Abstract
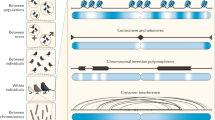
Recombination acts to shuffle the existing genetic variation within a population, leading to various approaches for detecting its action and estimating the rate at which it occurs. Here, we discuss the principal methodological and analytical approaches taken to understanding the distribution of recombination across the human genome. We first discuss the detection of recent crossover events in both well-characterised pedigrees and larger populations with extensive recent shared ancestry. We then describe approaches for learning about the fine-scale structure of recombination rate variation from patterns of genetic variation in unrelated individuals. Finally, we show how related approaches using individuals of admixed ancestry can provide an alternative approach to analysing recombination. Approaches differ not only in the statistical methods used, but also in the resolution of inference, the timescale over which recombination events are detected, and the extent to which inter-individual variation can be identified.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Broman, K.W., et al., Comprehensive human genetic maps: individual and sex-specific variation in recombination. Am J Hum Genet, 1998. 63(3): p. 861–9.
Kong, A., et al., A high-resolution recombination map of the human genome. Nat Genet, 2002. 31(3): p. 241–7.
The International HapMap Consortium, A haplotype map of the human genome. Nature, 2005. 437(7063): p. 1299–320.
McVean, G.A., et al., The fine-scale structure of recombination rate variation in the human genome. Science, 2004. 304(5670): p. 581–4.
Myers, S., et al., A fine-scale map of recombination rates and hotspots across the human genome. Science, 2005. 310(5746): p. 321–4.
Jeffreys, A.J., L. Kauppi, and R. Neumann, Intensely punctate meiotic recombination in the class II region of the major histocompatibility complex. Nat Genet, 2001. 29(2): p. 217–22.
Jeffreys, A.J., et al., Human recombination hotspots hidden in regions of strong marker association. Nat Genet, 2005. 37(6): p. 601–6.
Myers, S., et al., The distribution and causes of meiotic recombination in the human genome. Biochem Soc Trans, 2006. 34(Pt 4): p. 526–30.
Baudat, F., et al., PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science, 2010. 327(5967): p. 836–40.
Berg, I.L., et al., PRDM9 variation strongly influences recombination hot-spot activity and meiotic instability in humans. Nat Genet, 2010. 42(10): p. 859–63.
Myers, S., et al., Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science, 2010. 327(5967): p. 876–9.
Parvanov, E.D., P.M. Petkov, and K. Paigen, Prdm9 controls activation of mammalian recombination hotspots. Science, 2010. 327(5967): p. 835.
Marchini, J., et al., A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet, 2007. 39(7): p. 906–13.
Abecasis, G.R., D. Ghosh, and T.E. Nichols, Linkage disequilibrium: ancient history drives the new genetics. Hum Hered, 2005. 59(2): p. 118–24.
Price, A.L., et al., Sensitive detection of chromosomal segments of distinct ancestry in admixed populations. PLoS Genet, 2009. 5(6): p. e1000519.
McVean, G. and C.C. Spencer, Scanning the human genome for signals of selection. Curr Opin Genet Dev, 2006. 16(6): p. 624–9.
Nielsen, R., et al., Recent and ongoing selection in the human genome. Nat Rev Genet, 2007. 8(11): p. 857–68.
Myers, S., et al., A common sequence motif associated with recombination hotspots and genome instability in humans. Nat Genet, 2008. 40(9): p. 1124–9.
Stankiewicz, P. and J.R. Lupski, Genome architecture, rearrangements and genomic disorders. Trends Genet, 2002. 18(2): p. 74–82.
Kong, A., et al., Fine-scale recombination rate differences between sexes, populations and individuals. Nature, 2010. 467(7319): p. 1099–103.
Jeffreys, A.J., A. Ritchie, and R. Neumann, High resolution analysis of haplotype diversity and meiotic crossover in the human TAP2 recombination hotspot. Hum Mol Genet, 2000. 9(5): p. 725–33.
Jeffreys, A.J. and R. Neumann, Reciprocal crossover asymmetry and meiotic drive in a human recombination hot spot. Nat Genet, 2002. 31(3): p. 267–71.
Jeffreys, A.J. and R. Neumann, Factors influencing recombination frequency and distribution in a human meiotic crossover hotspot. Hum Mol Genet, 2005. 14(15): p. 2277–87.
Botstein, D., et al., Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet, 1980. 32(3): p. 314–31.
Coop, G., et al., High-resolution mapping of crossovers reveals extensive variation in fine-scale recombination patterns among humans. Science, 2008. 319(5868): p. 1395–8.
Elston, R.C. and J. Stewart, A general model for the genetic analysis of pedigree data. Hum Hered, 1971. 21(6): p. 523–42.
Lander, E.S. and P. Green, Construction of multilocus genetic linkage maps in humans. Proc Natl Acad Sci U S A, 1987. 84(8): p. 2363–7.
Kruglyak, L., et al., Parametric and nonparametric linkage analysis: a unified multipoint approach. Am J Hum Genet, 1996. 58(6): p. 1347–63.
Kong, A., et al., Detection of sharing by descent, long-range phasing and haplotype imputation. Nat Genet, 2008. 40(9): p. 1068–75.
Lewontin, R.C., The Interaction of Selection and Linkage. I. General Considerations; Heterotic Models. Genetics, 1964. 49(1): p. 49–67.
Hill, W.G. and A. Robertson, Linkage disequilibrium in finite populations. TAG Theoretical and Applied Genetics, 1968. 38(6): p. 226–231.
McVean, G., Linkage disequilibrium, recombination and selection, in The Handbook of Statistical Genetics, D.J. Balding, M. Bishop, and C. Cannings, Editors. 2008, Wiley. p. 909–940.
Ardlie, K.G., L. Kruglyak, and M. Seielstad, Patterns of linkage disequilibrium in the human genome. Nat Rev Genet, 2002. 3(4): p. 299–309.
Hudson, R.R. and N.L. Kaplan, Statistical properties of the number of recombination events in the history of a sample of DNA sequences. Genetics, 1985. 111(1): p. 147–64.
Myers, S., The Detection of Recombination Events Using DNA Sequence Data, in Department of Statistics, 2002, University of Oxford: Oxford.
Myers, S.R. and R.C. Griffiths, Bounds on the minimum number of recombination events in a sample history. Genetics, 2003. 163(1): p. 375–94.
Song, Y.S. and J. Hein, Constructing minimal ancestral recombination graphs. J Comput Biol, 2005. 12(2): p. 147–69.
Wakeley, J., Coalescent theory : an introduction, 2009, Greenwood Village, Colo.: Roberts & Co. Publishers. xii, 326 p.
Nordborg, M., Coalescent theory. 2000.
Hein, J., M.H. Schierup, and C. Wiuf, Gene genealogies, variation and evolution : a primer in coalescent theory, 2005, Oxford ; New York: Oxford University Press. xiii, 276 p.
Griffiths, R.C. and P. Marjoram, Ancestral inference from samples of DNA sequences with recombination. J Comput Biol, 1996. 3(4): p. 479–502.
The International HapMap Consortium, A second generation human haplotype map of over 3.1 million SNPs. Nature, 2007. 449(7164): p. 851–61.
Song, Y.S., R. Lyngso, and J. Hein, Counting all possible ancestral configurations of sample sequences in population genetics. IEEE/ACM Trans Comput Biol Bioinform, 2006. 3(3): p. 239–51.
Hudson, R.R., Two-locus sampling distributions and their application. Genetics, 2001. 159(4): p. 1805–17.
McVean, G., P. Awadalla, and P. Fearnhead, A coalescent-based method for detecting and estimating recombination from gene sequences. Genetics, 2002. 160(3): p. 1231–41.
Auton, A. and G. McVean, Recombination rate estimation in the presence of hotspots. Genome Res, 2007. 17(8): p. 1219–27.
Fearnhead, P., Consistency of estimators of the population-scaled recombination rate. Theor Popul Biol, 2003. 64(1): p. 67–79.
Hinch, A.G., et al., The landscape of recombination in African Americans. Nature, 2011. 476: p. 170–75.
Li, N. and M. Stephens, Modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics, 2003. 165(4): p. 2213–33.
Pfaff, C.L., et al., Population structure in admixed populations: effect of admixture dynamics on the pattern of linkage disequilibrium. Am J Hum Genet, 2001. 68(1): p. 198–207.
Patterson, N., et al., Methods for high-density admixture mapping of disease genes. Am J Hum Genet, 2004. 74(5): p. 979–1000.
Seldin, M.F., et al., Putative ancestral origins of chromosomal segments in individual african americans: implications for admixture mapping. Genome Res, 2004. 14(6): p. 1076–84.
Tian, C., et al., A genomewide single-nucleotide-polymorphism panel with high ancestry information for African American admixture mapping. Am J Hum Genet, 2006. 79(4): p. 640–9.
Kong, A., et al., Recombination rate and reproductive success in humans. Nat Genet, 2004. 36(11): p. 1203–6.
Ptak, S.E., et al., Fine-scale recombination patterns differ between chimpanzees and humans. Nat Genet, 2005. 37(4): p. 429–34.
Winckler, W., et al., Comparison of fine-scale recombination rates in humans and chimpanzees. Science, 2005. 308(5718): p. 107–11.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Auton, A., McVean, G. (2012). Estimating Recombination Rates from Genetic Variation in Humans. In: Anisimova, M. (eds) Evolutionary Genomics. Methods in Molecular Biology, vol 856. Humana Press. https://doi.org/10.1007/978-1-61779-585-5_9
Download citation
DOI: https://doi.org/10.1007/978-1-61779-585-5_9
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-61779-584-8
Online ISBN: 978-1-61779-585-5
eBook Packages: Springer Protocols