Abstract
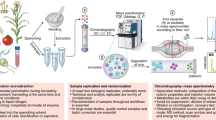
Metabolomics based on mass spectrometry can provide quantitative and qualitative information of the pool of metabolites (metabolome) present intracellularly or extracellularly in a given biological system. A typical metabolomics workflow requires several key steps such as quick and robust sample preparations with quenching of metabolism, chemical derivatization if needed, instrumental measurement, data-processing with/without database information and further statistical analysis and interpretation. Here, we introduce general metabolomics workflows for global and targeted analyses using gas chromatography or liquid chromatography coupled with mass spectrometers.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Nicholson JK, Lindon JC, Holmes E (1999) “Metabonomics”: understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica 29:1181–1189. https://doi.org/10.1080/004982599238047
Fiehn O (2002) Metabolomics – the link between genotypes and phenotypes. Plant Mol Biol 48:155–171. https://doi.org/10.1023/A:1013713905833
Zhang A, Sun H, Wang P et al (2012) Modern analytical techniques in metabolomics analysis. Analyst 137:293–300. https://doi.org/10.1039/C1AN15605E
Kyle JE, Casey CP, Stratton KG et al (2016) Comparing identified and statistically significant lipids and polar metabolites in 15-year old serum and dried blood spot samples for longitudinal studies. Rapid Commun Mass Spectrom. https://doi.org/10.1002/rcm.7808
Bennett BD, Yuan J, Kimball EH, Rabinowitz JD (2008) Absolute quantitation of intracellular metabolite concentrations by an isotope ratio-based approach. Nat Protoc 3:1299–1311. https://doi.org/10.1038/nprot.2008.107
Nakayasu ES, Nicora CD, Sims AC, et al (2016) MPLEx: a robust and universal protocol for single-sample integrative proteomic, metabolomic, and lipidomic analyses. mSystems 1(3). pii: e00043-16
Lai Z, Fiehn O (2016) Mass spectral fragmentation of trimethylsilylated small molecules. Mass Spectrom Rev. https://doi.org/10.1002/mas.21518
Coble JB, Fraga CG (2014) Comparative evaluation of preprocessing freeware on chromatography/mass spectrometry data for signature discovery. J Chromatogr A 1358:155–164. https://doi.org/10.1016/j.chroma.2014.06.100
Niu W, Knight E, Xia Q, McGarvey BD (2014) Comparative evaluation of eight software programs for alignment of gas chromatography–mass spectrometry chromatograms in metabolomics experiments. J Chromatogr A 1374:199–206. https://doi.org/10.1016/j.chroma.2014.11.005
Sumner LW, Amberg A, Barrett D et al (2007) Proposed minimum reporting standards for chemical analysis. Metabolomics 3:211–221. https://doi.org/10.1007/s11306-007-0082-2
Acknowledgment
This chapter is based upon work supported by the US Department of Energy, Office of Science, Office of Biological and Environmental Research, Genomic Science program, under Award Number DE- SC0008744. This is also enabled by the Environmental Molecular Sciences Laboratory (EMSL), a national scientific user facility sponsored by the US Department of Energy’s (DOE) Office of Biological and Environmental Research (OBER), and located at the Pacific Northwest National Laboratory (PNNL). PNNL is a multiprogram national laboratory operated by Battelle for the DOE under Contract DE-AC05-76RLO 1830.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Kim, YM., Heyman, H.M. (2018). Mass Spectrometry-Based Metabolomics. In: de Vries, R., Tsang, A., Grigoriev, I. (eds) Fungal Genomics. Methods in Molecular Biology, vol 1775. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7804-5_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7804-5_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7803-8
Online ISBN: 978-1-4939-7804-5
eBook Packages: Springer Protocols