Abstract
In nature, bacteria are primarily found as residents of surface-associated communities called biofilms. The formation of biofilms is a cyclical process that is initiated by single planktonic cells attaching to a surface, and comes full cycle when cells disperse from the mature biofilm to resume a planktonic lifestyle. Dispersion occurs in response to various signals and environmental cues, and results in surface-attached organisms liberating themselves from matrix-encased biofilms, apparent by single cells actively escaping from the biofilm, leaving behind eroded biofilms and microcolonies having central voids. Given the cyclic process of biofilm formation, it is not surprising that dispersion, like biofilm formation, is coincident with significant changes in the levels of the second messenger cyclic di-GMP. However, dispersion is not simply a reversion from the biofilm lifestyle to the planktonic mode of growth, as dispersed cells have been described as having a phenotype that is distinct from planktonic and biofilm cells. Using primarily the pathogen P. aeruginosa as example, this chapter provides an up-to-date compendium of cyclic di-GMP pathways connected to biofilm dispersion, including how sensing a diverse array of dispersion cues leads to the destruction of cyclic di-GMP, the escape from the biofilm matrix, and the appropriate phenotypic responses associated with dispersed cells.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Dispersion
- Cyclic di-GMP
- Motility
- Susceptibility
- Virulence
- cis-DA
- Extracellular dispersion cues
- Dispersion cue perception
- Signal relay
- Matrix degradative enzymes
1 Dispersion as a Flight Response
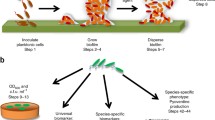
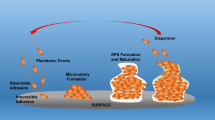
Biofilms are defined as a structured community of bacterial cells enclosed in a self-produced polymeric matrix and adherent to inert or living surfaces [1]. The ability to form a biofilm is a common trait of a diverse array of microbes, including lower order eukaryotes, with biofilms being the predominant mode of bacterial growth in nature [2]. The sessile lifestyle affords bacteria multiple protective advantages, allowing bacteria to remain within a favorable environmental niche or host. Compared to free-swimming bacteria, biofilms are better adapted to withstand nutrient deprivation, pH changes, oxygen radicals, biocides, and antimicrobial agents [3]. However, being in a biofilm is not only advantageous. As a biofilm grows in size, some cells will become increasingly separated from the bulk liquid interface and essential sources of energy or nutrients. Accumulation of waste products and toxins in the interior of biofilms pose additional challenges. Being trapped deep within a biofilm can, therefore, threaten cell survival. Thus, biofilm cells have evolved mechanisms which enable escaping the sessile mode of growth as a means of self-preservation, by liberating themselves from matrix-encased biofilms, and reverting back to the planktonic mode of growth. The transition to the planktonic mode of growth is referred to as dispersion [4,5,6,7]. Moreover, dispersion is considered a mechanism by which enables dissemination to new locales for colonization [4, 5, 8]. First described by Davies in 1999 [9], dispersion is apparent by single cells actively escaping from the biofilm, leaving behind eroded biofilms and microcolonies having central voids (Fig. 31.1) [4,5,6,7, 10,11,12,13,14,15]. Dispersion rarely involves the entire biofilm, with no more than 80% of the biofilm biomass being removed upon induction of dispersion [10, 13, 16,17,18]. Instead, selected microcolonies or areas within a biofilm will undergo a dispersion event at any particular time, in a manner often dependent on microcolony diameter [8].
2 Dispersion Induces a Switch in the Mode of Growth
Considering that dispersed cells escape from the biofilm as single cells suggests that dispersion is a way for bacteria to transition from the surface-associated to the planktonic mode of growth. Transition to and from the surface have been linked to the modulation of the intracellular signaling molecule bis-(3′-5′)-cyclic dimeric guanosine monophosphate (cyclic di-GMP). More specifically, biofilm formation or the sessile lifestyle have been associated with high levels of cyclic di-GMP, with elevated cyclic di-GMP levels, in turn, resulting in increased production of biofilm matrix components, adhesiveness/autoaggregation, and antimicrobial tolerance, but repressed motility [19,20,21,22,23,24,25,26]. In contrast, low cyclic di-GMP levels have been associated with a motile or planktonic existence. Levels of cyclic di-GMP are enzymatically modulated by diguanylate cyclases (DGCs), proteins containing a GGDEF domain, and phosphodiesterases (PDEs) harboring either an EAL or HD-GYP domain. In agreement with dispersion coinciding with single cells actively escaping from the biofilm and transitioning toward a motile mode of growth, dispersed cells are motile, characterized by increased expression of fliC (encoding flagellin type B), and cyclic di-GMP levels comparable to or lower than those found in planktonic cells [12, 13, 27, 28].
3 Translation of Dispersion Cue Perception into the Modulation of the Intracellular Cyclic di-GMP Pool
How is dispersion induced and how does dispersion result in a reduction in cyclic di-GMP levels? Dispersion occurs in response to a number of cues and signals including fatty acid signaling molecule belonging to the family of diffusible signaling factors (DSF), pH, ammonium chloride, heavy metals, and nitric oxide (NO), host factors such as bile salts, and availability of oxygen, iron, amino acids, and carbon sources (Table 31.1). The mechanism of dispersion cue perception has been determined in more detail for a select number of dispersion agents including fatty acid signals, carbon sources, and NO. In each case, dispersion cue sensing was found to require at minimum a membrane-bound sensory protein, and a protein involved in the modulation of cyclic di-GMP levels such as a phosphodiesterase or in the case of NO sensing, a bifunctional enzyme harboring GGDEF-EAL domains (Fig. 31.2). The components form a signal transduction cascade that upon dispersion cue perception likely initiate a phosphorelay to the cyclic di-GMP modulating enzyme, resulting in the activation of the phosphodiesterase activity, and thus, the reduction of cellular levels of cyclic di-GMP (Fig. 31.2). Our current understanding of selected dispersion signaling pathways resulting in the alteration in the cellular level of cyclic di-GMP is discussed in detail below.
Dispersion cue perception and relay resulting in the modulation of cyclic di-GMP levels. (a) Model of the DSF/Rpf signaling cascades in X. campesitris. RpfF synthesizes DSF. RpfC is a sensor kinase and the response regulator RpfG exhibits phosphodiesterase activity. The signaling cascade is shown at low (left) and high (right) DSF concentrations. (b) Model of the signaling cascade involved in sensing nutrient dispersion cues in P. aeruginosa. Conditions at low (left) and high (right) concentrations of nutrient dispersion cues are shown. (c) Nitric oxide-induced signaling events mediated by the heme-based sensor domain H-NOX. Left, nitric oxide-induced signaling events in Legionella pneumophila or Shewanella woodyi. NO sensing via the H-NOX protein results in reduced cyclic di-GMP levels by stimulating the PDE activity (EAL) domain. Right, in Vibrio cholerae or S. oneidensis, interaction of the NO-bound H-NOX domain with a coupled histidine kinase (H-NOK) controls the phosphorylation activity of the kinase. Specific phosphorylation events lead to a decrease in cyclic di-GMP levels, either by stimulating PDE or by controlling the transcriptional response through a dedicated transcription regulator (HTH). (d) Possible mechanism of NO-induced biofilm dispersal via the MHYT domain-containing NbdA protein in P. aeruginosa. While BdlA, DipA, and RbdA have been shown to be required for NO-induced dispersion, it is unclear whether they form a signaling cascade with NbdA. Binding of NO is supposed to occur via copper ions (Cu) located in the MHYT domain. . Conditions at high NO concentrations are shown. With the exception of NicD and NbdA, no domain resolution is shown. For BdlA, domains are indicated only to indicate non-processive cleavage for BdlA activation. DSF, cis-11-methyl-2-dodecenoic acid. PDE, phosphodiesterase. DGC, diguanylate cyclase. HTH, helix-turn-helix, transcription regulator. DISMED2, predicted periplasmic sensory domain. MHYT, predicted periplasmic sensory domain for diatomic gases. P, phosphorylation. Arrows indicate increased enzyme activity. Width of gray arrows indicates level of activity
3.1 Fatty Acids as Dispersion Signals
Akin to cell–cell signaling molecules, fatty acid signals are involved in intra-species, inter-species, and cross-kingdom communication where they regulate community-associated behavior including biofilm dispersion. The fatty acid acting as a dispersion autoinducer of P. aeruginosa biofilms has been identified as cis-2-decenoic acid (cis-DA) [10]. Additional dispersion autoinducer molecules include cis-11-methyl-2-dodecenoic acid (DSF) that has been shown to disaggregate flocs by Xanthomonas campesitris in liquid [46], and the Burkholderia cenocepatia cis-2-dodecenoic acid (BDSF) [47]. The dispersion response to cis-unsaturated fatty acids is fairly conserved, as cis-DA has been shown to induce dispersion of biofilms by Escherichia coli, Klebsiella pneumoniae, Proteus mirabilis, Streptococcus pyogenes, Bacillus subtilis, Staphylococcus aureus, and yeast Candida albicans biofilms [10], while the B. cenocepatia BDSF has been shown to trigger dispersion of Francisella novicida biofilms [47]. Amari et al. [51] demonstrated that production of cis-DA requires an enoyl-CoA synthetase encoded by dspI (PA14_54640, a PA0745 ortholog), with dspI inactivation resulting in significantly reduced dispersion and defective swarming motility. Microarray analysis furthermore suggested cis-DA to affect the expression of 666 genes encoding proteins involved in motility, chemotaxis, cell attachment, TCA cycle, exopolysaccharides and LPS synthesis and secretion, virulence, iron uptake, and respiration [52]. Rahmani-Badi et al. [52] furthermore predicted PA4982–PA4983 encoding a two-component system, to be involved in cis-DA signal perception. No experimental evidence, however, for cis-DA being sensed via PA4982–PA4983 exist. While mechanism by which cis-DA is perceived by P. aeruginosa and relayed to result in dispersion has yet to be elucidated, much is known about DSF signal sensing by X. campesitris (Fig. 31.2a) which requires the rpf gene cluster. While RpfF directs the synthesis of DSF, a two-component sensory transduction system comprising the hybrid sensor kinase RpfC and the response regulator RpfG has been implicated in the perception of the DSF signal and signal transduction [46]. DSF perception by RpfC is believed to lead to its autophosphorylation and subsequent phosphorelay to RpfG. RpfG is unique in that it contains no DNA-binding domain, but an HD-GYP domain that exhibits phosphodiesterase activity capable of degrading cyclic di-GMP to GMP [46, 53,54,55]. Phosphorylation is thought to activate RpfG for cyclic di-GMP degradation. In this way, RpfC/RpfG link perception of the cell–cell signal DSF to alteration in the cellular level of cyclic di-GMP.
3.2 Nutrient-Induced Dispersion
Both starvation and the sudden excess of nutrients have been shown to induce dispersion by Pseudomonas sp (Table 31.1). Nutrient cues include glucose, glutamate, succinate, citrate and induce dispersion when biofilms are exposed to a sudden increase in the carbon source concentration. Basu Roy and Sauer [11] demonstrated that while L-glutamate supported growth of P. aeruginosa, D-glutamate did not. However, both D- and L-glutamate were capable of inducing dispersion, indicating that nutrient cues are not metabolized in order to induce dispersion. Instead, in P. aeruginosa, nutrient cues including glutamate, citrate, and glucose are sensed by the diguanylate cyclase NicD belonging to a family of seven transmembrane (7TM) receptors (Fig. 31.2b). NicD directly interacts with BdlA and the phosphodiesterase DipA, with NicD contributing to the membrane association of the protein complex [11, 37]. Nutrient cue perception by NicD is believed to lead to dephosphorylation, with posttranslational modification coinciding with increased cyclase activity. Thus activated NicD contributes to the non-processive proteolysis and activation of the chemotaxis transducer protein BdlA, via phosphorylation and temporarily elevated cyclic di-GMP levels [11, 14]. BdlA activation requires an unusual, non-processive proteolytic cleavage found to be stimulated by increased cyclic di-GMP levels, and dependent on the protease ClpP, the chaperone ClpD, and BdlA phosphorylation [14, 37]. BdlA, in turn, activates the phosphodiesterase DipA, and recruits a second phosphodiesterase, RbdA, to ultimately reduce cellular cyclic di-GMP levels (Fig. 31.2b). An additional player is the diguanylate cyclase GcbA. GcbA contributes to BdlA cleavage during biofilm growth and has been shown to play an essential role in allowing biofilm cells to disperse in response to a variety of substances including carbohydrates, heavy metals, and NO [14, 15]. Likewise, BdlA and DipA appear to be required for P. aeruginosa biofilm dispersion in response to NO and heavy metals [12, 13], indicating GcbA, BdlA, and DipA to play a central role in the translation of a large variety of dispersion cues into the modulation of the intracellular cyclic di-GMP pool.
3.3 NO-Induced Dispersion
The diatomic gas nitric oxide (NO), a well-known signaling molecule in both prokaryotes and eukaryotes, is able to induce the dispersal of P. aeruginosa and other Gram-negative bacterial biofilms (Table 31.1). NO was first suggested by Webb et al. [56] to stimulate the release of planktonic cells from an established P. aeruginosa biofilm. The finding of NO serving as dispersion inducer was confirmed by Barraud et al. [18, 57] using several NO donors. Moreover, the studies linked NO to low cyclic di-GMP levels and changes in phosphodiesterase activity in a dose-dependent manner. In most species, NO is sensed by H-NOX (heme-nitric oxide/oxygen-binding) domain proteins, by NO binding to the heme moiety of H-NOX. H-NOX can directly interact with DGC to regulate cyclic di-GMP synthesis and degradation. In Legionella pneumophila or Shewanella woodyi, the H-NOX protein interacts upon NO binding with the bifunctional GGDEF-EAL (HaCE) protein, and lowers cyclic di-GMP levels by inhibiting the DGC activity but stimulation the PDE activity of the HaCE [58] (Fig. 31.2c). In Vibrio cholerae or S. oneidensis, interaction of the NO-bound H-NOX domain with a coupled histidine kinase (H-NOK) controls the phosphorylation activity of the kinase (Fig. 31.2c). Specific phosphorylation events lead to a decrease in cyclic di-GMP levels, either by stimulating the hydrolysis of cyclic di-GMP by a cognate PDE (via the fused REC domain) or by controlling the transcriptional response through a dedicated transcription regulator (HTH) [58]. In P. aeruginosa, NO sensing likewise involves the activation of cyclic di-GMP-specific phosphodiesterases in P. aeruginosa, ultimately leading to cyclic di-GMP decrease and biofilm dispersal (Fig. 31.2d). However, P. aeruginosa does not encode H-NOX proteins. Instead, NO sensing in P. aeruginosa was found to be linked to NbdA, an MHYT domain harboring phosphodiesterase [38, 58]. MHYT is a transmembrane domain of seven predicted membrane spanning helices and proposed to possess putative sensory function for diatomic gases like oxygen, carbon monoxide, or NO through protein-bound copper ions [59]. Considering that inactivation of bdlA, dipA, and gcbA impairs dispersion by P. aeruginosa biofilms in response to NO, the signaling cascade likely requires, in addition to NbdA, also BdlA, DipA, (Fig. 31.2d), and GcbA.
4 The Dispersion Phenotype
The cellular cyclic di-GMP levels noted upon dispersion cue sensing and induction of dispersion have been reported to be comparable to or lower than those found in planktonic cells [12, 13, 27, 28]. The low cyclic di-GMP levels explain much of the similarities found between dispersed and planktonic cells. Relative to biofilms, both are motile, and are susceptible to antimicrobial agents [16, 60,61,62]. However, dispersed cells are not identical to planktonic cells [4, 60, 61]. Instead, dispersed cells were found to be highly virulent when tested using various acute and chronic virulence models, to produce more matrix degrading enzymes, to be more primed to re-attach following egress from the biofilm, and to exhibit protein production and gene expression profiles that are distinct from planktonic cells and biofilms from which they escaped [4, 39, 60, 61, 63]. The distinct phenotype of dispersed cells, however, was found to be reversible and short-lived. Using qRT-PCR and antimicrobial susceptibility assays, Chambers et al. [60] demonstrated that in P. aeruginosa, differences between planktonic and dispersed cells remained for 2 h post-dispersion, with additional time being required for dispersed cells to display expression of genes indicative of exponential growth.
5 Cyclic di-GMP Levels and Downstream Pathways
The finding of dispersed cells being characterized by reduced cyclic di-GMP levels has led to the hypothesis that dispersed cells can be generated by reducing the intracellular cyclic di-GMP content through modulation of PDEs [28]. However, the cyclic di-GMP activated pathways have not been fully elucidated. Considering that dispersion coincides with biofilm erosion and single cells escaping the biofilm structure (Fig. 31.1), dispersion likely relies on factors that weaken the biofilm matrix. The biofilm matrix is composed of polysaccharides, eDNA, and adhesins [64], with Pseudomonas sp. using cyclic di-GMP regulated adhesins to reinforce the biofilm matrix. These adhesins have been identified in P. putida and P. fluorescens as the large outer-membrane protein LapA [27, 65,66,67], and CdrA in P. aeruginosa [67, 68]. Elevated cyclic di-GMP levels contribute to the localization of LapA to the cell surface, while low cyclic di-GMP levels result in LapA being released from the outer membrane via cleavage by the periplasmic cysteine protease LapG [66]. Gjermansen et al. [27] demonstrated that in P. putida, carbon starvation decrease the level of LapA, with LapA release resulting in biofilm dispersal, a response that was absent in ΔlapG mutant biofilms. Additionally, a plethora of matrix degrading factors such as proteases, deoxyribonucleases, and glycoside hydrolases have been linked to biofilm dispersal [39, 61, 69,70,71,72]. However, most studies have relied on inducing dispersion by the exogenous addition of these factors. For instance, PslG, a glycosyl hydrolase involved in the synthesis/degradation of a key biofilm matrix exopolysaccharide Psl in P. aeruginosa, disassembles existing biofilms within minutes at nanomolar concentrations when supplied exogenously [71]. However, as PslG is not predicted to be released from the cell, it is unlikely that PslG indeed contributes to matrix degradation during dispersion or activated in a low cyclic di-GMP environment. As for now, specific matrix degrading factors remain elusive.
Dispersion furthermore coincides with bacteria escaping from the biofilm being susceptible to antimicrobial agents. Recent findings suggested a link between cyclic di-GMP and drug susceptibility [60, 73, 74]. For instance, Gupta et al. [73] demonstrated that P. aeruginosa planktonic cells were rendered more resistant to antimicrobial agents upon increasing intracellular cyclic di-GMP, from 10–30 pmol/mg, to cyclic di-GMP levels more commonly found in biofilm cells (≥80 pmol/mg). Additionally, drug tolerance by P. aeruginosa biofilms and dispersed cells has been linked to the cyclic di-GMP-responsive transcriptional regulator BrlR [60, 75]. BrlR contributes to biofilm drug tolerance by activating the expression of multidrug efflux pumps and ABC transporters [76,77,78]. Low cyclic di-GMP levels, however, negatively impact BrlR levels and BrlR-DNA binding [26].
6 Concluding Remarks
Being a near-ubiquitous second messenger that coordinates diverse aspects of bacterial growth and behavior, it is not surprising that cyclic di-GMP has become known as the “second messenger extraordinaire” [79]. However, despite the large number of bacterial behavior and functional outputs that have been characterized since its discovery in the late 1980s, there is still much to learn, especially when it comes to the role of cyclic di-GMP in biofilm dispersion. While future experiments will be required to elucidate cyclic di-GMP dependent pathways leading to dispersion, indirect evidence suggests a role of AmrZ and FleQ. Originally described to inversely regulate alginate production and swimming motility in P. aeruginosa, AmrZ is now recognized as a global regulator of multiple virulence factors, including cyclic di-GMP, extracellular polysaccharide production including Pel and Psl polysaccharides, and flagella [80]. Support for AmrZ playing a role in dispersion stems from AmrZ affecting gcbA expression and inversely regulating exopolysaccharide production and motility [80]. Additionally, Chua et al. [61] demonstrated amrZ to be differentially expressed in dispersed relative to planktonic cells using RNA-seq. Similarly, FleQ may contribute to the cyclic di-GMP dependent pathways to induce dispersion. This cyclic di-GMP responsive transcriptional inversely contributes to the expression of pel genes required for Pel polysaccharide biosynthesis and of flagellar genes in response to cyclic di-GMP [81,82,83]. At high cyclic di-GMP levels, FleQ induces the expression of the pel operon while at low cyclic di-GMP levels, FleQ regulates the expression of flagellar genes but represses transcription of the pel operon required for Pel polysaccharide biosynthesis. It is of interest to note that FleQ is under the transcriptional control of AmrZ [80].
References
Costerton JW, Stewart PS, Greenberg EP (1999) Bacterial biofilms: a common cause of persistent infections. Science 284(5418):1318–1322
Geesey GG, Richardson WT, Yeomans HG, Irvin RT, Costerton JW (1977) Microscopic examination of natural sessile bacterial populations from an alpine stream. Can J Microbiol 23:1733–1736
Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM (1995) Microbial biofilms. Annu Rev Microbiol 49:711–745
Sauer K, Camper AK, Ehrlich GD, Costerton JW, Davies DG (2002) Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J Bacteriol 184(4):1140–1154
Stoodley P, Sauer K, Davies DG, Costerton JW (2002) Biofilms as complex differentiated communities. Annu Rev Microbiol 56(1):187–209. https://doi.org/10.1146/annurev.micro.56.012302.160705
Petrova OE, Sauer K (2016) Escaping the biofilm in more than one way: desorption, detachment or dispersion. Curr Opin Microbiol 30:67–78
Davies DG (2011) Biofilm dispersion. In: Biofilm highlights. Springer, Berlin, pp 1–28
Purevdorj-Gage B, Costerton WJ, Stoodley P (2005) Phenotypic differentiation and seeding dispersal in non-mucoid and mucoid Pseudomonas aeruginosa biofilms. Microbiology 151(5):1569–1576. https://doi.org/10.1099/mic.0.27536-0
Davies DG (1999) Regulation of matrix polymer in biofilm formation and dispersion. In: Wingender J, Neu TR, Flemming H-C (eds) Microbial extrapolymeric substances, characterization, structure and function. Springer, Berlin, pp 93–112
Davies DG, Marques CNH (2009) A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J Bacteriol 191(5):1393–1403. https://doi.org/10.1128/jb.01214-08
Basu Roy A, Sauer K (2014) Diguanylate cyclase NicD-based signalling mechanism of nutrient-induced dispersion by Pseudomonas aeruginosa. Mol Microbiol 94(4):771–793. https://doi.org/10.1111/mmi.12802
Basu Roy A, Petrova OE, Sauer K (2012) The phosphodiesterase DipA (PA5017) is essential for Pseudomonas aeruginosa biofilm dispersion. J Bacteriol 194:2904–2915. https://doi.org/10.1128/jb.05346-11
Morgan R, Kohn S, Hwang S-H, Hassett DJ, Sauer K (2006) BdlA, a chemotaxis regulator essential for biofilm dispersion in Pseudomonas aeruginosa. J Bacteriol 188(21):7335–7343
Petrova OE, Sauer K (2012) Dispersion by Pseudomonas aeruginosa requires an unusual posttranslational modification of BdlA. Proc Natl Acad Sci U S A 109(41):16690–16695
Petrova OE, Cherny KE, Sauer K (2015) The diguanylate cyclase GcbA facilitates Pseudomonas aeruginosa biofilm dispersion by activating BdlA. J Bacteriol 197(1):174–187. https://doi.org/10.1128/jb.02244-14
Sauer K, Cullen MC, Rickard AH, Zeef LAH, Davies DG, Gilbert P (2004) Characterization of nutrient-induced dispersion in Pseudomonas aeruginosa PAO1 biofilm. J Bacteriol 186(21):7312–7326. https://doi.org/10.1128/jb.186.21.7312-7326.2004
Barraud N, Storey MV, Moore ZP, Webb JS, Rice SA, Kjelleberg S (2009) Nitric oxide-mediated dispersal in single- and multi-species biofilms of clinically and industrially relevant microorganisms. Microb Biotechnol 2(3):370–378
Barraud N, Schleheck D, Klebensberger J, Webb JS, Hassett DJ, Rice SA, Kjelleberg S (2009) Nitric oxide signaling in Pseudomonas aeruginosa biofilms mediates phosphodiesterase activity, decreased cyclic di-GMP levels, and enhanced dispersal. J Bacteriol 191(23):7333–7342. https://doi.org/10.1128/jb.00975-09
Kuchma SL, Brothers KM, Merritt JH, Liberati NT, Ausubel FM, O’Toole GA (2007) BifA, a c-di-GMP phosphodiesterase, inversely regulates biofilm formation and swarming motility by Pseudomonas aeruginosa PA14. J Bacteriol 189:8165–8178. https://doi.org/10.1128/jb.00586-07
Merritt JH, Brothers KM, Kuchma SL, O’Toole GA (2007) SadC reciprocally influences biofilm formation and swarming motility via modulation of exopolysaccharide production and flagellar function. J Bacteriol 189(22):8154–8164. https://doi.org/10.1128/jb.00585-07
Römling U, Amikam D (2006) Cyclic di-GMP as a second messenger. Curr Opin Microbiol 9(2):218–228. https://doi.org/10.1016/j.mib.2006.02.010
Simm R, Morr M, Kader A, Nimtz M, Romling U (2004) GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol Microbiol 53(4):1123–1134. https://doi.org/10.1111/j.1365-2958.2004.04206.x
Thormann KM, Duttler S, Saville RM, Hyodo M, Shukla S, Hayakawa Y, Spormann AM (2006) Control of formation and cellular detachment from Shewanella oneidensis MR-1 biofilms by cyclic di-GMP. J Bacteriol 188(7):2681–2691. https://doi.org/10.1128/jb.188.7.2681-2691.2006
Romling U, Gomelsky M, Galperin MY (2005) C-di-GMP: the dawning of a novel bacterial signalling system. Mol Microbiol 57(3):629–639. https://doi.org/10.1111/j.1365-2958.2005.04697.x
Kirillina O, Fetherston JD, Bobrov AG, Abney J, Perry RD (2004) HmsP, a putative phosphodiesterase, and HmsT, a putative diguanylate cyclase, control Hms-dependent biofilm formation in Yersinia pestis. Mol Microbiol 54(1):75–88. https://doi.org/10.1111/j.1365-2958.2004.04253.x
Poudyal B, Sauer K (2018) PA3177 encodes an active diguanylate cyclase that contributes to the biofilm antimicrobial tolerance but not biofilm formation by P. aeruginosa. Antimicrob Agents Chemother 62(10):e01049–e01018. https://doi.org/10.1128/aac.01049-18
Gjermansen M, Nilsson M, Yang L, Tolker-Nielsen T (2010) Characterization of starvation-induced dispersion in Pseudomonas putida biofilms: genetic elements and molecular mechanisms. Mol Microbiol 75(4):815–826. https://doi.org/10.1111/j.1365-2958.2009.06793.x
Chua SL, Hultqvist LD, Yuan M, Rybtke M, Nielsen TE, Givskov M, Tolker-Nielsen T, Yang L (2015) In vitro and in vivo generation and characterization of Pseudomonas aeruginosa biofilm-dispersed cells via c-di-GMP manipulation. Nat Protoc 10(8):1165–1180. https://doi.org/10.1038/nprot.2015.067
An S, Wu J, Zhang L-H (2010) Modulation of Pseudomonas aeruginosa biofilm dispersal by a cyclic-di-GMP phosphodiesterase with a putative hypoxia-sensing domain. Appl Environ Microbiol 76(24):8160–8173. https://doi.org/10.1128/aem.01233-10
Thormann KM, Saville RM, Shukla S, Spormann AM (2005) Induction of rapid detachment in Shewanella oneidensis MR-1 biofilms. J Bacteriol 187(3):1014–1021. https://doi.org/10.1128/jb.187.3.1014-1021.2005
Stacy A, Everett J, Jorth P, Trivedi U, Rumbaugh KP, Whiteley M (2014) Bacterial fight-and-flight responses enhance virulence in a polymicrobial infection. Proc Natl Acad Sci U S A 111(21):7819–7824
Gjermansen M, Ragas P, Sternberg C, Molin S, Tolker-Nielsen T (2005) Characterization of starvation-induced dispersion in Pseudomonas putida biofilms. Environ Microbiol 7(6):894–904. https://doi.org/10.1111/j.1462-2920.2005.00775.x
Delaquis PJ, Caldwell DE, Lawrence JR, McCurdy AR (1989) Detachment of Pseudomonas fluorescens from biofilms on glass surfaces in response to nutrient stress. Microb Ecol 18(3):199–210
Delille A, Quiles F, Humbert F (2007) In situ monitoring of the nascent Pseudomonas fluorescens biofilm response to variations in the dissolved organic carbon level in low-nutrient water by attenuated total reflectance-Fourier transform infrared spectroscopy. Appl Environ Microbiol 73(18):5782–5788
Schleheck D, Barraud N, Klebensberger J, Webb JS, McDougald D, Rice SA, Kjelleberg S (2009) Pseudomonas aeruginosa PAO1 preferentially grows as aggregates in liquid batch cultures and disperses upon starvation. PLoS One 4(5):e5513
Huynh TT, McDougald D, Klebensberger J, Al Qarni B, Barraud N, Rice SA, Kjelleberg S, Schleheck D (2012) Glucose starvation-induced dispersal of Pseudomonas aeruginosa biofilms is cAMP and energy dependent. PLoS One 7(8):e42874. https://doi.org/10.1371/journal.pone.0042874
Petrova OE, Sauer K (2012) PAS domain residues and prosthetic group involved in BdlA-dependent dispersion response by Pseudomonas aeruginosa biofilms. J Bacteriol 194(21):5817–5828
Li Y, Heine S, Entian M, Sauer K, Frankenberg-Dinkel N (2013) NO-induced biofilm dispersion in Pseudomonas aeruginosa is mediated by a MHYT-domain coupled phosphodiesterase. J Bacteriol 195(16):3531–3542. https://doi.org/10.1128/jb.01156-12
Li Y, Petrova OE, Su S, Lau GW, Panmanee W, Na R, Hassett DJ, Davies DG, Sauer K (2014) BdlA, DipA and induced dispersion contribute to acute virulence and chronic persistence of Pseudomonas aeruginosa. PLoS Pathog 10(6):e1004168. https://doi.org/10.1371/journal.ppat.1004168
James GA, Korber DR, Caldwell DE, Costerton JW (1995) Digital image analysis of growth and starvation responses of a surface-colonizing Acinetobacter sp. J Bacteriol 177(4):907–915
Marks LR, Davidson BA, Knight PR, Hakansson AP (2013) Interkingdom signaling induces Streptococcus pneumoniae biofilm dispersion and transition from asymptomatic colonization to disease. MBio 4(4):e00438–e00413
Dimpy Kaliaa GM, Nakayamaa S, Zhenga Y, Zhoua J, Luoa Y, Guoa M, Roembkea BT, Sintim HO (2013) Nucleotide, c-di-GMP, c-di-AMP, cGMP, cAMP, (p)ppGpp signaling in bacteria and implications in pathogenesis. Chem Soc Rev 42:305–341
Carlson HK, Vance RE, Marletta MA (2010) H-NOX regulation of c-di-GMP metabolism and biofilm formation in Legionella pneumophila. Mol Microbiol 77(4):930–942. https://doi.org/10.1111/j.1365-2958.2010.07259.x
Liu N, Xu Y, Hossain S, Huang N, Coursolle D, Gralnick JA, Boon EM (2012) Nitric oxide regulation of cyclic di-GMP synthesis and hydrolysis in Shewanella woodyi. Biochemistry 51(10):2087–2099. https://doi.org/10.1021/bi201753f
Schmidt I, Steenbakkers PJ, op den Camp HJ, Schmidt K, Jetten MS (2004) Physiologic and proteomic evidence for a role of nitric oxide in biofilm formation by Nitrosomonas europaea and other ammonia oxidizers. J Bacteriol 186(9):2781–2788
Dow JM, Crossman L, Findlay K, He Y-Q, Feng J-X, Tang J-L (2003) Biofilm dispersal in Xanthomonas campestris is controlled by cell-cell signaling and is required for full virulence to plants. Proc Natl Acad Sci U S A 100(19):10995–11000. https://doi.org/10.1073/pnas.1833360100
Dean SN, Chung M-C, van Hoek ML (2015) Burkholderia diffusible signal factor signals to Francisella novicida to disperse biofilm and increase siderophore production. Appl Environ Microbiol 81(20):7057–7066
Musk DJ, Banko DA, Hergenrother PJ (2005) Iron salts perturb biofilm formation and disrupt existing biofilms of Pseudomonas aeruginosa. Chem Biol 12(7):789–796
Lanter BB, Sauer K, Davies DG (2014) Bacteria present in carotid arterial plaques are found as biofilm deposits which may contribute to enhanced risk of plaque rupture. mBio 5(3):01206–01214. https://doi.org/10.1128/mBio.01206-14
Hay AJ, Zhu J (2015) Host intestinal signal-promoted biofilm dispersal induces Vibrio cholerae colonization. Infect Immun 83(1):317–323
Amari DT, Marques CNH, Davies DG (2013) The putative enoyl-coenzyme a hydratase DspI is required for production of the Pseudomonas aeruginosa biofilm dispersion autoinducer cis-2-decenoic acid. J Bacteriol 195(20):4600–4610. https://doi.org/10.1128/jb.00707-13
Rahmani-Badi A, Sepehr S, Fallahi H, Heidari-Keshel S (2015) Dissection of the cis-2-decenoic acid signaling network in Pseudomonas aeruginosa using microarray technique. Front Microbiol 6:383. https://doi.org/10.3389/fmicb.2015.00383
Ryan RP, Fouhy Y, Lucey JF, Crossman LC, Spiro S, He Y-W, Zhang L-H, Heeb S, Camara M, Williams P, Dow JM (2006) Cell-cell signaling in Xanthomonas campestris involves an HD-GYP domain protein that functions in cyclic di-GMP turnover. Proc Natl Acad Sci U S A 103(17):6712–6717. https://doi.org/10.1073/pnas.0600345103
Ryan RP, McCarthy Y, Andrade M, Farah CS, Armitage JP, Dow JM (2010) Cell–cell signal-dependent dynamic interactions between HD-GYP and GGDEF domain proteins mediate virulence in Xanthomonas campestris. Proc Natl Acad Sci U S A 107(13):5989–5994. https://doi.org/10.1073/pnas.0912839107
Chin K-H, Lee Y-C, Tu Z-L, Chen C-H, Tseng Y-H, Yang J-M, Ryan RP, McCarthy Y, Dow JM, Wang AHJ, Chou S-H (2010) The cAMP receptor-like protein CLP is a novel c-di-GMP receptor linking cell–cell signaling to virulence gene expression in Xanthomonas campestris. J Mol Biol 396(3):646–662. https://doi.org/10.1016/j.jmb.2009.11.076
Webb JS, Thompson LS, James S, Charlton T, Tolker-Nielsen T, Koch B, Givskov M, Kjelleberg S (2003) Cell death in Pseudomonas aeruginosa biofilm development. J Bacteriol 185(15):4585–4592. https://doi.org/10.1128/jb.185.15.4585-4592.2003
Barraud N, Hassett DJ, Hwang S-H, Rice SA, Kjelleberg S, Webb JS (2006) Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J Bacteriol 188(21):7344–7353. https://doi.org/10.1128/jb.00779-06
Cutruzzolà F, Frankenberg-Dinkel N (2016) Origin and impact of nitric oxide in Pseudomonas aeruginosa biofilms. J Bacteriol 198(1):55–65. https://doi.org/10.1128/jb.00371-15
Galperin MY, Gaidenko TA, Mulkidjanian AY, Nakano M, Price CW (2001) MHYT, a new integral membrane sensor domain. FEMS Microbiol Lett 205(1):17–23
Chambers JR, Cherny KE, Sauer K (2017) Susceptibility of Pseudomonas aeruginosa dispersed cells to antimicrobial agents is dependent on the dispersion cue and class of the antimicrobial agent used. Antimicrob Agents Chemother 61(12):e00846–e00817. https://doi.org/10.1128/aac.00846-17
Chua SL, Liu Y, Yam JKH, Chen Y, Vejborg RM, Tan BGC, Kjelleberg S, Tolker-Nielsen T, Givskov M, Yang L (2014) Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyle. Nat Commun 5:4462. https://doi.org/10.1038/ncomms5462
Chua SL, Tan SY-Y, Rybtke MT, Chen Y, Rice SA, Kjelleberg S, Tolker-Nielsen T, Yang L, Givskov M (2013) Bis-(3′-5′)-cyclic dimeric GMP regulates antimicrobial peptide resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother 57(5):2066–2075
Fleming D, Rumbaugh K (2018) The consequences of biofilm dispersal on the host. Sci Rep 8(1):10738. https://doi.org/10.1038/s41598-018-29121-2
Flemming H-C (2016) EPS—then and now. Microorganisms 4(4):41
Hinsa SM, Espinosa-Urgel M, Ramos JL, O’Toole GA (2003) Transition from reversible to irreversible attachment during biofilm formation by Pseudomonas fluorescens WCS365 requires an ABC transporter and a large secreted protein. Mol Microbiol 49(4):905–918. https://doi.org/10.1046/j.1365-2958.2003.03615.x
Monds RD, Newell PD, Gross RH, O’Toole GA (2007) Phosphate-dependent modulation of c-di-GMP levels regulates Pseudomonas fluorescens Pf0-1 biofilm formation by controlling secretion of the adhesin LapA. Mol Microbiol 63(3):656–679. https://doi.org/10.1111/j.1365-2958.2006.05539.x
Rybtke M, Berthelsen J, Yang L, Høiby N, Givskov M, Tolker-Nielsen T (2015) The LapG protein plays a role in Pseudomonas aeruginosa biofilm formation by controlling the presence of the CdrA adhesin on the cell surface. Microbiology 4(6):917–930
Borlee BR, Goldman AD, Murakami K, Samudrala R, Wozniak DJ, Parsek MR (2010) Pseudomonas aeruginosa uses a cyclic-di-GMP-regulated adhesin to reinforce the biofilm extracellular matrix. Mol Microbiol 75(4):827–842. https://doi.org/10.1111/j.1365-2958.2009.06991.x
Kaplan JB (2010) Biofilm dispersal: mechanisms, clinical implications, and potential therapeutic uses. J Dent Res 89(3):205–218. https://doi.org/10.1177/0022034509359403
Kaplan JB, Ragunath C, Ramasubbu N, Fine DH (2003) Detachment of Actinobacillus actinomycetemcomitans biofilm cells by an endogenous β-hexosaminidase activity. J Bacteriol 185(16):4693–4698
Yu S, Su T, Wu H, Liu S, Wang D, Zhao T, Jin Z, Du W, Zhu M-J, Chua SL, Yang L, Zhu D, Gu L, Ma LZ (2015) PslG, a self-produced glycosyl hydrolase, triggers biofilm disassembly by disrupting exopolysaccharide matrix. Cell Res 25(12):1352–1367. https://doi.org/10.1038/cr.2015.129
Fleming D, Rumbaugh K (2017) Approaches to dispersing medical biofilms. Microorganisms 5(2):15
Gupta K, Liao J, Petrova OE, Cherny KE, Sauer K (2014) Elevated levels of the second messenger c-di-GMP contribute to antimicrobial resistance of Pseudomonas aeruginosa. Mol Microbiol 92(3):488–506. https://doi.org/10.1111/mmi.12587
Petrova OE, Gupta K, Liao J, Goodwine JS, Sauer K (2017) Divide and conquer: the Pseudomonas aeruginosa two-component hybrid SagS enables biofilm formation and recalcitrance of biofilm cells to antimicrobial agents via distinct regulatory circuits. Environ Microbiol 19(5):2005–2024. https://doi.org/10.1111/1462-2920.13719
Chambers JR, Liao J, Schurr MJ, Sauer K (2014) BrlR from Pseudomonas aeruginosa is a c-di-GMP-responsive transcription factor. Mol Microbiol 92(3):471–487. https://doi.org/10.1111/mmi.12562
Liao J, Schurr MJ, Sauer K (2013) The MerR-like regulator BrlR confers biofilm tolerance by activating multidrug-efflux pumps in Pseudomonas aeruginosa biofilms. J Bacteriol 195:3352–3363
Liao J, Sauer K (2012) The MerR-like transcriptional regulator BrlR contributes to Pseudomonas aeruginosa biofilm tolerance. J Bacteriol 194(18):4823–4836. https://doi.org/10.1128/jb.00765-12
Poudyal B, Sauer K (2018) The ABC of biofilm drug tolerance: the MerR-like regulator BrlR is an activator of ABC transport systems, with PA1874-77 contributing to the tolerance of Pseudomonas aeruginosa biofilms to tobramycin. Antimicrob Agents Chemother 62(2):e01981–e01917. https://doi.org/10.1128/aac.01981-17
Jenal U, Reinders A, Lori C (2017) Cyclic di-GMP: second messenger extraordinaire. Nat Rev Microbiol 15:271. https://doi.org/10.1038/nrmicro.2016.190
Jones CJ, Newsom D, Kelly B, Irie Y, Jennings LK, Xu B, Limoli DH, Harrison JJ, Parsek MR, White P, Wozniak DJ (2014) ChIP-Seq and RNA-Seq reveal an AmrZ-mediated mechanism for cyclic di-GMP synthesis and biofilm development by Pseudomonas aeruginosa. PLoS Pathog 10(3):e1003984. https://doi.org/10.1371/journal.ppat.1003984
Hickman JW, Harwood CS (2008) Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol Microbiol 69(2):376–389. https://doi.org/10.1111/j.1365-2958.2008.06281.x
Baraquet C, Murakami K, Parsek MR, Harwood CS (2012) The FleQ protein from Pseudomonas aeruginosa functions as both a repressor and an activator to control gene expression from the pel operon promoter in response to c-di-GMP. Nucleic Acids Res 40(15):7207–7218. https://doi.org/10.1093/nar/gks384
Baraquet C, Harwood CS (2013) Cyclic diguanosine monophosphate represses bacterial flagella synthesis by interacting with the Walker a motif of the enhancer-binding protein FleQ. Proc Natl Acad Sci U S A 110(46):18478–18483. https://doi.org/10.1073/pnas.1318972110
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Sauer, K. (2020). Cyclic di-GMP and the Regulation of Biofilm Dispersion. In: Chou, SH., Guiliani, N., Lee, V., Römling, U. (eds) Microbial Cyclic Di-Nucleotide Signaling. Springer, Cham. https://doi.org/10.1007/978-3-030-33308-9_31
Download citation
DOI: https://doi.org/10.1007/978-3-030-33308-9_31
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-33307-2
Online ISBN: 978-3-030-33308-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)