Abstract
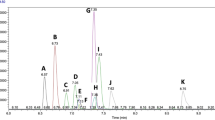
Untargeted metabolomics refers to the high-throughput analysis of the metabolic state of a biological system (e.g., tissue, biological fluid, cell culture) based on the concentration profile of all measurable free low molecular weight metabolites. Gas chromatography-mass spectrometry (GC-MS), being a highly sensitive and high-throughput analytical platform, has been proven a useful tool for untargeted studies of primary metabolism in a variety of applications. As an omic analysis, GC-MS metabolomics is a multistep procedure; thus, standardization of an untargeted GC-MS metabolomics protocol requires the integrated optimization of pre-analytical, analytical, and computational steps. The main difference of GC-MS metabolomics compared to other metabolomics analytical platforms, including liquid chromatography-MS, is the need for the derivatization of the metabolite extracts into volatile and thermally stable derivatives, the latter being quantified in the metabolic profiles. This analytical step requires special care in the optimization of the untargeted GC-MS metabolomics experimental protocol. Moreover, both the derivatization of the original sample and the compound fragmentation that takes place in GC-MS impose specialized GC-MS metabolomic data identification, quantification, normalization and filtering methods. In this chapter, we describe the integrated protocol of untargeted GC-MS metabolomics with both the analytical and computational steps, focusing on the GC-MS specific parts, and provide details on any sample depending differences.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Fiehn O (2002) Metabolomics–the link between genotypes and phenotypes. Plant Mol Biol 48(1–2):155–171. https://doi.org/10.1007/978-94-010-0448-0_11
Kanani H, Chrysanthopoulos PK, Klapa MI (2008) Standardizing GC-MS metabolomics. J Chromatogr B 871(2):191–201. https://doi.org/10.1016/j.jchromb.2008.04.049
Patti GJ, Yanes O, Siuzdak G (2012) Innovation: metabolomics: the apogee of the omics trilogy. Nat Rev Mol Cell Biol 13(4):263–269. https://doi.org/10.1038/nrm3314
Vasilopoulou CG, Margarity M, Klapa MI (2016) Metabolomic analysis in brain research: opportunities and challenges. Front Physiol 7:183. https://doi.org/10.3389/fphys.2016.00183
Spagou K, Theodoridis G, Wilson I et al (2011) A GC-MS metabolic profiling study of plasma samples from mice on low- and high-fat diets. J Chromatogr B 879(17–18):1467–1475. https://doi.org/10.1016/j.jchromb.2011.01.028
Kanani HH, Klapa MI (2007) Data correction strategy for metabolomics analysis using gas chromatography-mass spectrometry. Metab Eng 9(1):39–51. https://doi.org/10.1016/j.ymben.2006.08.001
Maga-Nteve C, Klapa MI (2016) Streamlining GC-MS metabolomic analysis using the M-IOLITE software suite. IFAC-PapersOnLine 49(26):286–288. https://doi.org/10.1016/j.ifacol.2016.12.140
Dutta B, Kanani H, Quackenbush J, Klapa MI (2009) Time-series integrated “omic” analyses to elucidate short-term stress-induced responses in plant liquid cultures. Biotechnol Bioeng 102(1):264–279. https://doi.org/10.1002/Bit.22036
Kanani H, Dutta B, Klapa MI (2010) Individual vs. combinatorial effect of elevated CO2 conditions and salinity stress on Arabidopsis thaliana liquid cultures: comparing the early molecular response using time-series transcriptomic and metabolomic analyses. BMC Syst Biol 4:177. https://doi.org/10.1186/1752-0509-4-177
Tooulakou G, Giannopoulos A, Nikolopoulos D et al (2016) “Alarm photosynthesis”: calcium oxalate crystals as an internal CO2 source in plants. Plant Physiol 171(4):2577–2585. https://doi.org/10.1104/pp.16.00111
Constantinou C, Chrysanthopoulos PK, Margarity M, Klapa MI (2011) GC-MS metabolomic analysis reveals significant alterations in cerebellar metabolic physiology in a mouse model of adult onset hypothyroidism. J Proteome Res 10(2):869–879. https://doi.org/10.1021/pr100699m
Maga-Nteve C, Vasilopoulou CG, Constantinou C et al (2017) Sex-comparative study of mouse cerebellum physiology under adult-onset hypothyroidism: the significance of GC-MS metabolomic data normalization in meta-analysis. J Chromatogr B 1041-1042:158–166. https://doi.org/10.1016/j.jchromb.2016.12.016
Chrysanthopoulos PK, Goudar CT, Klapa MI (2010) Metabolomics for high-resolution monitoring of the cellular physiological state in cell culture engineering. Metab Eng 12(3):212–222. https://doi.org/10.1016/j.ymben.2009.11.001
Vernardis SI, Goudar CT, Klapa MI (2013) Metabolic profiling reveals that time related physiological changes in mammalian cell perfusion cultures are bioreactor scale independent. Metab Eng 19:1–9. https://doi.org/10.1016/j.ymben.2013.04.005
Gkourogianni A, Kosteria I, Telonis AG et al (2014) Plasma metabolomic profiling suggests early indications for predisposition to latent insulin resistance in children conceived by ICSI. PLoS One 9(4):e94001. https://doi.org/10.1371/journal.pone.0094001
Saeed AI, Bhagabati NK, Braisted JC et al (2006) TM4 microarray software suite. Methods Enzymol 411:134–193. https://doi.org/10.1016/S0076-6879(06)11009-5
Saeed AI, Sharov V, White J et al (2003) TM4: a free, open-source system for microarray data management and analysis. BioTechniques 34(2):374–378
Allwood JW, Erban A, de Koning S et al (2009) Inter-laboratory reproducibility of fast gas chromatography-electron impact-time of flight mass spectrometry (GC-EI-TOF/MS) based plant metabolomics. Metabolomics 5(4):479–496. https://doi.org/10.1007/s11306-009-0169-z
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A 95(25):14863–14868. https://doi.org/10.1073/pnas.95.25.14863
Raychaudhuri S, Stuart JM, Altman RB (2000) Principal components analysis to summarize microarray experiments: application to sporulation time series. Pac Symp Biocomput 2000:455–466
Maitra S, Yan J (2008) Principle component analysis and partial least squares: two dimension reduction techniques for regression. In: 2008 Casualty actuarial society discussion paper program–applying multivariate statistical models. Casualty Actuarial Society, Quebec, pp 79–90
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A 98(9):5116–5121. https://doi.org/10.1073/pnas.091062498
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Papadimitropoulos, ME.P., Vasilopoulou, C.G., Maga-Nteve, C., Klapa, M.I. (2018). Untargeted GC-MS Metabolomics. In: Theodoridis, G., Gika, H., Wilson, I. (eds) Metabolic Profiling. Methods in Molecular Biology, vol 1738. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7643-0_9
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7643-0_9
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7642-3
Online ISBN: 978-1-4939-7643-0
eBook Packages: Springer Protocols