Abstract
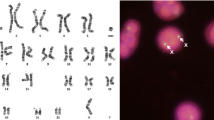
With the progress of array technologies and the enabled screening of individual human genomes, a new kind of polymorphism has been described – the so-called copy number variation (CNV) polymorphism. Copy number variants can be found in around 12% of the human genome sequence and have a size of up to several hundred kilobase pairs. These variants can not only differ between individuals, but also between corresponding alleles on homologous chromosomes. We recently developed a cytological assay for parental origin determination that relies on the design of CNV-based sets of probes for fluorescence in situ hybridization (POD-FISH). Here we describe an improved POD-FISH protocol that exploits “high frequency” variants for better discrimination of homologous chromosomes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lee, C. (2005) Vive la difference! Nat Genet 37, 660–661.
Schlötterer, C. (2004) The evolution of molecular markers-just a matter of fashion? Nat Rev Genet 5, 63–69.
Liehr, T., Ziegler, M., Starke, H., Heller, A., Kuechler, A., Kittner, G., Beensen, V., Seidel, J., Hassler, H., Musebeck, J., Claussen, U. (2003) Conspicuous GTG-banding results of the centromere-near region can be caused by alphoid DNA heteromorphism. Clin Genet 64, 166–167.
Müller, H., Klinger, H.P., Glasser, M. (1975) Chromosome polymorphism in a human newborn population II Potentials of polymorphic chromosome variants for characterizing the idiogram of an individual. Cytogenet Cell Genet 15, 239–255.
Gardner, R.G.M., Sutherland, G.R. (2004) Oxford Monographs on Medical Genetics No 46: Chromosome abnormalities and genetic counselling. 3rd edition Oxford University Press, Oxford, New York.
Shaffer, L.G., Tommerup, N. (eds) (2005) ISCN: An international System for Human Cytogenetic Nomenclature. S. Karger, Basel, Switzerland.
Liehr, T., Nietzel, A., Starke, H., Heller, A., Weise, A., Kuechler, A., Senger, G., Ebner, S., Martin, T., Stumm, M., Wegner, R., Tonnies, H., Hoppe, C., Claussen, U., von Eggeling, F. (2003) Characterization of small marker chromosomes (smc) by recently developed molecular cytogenetic approaches. J Assoc Genet Technol 29, 5–10.
Iafrate, A.J., Feuk, L., Rivera, M.N., Listewnik, M.L., Donahoe, P.K., Qi, Y., Scherer, S.W., Lee, C. (2004) Detection of large-scale variation in the human genome. Nat Genet 36, 949–951.
Sebat, J., Lakshmi, B., Troge, J., Alexander, J., Young, J., Lundin, P., Månér, S., Massa, H., Walker, M., Chi, M., Navin, N., Lucito, R., Healy, J., Hicks, J., Ye, K., Reiner, A., Gilliam, T.C., Trask, B., Patterson, N., Zetterberg, A., Wigler, M. (2004) Large-scale copy number polymorphism in the human genome. Science 23, 525–528.
Feuk, L., Marshall, C.R., Wintle, R.F., Scherer, S.W. (2006) Structural variants: changing the landscape of chromosomes and design of disease studies. Hum Mol Genet 15, 57–66.
Redon, R., Ishikawa, S., Fitch, K.R., Feuk, L., Perry, G.H., Andrews, T.D., Fiegler, H., Shapero, M.H., Carson, A.R., Chen, W., Cho, E.K., Dallaire, S., Freeman, J.L., González, J.R., Gratacòs, M., Huang, J., Kalaitzopoulos, D., Komura, D., MacDonald, J.R., Marshall, C.R., Mei, R., Montgomery, L., Nishimura, K., Okamura, K., Shen, F., Somerville, M.J., Tchinda, J., Valsesia, A., Woodwark, C., Yang, F., Zhang, J., Zerjal, T., Zhang, J., Armengol, L., Conrad, D.F., Estivill, X., Tyler-Smith, C., Carter, N.P., Aburatani, H., Lee, C., Jones, K.W., Scherer, S.W., Hurles, M.E. (2006) Global variation in copy number in the human genome. Nature 444, 444–454.
Wang, K., Li, M., Hadley, D., Liu, R., Glessner, J., Grant, S.F., Hakonarson, H., Bucan, M. (2007) PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res 17, 1665–1674.
Perry, G.H., Ben-Dor, A., Tsalenko, A., Sampas, N., Rodriguez-Revenga, L., Tran, C.W., Scheffer, A., Steinfeld, I., Tsang, P., Yamada, N.A., Park, H.S., Kim, J.I., Seo, J.S., Yakhini, Z., Laderman, S., Bruhn, L., Lee, C. (2008) The fine-scale and complex architecture of human copy-number variation. Am J Hum Genet 82, 685–695.
Weise, A., Gross, M., Mrasek, K., Mkrtchyan, H., Horsthemke, B., Jonsrud, C., Von Eggeling, F., Hinreiner, S., Witthuhn, V., Claussen, U., Liehr, T. (2008) Parental-origin-determination fluorescence in situ hybridization distinguishes homologous human chromosomes on a single-cell level. Int J Mol Med 21, 189–200.
Telenius, H., Carter, N.P., Bebb, C.E., Nordenskjöld, M., Ponder, B.A., Tunnacliffe, A. (1992) Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725.
Liehr, T., Claussen, U. (2002) FISH on chromosome preparations of peripheral blood, in: FISH-Technology (Rautenstrauss, B., Liehr, T., eds) Springer, Berlin, 73–81.
Acknowledgments
Supported in parts by the DFG (436 RUS 17/109/04, 436 RUS 17/22/06, WE 3617/2-1, 436 ARM 17/5/06, LI820/11-1, LI820/17-1), Boehringer Ingelheim Fonds, Evangelische Studienwerk e.V. Villigst, IZKF Jena (Start-up S16), IZKF together with the TMWFK (TP 3.7 and B307-04004), University Jena, Stiftung Leukämie, and Stefan-Morsch-Stiftung.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Weise, A., Gross, M., Hinreiner, S., Witthuhn, V., Mkrtchyan, H., Liehr, T. (2010). POD-FISH: A New Technique for Parental Origin Determination Based on Copy Number Variation Polymorphism. In: Bridger, J., Volpi, E. (eds) Fluorescence in situ Hybridization (FISH). Methods in Molecular Biology, vol 659. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-60761-789-1_22
Download citation
DOI: https://doi.org/10.1007/978-1-60761-789-1_22
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-60761-788-4
Online ISBN: 978-1-60761-789-1
eBook Packages: Springer Protocols