Abstract
The Notch signal transduction cascade requires cell-to-cell contact and results in the proteolytic processing of the Notch receptor and subsequent assembly of a transcriptional coactivator complex containing the Notch intracellular domain (NICD) and transcription factor RBPJ. In the absence of a Notch signal, RBPJ remains at Notch target genes and dampens transcriptional output. Like in other signaling pathways, RBPJ is able to switch from activation to repression by associating with corepressor complexes containing several chromatin-modifying enzymes. Here, we focus on the recent advances concerning RBPJ-corepressor functions, especially in regard to chromatin regulation. We put this into the context of one of the best-studied model systems for Notch, blood cell development. Alterations in the RBPJ-corepressor functions can contribute to the development of leukemia, especially in the case of acute myeloid leukemia (AML). The versatile role of transcription factor RBPJ in regulating pivotal target genes like c-MYC and HES1 may contribute to the better understanding of the development of leukemia.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Notch signaling is an evolutionary highly conserved pathway that plays a pivotal role in many cellular and developmental processes including T-cell development (Vijayaraghavan and Osborne 2018) and angiogenesis (Pitulescu et al. 2017; Tetzlaff and Fischer 2018). Although Notch was originally described as a neurogenic gene in Drosophila melanogaster, the first analysis of Drosophila embryos made it clear that Notch signals are pleiotropic, affecting many tissues. After the cloning and sequencing of the Notch gene in the 1980s, it became clear that Notch is a single-pass transmembrane receptor. Subsequently, the NOTCH1 gene was described to be a hotspot for chromosomal translocations in human T-cell acute lymphoblastic leukemia (T-ALL) (Ellisen et al. 1991). By now, we know that NOTCH1 mutations are found not only in human T-ALL (Weng et al. 2004) but also in other forms of human leukemia, for example, chronic lymphocytic leukemia (CLL) (Puente et al. 2011) as well as many other cancer types (Giaimo and Borggrefe 2018).
At the molecular level, Notch signal transduction bears some unique features not seen in other pathways like TGFβ, Wnt, or Hedgehog signaling [also reviewed in Borggrefe et al. 2016]. For example, the Notch pathway does not involve any second messengers. Notch signaling occurs through direct interactions between the Notch receptor and its ligand exposed on neighboring cells (Fig. 2.1). Upon ligand binding, the extracellular protease cleavage site of the receptor is exposed and cleaved by ADAM (a disintegrin and metalloproteinase) proteases. Subsequently, a second cleavage of the receptor is mediated by a γ-secretase-containing complex leading to the release of the Notch intracellular domain (NICD), which is itself a transcriptional coactivator (Fig. 2.1). The NICD migrates into the nucleus and functions as a transcriptional coactivator together with RBPJ and mastermind (MAM) [reviewed in Oswald and Kovall 2018]. The transcription factor RBPJ is a central molecular switch in the Notch pathway and mediates either transcriptional repression or activation of Notch target genes (Fig. 2.1).
The Notch signaling cascade. In absence of Notch signaling, the DNA-binding protein RBPJ is bound at the RBPJ-binding sites (RBS) where it recruits corepressors (CoR) preventing the expression of Notch target genes. The binding of ligands to Notch receptors induces a conformational change that allows their proteolytic cleavage by ADAM proteases producing an intermediate product known as NotchΔE. Subsequently, a γ-secretase-containing complex catalyzes a second cleavage of the Notch receptor releasing the Notch intracellular domain (NICD). The free NICD moves into the nucleus where it interacts with RBPJ and forms a trimeric complex together with Mastermind-like (MAML). This trimeric complex recruits additional coactivators (CoA) finally promoting expression of Notch target genes. Finally, proteasome-dependent degradation of the NICD terminates the signal, and the RBPJ-associated corepressor complex is reassembled at the RBS inducing repression of Notch target genes. Green and red balls indicate positive and negative histone marks, respectively
Transcription Factor RBPJ in Balancing Notch Target Gene Expression
Historically, RBPJ was discovered thirty years ago and was originally named RBPJκ [recombination signal binding protein for immunoglobulin kappa J region, (Hamaguchi et al. 1989)]. It also has different names, such as CBF1 (C promoter binding factor 1) or KBF2 [H-2K binding factor-2, (Brou et al. 1994)] and belongs to the CSL (Homo sapiens CBF1, Drosophila melanogaster Suppressor of Hairless and Caenorhabditis elegans Lag-1) protein family. The DNA binding sequence was identified as 5′-CGTGGGAA-3′ (Tun et al. 1994) and recent studies investigated the genome-wide distribution of RBPJ in several different tissues (Dieguez-Hurtado et al. 2019; Petrovic et al. 2019; Wang et al. 2011; Xie et al. 2016; Zhao et al. 2011). RBPJ shares some structural similarities with Rel Homology Domain proteins such as NF-κB1 (nuclear factor kappa B subunit 1) and NFAT [nuclear factor of activated T-cells, (Kovall and Hendrickson 2004)]. It is the centerpiece of transcriptional regulation in Notch signaling, acting as a molecular hub for interactions of either corepressor or coactivators. In the absence of a Notch signal, RBPJ interacts with the cofactor SHARP recruiting histone deacetylase-containing corepressor complexes. In the presence of a Notch signal, a ternary complex containing RBPJ, NICD and MAM-like (MAML) is assembled and expression of Notch target genes is induced (Fig. 2.1). The RBPJ/NICD/MAML-containing coactivator complex also recruits lysine acetyltransferases (KATs) such as KAT3B/Ep300 (lysine acetyltransferase 3B/E1A binding protein P300), KAT2B/PCAF (lysine acetyltransferase 2B/Ep300-CBP-associated factors) and KAT2A/GCN5 [lysine acetyltransferase 2A/ general control of amino acid synthesis protein 5-like 2, (Kurooka and Honjo 2000; Oswald et al. 2001)]. Interestingly, RBPJ was initially described as a repressor of transcription and its role as a molecular switch was further underscored by the finding that repression and activation via RBPJ involves the recruitment of distinct protein complexes [reviewed in (Borggrefe and Oswald 2009]. From these studies a model emerged (Fig. 2.1) stating that presence of NICD converts the RBPJ-corepressor to the RBPJ-NICD-coactivator complex (Borggrefe and Oswald 2009; Bray 2006).
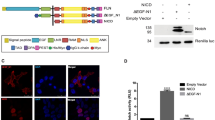
In the recent years, the RBPJ interactome has been extensively studied [(Borggrefe and Liefke 2012; Guruharsha et al. 2014; Ho et al. 2018; Yatim et al. 2012) and Table 2.1] in order to understand at the molecular level how gene repression and activation are regulated. As part of the corepressor complex, RBPJ can directly interact with corepressor SHARP [SMRT (silencing mediator for retinoid and thyroid receptor) and HDACs (histone deacetylases)-associated repressor protein], also known as mouse MINT (MSX2-interacting protein) or Spen [split ends (Oswald et al. 2002)]. Recently, we determined the binding surfaces of the RBPJ/SHARP interaction at atomic resolution using X-ray crystallography [Fig. 2.2 (Yuan et al. 2019)]. Based on the RBPJ/SHARP structure, we could design a dominant-negative form of SHARP in a Notch-OFF state (Giaimo et al. 2017a; Xu et al. 2017; Yuan et al. 2019). When overexpressing the wild-type (WT) form of the RBPJ-interacting domain (RBPID) of SHARP, derepression of Notch target genes was observed; however, this was not the case when, based on the crystal structure, we mutated two amino acids within this domain (Yuan et al. 2019). Previous studies linked the repressive activity of SHARP to HDACs (Oswald et al. 2002; Oswald et al. 2016), and, in line with these studies, we observed increased histone acetylation upon overexpression of the WT but not the mutant RBPID (Yuan et al. 2019). Importantly, RBPJ depletion leads to derepression of Notch target genes in the same setting. This phenotype is efficiently rescued by a WT RBPJ but not a mutant in which the residues required for its interaction with SHARP are mutated (Yuan et al. 2019).
Overview on the known crystal structures of RBPJ-associated complexes. (a) The structure of the Caenorhabditis elegans RBPJ/NICD/MAML ternary activation complex (PDBID: 2FO1). RBPJ, shown in cyan with a transparent white surface, consists of three major domains. The N-terminal domain (NTD) makes direct contacts primarily with MAML (red) and DNA (blue). The beta-trefoil domain (BTD) interacts with DNA and NICD (yellow). The C-terminal domain (CTD) interacts with MAML and NICD. (b) The structure of the Mus musculus RBPJ/SHARP repressor complex (PDBID: 6DKS). Like NICD, SHARP (orange) also binds the CTD and BTD of RBPJ. (c) Top: Structural representation of multiple corepressors that bind the BTD of RBPJ similarly. NICD (yellow) PDBID, 3 V79; SHARP (orange) PDBID, 6DKS; KyoT2/FHL1C (pink) PDBID, 4J2X; and RITA (light green) PDBID, 5EG6. Bottom: Multiple sequence alignments of coregulators that bind the BTD of RBPJ. Boxed in red is the highly conserved hydrophobic tetrapeptide seen in all four mammalian Notch isoforms as well as some corepressors. The blue boxes represent other highly conserved hydrophobic residues seen in multiple corepressors
SHARP is a protein of more than 400 kDa characterized by a highly conserved SPOC (Spen paralog and ortholog C-terminal) domain which has a strong transcriptional repressive activity that depends on CtIP/CtBP (CtBP-interacting protein/C-terminal-binding protein) (Oswald et al. 2005). To better dissect the mechanism of the RBPJ/SHARP-mediated transcriptional repression, we have recently characterized, by mass spectrometry (MS), the SPOC interactome (SPOCome) (Oswald et al. 2016). This approach identified the HDACs-containing NCoR (nuclear receptor corepressor) complex, explaining how HDACs are recruited to RBPJ-bound enhancer sites; however, it also identified the KMT2D (lysine-specific methyltransferase 2D) complex (Oswald et al. 2016). This finding was quite unexpected as the KMT2D complex is involved in transcriptional activation in contrast to the repressive activity of the SPOC domain of SHARP. Previous structural studies of SMRT, the ortholog of NCoR, in complex with the SPOC domain of SHARP unveiled that this interaction depends on the phosphorylation of highly conserved serine residues of SMRT by casein kinase II (CKII) (Mikami et al. 2013; Mikami et al. 2014). Of note, NCoR is also phosphorylated on serine residues at its C-terminus (Yoo et al. 2012, 2013), and we found that NCoR phosphorylation is required for its interaction with the SPOC domain and dependent on CKII [(Oswald et al. 2016) and Fig. 2.3]. KMT2D and NCoR are in competition for binding to SPOC, and phospho-NCoR displaces KMT2D leading to transcriptional repression [(Oswald et al. 2016) and Fig. 2.3]. These data support the hypothesis that SHARP, integrating different stimuli, acts as a poising factor for Notch target genes, balancing repressive and activating histone marks [(Giaimo et al. 2017b; Oswald et al. 2016) and Fig. 2.3].
Intermediate states involved in the transcriptional regulation of Notch target genes. The different RBPJ-associated corepressor complexes (SHARP, L3MBLT3, and KyoT2) are recruited at the same RBPJ-binding sites (RBS) in a well-defined temporal order and/or tissue-specific manner to promote repression of Notch target genes. Of note, SHARP can interact with the HDAC-containing NCoR corepressor complex when it is phosphorylated on two serine residues. NCoR recruitment moves the balance toward repression alternatively; NCoR interacts with the KMT2D-containing complex moving the balance toward gene activation. L3MBTL3 bridges the histone demethylase KDM1A/LSD1 (indicated as KDM1A) to RBPJ at RBS
The SPOC domain of SHARP directly interacts with ETO [eight-twenty-one, also known as MTG8 (myeloid translocation gene on 8q22)] which acts as a corepressor of Notch target genes by meaning of deacetylation (Salat et al. 2008). ETO is a member of the MTG family of corepressors of transcription which includes also MTGR1 (myeloid translocation gene-related protein 1) and MTG16 (myeloid translocation gene on chromosome 16 protein). Both MTGR1 and MTG16 interact with RBPJ; however, only MTG16 is displaced from RBPJ by the NOTCH1 intracellular domain 1 (NICD1) (Engel et al. 2010). In conclusion, the SPOC domain of SHARP is able to interact with several different proteins. It remains to be investigated how the varying interaction partners are recruited to enhancers. Our favorite working hypothesis is that posttranslational modifications (PTMs) such as phosphorylation determine specificity in terms of composition and strengths of recruitment of corepressors. In that regard, it is appealing that the highly conserved SPOC domain interacts with double-phosphorylated peptide (Oswald et al. 2016). This might be also the case for other SPOC-interaction partners. Interestingly, RBM15 [RNA-binding motif protein 15, also known as OTT (one twenty-two)], another SPOC domain-containing protein, was shown to modulate the Notch signaling pathway in a cell-type-specific fashion (Ma et al. 2007), marking the importance of the SPOC domain-containing proteins in the regulation of the Notch signaling pathway.
Another direct interactor of RBPJ is KyoT2 [also known as FHL1C (four-and-a-half LIM domain protein 1C) in human] which, competing with the NICD1, represses transcription of target genes (Collins et al. 2014; Taniguchi et al. 1998). The structure of RBPJ in complex with KyoT2/FHL1C reveals a good overlap with the RBPJ/SHARP and RBPJ/Notch binding surfaces [Fig. 2.2 (Collins et al. 2014; Yuan et al. 2019)]. One more isoform, KyoT3/FHL1B, is also able to interact with RBPJ and to promote gene repression (Liang et al. 2008), but this is not the case for the isoform KyoT1/FHL1A (Taniguchi et al. 1998). It remains to elucidate whether the repressive mechanism of KyoT2/FHL1C and KyoT3/FHL1B is exclusively based on their competition with the NICD or whether it depends on other cofactors, for example, a link between KyoT2/FHL1C and Polycomb has been proposed (Quin et al., PMID, 14999091, and Quin et al., 15,710,417).
As part of the corepressor complex, RBPJ recruits histone demethylase activities such as KDM5A/LID [lysine demethylase 5A/little imaginal discs (Di Stefano et al. 2011; Liefke et al. 2010; Moshkin et al. 2009)] and KDM1A/LSD1 [lysine demethylase 1A/lysine-specific demethylase 1 (Di Stefano et al. 2011; Mulligan et al. 2011; Xu et al. 2017; Yatim et al. 2012)]. KDM5A/LID directly interacts with RBPJ and demethylates H3K4me3 at RBPJ-bound enhancer sites promoting repression of Notch target genes (Liefke et al. 2010), while KDM1A/LSD1 indirectly interacts with RBPJ via L3MBTL3 [lethal (3) malignant brain tumor-like protein 3] (Xu et al. 2017). We identified L3MBTL3 in a screen for RBPJ interactors and observed that the L3MBTL3-RBPJ interaction is conserved in Drosophila melanogaster and Caenorhabditis elegans (Xu et al. 2017). Notably, L3MBTL3 and NICD1 bind to the same binding surface on RBPJ: While NICD1 displaces L3MBTL3 from RBPJ, the latter does not outcompete NICD1 for binding to RBPJ (Xu et al. 2017). The recruitment of KDM1A/LSD1 via L3MBTL3 is required to modulate H3K4 methylation states at RBPJ-bound enhancers promoting repression of target genes (Xu et al. 2017). In line with that, pharmacological inhibition of KDM1A/LSD1 leads to upregulation of Notch target genes (Augert et al. 2019); however it must be marked that, at least in lung cancer cells, KDM1A/LSD1 may indirectly regulate the Notch signaling pathway via a direct regulation of the expression of NOTCH1 (Augert et al. 2019). A previous study linked KDM1A/LSD1 to repression of Notch target genes, but the authors also observed that KDM1A/LSD1 associates with the NOTCH1 coactivator complex to modulate H3K9 methylation states and finally promoting expression of Notch target genes (Yatim et al. 2012). Altogether, these data suggest that KDM1A/LSD1 acts as both an activator and a repressor of the Notch-dependent gene expression program. Finally, the demethylase KDM7B/PHF8 (lysine demethylase 7B/PHD finger protein 8) is also part of the NOTCH1 coactivator complex and supports expression of Notch targets by modulating H3K27 methylation states (Yatim et al. 2012).
Based on the available structural and biophysical data, SHARP, KyoT2/FHL1C, and L3MBTL3 interact in a mutually exclusive fashion with RBPJ: Different intermediate complexes may be dynamically recruited at the same enhancer in a defined temporal order or in a tissue-specific manner to modulate the chromatin structure leading to gene repression (Fig. 2.3). Since the RBPJ-associated cofactors interactions are strong and the DNA-binding affinity of RBPJ is relatively weak, it can be assumed that the different RBPJ complexes are constantly exchanging, explaining how the different cofactors are recruited at a defined enhancer.
The activation of the Notch pathway leads to the release of the NICD from the cell membrane which, upon nuclear translocation, converts RBPJ from a repressor to an activator of transcription via the recruitment of additional coactivators (Fig. 2.1). One of the most important members of the coactivator complex is MAM which, together with RBPJ and NICD, forms a trimeric complex indispensable for activation of Notch target genes (Friedmann et al. 2008; Fryer et al. 2002; Kitagawa et al. 2001; Nam et al. 2006; Wilson and Kovall 2006; Wu et al. 2000, 2002). The human Mastermind family (Mastermind-like or MAML) consists of three members, all of them able to support Notch-dependent transcription (Lin et al. 2002). Probably, the most important function of MAML is to recruit the histone acetyltransferase (HAT) KAT3B/Ep300 to Notch target genes that supports gene expression via histone acetylation (Fryer et al. 2002; Jung et al. 2013; Oswald et al. 2001; Tottone et al. 2019; Wallberg et al. 2002). Additionally, KAT3B/Ep300 acetylates MAML leading to the recruitment of the coactivator NACK (Notch activation complex kinase) at Notch target genes (Jin et al. 2017; Weaver et al. 2014). MAML also recruits the cyclin C/CDK8 (Cyclin-dependent kinase 8) complex that phosphorylates the NICD leading to its proteasome-dependent degradation (Fryer et al. 2004). Another component of the Notch coactivator complex is the RNA helicase DDX5 (DEAD-box helicase 5) which, interacting with the long noncoding RNA SRA (steroid receptor coactivator), supports the recruitment of KAT3B/Ep300 and subsequent activation of Notch target genes (Jung et al. 2013; Lin et al. 2013). Furthermore, CARM1/PRMT4 (coactivator-associated arginine methyltransferase 1/protein arginine methyltransferase 4) promotes the activation of Notch target genes by arginine methylation of the NICD1 itself (Hein et al. 2015).
Recently, we characterized the interactome of the cleaved, active NICD1 in mouse progenitor cells (Giaimo et al. 2018). Using this approach, we identified the Ep400-KAT5/Tip60 (E1A-binding protein P400-lysine acetyltransferase 5/HIV-1 Tat-interactive protein, 60 kDa) complex (hereafter referred to as Ep400/Tip60 complex) as an NICD1 interactor. This complex attracted our attention as its subunits Ep400 and KAT5/Tip60 have been previously linked to deposition (Gevry et al. 2007) and acetylation (Kusch et al. 2004) of the histone variant H2A.Z, respectively. H2A.Z has been linked to several processes including heterochromatin regulation, DNA repair, and gene transcription both in a positive and negative fashion (Giaimo et al. 2019). We found that H2A.Z depletion leads to upregulation of Notch target genes, and this enhanced expression is associated with increased active marks, namely, H3K4me2 and H3K27ac, at Notch-dependent enhancer elements. These data suggest H2A.Z as a negative regulator of Notch target genes, and this conclusion is further supported by the observation that activation of Notch signaling leads to decreased H2A.Z occupancy at Notch-dependent enhancers (Giaimo et al. 2018). However, while H2A.Z occupancy negatively correlates with induction of Notch target genes, acetylation of H2A.Z (H2A.Zac) does it in a positive manner suggesting that H2A.Z is involved in both gene repression and activation and the difference between the two functions is obtained via its acetylation. Overexpression of H2A.Z leads to upregulation of Notch target gene Hairy and Enhancer of Split 1 (Hes1), while this upregulation is more modest when an acetylation-defective H2A.Z mutant is overexpressed (Giaimo et al. 2018). Our data further indicate that acetylation of H2A.Z is highly dynamic, which also reconciles previous contrasting results showing H2A.Z as a repressor or an activator of transcription (Gevry et al. 2007, 2009; Giaimo et al. 2019). We observed that Ep400 interacts with RBPJ and it is recruited to Notch-dependent enhancers in a Notch-dependent fashion (Giaimo et al. 2018). Furthermore, making use of a tethering approach, we could show that Tip60 promotes H2A.Zac supporting gene expression (Giaimo et al. 2018). In summary, our data suggest that in a Notch-OFF or poised state, the Ep400/Tip60 complex is recruited to Notch-dependent enhancer sites via an unstable interaction with RBPJ promoting loading of H2A.Z. Upon Notch activation, the interaction between the Ep400/Tip60 complex and RBPJ is stabilized via additional interactions with the NICD1 protein promoting acetylation of H2A.Z and finally gene expression (Fig. 2.4).
Model for the regulation of Notch target gene by the Ep400/Tip60 complex and histone variant H2A.Z. In the repressed (OFF) or poised state, RBPJ directly interacts with Ep400 leading to the recruitment of the Ep400/Tip60 complex. This interaction is unstable but sufficient to promote deposition of the histone variant H2A.Z. Upon activation of the Notch pathway (ON), the Notch intracellular domain (NICD) directly interacts with RBPJ and Ep400 leading to stabilization of the RBPJ-Ep400 interaction. In turn, this results in acetylation of H2A.Z (indicated with green balls) and finally gene activation
The classical model for the regulation of Notch target genes suggests that RBPJ is persistently bound to its cognate sequences promoting gene repression or activation based on the stimulation of the NOTCH receptor. However, recent studies challenged this model and suggest that RBPJ is weakly bound to its enhancers in absence of stimulus, while its genomic occupancy significantly increases upon activation of the Notch pathway and landing of the NICD at target enhancers (Fig. 2.5). Earlier studies in Drosophila melanogaster cell lines observed increased occupancy of Su(H) (Suppressor of Hairless), the Drosophila homolog of RBPJ, upon induction of Notch signaling (Krejci and Bray 2007). Recently, single-molecule tracking in vivo studies allowed to define that Su(H) transiently binds the DNA in the OFF state (Gomez-Lamarca et al. 2018). Upon Notch activation, the DNA binding of Su(H) significantly increases (Gomez-Lamarca et al. 2018).
New model for regulation of Notch target genes. (a) The classic model for regulation of Notch target genes is based on the binding of RBPJ at its cognate RBPJ-binding sites (RBS) in absence of Notch signaling. In this scenario, RBPJ recruits corepressors (CoR) preventing expression of target genes. (b) New data suggest that RBPJ transiently binds to the RBS in absence of Notch signaling promoting gene repression. Upon Notch activation, the cleaved Notch intracellular domain (NICD) interacts with RBPJ leading to increased DNA binding of RBPJ at the RBS. This event leads finally to activation of Notch target genes
Similarly to Drosophila, activation of the Notch pathway leads to increased RBPJ occupancy in mammalian cell lines (Castel et al. 2013; Wang et al. 2014; Yashiro-Ohtani et al. 2014); however, Castel and colleagues identified two different classes of RBPJ-binding sites: dynamic sites at which RBPJ is bound only upon Notch activation and static sites at which RBPJ is bound independently of the Notch activation (Castel et al. 2013). To note, NICD binding occurs exclusively at the dynamic but not static sites, and furthermore, Castel and colleagues observed that RBPJ depletion leads to derepression of few genes associated with static sites and about 50% of the genes associated with dynamic sites (Castel et al. 2013). However, this analysis uses different cell lines for ChIP-Seq (C2C12 cell lines) and gene expression analysis (quiescent satellite cells) (Castel et al. 2013). As a consequence, we do not know whether all the derepressed genes assumed to be associated with static or dynamic RBPJ sites are so. The DNA-binding strength of RBPJ does not seem to be regulated exclusively by the NICD, for example, a recent study proposed that HDAC1 and KDM5A/LID play a negative and positive role, respectively, in this process, at least in mitosis (Dreval et al. 2019). Similarly, the BRM complex promotes Su(H) binding in Drosophila (Pillidge and Bray 2019). The above findings implicate dynamic binding of RBPJ depending on dynamic coactivator binding.
Promoter specificity might also be achieved by the usage of mono- versus dimeric RBPJ-bound enhancers. In fact, several RBPJ enhancers are characterized by two correctly spaced and oriented binding motifs at which a dimeric NICD1/RBPJ/MAML complex is recruited (Hass et al. 2015; Nam et al. 2007; Severson et al. 2017). The structure of the dimeric NICD1/RBPJ/MAML complex has been solved (Arnett et al. 2010); however, we do not know the impact of PTMs of the NICD on the dimeric NICD1/RBPJ/MAML complex formation (Borggrefe et al. 2016).
The DNA binding of RBPJ is also dependent on other transcription factors (TFs). This is, for example, marked by the loss of function (LoF) of Lozenge (Lz), the Drosophila homolog of Runx (Runt-related transcription) factors, which results in reduced SuH occupancy (Terriente-Felix et al. 2013). Similarly, Lz overexpression leads to increased SuH occupancy and increased response to Notch activation (Skalska et al. 2015). In line with that, the Runx DNA-binding motif is found near the RBPJ-binding sites (Wang et al. 2011), and RBPJ and RUNX1 [(Runt-related transcription factor 1) also known as AML1 (acute myeloid leukemia 1)] colocalize genome-wide (Wang et al. 2014). In Drosophila, the basic helix-loop-helix (bHLH) TFs Twist and Dorsal prime RBPJ-dependent enhancers leading to synchronized and sustained enhancer activity (Falo-Sanjuan et al. 2019), marking the importance of tissue-specific TFs in the Notch response. Additionally, RBPJ colocalizes and interacts with the DNA-binding protein IKAROS which is required for repression of Notch target genes (Geimer Le Lay et al. 2014). However, the exact relationship between RBPJ and IKAROS is not clear, and we do not know whether their interaction is required to support the DNA binding of RBPJ and vice versa. This can be addressed by performing depletion of IKAROS followed by ChIP versus RBPJ and the other way round.
Notch Signaling in Hematopoiesis
The essential role for Notch signaling in inducing T-cell development has been extensively investigated [reviewed in Vijayaraghavan and Osborne 2018]. Inducible depletion of the Notch1 gene in bone marrow precursors results in a block of T-cell development associated with ectopic appearance of donor progenitor-derived B220+ immature B-cells in the thymus (Radtke et al. 1999; Wilson et al. 2001). Accordingly, RBPJ conditional knockout mice are characterized by a block in T-cell development associated with appearance of B-cells in the thymus (Han et al. 2002). Similar results were obtained by gain of function (GoF) of the Notch target gene Deltex1 (Izon et al. 2002) or by overexpression of a dominant-negative MAML1 (dnMAML1) (Maillard et al. 2004). These data suggest Notch signaling as the driver of the T-cell differentiation program, and in line with that, retroviral expression of the NICD in murine hematopoietic precursors, followed by transplantation into recipient mice, leads to an abnormal appearance of immature double-positive T-cells in the bone marrow and subsequent development of T-cell leukemia, while B-cell development is blocked (Pear et al. 1996; Pui et al. 1999). Furthermore, MTG16 knockout results in defects in T-cell differentiation both in mice and using MTG16−/− hematopoietic progenitors (Engel et al. 2010; Hunt et al. 2011).
The critical role of Notch signaling in activating the T-cell lineage differentiation program is also marked by the observation that the engagement of NOTCH receptors by DELTA-LIKE 1 (DLL1) ligand expressed on the surface of OP9 stromal cells leads to the differentiation of hematopoietic progenitor cells (HPCs) and embryonic stem cells (ESCs) into T-cells (Schmitt et al. 2004b; Schmitt and Zuniga-Pflucker 2002). The thymus represents a nonpermissive environment for the development of myeloid, natural killer (NK), and B-cells because the thymic epithelium offers the DLL1 and DLL4 ligands to the NOTCH1-expressing progenitor T-cells (Feyerabend et al. 2009; Schmitt et al. 2004a). Of note, Notch blocks the alternative differentiation pathways even if the cells are ectopically forced to express TFs required for differentiating versus other lineages (Franco et al. 2006; Laiosa et al. 2006).
Notch Signaling in Leukemia
Given the key role of Notch signaling in T-cell differentiation, it is not surprising to observe aberrant regulation of the Notch pathways in T-cell leukemias. Mutations of the NOTCH1 gene have been identified in T-ALL patients and cell lines (Breit et al. 2006; Larson Gedman et al. 2009; Mansour et al. 2006; Palomero et al. 2006; Weng et al. 2004). These mutations lead to increased activation of the pathway, and they can be classified into two different groups based on the molecular mechanism: mutations that lead to ligand-independent activation of the pathway and mutations that increase the half-life of the NICD proteins. The former include translocations that fuse the NICD-encoding sequences to another gene or mutations that influence the cleavage of the receptor, for example, mutations of the heterodimerization domain (HD) (Larson Gedman et al. 2009; Malecki et al. 2006; Weng et al. 2004). The latter are mutations that result in C-terminal truncated NICD proteins lacking the PEST (proline, glutamic acid, serine, and threonine) domain which is required for the turnover of the NICD proteins (Larson Gedman et al. 2009; Palomero et al. 2006; Weng et al. 2004). Mutations, although very rare, may also occur in the ankyrin repeats (ANKs) and in the transactivation domain (TAD) of NOTCH1 in T-ALL (Zhu et al. 2006).
Importantly, hyperactivation of the Notch pathway can also be achieved compromising the activity of its negative regulators. In fact, mutations of the NOTCH1 E3-ubiquitin-ligase F-Box and WD repeat domain-containing 7 (FBXW7)-encoding gene have been identified in T-ALL (Larson Gedman et al. 2009). Additionally, upregulation of positive regulators of the Notch pathway can also lead to hyperactivation of the pathway. In line with this, MAML2 is upregulated in B-cell-derived lymphomas (Kochert et al. 2011) and was described fused to the lysine-specific methyltransferase 2A (KMT2A) gene in T-ALL as result of a chromosomal inversion (Metzler et al. 2008). However, the Notch pathway, via HES1, seems to have a tumor-suppressive function in B-cell acute lymphoblastic leukemia [B-ALL (Kannan et al. 2011)].
Recently, activating NOTCH mutations of the PEST domain have been identified also in CLL (Fabbri et al. 2011; Puente et al. 2011). Interestingly, mutations of the 3´-UTR (untranslated region) of the NOTCH1 gene have also been identified in CLL leading to increased activation of the pathway (Puente et al. 2015). Activating NOTCH1 mutations significantly correlates with Richter transformation and chemorefractory CLL, and they have been proposed as predictors of poor survival (Fabbri et al. 2011). In line with a role for Notch signaling in CLL, γ-secretase inhibitor (GSI) treatment of B-CLL cells reduces their survival by meaning of apoptosis (Rosati et al. 2009). Similar conclusions were reached using antibodies directed against NOTCH receptors, and furthermore the same study observed that Notch signaling is involved in drug resistance (Nwabo Kamdje et al. 2012). Of note, EGR2 (early growth response 2) mutations are frequently associated with NOTCH1 or FBXW7 mutations in CLL patients (Young et al. 2017). The observation that active Notch signaling is detectable also in CLL cases that lack NOTCH1 mutations suggests that other mechanisms can be used to activate the pathway in this disease and imply Notch signaling as a more general deregulated pathway associated with CLL (Fabbri et al. 2017).
In mantle cell lymphomas (MCL), activating mutations of the NOTCH1 gene map to the HD- and PEST-encoding regions (Kridel et al. 2012), and recently a genome-wide study identified Notch targets and RBPJ-binding sites in MCL cell lines (Ryan et al. 2017). Similarly, truncating NOTCH2 mutations were detected in MCL and diffuse large B-cell lymphoma (DLBCL) with the latter also characterized by missense NOTCH2 mutations (Bea et al. 2013) and NOTCH1 mutations (Fabbri et al. 2011). Activating NOTCH2 and NOTCH1 mutations were also described in splenic marginal zone lymphomas (SMZL) as well as inactivating SHARP mutations and mutations of other components of the Notch pathway (Rossi et al. 2012).
Mutations of the Notch pathway similar to the leukemia-associated ones have also been identified in solid tumors (Giaimo and Borggrefe 2018) marking the importance to develop new therapies aimed to target the Notch pathway.
Aberrant regulation of the Notch signaling pathway was also linked to the AML characterized by the t(8;21)(q22/q22) translocation that fuses the AML1 (also known as RUNX1) gene to the ETO gene. AML1 encodes for a hematopoietic cell-specific TF which heterodimerizes with a non-DNA-binding protein called CBFβ [core-binding factor β (Ogawa et al. 1993a, b)], and it is essential for definitive hematopoietic development (Okada et al. 1998; Okuda et al. 1996; Wang et al. 1996). AML1 is characterized by an N-terminal Runt homology domain (RHD) which characterizes all members of the RUNX family (RUNX1, RUNX2, and RUNX3) and by a C-terminal TAD. Both DNA binding and heterodimerization with CBFβ are mediated through the RHD, and the function of CBFβ is to increase the stability and the DNA-binding affinity of AML1 (Huang et al. 2001; Tahirov et al. 2001). On the other side, the ETO gene, highly expressed in the brain (Miyoshi et al. 1993) and in hematopoietic cells (Erickson et al. 1996), is the homolog of the Drosophila Nervy in four regions protein (Feinstein et al. 1995); in fact, it encodes for a non-DNA-binding protein characterized by four evolutionarily conserved functional domains called nervy homology regions (NHR): the NHR2 forms an amphipathic helix and it is important for homodimerization (Lutterbach et al. 1998a) and heterodimerization with MTGR1 (Kitabayashi et al. 1998); the NHR4 contains two putative zinc fingers (ZnF) required for interactions with NCoR and SMRT which links ETO to HDACs (Gelmetti et al. 1998; Lutterbach et al. 1998b; Wang et al. 1998). Of note, the function of NHR4 is strongly dependent on NHR2 (Zhang et al. 2001).
The t(8;21)(q22/q22) translocation fuses the DNA encoding the first N-terminal 177 residues of AML1, which include the RHD, in frame with nearly all of ETO (Erickson et al. 1992; Kozu et al. 1993; Miyoshi et al. 1993; Nisson et al. 1992). This translocation leads to deletion of the C-terminal activation domain of AML1, and the resulting AML1/ETO (AE) protein acts as a dominant-negative form of AML1, which binds to AML1-binding sites (Gardini et al. 2008) repressing target genes (Frank et al. 1995; Liu et al. 2007). Of note, A/E expression requires additional mutations to induce leukemia in a murine in vivo model (Yuan et al. 2001), but this is not true for two different C-terminal truncated AE proteins [AML1/ETO 9a (AE9a) and AML1/ETO truncated (AEtr)] which are potent inducers of leukemia in mice (Yan et al. 2009; Yan et al. 2004; Yan et al. 2006). Interestingly, AE9a is deleted of NHR3 and NHR4, arguing against the well-accepted model that AE acts exclusively as a repressor of AML1 target genes (Heibert et al. 2001).
The first evidence about an aberrant regulation of the Notch signaling pathway in AML came out with the observation that overexpression of AE leads to upregulation of the Notch target gene Hes1 (Alcalay et al. 2003). The underlying mechanism was unveiled when ETO was identified as a component of the RBPJ/SHARP corepressor complex (Salat et al. 2008). In detail, ETO and, surprisingly, AE directly interact with SHARP, but while ETO is able to augment SHARP-mediated repression, this is not the case for AE. Furthermore, knockdown of ETO or overexpression of AE resulted in activation of Notch target genes, suggesting that AE is able to derepress their expression, probably contributing to the oncogenic potential of AE in AML (Salat et al. 2008). In line with that, MTG16 which was also found fused to AML1 in cases of secondary AML cases (Gamou et al. 1998) interacts with the RBPJ-associated corepressor complex, and this interaction is regulated in a Notch-dependent fashion (Engel et al. 2010).
The exact role of CBFβ in the transformation process driven by AE remained unclear for a long time as two different studies arrived to opposite conclusions (Kwok et al. 2009, 2010; Park et al. 2009; Roudaia et al. 2009). Kwok and colleagues observed that the AE/CBFβ interaction is dispensable for leukemic transformation when CBFβ-interacting deficient AE mutants were retrovirally transduced into primary bone marrow cells (Kwok et al. 2009, 2010). In contrast, the study from Roudaia and colleagues observed that this interaction is strongly required to induce leukemia (Park et al. 2009; Roudaia et al. 2009) and in support of that, inhibitors of the AE/CBFβ interaction reduce cell proliferation of the ME-1 cell line characterized by a chromosomal translocation that involves the CBFβ-encoding gene (Gorczynski et al. 2007).
Our recent study helped to clarify these contradicting results. We designed CBFβ-interacting defective AE9a mutants, and upon retroviral transduction into HoxB4-immortalized hematopoietic progenitors, we observed that the AE/CBFβ interaction is required to derepress Notch target genes but not to deregulate AML1 target genes (Thiel et al. 2017). Furthermore, the AE/CBFβ interaction is required for the colony-forming potential of transduced progenitors and to induce leukemia into recipient mice, and it must be noted that mice receiving the CBFβ-interacting defective AE9a mutant present only with myeloproliferative defects (Thiel et al. 2017). These data suggest that AE9a deregulate AML1 targets independently of CBFβ leading to a myeloproliferative disease; however, the AE9a/CBFβ interaction is required to deregulate Notch target genes and induce leukemia.
While these data suggest that derepression of Notch signaling has an oncogenic role in AML, other studies observed the opposite in fact: Notch signaling has a tumor-suppressive role in AML cases that are not associated with the t(8;21) translocation (Kannan et al. 2013; Lobry et al. 2013). In this case, the tumor-suppressive role of Notch signaling seems to be dependent on the repressive activity of HES1 (Kannan et al. 2013; Tian et al. 2015b). In line with that, HES1 expression correlates with a better prognosis in AML cases characterized by CBFβ alterations (Tian et al. 2015a), and pharmacological activation of Notch signaling has a tumor-suppressive function (Ye et al. 2016). In mouse models of MLL/AF9 (mixed-lineage leukemia/ALL1-fused gene from chromosome 9 protein)-induced AML, HES1 has a tumor-suppressive role by promoting repression of FLT3 [FMS-like tyrosine kinase 3 (Kato et al. 2015; Lobry et al. 2013)]. Interestingly, Lobry and colleagues observed that Notch activation, in an AE background, has a tumor-suppressive role (Lobry et al. 2013). The discrepancy observed between our study (Thiel et al. 2017) and the study from Lobry and colleagues (Lobry et al. 2013) may be due to the different approaches used: while we only overexpressed AE9a leading to derepression, Lobry and colleagues overexpressed both AE and NICD2. In this way, the activation levels may bring the difference(s) between oncogenic and tumor-suppressive role for Notch signaling with a weak activation (derepression) having an oncogenic role and a stronger activation (given by the AE and the NICD2 together) having a tumor-suppressive role.
Of note, also the fusion protein OTT/MAL [one twenty-two/megakaryocytic acute leukemia, also known as RBM15/MKL1 (RNA-binding motif protein 15/megakaryoblastic leukemia 1)], which is the result of the t(1;22)(p13;q13) translocation, was proposed to disturb the repressive function of RBPJ in acute megakaryoblastic leukemia [AMKL (Mercher et al. 2009)].
Conclusion and Outlook
The function of the transcription factor RBPJ not only in the presence but also in the absence of Notch activation is of major importance, since this affects chromatin regulation and hence target specificity. It would be highly desirable to have novel compounds that specifically disrupt the RBPJ-corepressor function in certain disease settings like AML. Similarly, compounds able to disrupt the activation function of RBPJ would be very helpful to avoid the serious off-target effects observed with γ-secretase inhibitors. In line with that, a recent study characterized a new RBPJ inhibitor that prevents both its repressive and activating function (Hurtado et al. 2019). Chromatin regulation is to be expected at the center of RBPJ-mediated repressive mechanisms.
Abbreviations
- ADAM:
-
A disintegrin and metalloproteinase
- AE:
-
AML1/ETO
- AE9a:
-
AML1/ETO 9a
- AEtr:
-
AML1/ETO truncated
- AF9:
-
ALL1-fused gene from chromosome 9 protein
- AMKL:
-
Acute megakaryoblastic leukemia
- AML:
-
Acute myeloid leukemia
- AML1:
-
Acute myeloid leukemia 1
- ANKs:
-
Ankyrin repeats
- B-ALL:
-
B-cell acute lymphoblastic leukemia
- CARM1:
-
Coactivator-associated arginine methyltransferase 1
- CBF1:
-
C promoter-binding factor 1
- CBFβ:
-
Core-binding factor β
- CDK8:
-
Cyclin-dependent kinase 8
- CKII:
-
Casein kinase II
- CLL:
-
Chronic lymphocytic leukemia
- CoA:
-
Coactivator
- CoR:
-
Corepressor
- CSL:
-
Homo sapiens CBF1, Drosophila melanogaster Suppressor of Hairless, and Caenorhabditis elegans Lag-1
- CtBP:
-
C-terminal-binding protein
- CtIP:
-
CtBP-interacting protein
- DDX5:
-
DEAD-box helicase 5
- DLBCL:
-
Diffuse large B-cell lymphoma
- DLL1:
-
DELTA-LIKE 1
- DLL4:
-
DELTA-LIKE 4
- dnMAML1:
-
dominant-negative MAML1
- EBNA2:
-
Epstein-Barr virus nuclear antigen 2
- EGR2 :
-
Early growth response 2
- Ep300:
-
E1A-binding protein P300
- Ep400:
-
E1A-binding protein P400
- ESCs:
-
Embryonic stem cells
- ETO:
-
Eight-twenty-one
- FBXW7:
-
F-Box and WD repeat domain-containing 7
- FHL1C:
-
Four-and-a-half LIM domain protein 1C
- FLT3:
-
FMS-like tyrosine kinase 3
- GCN5:
-
General control of amino acid synthesis protein 5-like 2
- GoF:
-
Gain of function
- GSI:
-
γ-secretase inhibitor
- H2A.Zac:
-
H2A.Z acetylation
- HAT:
-
Histone acetyltransferase
- HD:
-
heterodimerization domain
- HDACs:
-
Histone deacetylases
- Hes1 :
-
Hairy and Enhancer of Split 1
- HPCs:
-
Hematopoietic progenitor cells
- KAT:
-
lysine acetyltransferase
- KAT2A:
-
lysine acetyltransferase 2A
- KAT2B:
-
lysine acetyltransferase 2B
- KAT3B:
-
lysine acetyltransferase 3B
- KAT5:
-
lysine acetyltransferase 5
- KBF2:
-
H-2 K binding factor-2
- KDM1A:
-
lysine demethylase 1A
- KDM5A:
-
lysine demethylase 5A
- KDM7B:
-
lysine demethylase 7B
- KMT2A:
-
lysine-specific methyltransferase 2A
- KMT2D:
-
lysine-specific methyltransferase 2D
- L3MBTL3:
-
lethal (3) malignant brain tumor-like protein 3
- LID:
-
Little imaginal discs
- LoF:
-
Loss of function
- LSD1:
-
lysine-specific demethylase 1
- Lz:
-
Lozenge
- MAL:
-
Megakaryocytic acute leukemia
- MAM:
-
Mastermind
- MAML:
-
Mastermind-like
- MCL:
-
Mantle cell lymphomas
- MINT:
-
MSX2-interacting protein
- MKL1:
-
Megakaryoblastic leukemia 1
- MLL:
-
Mixed-lineage leukemia
- MS:
-
Mass spectrometry
- MTG16:
-
Myeloid translocation gene on chromosome 16 protein
- MTG8:
-
Myeloid translocation gene on 8q22
- MTGR1:
-
Myeloid translocation gene-related protein 1
- NACK:
-
Notch activation complex kinase
- NCoR:
-
Nuclear receptor corepressor
- NFAT:
-
Nuclear factor of activated T-cells
- NF-κB1:
-
Nuclear factor kappa B subunit 1
- NHR:
-
Nervy homology regions
- NICD:
-
NOTCH intracellular domain
- NICD1:
-
NOTCH1 intracellular domain 1
- NK:
-
Natural killer
- OTT:
-
One twenty-two
- PCAF:
-
Ep300-CBP-associated factors
- PEST:
-
Proline, glutamic acid, serine, and threonine
- PHF8:
-
PHD finger protein 8
- PRMT4:
-
Protein arginine methyltransferase 4
- PTMs:
-
Posttranslational modifications
- RBM15:
-
RNA-binding motif protein 15
- RBPID:
-
RBPJ-interacting domain
- RBPJ:
-
Recombination signal-binding protein for immunoglobulin kappa J region
- RBS:
-
RBPJ-binding sites
- RHD:
-
Runt homology domain
- Runx:
-
Runt-related transcription factor
- RUNX1:
-
Runt-related transcription factor 1
- SHARP:
-
SMRT and HDACs-associated repressor protein
- SMRT:
-
Silencing mediator for retinoid and thyroid receptor
- SMZL:
-
Splenic marginal zone lymphomas
- Spen:
-
split ends
- SPOC:
-
Spen paralog and ortholog C-terminal
- SPOCome:
-
SPOC interactome
- SRA :
-
Steroid receptor coactivator
- SuH:
-
Suppressor of Hairless
- TAD:
-
Transactivation domain
- T-ALL:
-
T-cell acute lymphoblastic leukemia
- TFs:
-
transcription factors
- Tip60:
-
HIV-1 Tat-interactive protein, 60 kDa
- UTR:
-
Untranslated region
- WT:
-
Wild type
- ZnF:
-
Zinc fingers
References
Alcalay M, Meani N, Gelmetti V, Fantozzi A, Fagioli M, Orleth A, Riganelli D, Sebastiani C, Cappelli E, Casciari C et al (2003) Acute myeloid leukemia fusion proteins deregulate genes involved in stem cell maintenance and DNA repair. J Clin Invest 112:1751–1761
Arnett KL, Hass M, McArthur DG, Ilagan MX, Aster JC, Kopan R, Blacklow SC (2010) Structural and mechanistic insights into cooperative assembly of dimeric Notch transcription complexes. Nat Struct Mol Biol 17:1312–1317
Augert A, Eastwood E, Ibrahim AH, Wu N, Grunblatt E, Basom R, Liggitt D, Eaton KD, Martins R, Poirier JT et al (2019) Targeting NOTCH activation in small cell lung cancer through LSD1 inhibition. Sci Signal 12
Bea S, Valdes-Mas R, Navarro A, Salaverria I, Martin-Garcia D, Jares P, Gine E, Pinyol M, Royo C, Nadeu F et al (2013) Landscape of somatic mutations and clonal evolution in mantle cell lymphoma. Proc Natl Acad Sci U S A 110:18250–18255
Borggrefe T, Liefke R (2012) Fine-tuning of the intracellular canonical Notch signaling pathway. Cell Cycle 11:264–276
Borggrefe T, Oswald F (2009) The Notch signaling pathway: transcriptional regulation at Notch target genes. Cell Mol Life Sci 66:1631–1646
Borggrefe T, Lauth M, Zwijsen A, Huylebroeck D, Oswald F, Giaimo BD (2016) The notch intracellular domain integrates signals from Wnt, Hedgehog, TGFbeta/BMP and hypoxia pathways. Biochim Biophys Acta 1863:303–313
Bray SJ (2006) Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 7:678–689
Breit S, Stanulla M, Flohr T, Schrappe M, Ludwig WD, Tolle G, Happich M, Muckenthaler MU, Kulozik AE (2006) Activating NOTCH1 mutations predict favorable early treatment response and long-term outcome in childhood precursor T-cell lymphoblastic leukemia. Blood 108:1151–1157
Brou C, Logeat F, Lecourtois M, Vandekerckhove J, Kourilsky P, Schweisguth F, Israel A (1994) Inhibition of the DNA-binding activity of Drosophila suppressor of hairless and of its human homolog, KBF2/RBP-J kappa, by direct protein-protein interaction with Drosophila hairless. Genes Dev 8:2491–2503
Castel D, Mourikis P, Bartels SJ, Brinkman AB, Tajbakhsh S, Stunnenberg HG (2013) Dynamic binding of RBPJ is determined by Notch signaling status. Genes Dev 27:1059–1071
Collins KJ, Yuan Z, Kovall RA (2014) Structure and function of the CSL-KyoT2 corepressor complex: a negative regulator of Notch signaling. Structure 22:70–81
Di Stefano L, Walker JA, Burgio G, Corona DF, Mulligan P, Naar AM, Dyson NJ (2011) Functional antagonism between histone H3K4 demethylases in vivo. Genes Dev 25:17–28
Dieguez-Hurtado R, Kato K, Giaimo BD, Nieminen-Kelha M, Arf H, Ferrante F, Bartkuhn M, Zimmermann T, Bixel MG, Eilken HM et al (2019) Loss of the transcription factor RBPJ induces disease-promoting properties in brain pericytes. Nat Commun 10:2817
Dreval K, Lake RJ, Fan HY (2019) HDAC1 negatively regulates selective mitotic chromatin binding of the Notch effector RBPJ in a KDM5A-dependent manner. Nucleic Acids Res 47:4521–4538
Ellisen LW, Bird J, West DC, Soreng AL, Reynolds TC, Smith SD, Sklar J (1991) TAN-1, the human homolog of the Drosophila notch gene, is broken by chromosomal translocations in T lymphoblastic neoplasms. Cell 66:649–661
Engel ME, Nguyen HN, Mariotti J, Hunt A, Hiebert SW (2010) Myeloid translocation gene 16 (MTG16) interacts with Notch transcription complex components to integrate Notch signaling in hematopoietic cell fate specification. Mol Cell Biol 30:1852–1863
Erickson P, Gao J, Chang KS, Look T, Whisenant E, Raimondi S, Lasher R, Trujillo J, Rowley J, Drabkin H (1992) Identification of breakpoints in t(8;21) acute myelogenous leukemia and isolation of a fusion transcript, AML1/ETO, with similarity to Drosophila segmentation gene, runt. Blood 80:1825–1831
Erickson PF, Dessev G, Lasher RS, Philips G, Robinson M, Drabkin HA (1996) ETO and AML1 phosphoproteins are expressed in CD34+ hematopoietic progenitors: implications for t(8;21) leukemogenesis and monitoring residual disease. Blood 88:1813–1823
Fabbri G, Rasi S, Rossi D, Trifonov V, Khiabanian H, Ma J, Grunn A, Fangazio M, Capello D, Monti S et al (2011) Analysis of the chronic lymphocytic leukemia coding genome: role of NOTCH1 mutational activation. J Exp Med 208:1389–1401
Fabbri G, Holmes AB, Viganotti M, Scuoppo C, Belver L, Herranz D, Yan XJ, Kieso Y, Rossi D, Gaidano G et al (2017) Common nonmutational NOTCH1 activation in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 114:E2911–E2919
Falo-Sanjuan J, Lammers NC, Garcia HG, Bray SJ (2019) Enhancer priming enables fast and sustained transcriptional responses to Notch signaling. Dev Cell 2019 Aug 19;50(4):411–425
Feinstein PG, Kornfeld K, Hogness DS, Mann RS (1995) Identification of homeotic target genes in Drosophila melanogaster including nervy, a proto-oncogene homologue. Genetics 140:573–586
Feyerabend TB, Terszowski G, Tietz A, Blum C, Luche H, Gossler A, Gale NW, Radtke F, Fehling HJ, Rodewald HR (2009) Deletion of Notch1 converts pro-T cells to dendritic cells and promotes thymic B cells by cell-extrinsic and cell-intrinsic mechanisms. Immunity 30:67–79
Fortini ME, Artavanis-Tsakonas S (1994) The suppressor of hairless protein participates in notch receptor signaling. Cell 79:273–282
Franco CB, Scripture-Adams DD, Proekt I, Taghon T, Weiss AH, Yui MA, Adams SL, Diamond RA, Rothenberg EV (2006) Notch/Delta signaling constrains reengineering of pro-T cells by PU.1. Proc Natl Acad Sci U S A 103:11993–11998
Frank R, Zhang J, Uchida H, Meyers S, Hiebert SW, Nimer SD (1995) The AML1/ETO fusion protein blocks transactivation of the GM-CSF promoter by AML1B. Oncogene 11:2667–2674
Friedmann DR, Wilson JJ, Kovall RA (2008) RAM-induced allostery facilitates assembly of a notch pathway active transcription complex. J Biol Chem 283:14781–14791
Fryer CJ, Lamar E, Turbachova I, Kintner C, Jones KA (2002) Mastermind mediates chromatin-specific transcription and turnover of the Notch enhancer complex. Genes Dev 16:1397–1411
Fryer CJ, White JB, Jones KA (2004) Mastermind recruits CycC:CDK8 to phosphorylate the Notch ICD and coordinate activation with turnover. Mol Cell 16:509–520
Gamou T, Kitamura E, Hosoda F, Shimizu K, Shinohara K, Hayashi Y, Nagase T, Yokoyama Y, Ohki M (1998) The partner gene of AML1 in t(16;21) myeloid malignancies is a novel member of the MTG8(ETO) family. Blood 91:4028–4037
Gardini A, Cesaroni M, Luzi L, Okumura AJ, Biggs JR, Minardi SP, Venturini E, Zhang DE, Pelicci PG, Alcalay M (2008) AML1/ETO oncoprotein is directed to AML1 binding regions and co-localizes with AML1 and HEB on its targets. PLoS Genet 4:e1000275
Geimer Le Lay AS, Oravecz A, Mastio J, Jung C, Marchal P, Ebel C, Dembele D, Jost B, Le Gras S, Thibault C et al (2014) The tumor suppressor Ikaros shapes the repertoire of notch target genes in T cells. Sci Signal 7:ra28
Gelmetti V, Zhang J, Fanelli M, Minucci S, Pelicci PG, Lazar MA (1998) Aberrant recruitment of the nuclear receptor corepressor-histone deacetylase complex by the acute myeloid leukemia fusion partner ETO. Mol Cell Biol 18:7185–7191
Gevry N, Chan HM, Laflamme L, Livingston DM, Gaudreau L (2007) p21 transcription is regulated by differential localization of histone H2A.Z. Genes Dev 21:1869–1881
Gevry N, Hardy S, Jacques PE, Laflamme L, Svotelis A, Robert F, Gaudreau L (2009) Histone H2A.Z is essential for estrogen receptor signaling. Genes Dev 23:1522–1533
Giaimo BD, Borggrefe T (2018) Introduction to molecular mechanisms in Notch signal transduction and disease pathogenesis. Adv Exp Med Biol 1066:3–30
Giaimo BD, Ferrante F, Borggrefe T (2017a) Chromatin immunoprecipitation (ChIP) in mouse T-cell lines. J Vis Exp
Giaimo BD, Oswald F, Borggrefe T (2017b) Dynamic chromatin regulation at Notch target genes. Transcription 8:61–66
Giaimo BD, Ferrante F, Vallejo DM, Hein K, Gutierrez-Perez I, Nist A, Stiewe T, Mittler G, Herold S, Zimmermann T et al (2018) Histone variant H2A.Z deposition and acetylation directs the canonical Notch signaling response. Nucleic Acids Res 46:8197–8215
Giaimo BD, Ferrante F, Herchenrother A, Hake SB, Borggrefe T (2019) The histone variant H2A.Z in gene regulation. Epigenetics Chromatin 12:37
Gomez-Lamarca MJ, Falo-Sanjuan J, Stojnic R, Abdul Rehman S, Muresan L, Jones ML, Pillidge Z, Cerda-Moya G, Yuan Z, Baloul S et al (2018) Activation of the notch signaling pathway in vivo elicits changes in CSL nuclear dynamics. Dev Cell 44:611–623 e617
Gorczynski MJ, Grembecka J, Zhou Y, Kong Y, Roudaia L, Douvas MG, Newman M, Bielnicka I, Baber G, Corpora T et al (2007) Allosteric inhibition of the protein-protein interaction between the leukemia-associated proteins Runx1 and CBFbeta. Chem Biol 14:1186–1197
Grossman SR, Johannsen E, Tong X, Yalamanchili R, Kieff E (1994) The Epstein-Barr virus nuclear antigen 2 transactivator is directed to response elements by the J kappa recombination signal binding protein. Proc Natl Acad Sci U S A 91:7568–7572
Guruharsha KG, Hori K, Obar RA, Artavanis-Tsakonas S (2014) Proteomic analysis of the Notch interactome. Methods Mol Biol 1187:181–192
Hamaguchi Y, Matsunami N, Yamamoto Y, Honjo T (1989) Purification and characterization of a protein that binds to the recombination signal sequence of the immunoglobulin J kappa segment. Nucleic Acids Res 17:9015–9026
Han H, Tanigaki K, Yamamoto N, Kuroda K, Yoshimoto M, Nakahata T, Ikuta K, Honjo T (2002) Inducible gene knockout of transcription factor recombination signal binding protein-J reveals its essential role in T versus B lineage decision. Int Immunol 14:637–645
Hass MR, Liow HH, Chen X, Sharma A, Inoue YU, Inoue T, Reeb A, Martens A, Fulbright M, Raju S et al (2015) SpDamID: marking DNA bound by protein complexes identifies Notch-dimer responsive enhancers. Mol Cell 59:685–697
Heibert SW, Lutterbach B, Durst K, Wang L, Linggi B, Wu S, Wood L, Amann J, King D, Hou Y (2001) Mechanisms of transcriptional repression by the t(8;21)-, t(12;21)-, and inv(16)-encoded fusion proteins. Cancer Chemother Pharmacol 48(Suppl 1):S31–S34
Hein K, Mittler G, Cizelsky W, Kuhl M, Ferrante F, Liefke R, Berger IM, Just S, Strang JE, Kestler HA et al (2015) Site-specific methylation of Notch1 controls the amplitude and duration of the Notch1 response. Sci Signal 8:ra30
Henkel T, Ling PD, Hayward SD, Peterson MG (1994) Mediation of Epstein-Barr virus EBNA2 transactivation by recombination signal-binding protein J kappa. Science 265:92–95
Ho DM, Guruharsha KG, Artavanis-Tsakonas S (2018) The Notch Interactome: complexity in signaling circuitry. Adv Exp Med Biol 1066:125–140
Huang G, Shigesada K, Ito K, Wee HJ, Yokomizo T, Ito Y (2001) Dimerization with PEBP2beta protects RUNX1/AML1 from ubiquitin-proteasome-mediated degradation. EMBO J 20:723–733
Hunt A, Fischer M, Engel ME, Hiebert SW (2011) Mtg16/Eto2 contributes to murine T-cell development. Mol Cell Biol 31:2544–2551
Hurtado C, Safarova A, Smith M, Chung R, Bruyneel AAN, Gomez-Galeno J, Oswald F, Larson CJ, Cashman JR, Ruiz-Lozano P et al (2019) Disruption of NOTCH signaling by a small molecule inhibitor of the transcription factor RBPJ. Sci Rep 9:10811
Izon DJ, Aster JC, He Y, Weng A, Karnell FG, Patriub V, Xu L, Bakkour S, Rodriguez C, Allman D, Pear WS (2002) Deltex1 redirects lymphoid progenitors to the B cell lineage by antagonizing Notch1. Immunity 16:231–243
Jin K, Zhou W, Han X, Wang Z, Li B, Jeffries S, Tao W, Robbins DJ, Capobianco AJ (2017) Acetylation of mastermind-like 1 by p300 drives the recruitment of NACK to initiate Notch-dependent transcription. Cancer Res 77:4228–4237
Jung C, Mittler G, Oswald F, Borggrefe T (2013) RNA helicase Ddx5 and the noncoding RNA SRA act as coactivators in the Notch signaling pathway. Biochim Biophys Acta 1833:1180–1189
Kannan S, Fang W, Song G, Mullighan CG, Hammitt R, McMurray J, Zweidler-McKay PA (2011) Notch/HES1-mediated PARP1 activation: a cell type-specific mechanism for tumor suppression. Blood 117:2891–2900
Kannan S, Sutphin RM, Hall MG, Golfman LS, Fang W, Nolo RM, Akers LJ, Hammitt RA, McMurray JS, Kornblau SM et al (2013) Notch activation inhibits AML growth and survival: a potential therapeutic approach. J Exp Med 210:321–337
Kato T, Sakata-Yanagimoto M, Nishikii H, Ueno M, Miyake Y, Yokoyama Y, Asabe Y, Kamada Y, Muto H, Obara N et al (2015) Hes1 suppresses acute myeloid leukemia development through FLT3 repression. Leukemia 29:576–585
Kitabayashi I, Ida K, Morohoshi F, Yokoyama A, Mitsuhashi N, Shimizu K, Nomura N, Hayashi Y, Ohki M (1998) The AML1-MTG8 leukemic fusion protein forms a complex with a novel member of the MTG8(ETO/CDR) family, MTGR1. Mol Cell Biol 18:846–858
Kitagawa M, Oyama T, Kawashima T, Yedvobnick B, Kumar A, Matsuno K, Harigaya K (2001) A human protein with sequence similarity to Drosophila mastermind coordinates the nuclear form of notch and a CSL protein to build a transcriptional activator complex on target promoters. Mol Cell Biol 21:4337–4346
Kochert K, Ullrich K, Kreher S, Aster JC, Kitagawa M, Johrens K, Anagnostopoulos I, Jundt F, Lamprecht B, Zimber-Strobl U et al (2011) High-level expression of Mastermind-like 2 contributes to aberrant activation of the NOTCH signaling pathway in human lymphomas. Oncogene 30:1831–1840
Kovall RA, Hendrickson WA (2004) Crystal structure of the nuclear effector of Notch signaling, CSL, bound to DNA. EMBO J 23:3441–3451
Kozu T, Miyoshi H, Shimizu K, Maseki N, Kaneko Y, Asou H, Kamada N, Ohki M (1993) Junctions of the AML1/MTG8(ETO) fusion are constant in t(8;21) acute myeloid leukemia detected by reverse transcription polymerase chain reaction. Blood 82:1270–1276
Krejci A, Bray S (2007) Notch activation stimulates transient and selective binding of Su(H)/CSL to target enhancers. Genes Dev 21:1322–1327
Kridel R, Meissner B, Rogic S, Boyle M, Telenius A, Woolcock B, Gunawardana J, Jenkins C, Cochrane C, Ben-Neriah S et al (2012) Whole transcriptome sequencing reveals recurrent NOTCH1 mutations in mantle cell lymphoma. Blood 119:1963–1971
Kurooka H, Honjo T (2000) Functional interaction between the mouse notch1 intracellular region and histone acetyltransferases PCAF and GCN5. J Biol Chem 275:17211–17220
Kusch T, Florens L, Macdonald WH, Swanson SK, Glaser RL, Yates JR 3rd, Abmayr SM, Washburn MP, Workman JL (2004) Acetylation by Tip60 is required for selective histone variant exchange at DNA lesions. Science 306:2084–2087
Kwok C, Zeisig BB, Qiu J, Dong S, So CW (2009) Transforming activity of AML1-ETO is independent of CBFbeta and ETO interaction but requires formation of homo-oligomeric complexes. Proc Natl Acad Sci U S A 106:2853–2858
Kwok C, Zeisig BB, Dong S, So CW (2010) The role of CBFbeta in AML1-ETO’s activity. Blood 115:3176–3177
Laiosa CV, Stadtfeld M, Xie H, de Andres-Aguayo L, Graf T (2006) Reprogramming of committed T cell progenitors to macrophages and dendritic cells by C/EBP alpha and PU.1 transcription factors. Immunity 25:731–744
Larson Gedman A, Chen Q, Kugel Desmoulin S, Ge Y, LaFiura K, Haska CL, Cherian C, Devidas M, Linda SB, Taub JW, Matherly LH (2009) The impact of NOTCH1, FBW7 and PTEN mutations on prognosis and downstream signaling in pediatric T-cell acute lymphoblastic leukemia: a report from the Children’s Oncology Group. Leukemia 23:1417–1425
Liang Y, Chang J, Lynch SJ, Lukac DM, Ganem D (2002) The lytic switch protein of KSHV activates gene expression via functional interaction with RBP-Jkappa (CSL), the target of the Notch signaling pathway. Genes Dev 16:1977–1989
Liang L, Zhang HW, Liang J, Niu XL, Zhang SZ, Feng L, Liang YM, Han H (2008) KyoT3, an isoform of murine FHL1, associates with the transcription factor RBP-J and represses the RBP-J-mediated transactivation. Biochim Biophys Acta 1779:805–810
Liefke R, Oswald F, Alvarado C, Ferres-Marco D, Mittler G, Rodriguez P, Dominguez M, Borggrefe T (2010) Histone demethylase KDM5A is an integral part of the core Notch-RBP-J repressor complex. Genes Dev 24:590–601
Lin SE, Oyama T, Nagase T, Harigaya K, Kitagawa M (2002) Identification of new human mastermind proteins defines a family that consists of positive regulators for notch signaling. J Biol Chem 277:50612–50620
Lin S, Tian L, Shen H, Gu Y, Li JL, Chen Z, Sun X, You MJ, Wu L (2013) DDX5 is a positive regulator of oncogenic NOTCH1 signaling in T cell acute lymphoblastic leukemia. Oncogene 32:4845–4853
Ling PD, Rawlins DR, Hayward SD (1993) The Epstein-Barr virus immortalizing protein EBNA-2 is targeted to DNA by a cellular enhancer-binding protein. Proc Natl Acad Sci U S A 90:9237–9241
Liu Y, Chen W, Gaudet J, Cheney MD, Roudaia L, Cierpicki T, Klet RC, Hartman K, Laue TM, Speck NA, Bushweller JH (2007) Structural basis for recognition of SMRT/N-CoR by the MYND domain and its contribution to AML1/ETO’s activity. Cancer Cell 11:483–497
Lobry C, Ntziachristos P, Ndiaye-Lobry D, Oh P, Cimmino L, Zhu N, Araldi E, Hu W, Freund J, Abdel-Wahab O et al (2013) Notch pathway activation targets AML-initiating cell homeostasis and differentiation. J Exp Med 210:301–319
Lutterbach B, Sun D, Schuetz J, Hiebert SW (1998a) The MYND motif is required for repression of basal transcription from the multidrug resistance 1 promoter by the t(8;21) fusion protein. Mol Cell Biol 18:3604–3611
Lutterbach B, Westendorf JJ, Linggi B, Patten A, Moniwa M, Davie JR, Huynh KD, Bardwell VJ, Lavinsky RM, Rosenfeld MG et al (1998b) ETO, a target of t(8;21) in acute leukemia, interacts with the N-CoR and mSin3 corepressors. Mol Cell Biol 18:7176–7184
Ma X, Renda MJ, Wang L, Cheng EC, Niu C, Morris SW, Chi AS, Krause DS (2007) Rbm15 modulates Notch-induced transcriptional activation and affects myeloid differentiation. Mol Cell Biol 27:3056–3064
Maillard I, Weng AP, Carpenter AC, Rodriguez CG, Sai H, Xu L, Allman D, Aster JC, Pear WS (2004) Mastermind critically regulates Notch-mediated lymphoid cell fate decisions. Blood 104:1696–1702
Malecki MJ, Sanchez-Irizarry C, Mitchell JL, Histen G, Xu ML, Aster JC, Blacklow SC (2006) Leukemia-associated mutations within the NOTCH1 heterodimerization domain fall into at least two distinct mechanistic classes. Mol Cell Biol 26:4642–4651
Mansour MR, Linch DC, Foroni L, Goldstone AH, Gale RE (2006) High incidence of Notch-1 mutations in adult patients with T-cell acute lymphoblastic leukemia. Leukemia 20:537–539
Mercher T, Raffel GD, Moore SA, Cornejo MG, Baudry-Bluteau D, Cagnard N, Jesneck JL, Pikman Y, Cullen D, Williams IR et al (2009) The OTT-MAL fusion oncogene activates RBPJ-mediated transcription and induces acute megakaryoblastic leukemia in a knockin mouse model. J Clin Invest 119:852–864
Metzler M, Staege MS, Harder L, Mendelova D, Zuna J, Fronkova E, Meyer C, Flohr T, Bednarova D, Harbott J et al (2008) Inv(11)(q21q23) fuses MLL to the Notch co-activator mastermind-like 2 in secondary T-cell acute lymphoblastic leukemia. Leukemia 22:1807–1811
Mikami S, Kanaba T, Ito Y, Mishima M (2013) NMR assignments of SPOC domain of the human transcriptional corepressor SHARP in complex with a C-terminal SMRT peptide. Biomol NMR Assign 7:267–270
Mikami S, Kanaba T, Takizawa N, Kobayashi A, Maesaki R, Fujiwara T, Ito Y, Mishima M (2014) Structural insights into the recruitment of SMRT by the corepressor SHARP under phosphorylative regulation. Structure 22:35–46
Miyoshi H, Kozu T, Shimizu K, Enomoto K, Maseki N, Kaneko Y, Kamada N, Ohki M (1993) The t(8;21) translocation in acute myeloid leukemia results in production of an AML1-MTG8 fusion transcript. EMBO J 12:2715–2721
Moshkin YM, Kan TW, Goodfellow H, Bezstarosti K, Maeda RK, Pilyugin M, Karch F, Bray SJ, Demmers JA, Verrijzer CP (2009) Histone chaperones ASF1 and NAP1 differentially modulate removal of active histone marks by LID-RPD3 complexes during NOTCH silencing. Mol Cell 35:782–793
Mulligan P, Yang F, Di Stefano L, Ji JY, Ouyang J, Nishikawa JL, Toiber D, Kulkarni M, Wang Q, Najafi-Shoushtari SH et al (2011) A SIRT1-LSD1 corepressor complex regulates Notch target gene expression and development. Mol Cell 42:689–699
Nam Y, Sliz P, Song L, Aster JC, Blacklow SC (2006) Structural basis for cooperativity in recruitment of MAML coactivators to Notch transcription complexes. Cell 124:973–983
Nam Y, Sliz P, Pear WS, Aster JC, Blacklow SC (2007) Cooperative assembly of higher-order Notch complexes functions as a switch to induce transcription. Proc Natl Acad Sci U S A 104:2103–2108
Nisson PE, Watkins PC, Sacchi N (1992) Transcriptionally active chimeric gene derived from the fusion of the AML1 gene and a novel gene on chromosome 8 in t(8;21) leukemic cells. Cancer Genet Cytogenet 63:81–88
Nwabo Kamdje AH, Bassi G, Pacelli L, Malpeli G, Amati E, Nichele I, Pizzolo G, Krampera M (2012) Role of stromal cell-mediated Notch signaling in CLL resistance to chemotherapy. Blood Cancer J 2:e73
Ogawa E, Inuzuka M, Maruyama M, Satake M, Naito-Fujimoto M, Ito Y, Shigesada K (1993a) Molecular cloning and characterization of PEBP2 beta, the heterodimeric partner of a novel Drosophila runt-related DNA binding protein PEBP2 alpha. Virology 194:314–331
Ogawa E, Maruyama M, Kagoshima H, Inuzuka M, Lu J, Satake M, Shigesada K, Ito Y (1993b) PEBP2/PEA2 represents a family of transcription factors homologous to the products of the Drosophila runt gene and the human AML1 gene. Proc Natl Acad Sci U S A 90:6859–6863
Okada H, Watanabe T, Niki M, Takano H, Chiba N, Yanai N, Tani K, Hibino H, Asano S, Mucenski ML et al (1998) AML1(−/−) embryos do not express certain hematopoiesis-related gene transcripts including those of the PU.1 gene. Oncogene 17:2287–2293
Okuda T, van Deursen J, Hiebert SW, Grosveld G, Downing JR (1996) AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell 84:321–330
Oswald F, Kovall RA (2018) CSL-associated corepressor and coactivator complexes. Adv Exp Med Biol 1066:279–295
Oswald F, Tauber B, Dobner T, Bourteele S, Kostezka U, Adler G, Liptay S, Schmid RM (2001) p300 acts as a transcriptional coactivator for mammalian Notch-1. Mol Cell Biol 21:7761–7774
Oswald F, Kostezka U, Astrahantseff K, Bourteele S, Dillinger K, Zechner U, Ludwig L, Wilda M, Hameister H, Knochel W et al (2002) SHARP is a novel component of the Notch/RBP-Jkappa signalling pathway. EMBO J 21:5417–5426
Oswald F, Winkler M, Cao Y, Astrahantseff K, Bourteele S, Knochel W, Borggrefe T (2005) RBP-Jkappa/SHARP recruits CtIP/CtBP corepressors to silence Notch target genes. Mol Cell Biol 25:10379–10390
Oswald F, Rodriguez P, Giaimo BD, Antonello ZA, Mira L, Mittler G, Thiel VN, Collins KJ, Tabaja N, Cizelsky W et al (2016) A phospho-dependent mechanism involving NCoR and KMT2D controls a permissive chromatin state at Notch target genes. Nucleic Acids Res 44:4703–4720
Palomero T, Barnes KC, Real PJ, Glade Bender JL, Sulis ML, Murty VV, Colovai AI, Balbin M, Ferrando AA (2006) CUTLL1, a novel human T-cell lymphoma cell line with t(7;9) rearrangement, aberrant NOTCH1 activation and high sensitivity to gamma-secretase inhibitors. Leukemia 20:1279–1287
Park S, Speck NA, Bushweller JH (2009) The role of CBFbeta in AML1-ETO’s activity. Blood 114:2849–2850
Pear WS, Aster JC, Scott ML, Hasserjian RP, Soffer B, Sklar J, Baltimore D (1996) Exclusive development of T cell neoplasms in mice transplanted with bone marrow expressing activated Notch alleles. J Exp Med 183:2283–2291
Petrovic J, Zhou Y, Fasolino M, Goldman N, Schwartz GW, Mumbach MR, Nguyen SC, Rome KS, Sela Y, Zapataro Z et al (2019) Oncogenic Notch promotes long-range regulatory interactions within Hyperconnected 3D cliques. Mol Cell 2019 Mar 21;73(6):1174–1190
Pillidge Z, Bray SJ (2019) SWI/SNF chromatin remodeling controls Notch-responsive enhancer accessibility. EMBO Rep 2019 May;20(5):e46944
Pitulescu ME, Schmidt I, Giaimo BD, Antoine T, Berkenfeld F, Ferrante F, Park H, Ehling M, Biljes D, Rocha SF et al (2017) Dll4 and notch signalling couples sprouting angiogenesis and artery formation. Nat Cell Biol 19:915–927
Puente XS, Pinyol M, Quesada V, Conde L, Ordonez GR, Villamor N, Escaramis G, Jares P, Bea S, Gonzalez-Diaz M et al (2011) Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 475:101–105
Puente XS, Bea S, Valdes-Mas R, Villamor N, Gutierrez-Abril J, Martin-Subero JI, Munar M, Rubio-Perez C, Jares P, Aymerich M et al (2015) Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526:519–524
Pui JC, Allman D, Xu L, DeRocco S, Karnell FG, Bakkour S, Lee JY, Kadesch T, Hardy RR, Aster JC, Pear WS (1999) Notch1 expression in early lymphopoiesis influences B versus T lineage determination. Immunity 11:299–308
Radtke F, Wilson A, Stark G, Bauer M, van Meerwijk J, MacDonald HR, Aguet M (1999) Deficient T cell fate specification in mice with an induced inactivation of Notch1. Immunity 10:547–558
Rosati E, Sabatini R, Rampino G, Tabilio A, Di Ianni M, Fettucciari K, Bartoli A, Coaccioli S, Screpanti I, Marconi P (2009) Constitutively activated Notch signaling is involved in survival and apoptosis resistance of B-CLL cells. Blood 113:856–865
Rossi D, Trifonov V, Fangazio M, Bruscaggin A, Rasi S, Spina V, Monti S, Vaisitti T, Arruga F, Fama R et al (2012) The coding genome of splenic marginal zone lymphoma: activation of NOTCH2 and other pathways regulating marginal zone development. J Exp Med 209:1537–1551
Roudaia L, Cheney MD, Manuylova E, Chen W, Morrow M, Park S, Lee CT, Kaur P, Williams O, Bushweller JH, Speck NA (2009) CBFbeta is critical for AML1-ETO and TEL-AML1 activity. Blood 113:3070–3079
Ryan RJH, Petrovic J, Rausch DM, Zhou Y, Lareau CA, Kluk MJ, Christie AL, Lee WY, Tarjan DR, Guo B et al (2017) A B cell Regulome links Notch to downstream oncogenic pathways in small B cell lymphomas. Cell Rep 21:784–797
Salat D, Liefke R, Wiedenmann J, Borggrefe T, Oswald F (2008) ETO, but not leukemogenic fusion protein AML1/ETO, augments RBP-Jkappa/SHARP-mediated repression of notch target genes. Mol Cell Biol 28:3502–3512
Schmitt TM, Zuniga-Pflucker JC (2002) Induction of T cell development from hematopoietic progenitor cells by delta-like-1 in vitro. Immunity 17:749–756
Schmitt TM, Ciofani M, Petrie HT, Zuniga-Pflucker JC (2004a) Maintenance of T cell specification and differentiation requires recurrent notch receptor-ligand interactions. J Exp Med 200:469–479
Schmitt TM, de Pooter RF, Gronski MA, Cho SK, Ohashi PS, Zuniga-Pflucker JC (2004b) Induction of T cell development and establishment of T cell competence from embryonic stem cells differentiated in vitro. Nat Immunol 5:410–417
Severson E, Arnett KL, Wang H, Zang C, Taing L, Liu H, Pear WS, Shirley Liu X, Blacklow SC, Aster JC (2017) Genome-wide identification and characterization of Notch transcription complex-binding sequence-paired sites in leukemia cells. Sci Signal 2017 May 2;10(477):eaag1598
Skalska L, Stojnic R, Li J, Fischer B, Cerda-Moya G, Sakai H, Tajbakhsh S, Russell S, Adryan B, Bray SJ (2015) Chromatin signatures at Notch-regulated enhancers reveal large-scale changes in H3K56ac upon activation. EMBO J 34:1889–1904
Tabaja N, Yuan Z, Oswald F, Kovall RA (2017) Structure-function analysis of RBP-J-interacting and tubulin-associated (RITA) reveals regions critical for repression of Notch target genes. J Biol Chem 292:10549–10563
Tahirov TH, Inoue-Bungo T, Morii H, Fujikawa A, Sasaki M, Kimura K, Shiina M, Sato K, Kumasaka T, Yamamoto M et al (2001) Structural analyses of DNA recognition by the AML1/Runx-1 Runt domain and its allosteric control by CBFbeta. Cell 104:755–767
Taniguchi Y, Furukawa T, Tun T, Han H, Honjo T (1998) LIM protein KyoT2 negatively regulates transcription by association with the RBP-J DNA-binding protein. Mol Cell Biol 18:644–654
Terriente-Felix A, Li J, Collins S, Mulligan A, Reekie I, Bernard F, Krejci A, Bray S (2013) Notch cooperates with Lozenge/Runx to lock haemocytes into a differentiation programme. Development 140:926–937
Tetzlaff F, Fischer A (2018) Control of blood vessel formation by Notch signaling. Adv Exp Med Biol 1066:319–338
Thiel VN, Giaimo BD, Schwarz P, Soller K, Vas V, Bartkuhn M, Blatte TJ, Dohner K, Bullinger L, Borggrefe T et al (2017) Heterodimerization of AML1/ETO with CBFbeta is required for leukemogenesis but not for myeloproliferation. Leukemia 31:2491–2502
Tian C, Tang Y, Wang T, Yu Y, Wang X, Wang Y, Zhang Y (2015a) HES1 is an independent prognostic factor for acute myeloid leukemia. Onco Targets Ther 8:899–904
Tian C, Yu Y, Jia Y, Zhu L, Zhang Y (2015b) HES1 activation suppresses proliferation of leukemia cells in acute myeloid leukemia. Ann Hematol 94:1477–1483
Tottone L, Zhdanovskaya N, Carmona Pestana A, Zampieri M, Simeoni F, Lazzari S, Ruocco V, Pelullo M, Caiafa P, Felli MP et al (2019) Histone modifications drive aberrant Notch3 expression/activity and growth in T-ALL. Front Oncol 9:198
Tun T, Hamaguchi Y, Matsunami N, Furukawa T, Honjo T, Kawaichi M (1994) Recognition sequence of a highly conserved DNA binding protein RBP-J kappa. Nucleic Acids Res 22:965–971
Vijayaraghavan J, Osborne BA (2018) Notch and T cell function – a complex tale. Adv Exp Med Biol 1066:339–354
Wacker SA, Alvarado C, von Wichert G, Knippschild U, Wiedenmann J, Clauss K, Nienhaus GU, Hameister H, Baumann B, Borggrefe T et al (2011) RITA, a novel modulator of Notch signalling, acts via nuclear export of RBP-J. EMBO J 30:43–56
Wallberg AE, Pedersen K, Lendahl U, Roeder RG (2002) p300 and PCAF act cooperatively to mediate transcriptional activation from chromatin templates by notch intracellular domains in vitro. Mol Cell Biol 22:7812–7819
Waltzer L, Logeat F, Brou C, Israel A, Sergeant A, Manet E (1994) The human J kappa recombination signal sequence binding protein (RBP-J kappa) targets the Epstein-Barr virus EBNA2 protein to its DNA responsive elements. EMBO J 13:5633–5638
Wang Q, Stacy T, Binder M, Marin-Padilla M, Sharpe AH, Speck NA (1996) Disruption of the Cbfa2 gene causes necrosis and hemorrhaging in the central nervous system and blocks definitive hematopoiesis. Proc Natl Acad Sci U S A 93:3444–3449
Wang J, Hoshino T, Redner RL, Kajigaya S, Liu JM (1998) ETO, fusion partner in t(8;21) acute myeloid leukemia, represses transcription by interaction with the human N-CoR/mSin3/HDAC1 complex. Proc Natl Acad Sci U S A 95:10860–10865
Wang H, Zou J, Zhao B, Johannsen E, Ashworth T, Wong H, Pear WS, Schug J, Blacklow SC, Arnett KL et al (2011) Genome-wide analysis reveals conserved and divergent features of Notch1/RBPJ binding in human and murine T-lymphoblastic leukemia cells. Proc Natl Acad Sci U S A 108:14908–14913
Wang H, Zang C, Taing L, Arnett KL, Wong YJ, Pear WS, Blacklow SC, Liu XS, Aster JC (2014) NOTCH1-RBPJ complexes drive target gene expression through dynamic interactions with superenhancers. Proc Natl Acad Sci U S A 111:705–710
Weaver KL, Alves-Guerra MC, Jin K, Wang Z, Han X, Ranganathan P, Zhu X, DaSilva T, Liu W, Ratti F et al (2014) NACK is an integral component of the Notch transcriptional activation complex and is critical for development and tumorigenesis. Cancer Res 74:4741–4751
Weng AP, Ferrando AA, Lee W, Morris JP t, Silverman LB, Sanchez-Irizarry C, Blacklow SC, Look AT, Aster JC (2004) Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306:269–271
Wilson JJ, Kovall RA (2006) Crystal structure of the CSL-Notch-Mastermind ternary complex bound to DNA. Cell 124:985–996
Wilson A, MacDonald HR, Radtke F (2001) Notch 1-deficient common lymphoid precursors adopt a B cell fate in the thymus. J Exp Med 194:1003–1012
Wu L, Aster JC, Blacklow SC, Lake R, Artavanis-Tsakonas S, Griffin JD (2000) MAML1, a human homologue of Drosophila mastermind, is a transcriptional co-activator for NOTCH receptors. Nat Genet 26:484–489
Wu L, Sun T, Kobayashi K, Gao P, Griffin JD (2002) Identification of a family of mastermind-like transcriptional coactivators for mammalian notch receptors. Mol Cell Biol 22:7688–7700
Xie Q, Wu Q, Kim L, Miller TE, Liau BB, Mack SC, Yang K, Factor DC, Fang X, Huang Z et al (2016) RBPJ maintains brain tumor-initiating cells through CDK9-mediated transcriptional elongation. J Clin Invest 126:2757–2772
Xu T, Park SS, Giaimo BD, Hall D, Ferrante F, Ho DM, Hori K, Anhezini L, Ertl I, Bartkuhn M et al (2017) RBPJ/CBF1 interacts with L3MBTL3/MBT1 to promote repression of Notch signaling via histone demethylase KDM1A/LSD1. EMBO J 36:3232–3249
Yan M, Burel SA, Peterson LF, Kanbe E, Iwasaki H, Boyapati A, Hines R, Akashi K, Zhang DE (2004) Deletion of an AML1-ETO C-terminal NcoR/SMRT-interacting region strongly induces leukemia development. Proc Natl Acad Sci U S A 101:17186–17191
Yan M, Kanbe E, Peterson LF, Boyapati A, Miao Y, Wang Y, Chen IM, Chen Z, Rowley JD, Willman CL, Zhang DE (2006) A previously unidentified alternatively spliced isoform of t(8;21) transcript promotes leukemogenesis. Nat Med 12:945–949
Yan M, Ahn EY, Hiebert SW, Zhang DE (2009) RUNX1/AML1 DNA-binding domain and ETO/MTG8 NHR2-dimerization domain are critical to AML1-ETO9a leukemogenesis. Blood 113:883–886
Yashiro-Ohtani Y, Wang H, Zang C, Arnett KL, Bailis W, Ho Y, Knoechel B, Lanauze C, Louis L, Forsyth KS et al (2014) Long-range enhancer activity determines Myc sensitivity to Notch inhibitors in T cell leukemia. Proc Natl Acad Sci U S A 111:E4946–E4953
Yatim A, Benne C, Sobhian B, Laurent-Chabalier S, Deas O, Judde JG, Lelievre JD, Levy Y, Benkirane M (2012) NOTCH1 nuclear interactome reveals key regulators of its transcriptional activity and oncogenic function. Mol Cell 48:445–458
Ye Q, Jiang J, Zhan G, Yan W, Huang L, Hu Y, Su H, Tong Q, Yue M, Li H et al (2016) Small molecule activation of NOTCH signaling inhibits acute myeloid leukemia. Sci Rep 6:26510
Yoo JY, Choi HK, Choi KC, Park SY, Ota I, Yook JI, Lee YH, Kim K, Yoon HG (2012) Nuclear hormone receptor corepressor promotes esophageal cancer cell invasion by transcriptional repression of interferon-gamma-inducible protein 10 in a casein kinase 2-dependent manner. Mol Biol Cell 23:2943–2954
Yoo JY, Lim BJ, Choi HK, Hong SW, Jang HS, Kim C, Chun KH, Choi KC, Yoon HG (2013) CK2-NCoR signaling cascade promotes prostate tumorigenesis. Oncotarget 4:972–983
Young E, Noerenberg D, Mansouri L, Ljungstrom V, Frick M, Sutton LA, Blakemore SJ, Galan-Sousa J, Plevova K, Baliakas P et al (2017) EGR2 mutations define a new clinically aggressive subgroup of chronic lymphocytic leukemia. Leukemia 31:1547–1554
Yuan Y, Zhou L, Miyamoto T, Iwasaki H, Harakawa N, Hetherington CJ, Burel SA, Lagasse E, Weissman IL, Akashi K, Zhang DE (2001) AML1-ETO expression is directly involved in the development of acute myeloid leukemia in the presence of additional mutations. Proc Natl Acad Sci U S A 98:10398–10403
Yuan Z, VanderWielen BD, Giaimo BD, Pan L, Collins CE, Turkiewicz A, Hein K, Oswald F, Borggrefe T, Kovall RA (2019) Structural and functional studies of the RBPJ-SHARP complex reveal a conserved corepressor binding site. Cell Rep 26:845–854 e846
Zhang J, Hug BA, Huang EY, Chen CW, Gelmetti V, Maccarana M, Minucci S, Pelicci PG, Lazar MA (2001) Oligomerization of ETO is obligatory for corepressor interaction. Mol Cell Biol 21:156–163
Zhao B, Zou J, Wang H, Johannsen E, Peng CW, Quackenbush J, Mar JC, Morton CC, Freedman ML, Blacklow SC et al (2011) Epstein-Barr virus exploits intrinsic B-lymphocyte transcription programs to achieve immortal cell growth. Proc Natl Acad Sci U S A 108:14902–14907
Zhu YM, Zhao WL, Fu JF, Shi JY, Pan Q, Hu J, Gao XD, Chen B, Li JM, Xiong SM et al (2006) NOTCH1 mutations in T-cell acute lymphoblastic leukemia: prognostic significance and implication in multifactorial leukemogenesis. Clin Cancer Res 12:3043–3049
Zimber-Strobl U, Strobl LJ, Meitinger C, Hinrichs R, Sakai T, Furukawa T, Honjo T, Bornkamm GW (1994) Epstein-Barr virus nuclear antigen 2 exerts its transactivating function through interaction with recombination signal binding protein RBP-J kappa, the homologue of Drosophila Suppressor of Hairless. EMBO J 13:4973–4982
Acknowledgments
This work was supported by the collaborative research grant TRR81 and the Heisenberg program (BO 1639/5-1) by the DFG (German Research Foundation) and the Excellence Cluster for Cardio Pulmonary System (ECCPS) in Giessen to T.B. Funding for open access charge was provided by the DFG collaborative research TRR81. B.D.G. is supported by a Research Grant of the University Medical Center Giessen and Marburg (UKGM).
Competing Interests
The authors declare that they have no competing interests.
Consent for Publication
All authors have approved the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Giaimo, B.D., Gagliani, E.K., Kovall, R.A., Borggrefe, T. (2021). Transcription Factor RBPJ as a Molecular Switch in Regulating the Notch Response. In: Reichrath, J., Reichrath, S. (eds) Notch Signaling in Embryology and Cancer. Advances in Experimental Medicine and Biology, vol 1287. Springer, Cham. https://doi.org/10.1007/978-3-030-55031-8_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-55031-8_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-55030-1
Online ISBN: 978-3-030-55031-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)