Abstract
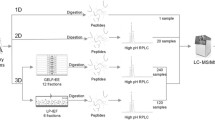
Urine has garnered tremendous interest over the past decade as a potential source of protein biomarkers for various pathologies. However, due to its low protein concentration and the presence of interfering compounds, urine constitutes a challenging analyte in proteomics. In the context of a project aimed at the discovery and evaluation of new candidate biomarkers of bladder cancer in urine, our laboratory has implemented and evaluated an array of preparation techniques for urinary proteome analysis. We present here the protocol that, in our hands, yielded the best overall proteome coverage with the lowest analytical effort. It begins with protein precipitation using trichloroacetic acid, in solution digestion and RP-C18 cartridge desalting of the resulting peptides mixture, and is followed by peptide fractionation by gel-free isoelectric focusing, and nano-LC-MS/MS for database compilation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Adachi J, Kumar C, Zhang Y et al (2006) The human urinary proteome contains more than 1500 proteins, including a large proportion of membrane proteins. Genome Biol 7:R80
Kentsis A, Monigatti F, Dorff K et al (2009) Urine proteomics for profiling of human disease using high accuracy mass spectrometry. Proteomics Clin Appl 3:1052–1061
Marimuthu A, O’Meally RN, Chaerkady R et al (2011) A comprehensive map of the human urinary proteome. J Proteome Res 10:2734–2743
Kim KH, Moon MH (2009) High speed two-dimensional protein separation without gel by isoelectric focusing-asymmetrical flow field flow fractionation: application to urinary proteome. J Proteome Res 8:4272–4278
Smith G, Barratt D, Rowlinson R et al (2005) Development of a high-throughput method for preparing human urine for two-dimensional electrophoresis. Proteomics 5:2315–2318
Candiano G, Santucci L, Petretto A et al (2010) 2D-electrophoresis and the urine proteome map: where do we stand? J Proteomics 73:829–844
Pasa-Tolić L, Lipton MS, Masselon CD et al (2002) Gene expression profiling using advanced mass spectrometric approaches. J Mass Spectrom 37:1185–1198
Court M, Selevsek N, Matondo M et al (2011) Toward a standardized urine proteome analysis methodology. Proteomics 11:1160–1171
Elias JE, Gygi SP (2007) Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat Methods 4:207–214
Dupierris V, Masselon C, Court M et al (2009) A toolbox for validation of mass spectrometry peptides identification and generation of database: IRMa. Bioinformatics 25:1980–1981
Monroe ME, Tolić N, Jaitly N et al (2007) VIPER: an advanced software package to support high-throughput LC-MS peptide identification. Bioinformatics 23:2021–2023
Acknowledgements
Dr. Dimitri Vordos, Edwige Lopes, Yves Allory, Hopital Mondor in Paris, and Nuria Mallats CNIO in Madrid are gratefully acknowledged for their assistance in specimen collection. The authors thank Dr. Christophe Bruley and Veronique Dupierris, CEA Grenoble, for the IRMa software and his help with AMT tag database compilation, and Madalen LeGorrec, CEA Grenoble, for the protein dosage and the pools constitution. The authors thank Dr. Elodie Duriez from CRP-Santé in Luxembourg for her comments and helpful discussion. The authors also express gratitude to all the members of the DECanBio consortium for their help and advice. This work was funded through the EU FP7 program DECanBio (www.decanbio.eu) [Grant Agreement #201333]. The Decon-2LS and VIPER software used for data processing were provided by the W. R. Wiley Environmental Molecular Science Laboratory, a national scientific user facility sponsored by the US Department of Energy’s Office of Biological and Environmental Research, located at PNNL. PNNL is operated by Battelle Memorial Institute for the US Department of Energy under contract DE-AC05-76RL0 1830.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Court, M., Garin, J., Masselon, C.D. (2015). Urine Sample Preparation and Fractionation for Global Proteome Profiling by LC-MS. In: Vlahou, A., Makridakis, M. (eds) Clinical Proteomics. Methods in Molecular Biology, vol 1243. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1872-0_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1872-0_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1871-3
Online ISBN: 978-1-4939-1872-0
eBook Packages: Springer Protocols