Abstract
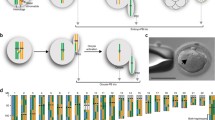
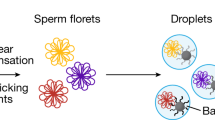
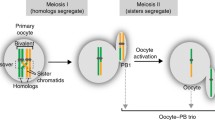
During meiosis, homologous chromosomes (homologs) undergo recombinational interactions, resulting in the formation of crossovers (COs) or noncrossovers (NCOs). Both COs and NCOs are initiated by the same event: programmed double-strand DNA breaks (DSBs), which occur preferentially at hotspots throughout the genome. COs contribute to the genetic diversity of gametes and are needed to promote proper meiotic chromosome segregation. Accordingly, their formation is tightly controlled. In the mouse, the sites of preferred CO formation differ between male and female chromosomes, both on a regional level and on the level of individual hotspots. Sperm typing using (half-sided) allele-specific PCR has proven a powerful technique to characterize COs and all detectable NCOs at hotspots on male human and mouse chromosomes. In contrast, very little is known about the properties of hotspots in female meiosis. This chapter describes an adaptation of sperm typing to analyze recombinants in a hotspot, using DNA isolated from an ovary cell suspension enriched for oocytes.
An erratum to this chapter is available at http://dx.doi.org/10.1007/978-1-62703-191-2_21
An erratum to this chapter can be found at http://dx.doi.org/10.1007/978-1-62703-191-2_21
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Keeney S (2007) Spo11 and the formation of DNA double-strand breaks in meiosis. In: Lankenau DH (ed) Recombination and meiosis. Springer, Heidelberg, Germany, pp 81–123
Allers T, Lichten M (2001) Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106:47–57
Hunter N, Kleckner N (2001) The single-end invasion: an asymmetric intermediate at the double-strand break to double-holliday junction transition of meiotic recombination. Cell 106:59–70
Kauppi L, Jeffreys AJ, Keeney S (2004) Where the crossovers are: recombination distributions in mammals. Nat Rev Genet 5:413–424
Jeffreys AJ, Kauppi L, Neumann R (2001) Intensely punctate meiotic recombination in the class II region of the major histocompatibility complex. Nat Genet 29:217–222
Guillon H, de Massy B (2002) An initiation site for meiotic crossing-over and gene conversion in the mouse. Nat Genet 32:296–299
Jeffreys AJ, May CA (2004) Intense and highly localized gene conversion activity in human meiotic crossover hot spots. Nat Genet 36:151–156
Guillon H, Baudat F, Grey C, Liskay RM, de Massy B (2005) Crossover and noncrossover pathways in mouse meiosis. Mol Cell 20:563–573
Holloway K, Lawson VE, Jeffreys AJ (2006) Allelic recombination and de novo deletions in sperm in the human beta-globin gene region. Hum Mol Genet 15:1099–1111
Cole F, Keeney S, Jasin M (2010) Comprehensive, fine-scale dissection of homologous recombination outcomes at a hot spot in mouse meiosis. Mol Cell 39:700–710
Paigen K et al (2008) The recombinational anatomy of a mouse chromosome. PLoS Genet 4(7):e1000119
Billings T et al (2010) Patterns of recombination activity on mouse chromosome 11 revealed by high resolution mapping. PLoS ONE 5(12):e15340
de Boer E et al (2006) Two levels of interference in mouse meiotic recombination. Proc Natl Acad Sci U S A 103:9607–9612
Bois PR (2007) A highly polymorphic meiotic recombination mouse hot spot exhibits incomplete repair. Mol Cell Biol 27:7053–7062
Smagulova F et al (2011) Genome-wide analysis reveals novel molecular features of mouse recombination hotspots. Nature 472:375–378
Jeffreys AJ, Murray J, Neumann R (1998) High-resolution mapping of crossovers in human sperm defines a minisatellite-associated recombination hotspot. Mol Cell 2:267–273
Hubert R et al (1994) High resolution localization of recombination hot spots using sperm typing. Nat Genet 7:420–424
Kauppi L, May CA, Jeffreys AJ (2009) Analysis of meiotic recombination products from human sperm. Methods Mol Biol 557:323–355
Cole F, Jasin M (2011) Isolation of meiotic recombinants from mouse sperm. Methods Mol Biol 745:251–282
Baudat F, de Massy B (2009) Parallel detection of crossovers and non-crossovers in mouse germ cells. Methods Mol Biol 557:305–322
Eppig JJ, Schroeder AC (1989) Capacity of mouse oocytes from preantral follicles to undergo embryogenesis and development to live young after growth, maturation, and fertilization in vitro. Biol Reprod 41:268–276
McClellan KA, Gosden R, Taketo T (2003) Continuous loss of oocytes throughout meiotic prophase in the normal mouse ovary. Dev Biol 258:334–348
Wood WI et al (1985) Base composition-independent hybridization in tetramethylammonium chloride: a method for oligonucleotide screening of highly complex gene libraries. Proc Natl Acad Sci USA 82:1585–1588
Jeffreys AJ, Neumann R (2002) Reciprocal crossover asymmetry and meiotic drive in a human recombination hot spot. Nat Genet 31:267–271
Acknowledgements
We thank Liisa Kauppi and Francesca Cole for advice on allele-specific PCR. This work was supported by a Netherlands Organization for Scientific Research Rubicon Grant 825.07.006 (E.B.) and a National Institutes of Health Grant R01 HD53855 (S.K. and M.J.).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
de Boer, E., Jasin, M., Keeney, S. (2013). Analysis of Recombinants in Female Mouse Meiosis. In: Homer, H. (eds) Mammalian Oocyte Regulation. Methods in Molecular Biology, vol 957. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-191-2_2
Download citation
DOI: https://doi.org/10.1007/978-1-62703-191-2_2
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-190-5
Online ISBN: 978-1-62703-191-2
eBook Packages: Springer Protocols