Abstract
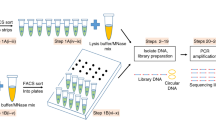
MNase-seq allows the genome-wide examination of the nucleosome landscape by determination of nucleosome positioning and occupancy. Typically, native or formaldehyde fixed chromatin is subjected to digestion by micrococcal nuclease (MNase), which degrades linker DNA and yields mainly mono-nucleosomes. The resulting material can be processed directly or can be subjected to an optional chromatin immunoprecipitation step (MNase-ChIP-seq). De-crosslinked and purified DNA is then subjected to next-generation sequencing. The protocol presented here has been tailored for the analysis of nucleosome landscape in the malaria parasite, Plasmodium falciparum, but most steps are directly applicable to other cell types. We also discuss general considerations for experimental design and computational analysis, which are crucial for accurate investigation of the nucleosome landscape.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lieleg C, Krietenstein N, Walker M, Korber P (2015) Nucleosome positioning in yeasts: methods, maps, and mechanisms. Chromosoma 124:131–151
Hughes AL, Rando OJ (2014) Mechanisms underlying nucleosome positioning in vivo. Annu Rev Biophys 43:41–63
Clark RJ, Felsenfeld G (1971) Structure of chromatin. Nat New Biol 229:101–106
Heins JN, Suriano JR, Taniuchi H, Anfinsen CB (1967) Characterization of a nuclease produced by Staphylococcus aureus. J Biol Chem 242:1016–1020
Cui K, Zhao K (2012) Genome-wide approaches to determining nucleosome occupancy in metazoans using MNase-Seq. Methods Mol Biol 833:413–419
Platt JL, Kent NA, Harwood AJ, Kimmel AR (2013) Analysis of chromatin organization by deep sequencing technologies. Methods Mol Biol 983:173–183
Ishii H, Kadonaga JT, Ren B (2015) MPE-seq, a new method for the genome-wide analysis of chromatin structure. Proc Natl Acad Sci U S A 112:E3457–E3465
Rhee HS, Bataille AR, Zhang L, Pugh BF (2014) Subnucleosomal structures and nucleosome asymmetry across a genome. Cell 159:1377–1388
Brogaard K, Xi L, Wang JP, Widom J (2012) A map of nucleosome positions in yeast at base-pair resolution. Nature 486:496–501
Allan J, Fraser RM, Owen-Hughes T, Keszenman-Pereyra D (2012) Micrococcal nuclease does not substantially bias nucleosome mapping. J Mol Biol 417:152–164
Kensche PR, Hoeijmakers WA, Toenhake CG, Bras M, Chappell L, Berriman M, Bartfai R (2016) The nucleosome landscape of Plasmodium falciparum reveals chromatin architecture and dynamics of regulatory sequences. Nucleic Acids Res 44:2110–2124
Orlando V, Strutt H, Paro R (1997) Analysis of chromatin structure by in vivo formaldehyde cross-linking. Methods 11:205–214
Mieczkowski J, Cook A, Bowman SK, Mueller B, Alver BH, Kundu S, Deaton AM, Urban JA, Larschan E, Park PJ, Kingston RE, Tolstorukov MY (2016) MNase titration reveals differences between nucleosome occupancy and chromatin accessibility. Nat Commun 7:11485
Rizzo JM, Bard JE, Buck MJ (2012) Standardized collection of MNase-seq experiments enables unbiased dataset comparisons. BMC Mol Biol 13:15
Horz W, Altenburger W (1981) Sequence specific cleavage of DNA by micrococcal nuclease. Nucleic Acids Res 9:2643–2658
Kaplan N, Hughes TR, Lieb JD, Widom J, Segal E (2010) Contribution of histone sequence preferences to nucleosome organization: proposed definitions and methodology. Genome Biol 11:140
Nikitina T, Wang D, Gomberg M, Grigoryev SA, Zhurkin VB (2013) Combined micrococcal nuclease and exonuclease III digestion reveals precise positions of the nucleosome core/linker junctions: implications for high-resolution nucleosome mapping. J Mol Biol 425:1946–1960
Meyer CA, Liu XS (2014) Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. Nat Rev Genet 15:709–721
Oyola SO, Otto TD, Gu Y, Maslen G, Manske M, Campino S, Turner DJ, Macinnis B, Kwiatkowski DP, Swerdlow HP, Quail MA (2012) Optimizing Illumina next-generation sequencing library preparation for extremely AT-biased genomes. BMC Genomics 13:1
Hoeijmakers WA, Bartfai R, Francoijs KJ, Stunnenberg HG (2010) Linear amplification for deep sequencing. Nat Protoc 6:1026–1036
Kivioja T, Vaharautio A, Karlsson K, Bonke M, Enge M, Linnarsson S, Taipale J (2012) Counting absolute numbers of molecules using unique molecular identifiers. Nat Methods 9:72–74
Quintales L, Vazquez E, Antequera F (2014) Comparative analysis of methods for genome-wide nucleosome cartography. Brief Bioinform 16:576–587
Teif VB (2015) Nucleosome positioning: resources and tools online. Brief Bioinform 17:745–757
Henikoff JG, Belsky JA, Krassovsky K, MacAlpine DM, Henikoff S (2011) Epigenome characterization at single base-pair resolution. Proc Natl Acad Sci U S A 108:18318–18323
Hoeijmakers WA, Bartfai R, Francoijs KJ, Stunnenberg HG (2011) Linear amplification for deep sequencing. Nat Protoc 6:1026–1036
Acknowledgments
The research leading to this protocol has received funding from The Netherlands Organization for Scientific Research (NWO-Vidi 864.11.007 to R.B.) and The National Institutes of Health (EuPathDB-Driving Biological Project subaward # 553539 to R.B.). We would like to acknowledge Dr. Kensche for valuable input and discussions concerning data analysis and Christa Toenhake for proofreading of the manuscript. Furthermore, we would like to thank our colleagues at the Department of Molecular Biology, the Department of Molecular Developmental Biology, and the Department of Medical Microbiology of Radboud University and St. Radboud UMC for support and advice.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Hoeijmakers, W.A.M., Bártfai, R. (2018). Characterization of the Nucleosome Landscape by Micrococcal Nuclease-Sequencing (MNase-seq). In: Visa, N., Jordán-Pla, A. (eds) Chromatin Immunoprecipitation. Methods in Molecular Biology, vol 1689. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7380-4_8
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7380-4_8
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7379-8
Online ISBN: 978-1-4939-7380-4
eBook Packages: Springer Protocols