm. J Bacteriol 186:7312–7326
Serra DO, Hengge R (2014) Stress responses go three dimensional—the spatial order of physiological differentiation in bacterial macrocolony biofilms. Environ Microbiol 16:1455–1471
Simm R, Morr M, Kader A, Nimtz M, Romling U (2004) GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol Microbiol 53:1123–1134
Soren O, Rineh A, Silva DG, Cai Y, Howlin RP, Allan RN, Feelisch M, Davies JC, Connett GJ, Faust SN (2020) Cephalosporin nitric oxide-donor prodrug DEA-C3D disperses biofilms formed by clinical cystic fibrosis isolates of Pseudomonas aeruginosa. J Antimicrob Chemother 75:117–125
Starkey M, Hickman JH, Ma L, Zhang N, De Long S, Hinz A, Palacios S, Manoil C, Kirisits MJ, Starner TD, Wozniak DJ, Harwood CS, Parsek MR (2009) Pseudomonas aeruginosa rugose small-colony variants have adaptations that likely promote persistence in the Cystic fibrosis lung. J Bacteriol 191:3492–3503
Stoodley P, Sauer K, Davies DG, Costerton JW (2002) Biofilms as complex differentiated communities. Annu Rev Microbiol 56:187–209
Strempel N, Nusser M, Neidig A, Brenner-Weiss G, Overhage J (2017) The oxidative stress agent hypochlorite stimulates c-di-GMP synthesis and biofilm formation in Pseudomonas aeruginosa. Front Microbiol 8
Toutain CM, Caizza NC, Zegans ME, O’Toole GA (2007) Roles for flagellar stators in biofilm formation by Pseudomonas aeruginosa. Res Microbiol 158:471–477
Ueda A, Wood TK (2010) Tyrosine phosphatase TpbA of Pseudomonas aeruginosa controls extracellular DNA via cyclic diguanylic acid concentrations. Environ Microbiol Rep 2:449–455
Valentini M, Filloux A (2016) Biofilms and cyclic di-GMP (c-di-GMP) signaling: lessons from Pseudomonas aeruginosa and other bacteria. J Biol Chem 291:12547–12555
Valentini M, Laventie B-J, Moscoso J, Jenal U, Filloux A (2016) The diguanylate cyclase HsbD intersects with the HptB regulatory cascade to control Pseudomonas aeruginosa biofilm and motility. PLoS Genet 12:e1006354
Webster SS, Lee CK, Schmidt WC, Wong GC, O’Toole GA (2021) Interaction between the type 4 pili machinery and a diguanylate cyclase fine-tune c-di-GMP levels during early biofilm formation. Proc Natl Acad Sci USA 118
Wei Q, Leclercq S, Bhasme P, Xu A, Zhu B, Zhang Y, Zhang M, Wang S, Ma LZ, Kivisaar M (2019) Diguanylate cyclases and phosphodiesterases required for basal-level c-di-GMP in Pseudomonas aeruginosa as revealed by systematic phylogenetic and transcriptomic analyses. Appl Environ Microbiol 85:e01194–e01119
Whitchurch CB, Leech AJ, Young MD, Kennedy D, Sargent JL, Bertrand JJ, Semmler AB, Mellick AS, Martin PR, Alm RA (2004) Characterization of a complex chemosensory signal transduction system which controls twitching motility in Pseudomonas aeruginosa. Mol Microbiol 52:873–893
Williamson KS, Richards LA, Perez-Osorio AC, Pitts B, McInnerney K, Stewart PS, Franklin MJ (2012) Heterogeneity in Pseudomonas aeruginosa biofilms includes expression of ribosome hibernation factors in the antibiotic-tolerant subpopulation and hypoxia-induced stress response in the metabolically active population. J Bacteriol 194:2062–2073
Yin W, Xu S, Wang Y, Zhang Y, Chou S-H, Galperin MY, He J (2021) Ways to control harmful biofilms: prevention, inhibition, and eradication. Crit Rev Microbiol 47:57–78
Zhang Y, Guo J, Zhang N, Yuan W, Lin Z, Huang W (2019) Characterization and analysis of a novel diguanylate cyclase PA0847 from Pseudomonas aeruginosa PAO1. Infect Drug Resist 12:655
Zhao K, Tseng BS, Beckerman B, Jin F, Gibiansky ML, Harrison JJ, Luijten E, Parsek MR, Wong GC (2013) Psl trails guide exploration and microcolony formation in Pseudomonas aeruginosa biofilms. Nature 497:388–391
Zheng Y, Tsuji G, Opoku-Temeng C, Sintim HO (2016) Inhibition of P. aeruginosa c-di-GMP phosphodiesterase RocR and swarming motility by a benzoisothiazolinone derivative. Chem Sci 7:6238–6244
Zhou J, Watt S, Wang J, Nakayama S, Sayre DA, Lam Y-f, Lee VT, Sintim HO (2013) Potent suppression of c-di-GMP synthesis via I-site allosteric inhibition of diguanylate cyclases with 2′-Fc-di-GMP. Bioorg Med Chem 21:4396–4404
Acknowledgments
This work was supported by grants from the National Institutes of Health (2R01AI080710, R01AI150761).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Park, S., Sauer, K. (2022). Controlling Biofilm Development Through Cyclic di-GMP Signaling. In: Filloux, A., Ramos, JL. (eds) Pseudomonas aeruginosa. Advances in Experimental Medicine and Biology, vol 1386. Springer, Cham. https://doi.org/10.1007/978-3-031-08491-1_3
Download citation
DOI: https://doi.org/10.1007/978-3-031-08491-1_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-08490-4
Online ISBN: 978-3-031-08491-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)