Abstract
Biomarkers are measurable changes associated with the disease. Urine can reflect the changes of the body while blood is under control of the homeostatic mechanisms; thus, urine is considered an important source for early and sensitive disease biomarker discovery. A comprehensive profile of the urinary proteome will provide a basic understanding of urinary proteins. In this paper, we present an in-depth analysis of the urinary proteome based on different separation strategies, including direct one dimensional liquid chromatography–tandem mass spectrometry (LC/MS/MS), two dimensional LC/MS/MS, and gel-eluted liquid fraction entrapment electrophoresis/liquid-phase isoelectric focusing followed by two dimensional LC/MS/MS. A total of 6085 proteins were identified in healthy urine, of which 2001 were not reported in previous studies and the concentrations of 2571 proteins were estimated (spanning a magnitude of 106) with an intensity-based absolute quantification algorithm. The urinary proteins were annotated by their tissue distribution. Detailed information can be accessed at the “Human Urine Proteome Database” (www.urimarker.com/urine).
Similar content being viewed by others
Introduction
Urine is associated with glomerular filtration, tubular reabsorption and secretion1. Biomarkers are measurable changes associated with the disease2. Because urine can accumulate changes from the body2, 3, it is considered to be one of the most attractive sources for early and sensitive biomarker discovery. Urinary proteomic studies have identified many candidate biomarkers for various urogenital diseases, such as acute kidney injury, bladder cancer and diabetic nephropathy4,5,6. As urinary proteins are composed largely of filtered plasma proteins, the urinary proteome is also considered to be valuable for detecting a broad range of complex disorders, such as encephalopathy, heart failure and intestinal ischemia7,8,9.
In the biomarker discovery process, it is essential to comprehensively profile the normal urinary proteome as a baseline reference. With the rapid development of mass spectrometry (MS), larger numbers of urinary proteins were identified by various strategies. In 2001, Patterson et al. first identified 124 urinary proteins10, 11. In 2005, Sun et al. identified 226 proteins in normal urine with quality control LC/MS/MS data3. In 2006, Adachi et al. reported the first urinary proteome result (1543 proteins) from high resolution mass spectrometry12. In 2011, 1823 urinary proteins were identified by high resolution MS and MS/MS13. Many efforts have been made to identify more urinary proteins in recent years14,15,16. Currently, the human urine PeptideAtlas database contains a total of 23,739 peptides corresponding to 2487 proteins17.
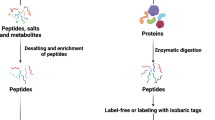
In 2014, two large-scale MS-based drafts of the human proteome identified 17,294 and 18,097 human gene products from 30 and 44 tissues and body fluids, respectively18, 19. In each study, the number of identified proteins was quite large and even approached the number of protein-coding genes in the complete human genome analysis20. Compared with the depth of the human tissue proteome, the urinary proteome has been relatively less studied. We are curious about how many proteins could be identified in human urinary proteome. Therefore, we performed an in-depth urinary proteomic analysis using normal human urine samples. And to achieve maximal urinary proteome coverage one-, two- and three-dimensional separation strategies (Fig. 1) were employed in this study. By in-depth analysis, a readily obtainable source for the human urinary proteome, “Human Urinary Proteome Database” could be provided. The comparison of three separation strategies could provide detailed information about the potential application of different separation methods. The detailed workflow was as followings: In one-dimensional (1D) separation, digested urinary peptides were directly analyzed by 1D liquid chromatography-tandem mass spectrometry (LC/MS/MS). In two-dimensional (2D) separation, urinary peptides were fractionated by offline high-pH reverse-phase liquid chromatography (RPLC) prior to analysis by 1DLC/MS/MS. In three-dimensional (3D) separation, urinary proteins were first fractionated by gel-eluted liquid fraction entrapment electrophoresis (GELFrEE) or liquid-phase isoelectric focusing (LP-IEF) and urinary peptides digested from GELFrEE/LP-IEF fractions were fractionated by RPLC as performed for 2D separation and finally analyzed by 1DLC/MS/MS. In total, 383 LC/MS/MS runs were analyzed by hybrid quadrupole-time-of-flight mass spectrometry (TripleTOF 5600).
The workflow of urinary proteome analysis. Pooled urine from 24 humans was analyzed using three separation strategies. 1D: Urinary peptides were directly analyzed via 1DLC/MS/MS without fractionation. 2D: Urinary peptides were analyzed via offline RPLC and 1DLC/MS/MS. 3D: Urinary proteins were first fractionated by GELFrEE/LP-IEF prior to offline RPLC. A total of 383 fractions were analyzed by LC/MS/MS using high-resolution TripleTOF 5600 MS. A urine proteome database was then constructed based on bioinformatics analyses.
Results and Discussion
Comprehensive identification of urinary proteome
In this study, pooled urine samples were used to establish a large database of urinary proteins. The following filters were used to select the final protein identification list (1). The FDR at the protein level was set to <1%, and (2) each protein should include at least two unique peptides. When identified peptides were shared between two proteins, they were combined and reported as one protein group. The results from 1DLC, 2DLC and 3DLC yielded average FDRs of 0.10%, 0.26% and 1% at the spectrum, peptide and protein levels, respectively (Supplemental File 1). Then the datasets were combined together with Scaffold perSPECtives.
In 1D analysis, 808 protein groups were identified in three technical replicates, and the protein-overlapping rate was 86.3%, indicating the superior reproducibility of LC/MS analysis. In 2D analysis, a total of 3162 protein groups were identified. In 3D analysis, urinary proteins were first separated by GELFrEE/LP-IEF (Fig. 2A,B). GELFrEE enables mass range proteome separations based on molecular weight (MW), and IEF fractionates proteins according to isoelectric point (pI)21, 22. The GELFrEE and LP-IEF fractions were then further separated by RPLC, and a total of 6085 protein groups were identified. The overlap among the proteins identified in the 1D, 2D and 3D analyses is displayed in Fig. 2C. Almost all proteins from the 1D and 2D analyses were included in the 3D results except for 9 and 15 proteins from the 1D and 2D results, respectively. The possible reasons why these proteins cannot be identified in the 3D methods were still unknown. Maybe these proteins were lost during 2D or 3D separate by high pH RPLC or IEF/GELFrEE. It may be also as a result of the random sampling of DDA detection modes. Therefore, we are not sure whether these proteins were false positive identification or not. Then these proteins (Supplemental File 2) were removed from the subsequent analysis to ensure data accuracy and reliability. Thus, the whole urine proteome dataset eventually contained 6085 protein groups (Supplemental Table 1).
The results from three separation strategies. (A) A Coomassie-stained Bis-Tris gel image of 12 GELFrEE fractions over a broad mass range. (B) Coomassie-stained Bis-Tris gel image of 10 LP-IEF fractions over a pI range from 3 to 10. (C) Venn diagram of proteins identified by three separation strategies. (D) Venn diagram of proteins identified from this study as well as previous urine and exosome proteome studies. (E) Comparative analysis of the urine, kidney and plasma proteome.
Several studies have been conducted to characterize the normal human urinary proteome. Table 1 summarizes the current largest-scale studies of human urine and urinary exosomes using high-resolution MS12,13,14,15,16, 23,24,25. The protein accessions in each dataset were mapped to the corresponding gene IDs26, 27. Total nine large-scale urinary and exosome proteomic analyses were performed in recent years. When all of the data from these nine studies were combined, a total of 8021 gene products were detected in the human urinary proteome (Supplemental Table 2). When comparing previous data with our results (Fig. 2D), total 2001 gene products were uniquely identified in this study. The possible reasons of differences in urine proteome between different studies may be genetic factors, individual variations, different separate methods and MS preference.
Urinary proteins, which are considered to represent the protein composition of the output of the kidneys28, are primarily composed of proteins derived from plasma filtration and urinary tract system secretion. A comparative analysis of the urine, plasma and kidney proteome would provide a more concrete link to determine how many plasma- and kidney-related proteins could be detected in urine. The PeptideAtlas builds yielded 3553 and 4005 non-redundant proteins at 1% FDR for the plasma and kidney proteomes29. In contrast, a total of 2940 (47.7%) and 3238 (52.6%) of the gene products identified in this urinary proteome study were common to the gene products (Fig. 2E) that were reported in the plasma (81.1%) and kidney proteomes (75.6%), respectively. According to previous report, approximately 30% of urinary proteins originate from the plasma proteins, whereas 70% comes from the kidney and the urinary tract30. From our study, maybe the difference between plasma and urine is smaller than expected. And it might be possible that more common proteins will be identified with the development of MS in the future. By comparison with kidney proteome, we want to show the overlap between urine and kidney proteome. The large overlap may give evidences that urine can better reflect the functions of kidney.
Quantitative analysis of urinary proteins
Quantitation of urinary proteins will improve our understanding of the urinary proteome and will facilitate the development of urinary biomarkers. Accordingly, we aimed to quantify each protein using the iBAQ algorithm, which provides a rough indication of actual protein levels31. In 3D analysis, equal amounts of protein from each GELFrEE/LP-IEF fraction were used for LC/MS/MS analysis, which thus cannot provide an accurate quantitative analysis. Therefore, data from the 2D strategy were used, and a total of 2571 proteins were quantified with the iBAQ algorithm. The dynamic range of relative abundance spanned six orders of magnitude (Fig. 3A), which was consistent with previous observations32. Considering that more than 3000 proteins in the 3D analysis were not quantified, the dynamic range is expected to be even bigger. In the present analysis, serum albumin and uromodulin were the most abundant urinary proteins.
Quantitative analysis of urinary proteins by the iBAQ method. (A) The relative expression and concentrations of 2,571 proteins in the 2D analysis were estimated by iBAQ. The left y-axis represents relative abundance, and the right y axis represents estimated concentration (pg/mL). (1) ALB: albumin; UROM: uromodulin, the two most abundant proteins. (2) RARS: arginine-tRNA ligase, the least abundant protein in 2D analysis; (3) RNASE 6: ribonuclease K6, the least abundant protein in 1D analysis. (B) Correlation plot between estimated concentrations and immunoassays results.
The average concentration of urinary albumin, which was one of the most easily detected urinary proteins, was approximately 2.2–3.3 µg/mL12, 33 in normal human urine. With the iBAQ value ratios, concentrations of the other 2570 proteins could subsequently be estimated (Supplemental Table 3). The estimated concentration of arginine-tRNA ligase (RARS), which had the lowest relative abundance in the 2D analysis, was 0.68~1.02 pg/mL. As the 2D results contained almost all of the proteins from the 1D separation, concentrations of the 753 proteins from the 1D analysis could be inferred. Among them, ribonuclease K6 (RNASE 6) was the least abundant protein in the 1D analysis with an estimated concentration of 5.58~8.37 pg/mL.
To evaluate the accuracy of estimated concentrations with the iBAQ algorithm and the corresponding application to other samples in different labs, the estimated concentrations were compared with the results from immunoassay screening in a previous urinary candidate biomarker study34. A total of 89 proteins were commonly evaluated in both studies (Fig. 3B and Supplemental Table 4, R2 = 0.59).
Functional annotation of three separation strategies
Functional annotations of urinary proteins based on the degree of analysis depth may be helpful in providing insight into the analysis approach difference in protein composition, reflecting pathophysiological states and determining suitable separation methods for some diseases. To analyze the protein identification data from the three separation strategies, 6085 proteins were divided into three groups as follows: Group 1D, proteins identified in 1D analysis (799 proteins); Group 2D, proteins identified in 2D analysis, excluding those identified in the 1D analysis (2362 proteins); and Group 3D, proteins identified in 3D analysis, excluding those identified in both the 1D and 2D analyses (2924 proteins).
IPA analysis was performed to provide insight into the functions of the three groups (Fig. 4 and Supplemental Table 5). Extracellular proteins and plasma membrane proteins were enriched in Group 1D (56%), as previously reported12. The most significant pathways in Group 1D were functionally similar to plasma components, such as inflammatory responses, coagulation and glucose metabolism. Acute phase response signaling, which is one of the top pathways for Group 1D, is a rapid inflammatory response that provides protection against some infections by nonspecific defenses. It consists of an increase in inflammatory factors (such as IL-1) and a change in the levels of several plasma proteins (such as ALB and APOA1/2). For example, alpha-1-acid glycoprotein 1 (ORM1), an extracellular protein, is involved in the acute phase response. Overexpression of ORM1 in urine was associated with acute pediatric appendicitis35. As Group 2D demonstrated considerable enrichment of intracellular proteins (58%), most of the pathways were involved in cellular signaling such as EIF2 Signaling and Regulation of eIF4 and p70S6K signaling. Proteins in Group 3D were also over-represented in the cytoplasm and nucleus (63%). Most of the canonical pathways in Group 3D were closely related to interleukin signaling.
Considering the above results, we assumed that proteins in the three groups were functionally different. If the purpose of research was to study basic physiological activities, such as cell movement and proliferation, maybe the urinary proteome can be analyzed without further separation in most cases. If aim at intracellular activities and functions of organs, maybe the in-depth analysis is necessary.
Landscape of proteins detected in urine
Previous studies reported that urine might reflect kidney function and identified some potential biomarkers of kidney disease36. According to previous analyses, the Human Urinary Proteome Database contains proteins localized in the glomeruli of nephron segments (Table 2) and molecules to detect injures to specific tubules of nephron segments (Table 3). Extracellular macromolecular laminin, type IV collagen α3α4α5, heparan sulfate proteoglycan agrin, and nidogen were the main components of the glomerular basement membrane (GBM)37 and could all be identified by the one-dimensional method without fractionation. Nephrin and podocin are both specifically expressed in the slit diaphragm, which is pivotal in maintaining the selective permeability of podocytes in the glomerular filtration barrier38. The cytoplasmic protein CD2-associated protein (CD2AP) localizes to the podocyte slit diaphragm where it has been shown to bind to nephrin and podocin39. The above three podocyte-related proteins could be identified in Group 2D. The fatty acid-binding proteins (FABPs) in Group 2D are a class of small intracellular proteins that bind long chain fatty acids. Liver-type FABP is mainly present in the cells of the proximal tubules, while heart-type FABP is predominantly localized in the distal tubules40. The above results showed that both glomerulus and tubules-related proteins could be found in the urine, which indicated that the urine proteome might reflect changes of kidney function.
Some tissue or serum biomarkers of kidney diseases could also be detected in our urine proteome database. For example, the phospholipase A2 receptor (PLA2R), a plasma membrane glycoprotein located on normal podocytes, was a major target antigen in idiopathic membranous nephropathy41. PLA2R could be detected in Groups 2D and 3D. Urokinase plasminogen activator surface receptor (uPAR) is a glycosylphosphatidylinoisitol -anchored three-domain protein and is expressed in human glomerular cells. Serum concentrations of soluble uPAR are significantly elevated in most subjects with primary focal segmental glomerulosclerosis (FSGS)42. If these tissue or serum biomarkers could be confirmed as urinary biomarkers, the human urinary proteome database would provide a convenient way to discover noninvasive urinary candidate biomarkers. In addition to kidney diseases, previous studies also reported that some other diseases, such as acute pancreatitis43, might possess urinary biomarkers. The human urinary proteome database provides brief information on known biomarkers for predicting various types of organ injury (Supplemental Table 6).
Moreover, these proteins detected in urine were annotated by their tissue distribution based on an integrated omics approach that involves quantitative transcriptomics and tissue microarray–based immunohistochemistry in previous studies44. The detailed annotation data of each protein were shown in the following database. The tissue with maximum numbers of highly expressed proteins detected in urine both at protein and mRNA levels was brain (Fig. 5A, Supplemental Figure 1). Other tissues with more highly expressed proteins were mostly digestive organs such as colon and stomach. As expected, more tissue-related proteins could be detected in Group 2D and 3D than in Group 1D (Fig. 5B).
Tissue distribution of urinary proteins at protein level. (A) Urinary proteome distributions across 44 tissues. The numbers in the bracket denote the number of highly expressed proteins of the tissue detected in urine. (B) The distribution of tissue-related proteins and the corresponding separation strategy for top ten tissues.
The Human Urinary Proteome Database
To provide a readily obtainable source for the human urinary proteome, the “Human Urinary Proteome Database” was constructed (Fig. 6) based on the above analyses. The database was constructed using open source technologies and is freely available at www.urimarker.com/urine. A total of 3048648 spectra, 68151 unique peptides and 6085 proteins are included, along with detailed information such as the protein name, accession number, peptide sequence, sequence coverage and unique peptide number.
An overview of the human urinary proteome database. (A) The protein level results include the unique peptide count, total peptide count and relative quantitation and estimated concentration. Proteins are linked to the UniProt website by clicking the accessions. (B) The peptide level results include peptide sequences and observed m/z values. (C) The database provides the experimental pI and MW distribution of all identified proteins. (D) The “MW-PI” section provides a succinct figure summarizing the theoretical MW and pI information for each protein. (E) Biomarker application of all identified proteins.
Each protein is featured with annotated data, including relative quantitative information, estimated concentrations, theoretical and experimental MW and pI. Remarkably, some high-abundance proteins were observed spanning multiple fractions in both the GELFrEE and LP-IEF separations. It is generally accepted that mass/pI deviation may occur due to the presence of fragments, protein polymers, isoforms, protein degradation, post-translational modifications and low focusing quality in the basic region of the immobilized pH-gradient strips, as well as due to the pI prediction algorithm used45,46,47,48. Moreover, a novel section labeled ‘MW-PI image’ provides a succinct figure indicating the significant MW and pI information for all of the identified urinary proteins, which might be helpful for generating a brief scan of proteins in a pI and MW range of interest. For biomarker studies, the “Biomarker” section also yields potential biomarkers for applications in diagnosis, disease progression and prognosis.
The Human Urinary Proteome Database serves as a reference repository for urinary proteins, as it offers the largest number of such proteins to date. All of the data retrieved from three separations not only detail the normal human urinary proteome but also categorize all proteins by different separation methods. Moreover, the database can be used for targeted proteomics that rely on the proper selection of peptides and transitions to guide the selection of proteotypic peptides for candidate proteins49.
Materials and Methods
Ethics statement
Prior to study enrollment, all of the healthy volunteers were given a verbal explanation of the study and each participant signed an informed consent document. The consent procedure and the research protocol were approved by the Medical Ethics Committee of Peking Union Medical College. All methods in this study were performed in accordance with the guidelines and regulations.
Experimental design and statistical rationale
Twenty-four healthy volunteers (38 ± 11 years old), including twelve males and twelve females, were enrolled. Exclusion criteria included the following conditions: menstrual bleeding, any prescription drug use and acute or chronic medical illness. The age, sex and smoking habits of the healthy subjects were recorded (Supplemental File 3).
After random urine collection, all of the samples were immediately centrifuged for 30 min at 3,500 g. After precipitate removal, urinary proteins were extracted by acetone precipitation. Lysis buffer (7 M urea, 2 M thiourea, 25 mM dithiothreitol and 50 mM Tris) was used to re-dissolve urinary proteins. The twenty-four urinary protein samples were pooled with equal amounts of protein into one sample for 1D, 2D and 3D analyses (Supplemental File 4).
GELFrEE and LP-IEF fractionation
For GELFrEE separation, urine samples were prepared using a protocol by Tran et al.45. Briefly, the pooled sample was fractionated in parallel using an eight-channel multiplexed GELFrEE 8100 Fractionation system (Protein Discovery, Knoxville, TN, USA). Application of 50 V for approximately 75 min and then 100 V for 105 min resulted in twelve GELFrEE fractions. The volume of each fraction was concentrated to approximately 125 μL using a SpeedVac Concentrator (Thermo Fisher Scientific, Asheville, NC, USA). Next, the samples underwent SDS removal using Pierce Detergent Removal Spin Columns (Pierce, Rockford, IL, USA).
For LP-IEF fractionation, urinary proteins were desalted and cleaned using Amicon Ultrafiltration devices with a 10-kDa molecular weight cutoff (Merck Millipore Inc., Billerica, MA, USA). Then, the desalted urinary proteins were focused (approximately 2.5 h at 1 W) using a ten-chamber Microrotofor LP-IEF system (Bio-Rad, Hemel Hempstead, UK). Ten IEF fractions were collected; few protein bands appeared in fractions 7–10. Then fractions 6–10 were pooled into one sample.
Protein digestion
Urinary proteins were digested with trypsin (Trypsin Gold, mass spec grade, Promega, WI, USA) using filter-aided sample preparation methods50. Proteins were loaded onto 10-kDa filter devices (Pall, Port Washington, NY, USA), and 8 M urea in 0.1 M Tris-HCl (pH 8.5) was added to wash the samples. The proteins were denatured by incubation with 50 mM dithiothreitol at 56 °C for 1 h and then alkylated in the dark for 45 min in 55 mM iodoacetamide. Trypsin was added (enzyme to protein ratio of 1:50), and the samples were incubated at 37 °C overnight. After digestion, the peptide mixtures were desalted on Oasis HLB cartridges (Waters, Milford, USA) and lyophilized for high-performance liquid chromatography separation.
Offline high-pH RPLC separation
In total, nineteen samples, including eighteen fractions that were separated by GELFrEE and LP-IEF and a pooled urine sample, were fractionated by offline high-pH RPLC columns (4.6 mm × 250 mm, C18, 3 μm; Waters Corp, Milford, USA). The samples were loaded onto the column in buffer A1 (10 mM NH4FA in H2O, pH = 10). The elution gradient was 5–30% buffer B1 (10 mM NH4FA in 90% acetonitrile, pH = 10; flow rate = 1 mL/min) for 60 min. The eluted peptides were collected at one fraction per minute. After lyophilization, the 60 fractions were re-suspended in 0.1% formic acid and concatenated into 20 fractions by combining fractions 1, 21, 41 and so on51.
Online LC-MS/MS analysis
Each sample was analyzed on a reverse-phase C18 self-packed capillary LC column (75 μm × 100 mm, 3 μm). The elution gradient was 5–30% buffer B2 (0.1% formic acid, 99.9% acetonitrile; flow rate = 0.3 μL/min) for 100 min. A TripleTOF 5600 coupled with UPLC system was used to analyze the sample, and the MS data were acquired in a high-sensitivity mode using the following parameters: 30 data-dependent MS/MS scans per full scan; full scans were acquired at a resolution of 40,000 and MS/MS scans were acquired at 20,000; rolling collision energy; charge state screening (including precursors with +2 to +4 charge state); dynamic exclusion (exclusion duration 15 s); MS/MS scan range of 250–1800 m/z; and scan time of 50 ms. For 1D separation, the pooled urine sample was analyzed with three technical replicates.
Data processing
The MS/MS data were processed using Mascot software (version 2.3.02, Matrix Science, London, UK) and searched against the SwissProt database (Homo sapiens, 20,267 sequences, 2013_07 version). The search allowed two missed cleavage sites in the trypsin digestion, cysteine carbamidomethylation was set as a fixed modification and both parent and fragment ion mass tolerances were set to 0.05 Da. Mascot search results were filtered using the decoy database method in Scaffold (version 4.3.2, Proteome Software Inc., Portland, OR). Peptide identifications were accepted if they could be shown to achieve a false discovery rate (FDR) of less than 1.0% by the Scaffold Local FDR algorithm. Protein identifications were accepted if they could be shown to achieve a FDR of less than 1.0% and contained at least 2 unique identified peptides. Protein probabilities were assigned by the Protein Prophet algorithm52. Proteins that contained similar peptides and could not be differentiated based on MS/MS analysis alone were grouped to satisfy the principles of parsimony. Proteins sharing significant peptide evidence were grouped into clusters.
Total 20 results from 1DLC, 2DLC and 3DLC (12 GELFrEE fractions and 6 LP-IEF fractions) were filtered by Scaffold with the above parameters and yielded average FDRs of 0.10%, 0.26% and 1% at the spectrum, peptide and protein levels, respectively. Then, the 20 datasets were combined together with Scaffold perSPECtives (version 2.0.4, Proteome Software Inc., Portland, OR).
To rank the relative abundance of different proteins, an intensity-based absolute quantification (iBAQ) algorithm was used53. The protein intensities were first computed by Progenesis LC–MS (version 2.6, Nonlinear Dynamics, UK)54 as the sum of all identified peptide intensities (maximum peak intensities of the peptide elution profile, including all peaks in the isotope cluster). The iBAQ result was obtained as the peptide intensities divided by the number of theoretically observable peptides of the protein (calculated by in silico protein digestion; all fully tryptic peptides between 6 and 30 amino acids were counted).
For functional analysis, ingenuity pathway analysis (IPA) software (Ingenuity Systems, www.ingenuity.com) was used to analyze cellular components, canonical gene pathways, functions and candidate biomarkers.
References
Casey, H. W., Ayers, K. N. & Robinson, F. In Pathology of laboratory animals 115–173 (Springer, 1978).
Gao, Y. Urine-an untapped goldmine for biomarker discovery? Science China. Life sciences 56, 1145–1146, doi:10.1007/s11427-013-4574-1 (2013).
Sun, W. et al. Human urine proteome analysis by three separation approaches. Proteomics 5, 4994–5001, doi:10.1002/pmic.200401334 (2005).
Ho, J. et al. Mass spectrometry-based proteomic analysis of urine in acute kidney injury following cardiopulmonary bypass: a nested case-control study. American journal of kidney diseases: the official journal of the National Kidney Foundation 53, 584–595, doi:10.1053/j.ajkd.2008.10.037 (2009).
Orenes-Pinero, E. et al. Searching urinary tumor markers for bladder cancer using a two-dimensional differential gel electrophoresis (2D-DIGE) approach. Journal of proteome research 6, 4440–4448, doi:10.1021/pr070368w (2007).
Zubiri, I. et al. Diabetic nephropathy induces changes in the proteome of human urinary exosomes as revealed by label-free comparative analysis. Journal of proteomics 96, 92–102, doi:10.1016/j.jprot.2013.10.037 (2014).
Thuijls, G. et al. Early diagnosis of intestinal ischemia using urinary and plasma fatty acid binding proteins. Annals of surgery 253, 303–308, doi:10.1097/SLA.0b013e318207a767 (2011).
Liu, L. et al. Evaluation of urinary S100B protein level and lactate/creatinine ratio for early diagnosis and prognostic prediction of neonatal hypoxic-ischemic encephalopathy. Neonatology 97, 41–44, doi:10.1159/000227292 (2010).
Chugh, S. et al. Pilot study identifying myosin heavy chain 7, desmin, insulin-like growth factor 7, and annexin A2 as circulating biomarkers of human heart failure. Proteomics 13, 2324–2334, doi:10.1002/pmic.201200455 (2013).
Davis, M. T. et al. Towards defining the urinary proteome using liquid chromatography-tandem mass spectrometry. II. Limitations of complex mixture analyses. Proteomics 1, 108–117, doi:10.1002/1615-9861(200101)1:1<108::AID-PROT108>3.0.CO;2-5 (2001).
Spahr, C. S. et al. Towards defining the urinary proteome using liquid chromatography-tandem mass spectrometry. I. Profiling an unfractionated tryptic digest. Proteomics 1, 93–107, doi:10.1002/1615-9861(200101)1:1<93::AID-PROT93>3.0.CO;2-3 (2001).
Adachi, J., Kumar, C., Zhang, Y., Olsen, J. V. & Mann, M. The human urinary proteome contains more than 1500 proteins, including a large proportion of membrane proteins. Genome biology 7, R80, doi:10.1186/gb-2006-7-9-R80 (2006).
Marimuthu, A. et al. A comprehensive map of the human urinary proteome. Journal of proteome research 10, 2734–2743, doi:10.1021/pr2003038 (2011).
Li, Q. R. et al. A comprehensive and non-prefractionation on the protein level approach for the human urinary proteome: touching phosphorylation in urine. Rapid communications in mass spectrometry: RCM 24, 823–832, doi:10.1002/rcm.4441 (2010).
Zheng, J., Liu, L., Wang, J. & Jin, Q. Urinary proteomic and non-prefractionation quantitative phosphoproteomic analysis during pregnancy and non-pregnancy. BMC genomics 14, 777, doi:10.1186/1471-2164-14-777 (2013).
Santucci, L. et al. From hundreds to thousands: Widening the normal human Urinome (1). Journal of proteomics 112C, 53–62, doi:10.1016/j.jprot.2014.07.021 (2014).
Desiere, F. et al. The PeptideAtlas project. Nucleic acids research 34, D655–658, doi:10.1093/nar/gkj040 (2006).
Kim, M. S. et al. A draft map of the human proteome. Nature 509, 575–581, doi:10.1038/nature13302 (2014).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587, doi:10.1038/nature13319 (2014).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74, doi:10.1038/nature11247 (2012).
Tran, J. C. & Doucette, A. A. Gel-eluted liquid fraction entrapment electrophoresis: an electrophoretic method for broad molecular weight range proteome separation. Analytical chemistry 80, 1568–1573, doi:10.1021/ac702197w (2008).
Ahmed, F. E. Sample preparation and fractionation for proteome analysis and cancer biomarker discovery by mass spectrometry. Journal of separation science 32, 771–798, doi:10.1002/jssc.200800622 (2009).
Gonzales, P. A. et al. Large-scale proteomics and phosphoproteomics of urinary exosomes. Journal of the American Society of Nephrology: JASN 20, 363–379, doi:10.1681/ASN.2008040406 (2009).
Wang, Z., Hill, S., Luther, J. M., Hachey, D. L. & Schey, K. L. Proteomic analysis of urine exosomes by multidimensional protein identification technology (MudPIT). Proteomics 12, 329–338, doi:10.1002/pmic.201100477 (2012).
Hogan, M. C. et al. Subfractionation, characterization, and in-depth proteomic analysis of glomerular membrane vesicles in human urine. Kidney international 85, 1225–1237, doi:10.1038/ki.2013.422 (2014).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols 4, 44–57, doi:10.1038/nprot.2008.211 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic acids research 37, 1–13, doi:10.1093/nar/gkn923 (2009).
Jia, L. et al. An attempt to understand kidney’s protein handling function by comparing plasma and urine proteomes. PloS one 4, e5146, doi:10.1371/journal.pone.0005146 (2009).
Farrah, T. et al. State of the human proteome in 2013 as viewed through PeptideAtlas: comparing the kidney, urine, and plasma proteomes for the biology- and disease-driven Human Proteome Project. Journal of proteome research 13, 60–75, doi:10.1021/pr4010037 (2014).
Pieper, R. et al. Characterization of the human urinary proteome: a method for high-resolution display of urinary proteins on two-dimensional electrophoresis gels with a yield of nearly 1400 distinct protein spots. Proteomics 4, 1159–1174, doi:10.1002/pmic.200300661 (2004).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Molecular & cellular proteomics: MCP 11, M111 014050, doi:10.1074/mcp.M111.014050 (2012).
Nagaraj, N. & Mann, M. Quantitative analysis of the intra- and inter-individual variability of the normal urinary proteome. Journal of proteome research 10, 637–645, doi:10.1021/pr100835s (2011).
Dyer, A. R. et al. Evaluation of measures of urinary albumin excretion in epidemiologic studies. American journal of epidemiology 160, 1122–1131, doi:10.1093/aje/kwh326 (2004).
Nolen, B. M. et al. An extensive targeted proteomic analysis of disease-related protein biomarkers in urine from healthy donors. PloS one 8, e63368, doi:10.1371/journal.pone.0063368 (2013).
Kentsis, A. et al. Discovery and validation of urine markers of acute pediatric appendicitis using high-accuracy mass spectrometry. Annals of emergency medicine 55, 62–70 e64, doi:10.1016/j.annemergmed.2009.04.020 (2010).
Shao, C., Wang, Y. & Gao, Y. Applications of urinary proteomics in biomarker discovery. Science China. Life sciences 54, 409–417, doi:10.1007/s11427-011-4162-1 (2011).
Suh, J. H. & Miner, J. H. The glomerular basement membrane as a barrier to albumin. Nature reviews. Nephrology 9, 470–477, doi:10.1038/nrneph.2013.109 (2013).
Welsh, G. I. & Saleem, M. A. The podocyte cytoskeleton–key to a functioning glomerulus in health and disease. Nature reviews. Nephrology 8, 14–21, doi:10.1038/nrneph.2011.151 (2012).
Schwarz, K. et al. Podocin, a raft-associated component of the glomerular slit diaphragm, interacts with CD2AP and nephrin. The Journal of clinical investigation 108, 1621–1629, doi:10.1172/JCI12849 (2001).
Vaidya, V. S., Ferguson, M. A. & Bonventre, J. V. Biomarkers of acute kidney injury. Annual review of pharmacology and toxicology 48, 463–493, doi:10.1146/annurev.pharmtox.48.113006.094615 (2008).
Beck, L. H. Jr. et al. M-type phospholipase A2 receptor as target antigen in idiopathic membranous nephropathy. The New England journal of medicine 361, 11–21, doi:10.1056/NEJMoa0810457 (2009).
Wei, C. et al. Circulating urokinase receptor as a cause of focal segmental glomerulosclerosis. Nature medicine 17, 952–960, doi:10.1038/nm.2411 (2011).
Flint, R. S. et al. Probing the urinary proteome of severe acute pancreatitis. HPB: the official journal of the International Hepato Pancreato Biliary Association 9, 447–455, doi:10.1080/13651820701721744 (2007).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419, doi:10.1126/science.1260419 (2015).
Tran, J. C. et al. Mapping intact protein isoforms in discovery mode using top-down proteomics. Nature 480, 254–258, doi:10.1038/nature10575 (2011).
Cargile, B. J., Sevinsky, J. R., Essader, A. S., Eu, J. P. & Stephenson, J. L. Jr. Calculation of the isoelectric point of tryptic peptides in the pH 3.5–4.5 range based on adjacent amino acid effects. Electrophoresis 29, 2768–2778, doi:10.1002/elps.200700701 (2008).
Rabilloud, T. et al. Power and limitations of electrophoretic separations in proteomics strategies. Mass spectrometry reviews 28, 816–843, doi:10.1002/mas.20204 (2009).
Vaezzadeh, A. R., Briscoe, A. C., Steen, H. & Lee, R. S. One-step sample concentration, purification, and albumin depletion method for urinary proteomics. Journal of proteome research 9, 6082–6089, doi:10.1021/pr100924s (2010).
Guldbrandsen, A. et al. In-depth characterization of the cerebrospinal fluid (CSF) proteome displayed through the CSF proteome resource (CSF-PR). Molecular & cellular proteomics: MCP 13, 3152–3163, doi:10.1074/mcp.M114.038554 (2014).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nature methods 6, 359–362, doi:10.1038/nmeth.1322 (2009).
Wang, Y. et al. Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human MCF10A cells. Proteomics 11, 2019–2026, doi:10.1002/pmic.201000722 (2011).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Analytical chemistry 75, 4646–4658 (2003).
Schwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342, doi:10.1038/nature10098 (2011).
Hauck, S. M. et al. Deciphering membrane-associated molecular processes in target tissue of autoimmune uveitis by label-free quantitative mass spectrometry. Molecular & cellular proteomics: MCP 9, 2292–2305, doi:10.1074/mcp.M110.001073 (2010).
Satoskar, A. A. et al. Characterization of glomerular diseases using proteomic analysis of laser capture microdissected glomeruli. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc 25, 709–721, doi:10.1038/modpathol.2011.205 (2012).
Skoberne, A., Konieczny, A. & Schiffer, M. Glomerular epithelial cells in the urine: what has to be done to make them worthwhile? American journal of physiology. Renal physiology 296, F230–241, doi:10.1152/ajprenal.90507.2008 (2009).
Fukuda, H. et al. Podocin is translocated to cytoplasm in puromycin aminonucleoside nephrosis rats and in poor-prognosis patients with IgA nephropathy. Cell and tissue research 360, 391–400, doi:10.1007/s00441-014-2100-9 (2015).
Mansour, H. et al. T-cell transcriptome analysis points up a thymic disorder in idiopathic nephrotic syndrome. Kidney international 67, 2168–2177, doi:10.1111/j.1523-1755.2005.00322.x (2005).
Kimura, M. et al. Expression of alpha-actinin-4 in human diabetic nephropathy. Internal medicine 47, 1099–1106 (2008).
Kaplan, J. M. et al. Mutations in ACTN4, encoding alpha-actinin-4, cause familial focal segmental glomerulosclerosis. Nature genetics 24, 251–256, doi:10.1038/73456 (2000).
Kubiak-Wlekly, A. et al. The Comparison of the Podocyte Expression of Synaptopodin, CR1 and Neprilysin in Human Glomerulonephritis: Could the Expression of CR1 be Clinically Relevant? International journal of biomedical science: IJBS 5, 28–36 (2009).
Johnstone, D. B. et al. Podocyte-specific deletion of Myh9 encoding nonmuscle myosin heavy chain 2A predisposes mice to glomerulopathy. Molecular and cellular biology 31, 2162–2170, doi:10.1128/MCB.05234-11 (2011).
Yard, B. A. et al. Decreased glomerular expression of agrin in diabetic nephropathy and podocytes, cultured in high glucose medium. Experimental nephrology 9, 214–222, doi:52614 (2001).
Joosten, S. A. et al. Antibody response against the glomerular basement membrane protein agrin in patients with transplant glomerulopathy. American journal of transplantation: official journal of the American Society of Transplantation and the American Society of Transplant Surgeons 5, 383–393, doi:10.1111/j.1600-6143.2005.00690.x (2005).
Kashtan, C. E. & Kim, Y. Distribution of the alpha 1 and alpha 2 chains of collagen IV and of collagens V and VI in Alport syndrome. Kidney international 42, 115–126 (1992).
Kim, Y. et al. Differential expression of basement membrane collagen chains in diabetic nephropathy. The American journal of pathology 138, 413–420 (1991).
Kim, Y. et al. Differential expression of basement membrane collagen in membranous nephropathy. The American journal of pathology 139, 1381–1388 (1991).
Setty, S. et al. Differential expression of laminin isoforms in diabetic nephropathy and other renal diseases. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc 25, 859–868, doi:10.1038/modpathol.2011.216 (2012).
Jim, B. et al. Dysregulated nephrin in diabetic nephropathy of type 2 diabetes: a cross sectional study. PloS one 7, e36041, doi:10.1371/journal.pone.0036041 (2012).
Kim, J. M. et al. CD2-associated protein haploinsufficiency is linked to glomerular disease susceptibility. Science 300, 1298–1300, doi:10.1126/science.1081068 (2003).
Ye, H. et al. Urinary podocalyxin positive-element occurs in the early stage of diabetic nephropathy and is correlated with a clinical diagnosis of diabetic nephropathy. Journal of diabetes and its complications 28, 96–100, doi:10.1016/j.jdiacomp.2013.08.006 (2014).
Tufro, A. & Veron, D. VEGF and podocytes in diabetic nephropathy. Seminars in nephrology 32, 385–393, doi:10.1016/j.semnephrol.2012.06.010 (2012).
El-Koraie, A. F., Baddour, N. M., Adam, A. G., El-Kashef, E. H. & El Nahas, A. M. Cytoskeletal protein expression and regenerative markers in schistosomal nephropathy. Nephrology, dialysis, transplantation: official publication of the European Dialysis and Transplant Association - European Renal Association 17, 803–812 (2002).
Birmingham, D. J. et al. Relationship of Circulating Anti-C3b and Anti-C1q IgG to Lupus Nephritis and Its Flare. Clinical journal of the American Society of Nephrology: CJASN 11, 47–53, doi:10.2215/CJN.03990415 (2016).
Ilhan, N., Kahraman, N., Seckin, D., Ilhan, N. & Colak, R. Apo E gene polymorphism on development of diabetic nephropathy. Cell biochemistry and function 25, 527–532, doi:10.1002/cbf.1348 (2007).
Fugmann, T. et al. Proteomic identification of vanin-1 as a marker of kidney damage in a rat model of type 1 diabetic nephropathy. Kidney international 80, 272–281, doi:10.1038/ki.2011.116 (2011).
Wang, Q. Z. et al. Cofilin1 is involved in hypertension-induced renal damage via the regulation of NF-kappaB in renal tubular epithelial cells. Journal of translational medicine 13, 323, doi:10.1186/s12967-015-0685-8 (2015).
Castelletti, F. et al. Mutations in FN1 cause glomerulopathy with fibronectin deposits. Proceedings of the National Academy of Sciences of the United States of America 105, 2538–2543, doi:10.1073/pnas.0707730105 (2008).
Hanamura, K. et al. Detection of myeloperoxidase in membranous nephropathy-like deposits in patients with anti-neutrophil cytoplasmic antibody-associated glomerulonephritis. Human pathology 42, 649–658, doi:10.1016/j.humpath.2010.08.020 (2011).
Bonventre, J. V., Vaidya, V. S., Schmouder, R., Feig, P. & Dieterle, F. Next-generation biomarkers for detecting kidney toxicity. Nature biotechnology 28, 436–440, doi:10.1038/nbt0510-436 (2010).
Goligorsky, M. S., Addabbo, F. & O’Riordan, E. Diagnostic potential of urine proteome: a broken mirror of renal diseases. Journal of the American Society of Nephrology: JASN 18, 2233–2239, doi:10.1681/ASN.2006121399 (2007).
Vaidya, V. S. & Bonventre, J. V. Mechanistic biomarkers for cytotoxic acute kidney injury. Expert opinion on drug metabolism & toxicology 2, 697–713, doi:10.1517/17425255.2.5.697 (2006).
Trof, R. J., Di Maggio, F., Leemreis, J. & Groeneveld, A. B. Biomarkers of acute renal injury and renal failure. Shock 26, 245–253, doi:10.1097/01.shk.0000225415.5969694.ce (2006).
Gal-Yam, E. N., Saito, Y., Egger, G. & Jones, P. A. Cancer epigenetics: modifications, screening, and therapy. Annual review of medicine 59, 267–280, doi:10.1146/annurev.med.59.061606.095816 (2008).
Thrailkill, K. M. et al. Microalbuminuria in type 1 diabetes is associated with enhanced excretion of the endocytic multiligand receptors megalin and cubilin. Diabetes care 32, 1266–1268, doi:10.2337/dc09-0112 (2009).
Schaub, J. A. et al. Perioperative heart-type fatty acid binding protein is associated with acute kidney injury after cardiac surgery. Kidney international 88, 576–583, doi:10.1038/ki.2015.104 (2015).
Wu, I. & Parikh, C. R. Screening for kidney diseases: older measures versus novel biomarkers. Clinical journal of the American Society of Nephrology: CJASN 3, 1895–1901, doi:10.2215/CJN.02030408 (2008).
Ricci, Z. & Ronco, C. Year in review: Critical Care 2004 - nephrology. Critical care 9, 523–527, doi:10.1186/cc3791 (2005).
Iida, T. et al. Decreased urinary calbindin 1 levels in proteinuric rats and humans with distal nephron segment injuries. Clinical and experimental nephrology 18, 432–443, doi:10.1007/s10157-013-0835-3 (2014).
Sawai, K. et al. Expression of CCN1 (CYR61) in developing, normal, and diseased human kidney. American journal of physiology. Renal physiology 293, F1363–1372, doi:10.1152/ajprenal.00205.2007 (2007).
Lameire, N., Van Biesen, W. & Vanholder, R. Acute renal failure. Lancet 365, 417–430, doi:10.1016/S0140-6736(05)17831-3 (2005).
Nuyts, G. D. et al. Human urinary intestinal alkaline phosphatase as an indicator of S3-segment-specific alterations in incipient diabetic nephropathy. Nephrology, dialysis, transplantation: official publication of the European Dialysis and Transplant Association - European Renal Association 9, 377–381 (1994).
Adiyanti, S. S. & Loho, T. Acute Kidney Injury (AKI) biomarker. Acta medica Indonesiana 44, 246–255 (2012).
Ramesh, G., Kwon, O. & Ahn, K. Netrin-1: a novel universal biomarker of human kidney injury. Transplantation proceedings 42, 1519–1522, doi:10.1016/j.transproceed.2009.11.040 (2010).
Mishra, J. et al. Neutrophil gelatinase-associated lipocalin (NGAL) as a biomarker for acute renal injury after cardiac surgery. Lancet 365, 1231–1238, doi:10.1016/S0140-6736(05)74811-X (2005).
Yu, X. Q. et al. Osteopontin expression in progressive renal injury in remnant kidney: role of angiotensin II. Kidney international 58, 1469–1480, doi:10.1046/j.1523-1755.2000.00309.x (2000).
Liu, Y. et al. Urinary interleukin 18 for detection of acute kidney injury: a meta-analysis. American journal of kidney diseases: the official journal of the National Kidney Foundation 62, 1058–1067, doi:10.1053/j.ajkd.2013.05.014 (2013).
Varghese, S. A. et al. Identification of diagnostic urinary biomarkers for acute kidney injury. Journal of investigative medicine: the official publication of the American Federation for Clinical Research 58, 612–620, doi:10.231/JIM.0b013e3181d473e7 (2010).
Acknowledgements
This work was supported by the National Basic Research Program of China (No. 2013CB530805, No. 2014CBA02005), National Key Research and Development Program of China (No. 2016 YFC 1306300), the National Natural Science Foundation of China (No. 31200614, No. 31400669), the Beijing Natural Science Foundation (No. 5132028, 7173264, 7172076), the Fundamental Research Funds for the Central Universities (2015KJJCB21), Beijing Normal University (No. 11100704) and Biologic Medicine Information Center of China, National Scientific Data Sharing Platform for Population and Health.
Author information
Authors and Affiliations
Contributions
M.Z., W.S. and Y.G. prepared the first draft. M.L. (Menglin Li) and W.S. conceived and designed the experiments. M.L. (Menglin Li), and Z.G. performed the experiments. M.Z., Y.Y. and C.S. analyzed the data. Y.S. and M.L. (Mingxi Li) contributed to collect clinical samples. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, M., Li, M., Yang, Y. et al. A comprehensive analysis and annotation of human normal urinary proteome. Sci Rep 7, 3024 (2017). https://doi.org/10.1038/s41598-017-03226-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03226-6
- Springer Nature Limited
This article is cited by
-
Proteomics of prostate cancer serum and plasma using low and high throughput approaches
Clinical Proteomics (2024)
-
Recent progress in mass spectrometry-based urinary proteomics
Clinical Proteomics (2024)
-
Identification and characterization of intact glycopeptides in human urine
Scientific Reports (2024)
-
Urinary complement factor D is increased in primary malignant hypertension: a single-center, cross-sectional study
Scientific Reports (2024)
-
Proteomic profiling of cerebrospinal fluid reveals TKT as a potential biomarker for medulloblastoma
Scientific Reports (2024)