Abstract
Several strategies have been developed to generate targeted gene disruptions in zebrafish.
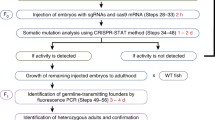
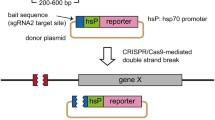
Here we developed a simple targeted gene inactivation strategy in zebrafish using a clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein (Cas) system. By injecting two simple in vitro-synthesized components [Cas9 mRNA and single guide (sgRNA)] into one-cell-stage embryos, mutations of the target gene could be efficiently generated. We used a codon-optimized version of Cas9 to improve its translation efficiency in zebrafish. In addition, we designed a cloning-free strategy to facilitate the synthesis of sgRNA. The system allows biallelic inactivation of multiple genes simultaneously by co-injecting a mix of sgRNAs with a single Cas9 construct. This flexible strategy of gene inactivation provides an efficient way to interrogate gene functions and genetic interactions in zebrafish.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Barnes DE (2001) Non-homologous end joining as a mechanism of DNA repair. Curr Biol 11:R455–R457
Mani M, Smith J, Kandavelou K et al (2005) Binding of two zinc finger nuclease monomers to two specific sites is required for effective double-strand DNA cleavage. Biochem Biophys Res Commun 334:1191–1197
Urnov FD, Miller JC, Lee YL et al (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651
Miller JC, Tan S, Qiao G et al (2011) A TALE nuclease architecture for efficient genome editing. Nat Biotechnol 29:143–148
Boch J, Scholze H, Schornack S et al (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326:1509–1512
Christian M, Cermak T, Doyle EL et al (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186:757–761
Jinek M, Chylinski K, Fonfara I et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Cong L, Ran FA, Cox D et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Mali P, Yang L, Esvelt KM et al (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Jinek M, East A, Cheng A et al (2013) RNA-programmed genome editing in human cells. Elife 2:e00471
Doyon Y, McCammon JM, Miller JC et al (2008) Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat Biotechnol 26:702–708
Meng X, Noyes MB, Zhu LJ et al (2008) Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat Biotechnol 26:695–701
Huang P, Xiao A, Zhou M et al (2011) Heritable gene targeting in zebrafish using customized TALENs. Nat Biotechnol 29:699–700
Sander JD, Cade L, Khayter C et al (2011) Targeted gene disruption in somatic zebrafish cells using engineered TALENs. Nat Biotechnol 29:697–698
Hruscha A, Krawitz P, Rechenberg A et al (2013) Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development 140:4982–4987
Hwang WY, Fu Y, Reyon D et al (2013) Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 31:227–229
Jao LE, Wente SR, Chen W (2013) Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc Natl Acad Sci U S A 110:13904–13909
Chang N, Sun C, Gao L et al (2013) Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res 23:465–472
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157:1262–1278
Brouns SJ, Jore MM, Lundgren M et al (2008) Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321:960–964
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845
Nakata A, Amemura M, Makino K (1989) Unusual nucleotide arrangement with repeated sequences in the Escherichia coli K-12 chromosome. J Bacteriol 171:3553–3556
Jansen R, Embden JD, Gaastra W et al (2002) Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 43:1565–1575
Mojica FJ, Diez-Villasenor C, Garcia-Martinez J et al (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60:174–182
Pourcel C, Salvignol G, Vergnaud G (2005) CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 151:653–663
Bolotin A, Quinquis B, Sorokin A et al (2005) Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151:2551–2561
Barrangou R, Fremaux C, Deveau H et al (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Deltcheva E, Chylinski K, Sharma CM et al (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471:602–607
Makarova KS, Haft DH, Barrangou R et al (2011) Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 9:467–477
Ran FA, Hsu PD, Wright J et al (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308
Gagnon JA, Valen E, Thyme SB et al (2014) Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS One 9:e98186
Guschin DY, Waite AJ, Katibah GE et al (2010) A rapid and general assay for monitoring endogenous gene modification. Methods Mol Biol 649:247–256
Hill JT, Demarest BL, Bisgrove BW et al (2014) Poly peak parser: method and software for identification of unknown indels using Sanger sequencing of polymerase chain reaction products. Dev Dyn 243:1632
Varshney GK, Pei W, LaFave MC et al (in press) High-throughput gene targeting and phenotyping in zebrafish using CRISPR/Cas9. Genome Res
Acknowledgements
We thank Shawn Burgess for communicating unpublished data. This work was supported by R01DK088686 (to W.C.) from the NIH.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Yin, L., Jao, LE., Chen, W. (2015). Generation of Targeted Mutations in Zebrafish Using the CRISPR/Cas System. In: Fiedler, L. (eds) VEGF Signaling. Methods in Molecular Biology, vol 1332. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2917-7_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2917-7_16
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2916-0
Online ISBN: 978-1-4939-2917-7
eBook Packages: Springer Protocols