Abstract
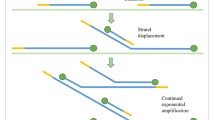
The here described method of isothermal whole genome amplification (iWGA) uses a Phi29 DNA polymerase-based kit (Illustra GenomiPhi V2 DNA Amplification Kit) that amplifies minute quantities of DNA by multiple strand displacement upon random hexamer primer binding. Starting from genomic DNA or single cells this amplification yields up to 5 μg of iWGA product with fragment lengths of 10 kb and longer. As this amplification lacks the need of fragmenting DNA, its products are well suited for many downstream applications (e.g. sequencing and DNA profiling). On the contrary, degraded DNA samples are not supported by the nature of the amplification and are not well suited.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Blanco L, Bernad A, Lazaro JM, Martin G, Garmendia C, Salas M (1989) Highly efficient DNA synthesis by the phage phi 29 DNA polymerase. Symmetrical mode of DNA replication. J Biol Chem 264(15):8935–8940
Esteban JA, Salas M, Blanco L (1993) Fidelity of phi 29 DNA polymerase. Comparison between protein-primed initiation and DNA polymerization. J Biol Chem 268(4):2719–2726
Paez JG, Lin M, Beroukhim R, Lee JC, Zhao X, Richter DJ, Gabriel S, Herman P, Sasaki H, Altshuler D, Li C, Meyerson M, Sellers WR (2004) Genome coverage and sequence fidelity of phi29 polymerase-based multiple strand displacement whole genome amplification. Nucleic Acids Res 32(9), e71. doi:10.1093/nar/gnh069
Dean FB, Nelson JR, Giesler TL, Lasken RS (2001) Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res 11(6):1095–1099. doi:10.1101/gr.180501
Lizardi PM, Huang X, Zhu Z, Bray-Ward P, Thomas DC, Ward DC (1998) Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat Genet 19(3):225–232. doi:10.1038/898
Spits C, Le Caignec C, De Rycke M, Van Haute L, Van Steirteghem A, Liebaers I, Sermon K (2006) Whole-genome multiple displacement amplification from single cells. Nat Protoc 1(4):1965–1970. doi:10.1038/nprot.2006.326
Kroneis T, Geigl JB, El-Heliebi A, Auer M, Ulz P, Schwarzbraun T, Dohr G, Sedlmayr P (2011) Combined Molecular Genetic and Cytogenetic Analysis from Single Cells after Isothermal Whole-Genome Amplification. Clin Chem 57(7):1032–1041
Konings P, Vanneste E, Jackmaert S, Ampe M, Verbeke G, Moreau Y, Vermeesch JR, Voet T (2012) Microarray analysis of copy number variation in single cells. Nat Protoc 7(2):281–310. doi:10.1038/nprot.2011.426
Cheng J, Vanneste E, Konings P, Voet T, Vermeesch JR, Moreau Y (2011) Single-cell copy number variation detection. Genome Biol 12(8):R80. doi:10.1186/gb-2011-12-8-r80
Van der Aa N, Cheng J, Mateiu L, Zamani Esteki M, Kumar P, Dimitriadou E, Vanneste E, Moreau Y, Vermeesch JR, Voet T (2013) Genome-wide copy number profiling of single cells in S-phase reveals DNA-replication domains. Nucleic Acids Res 41(6), e66. doi:10.1093/nar/gks1352
Lage JM, Leamon JH, Pejovic T, Hamann S, Lacey M, Dillon D, Segraves R, Vossbrinck B, Gonzalez A, Pinkel D, Albertson DG, Costa J, Lizardi PM (2003) Whole genome analysis of genetic alterations in small DNA samples using hyperbranched strand displacement amplification and array-CGH. Genome Res 13(2):294–307. doi:10.1101/gr.377203
Voet T, Kumar P, Van Loo P, Cooke SL, Marshall J, Lin ML, Zamani Esteki M, Van der Aa N, Mateiu L, McBride DJ, Bignell GR, McLaren S, Teague J, Butler A, Raine K, Stebbings LA, Quail MA, D'Hooghe T, Moreau Y, Futreal PA, Stratton MR, Vermeesch JR, Campbell PJ (2013) Single-cell paired-end genome sequencing reveals structural variation per cell cycle. Nucleic Acids Res 41(12):6119–6138. doi:10.1093/nar/gkt345
Maciejewska A, Jakubowska J, Pawlowski R (2013) Whole genome amplification of degraded and nondegraded DNA for forensic purposes. Int J Legal Med 127(2):309–319. doi:10.1007/s00414-012-0764-9
Maciejewska A, Jakubowska J, Pawlowski R (2014) Different whole-genome amplification methods as a preamplification tool in Y-chromosome Loci analysis. Am J Forensic Med Pathol 35(2):140–144. doi:10.1097/PAF.0000000000000093
Kumar G, Garnova E, Reagin M, Vidali A (2008) Improved multiple displacement amplification with phi29 DNA polymerase for genotyping of single human cells. Biotechniques 44(7):879–890. doi:10.2144/000112755
Mohlendick B, Stoecklein NH (2014) Analysis of copy-number alterations in single cells using microarray-based comparative genomic hybridization (aCGH). Curr Protoc Cell Biol 65:22.19.1–22.19.23. doi:10.1002/0471143030.cb2219s65
Acknowledgement
This work was supported by the EU SAFE Network of Excellence (LSHB-CT-2004-503243, EU 6th Framework Package), the Austrian Federal Ministry for Transport, Innovation and Technology together with the Austrian Science Fund (grant number TRP 17-B18) and the County of Styria, Austria.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Kroneis, T., El-Heliebi, A. (2015). Whole Genome Amplification by Isothermal Multiple Strand Displacement Using Phi29 DNA Polymerase. In: Kroneis, T. (eds) Whole Genome Amplification. Methods in Molecular Biology, vol 1347. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2990-0_8
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2990-0_8
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2989-4
Online ISBN: 978-1-4939-2990-0
eBook Packages: Springer Protocols