Abstract
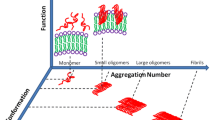
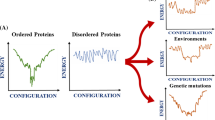
Proteins exhibit a remarkable structural plasticity and may undergo conformational changes resulting in protein misfolding both in a biological context and upon perturbing physiopathological conditions. Such nonfunctional protein conformers, including misfolded states and aggregates, are often associated to protein folding diseases. Understanding the biology of protein folding diseases thus requires tools that allow the structural characterization of nonnative conformations of proteins and their interconversions. Here we present detailed procedures to monitor protein conformational changes and aggregation based on spectroscopic and biophysical methods that include circular dichroism, ATR-Fourier-transformed infrared spectroscopy, fluorescence spectroscopy and dynamic light scattering. To illustrate the application of these methods we report to our previous studies on misfolding, aggregation and amyloid fibril formation by superoxide dismutase 1 (SOD1), a protein whose toxic deposition is implicated in the neurodegenerative disease amyotrophic lateral sclerosis (ALS).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Abbreviations
- CD:
-
Circular dichroism
- FTIR:
-
Fourier-transformed infrared spectroscopy
- DLS:
-
Dynamic light scattering
- SOD1:
-
Superoxide dismutase 1
- 1,8-ANS:
-
8-Anilinonaphthalene-1-sulfonic acid
- ThT:
-
Thioflavin T
References
Rowland LP, Shneider NA (2001) Amyotrophic lateral sclerosis. N Engl J Med 344(22):1688–1700
Nordlund A, Leinartaite L, Saraboji K, Aisenbrey C, Grobner G, Zetterstrom P, Danielsson J, Logan DT, Oliveberg M (2009) Functional features cause misfolding of the ALS-provoking enzyme SOD1. Proc Natl Acad Sci U S A 106(24):9667–9672. https://doi.org/10.1073/pnas.0812046106
Cristovao JS, Santos R, Gomes CM (2016) Metals and neuronal metal binding proteins implicated in Alzheimer's disease. Oxidative Med Cell Longev 2016:9812178. https://doi.org/10.1155/2016/9812178
Leal SS, Botelho HM, Gomes CM (2012) Metal ions as modulators of protein conformation and misfolding in neurodegeneration. Coordin Chem Rev 256(19–20):2253–2270. https://doi.org/10.1016/j.ccr.2012.04.004
Leal SS, Cardoso I, Valentine JS, Gomes CM (2013) Calcium ions promote superoxide dismutase 1 (SOD1) aggregation into non-fibrillar amyloid: a link to toxic effects of calcium overload in amyotrophic lateral sclerosis (ALS)? J Biol Chem 288(35):25219–25228. https://doi.org/10.1074/jbc.M113.470740
Leal SS, Cristovao JS, Biesemeier A, Cardoso I, Gomes CM (2015) Aberrant zinc binding to immature conformers of metal-free copper-zinc superoxide dismutase triggers amorphous aggregation. Metallomics 7(2):333–346. https://doi.org/10.1039/c4mt00278d
Leal SS, Gomes CM (2015) Calcium dysregulation links ALS defective proteins and motor neuron selective vulnerability. Front Cell Neurosci 9:225. https://doi.org/10.3389/fncel.2015.00225
Estacio SG, Leal SS, Cristovao JS, Faisca PF, Gomes CM (2015) Calcium binding to gatekeeper residues flanking aggregation-prone segments underlies non-fibrillar amyloid traits in superoxide dismutase 1 (SOD1). Biochim Biophys Acta 1854(2):118–126. https://doi.org/10.1016/j.bbapap.2014.11.005
Greenfield NJ (2006) Using circular dichroism spectra to estimate protein secondary structure. Nat Protoc 1(6):2876–2890. https://doi.org/10.1038/nprot.2006.202
Henriques BJ, Lucas TG, Rodrigues JV, Frederiksen JH, Teixeira MS, Tiranti V, Bross P, Gomes CM (2014) Ethylmalonic encephalopathy ETHE1 R163W/R163Q mutations alter protein stability and redox properties of the iron Centre. PLoS One 9(9):e107157. https://doi.org/10.1371/journal.pone.0107157
Kelly SM, Price NC (2006) Circular dichroism to study protein interactions. Curr Protoc Protein Sci. Chapter 20:Unit 20 10. doi:https://doi.org/10.1002/0471140864.ps2010s46
Clarke DT (2012) Circular dichroism in protein folding studies. Curr Protoc Protein Sci. Chapter 28:Unit 28 23. doi:https://doi.org/10.1002/0471140864.ps2803s70
Kelly SM, Jess TJ, Price NC (2005) How to study proteins by circular dichroism. Biochim Biophys Acta 1751(2):119–139. https://doi.org/10.1016/j.bbapap.2005.06.005
Radko SP, Khmeleva SA, Suprun EV, Kozin SA, Bodoev NV, Makarov AA, Archakov AI, Shumyantseva VV (2015) Physico-chemical methods for studying amyloid-beta aggregation. Biochem Mosc Suppl S 9(3):258–274. https://doi.org/10.1134/S1990750815030075
Barth A (2007) Infrared spectroscopy of proteins. Biochim Biophys Acta 1767(9):1073–1101. https://doi.org/10.1016/j.bbabio.2007.06.004
Sarroukh R, Goormaghtigh E, Ruysschaert JM, Raussens V (2013) ATR-FTIR: a "rejuvenated" tool to investigate amyloid proteins. Biochim Biophys Acta 1828(10):2328–2338. https://doi.org/10.1016/j.bbamem.2013.04.012
Ladokhin AS (2006) Fluorescence spectroscopy in peptide and protein analysis. In: Encyclopedia of analytical chemistry. John Wiley & Sons, Ltd, Hoboken, NJ. https://doi.org/10.1002/9780470027318.a1611
Hawe A, Sutter M, Jiskoot W (2008) Extrinsic fluorescent dyes as tools for protein characterization. Pharm Res 25(7):1487–1499. https://doi.org/10.1007/s11095-007-9516-9
Leal SS, Gomes CM (2007) Studies of the molten globule state of ferredoxin: structural characterization and implications on protein folding and iron-sulfur center assembly. Proteins 68(3):606–616. https://doi.org/10.1002/prot.21448
Henriques BJ, Saraiva LM, Gomes CM (2005) Probing the mechanism of rubredoxin thermal unfolding in the absence of salt bridges by temperature jump experiments. Biochem Biophys Res Commun 333(3):839–844. https://doi.org/10.1016/j.bbrc.2005.06.004
Hagmeyer S, Cristovao JS, Mulvihill JJE, Boeckers TM, Gomes CM, Grabrucker AM (2017) Zinc binding to S100B affords regulation of trace metal homeostasis and excitotoxicity in the brain. Front Mol Neurosci 10:456. https://doi.org/10.3389/fnmol.2017.00456
Biancalana M, Koide S (2010) Molecular mechanism of Thioflavin-T binding to amyloid fibrils. Biochim Biophys Acta 1804(7):1405–1412. https://doi.org/10.1016/j.bbapap.2010.04.001
Cristovao JS, Leal SS, Cardoso I, Gomes CM (2013) Small molecules present in the cerebrospinal fluid metabolome influence superoxide dismutase 1 aggregation. Int J Mol Sci 14(9):19128–19145. https://doi.org/10.3390/ijms140919128
Botelho HM, Leal SS, Cardoso I, Yanamandra K, Morozova-Roche LA, Fritz G, Gomes CM (2012) S100A6 amyloid fibril formation is calcium-modulated and enhances superoxide dismutase-1 (SOD1) aggregation. J Biol Chem 287(50):42233–42242. https://doi.org/10.1074/jbc.M112.396416
Matos AM, Cristovao JS, Yashunsky DV, Nifantiev NE, Viana AS, Gomes CM, Rauter AP (2017) Synthesis and effects of flavonoid structure variation on amyloid-beta aggregation. Pure Appl Chem 89(9):1305–1320. https://doi.org/10.1515/pac-2017-0201
Klingstedt T, Aslund A, Simon RA, Johansson LBG, Mason JJ, Nystrom S, Hammarstrom P, Nilsson KPR (2011) Synthesis of a library of oligothiophenes and their utilization as fluorescent ligands for spectral assignment of protein aggregates. Org Biomol Chem 9(24):8356–8370. https://doi.org/10.1039/c1ob05637a
Younan ND, Viles JH (2015) A comparison of three fluorophores for the detection of amyloid fibers and prefibrillar oligomeric assemblies. ThT (Thioflavin T); ANS (1-Anilinonaphthalene-8-sulfonic acid); and bisANS (4,4′-Dianilino-1,1′-binaphthyl-5,5′-disulfonic acid). Biochemistry 54(28):4297–4306. https://doi.org/10.1021/acs.biochem.5b00309
Royer CA (2006) Probing protein folding and conformational transitions with fluorescence. Chem Rev 106(5):1769–1784. https://doi.org/10.1021/cr0404390
Hassan PA, Rana S, Verma G (2015) Making sense of Brownian motion: colloid characterization by dynamic light scattering. Langmuir 31(1):3–12. https://doi.org/10.1021/la501789z
Lorber B, Fischer F, Bailly M, Roy H, Kern D (2012) Protein analysis by dynamic light scattering: methods and techniques for students. Biochemistry and molecular biology education : a bimonthly publication of the international union of. Biochem Mol Biol 40(6):372–382. https://doi.org/10.1002/bmb.20644
Khodabandehloo A, Chen DD (2017) Particle sizing methods for the detection of protein aggregates in biopharmaceuticals. Bioanalysis 9(3):313–326. https://doi.org/10.4155/bio-2016-0269
den Engelsman J, Garidel P, Smulders R, Koll H, Smith B, Bassarab S, Seidl A, Hainzl O, Jiskoot W (2011) Strategies for the assessment of protein aggregates in pharmaceutical biotech product development. Pharm Res 28(4):920–933. https://doi.org/10.1007/s11095-010-0297-1
Filipe V, Hawe A, Jiskoot W (2010) Critical evaluation of nanoparticle tracking analysis (NTA) by NanoSight for the measurement of nanoparticles and protein aggregates. Pharm Res 27(5):796–810. https://doi.org/10.1007/s11095-010-0073-2
Ahl IM, Lindberg MJ, Tibell LA (2004) Coexpression of yeast copper chaperone (yCCS) and CuZn-superoxide dismutases in Escherichia coli yields protein with high copper contents. Protein Expr Purif 37(2):311–319. https://doi.org/10.1016/j.pep.2004.06.006
Sabel CE, Neureuther JM, Siemann S (2010) A spectrophotometric method for the determination of zinc, copper, and cobalt ions in metalloproteins using Zincon. Anal Biochem 397(2):218–226. https://doi.org/10.1016/j.ab.2009.10.037
Acknowledgments
This work was partially supported by the Fundação para a Ciência e Tecnologia (FCT/MCTES, Portugal) through fellowships to J.S.C. (SFRH/BD/101171/2014) and B.J.H. (SFRH/BPD/74475/2010), and grant PTDC/BBB-BQB/5366/2014 (to B.J.H.) and PTDC/NEU-NMC/2138/2014 (to C.M.G.). The Gomes laboratory is partly supported by grant UID/MULTI/04046/2013 from FCT/MCTES/PIDDAC (to BioISI). Bial Foundation is acknowledged through grant PT/FB/BL-2014-343 (to CMG). Joana S. Cristóvão and Bárbara J. Henriques contributed equally to this work.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Cristóvão, J.S., Henriques, B.J., Gomes, C.M. (2019). Biophysical and Spectroscopic Methods for Monitoring Protein Misfolding and Amyloid Aggregation. In: Gomes, C. (eds) Protein Misfolding Diseases. Methods in Molecular Biology, vol 1873. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8820-4_1
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8820-4_1
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8819-8
Online ISBN: 978-1-4939-8820-4
eBook Packages: Springer Protocols