Abstract
Chemically synthesized short interfering RNA (siRNA) has ushered a new era in the application of RNA interference (RNAi) against viral genes. We have paid particular attention to respiratory viruses that wreak heavy morbidity and mortality worldwide. The clinically significant ones include respiratory syncytial virus (RSV), parainfluenza virus (PIV) (two Paramyxoviruses), and influenza virus (an Orthomyxovirus). As the infection by these viruses is clinically restricted to the respiratory tissues, mainly the lungs, the logical route for the application of the siRNA was also the same, i.e., via the nasal route. Following the initial success of single intranasal siRNA against RSV, we now offer two new strategies: (1) second-generation siRNAs, used against the paramyxoviral RNA polymerase large subunit (L), (2) siRNA cocktail with a novel transfection reagent, used against influenza virus. Based on these results, we propose the following consensus for designing intranasal antiviral siRNAs: (a) modified 19–27 nt-long double-stranded siRNAs are functional in the lung, (b) excessive 2′-OMe and 2′-F modifications in either or both strands of these siRNAs reduce efficacy, (c) limited modifications in the sense strand are beneficial, although their precise efficacy may be position-dependent, (d) cocktail of multiple siRNAs can be highly effective against multiple viral strains and subtypes.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
Short interfering RNAs (siRNAs) trigger posttranscriptional gene silencing in a variety of metazoan cells and tissues [1]. Successful use of synthetic double-stranded siRNA in 2001 [1, 2] targeting cellular and viral genes in cell culture, opened the door to siRNA as prospective antivirals and drugs for gene therapy. The “first-generation” siRNAs were designed to mimic the products of the Dicer endonuclease cleavage; they were 19 nt-long duplexes with 3′-terminal 2 nt overhangs and contained natural, unmodified ribose and bases. As reported by us [3, 4] and several other laboratories [5–7], these siRNAs proved effective both in cell culture and in animals.
The advantages of siRNA-based therapy over traditional organic small molecule drugs are as follows: (1) The siRNAs can be rapidly synthesized and scaled up for production; (2) In the event of viral resistance to the existing siRNA, one can shift to a different target sequence; (3) All siRNAs need the same synthetic chemistry and hence, can be manufactured in an established national facility; (4) siRNAs are water-soluble; properly formulated lyophilized preparations can have a long shelf life, and can be easily reconstituted with sterile water.
Of particular mention is the potent antiviral activity of siRNAs, first documented against respiratory syncytial virus (RSV) as proof-of-concept [3]. In this study, synthetic first-generation siRNAs, designed against essential RSV genes, prevented RSV growth in cultured lung epithelial cells [3]. To translate this success to an animal model [4], we reasoned that the most logical route for delivery of these siRNAs should be the nose (intranasal). First, it is noninvasive. Second, it should involve relatively simple application procedures, such as nasal drops or aerosol inhalation. Third, the siRNA drug will follow the same route as the virus and reach the same tissues, including the lungs. Lastly, our subsequent results showed that the intranasally administered siRNA remains in the lung and does not leach into blood or other tissues, minimizing the risk of systemic side effects (data not shown). This is particularly important since, unlike cell-specific delivery reagents [5], the reagents used here (TransIT-TKO or HKP) are universally applicable, and therefore, intranasal delivery serves to have the cargo localized within the lung, providing specificity against respiratory infections. In this chapter, we have, therefore, focused on the intranasal delivery of anti-respiratory viral siRNAs.
We first summarize the progress made in four major respiratory viruses: RSV, parainfluenza virus (PIV), influenza virus (Flu), and severe acute respiratory syndrome (SARS) coronavirus (SARS-CoV) [4, 6, 8–11]. The first three viruses are clinically the most significant, claiming a large number of human lives each year throughout the world. Annual flu epidemics alone affect 10–20 % of the U.S. population, averaging about 114,000 hospitalizations. Currently, two strategies, vaccines and small molecule therapeutics, are utilized to control the spread of flu. However, the high mutation rate of RNA genomes in general and the reassortment of flu virus genome segments in particular are major hurdles in flu prevention and treatment. The rapid emergence of new flu strains that are resistant against the M2 ion channel inhibitors (amantadine, rimantadine) as well as the viral neuraminidase (NA) inhibitors (zanamivir, oseltamivir) is a significant global concern [12]. RSV is the most significant cause of severe lower respiratory tract infection in infants and an important cause of acute respiratory illness in the elderly as well [13, 14]. In pediatric infection, PIV is second only to RSV [15]. Ribavirin and IFN, although partially effective in treating these infections, are both relatively nonspecific and toxic [14]. Regarding the molecular features, RSV and PIV are nonsegmented negative-strand RNA viruses, belonging to different genera of the Paramyxoviridae family [6]. In contrast, flu is an Orthomyxovirus and contains segmented negative-strand RNA genome [9, 10]. Coronoviruses such as SARS-CoV contain positive-strand RNA genomes [11]. A commonality among these RNA viruses is that they encode genes for RNA-dependent RNA polymerase (RdRP) to transcribe and replicate their own genomes, because the host animal cells are devoid of such activity. Many of the potent antiviral siRNAs have, therefore, targeted viral genes coding for RdRP subunits or proteins that are related to RdRP function [3, 4, 6, 9–11]. These early studies targeted: P (Phosphoprotein), L (Large) (see Table 1), and N (Nucleocapsid) genes of RSV; P and L genes of PIV; PA and NP genes of Flu; and the replicase gene of SARS-CoV. In all cases, siRNAs were designed against essential viral gene(s), optimized in cell culture and then used intranasally in the appropriate animal model. Antiviral activity was shown against RSV and parainfluenza virus (PIV) using the BALB/c mouse model as well [4]. In those studies, optimal results were obtained with siRNAs delivered in complex with either Oligofectamine (Invitrogen, Carlsbad, CA) or Mirus Transit TKO reagent (see Subheading 2), although uncomplexed intranasal siRNA also showed some efficacy [4]. Intranasal siRNA, complexed with oligofectamine or polyethyleneimine (PEI), was also protective against highly pathogenic influenza A viruses of the H5 and H7 subtypes in mice [9, 10]. Notwithstanding their success, the activity of these siRNAs was transient, lasting only a few days. Therefore, enhancement of the intracellular and extracellular stability of synthetic siRNAs while increasing (or without compromising) their RNAi activity is a continuing goal for therapeutic translation of RNAi.
A variety of chemical modifications, including terminal and internal ones, have been added to the first-generation siRNA sequences to improve stability and delivery, leading to what we call “second-generation” siRNAs. Advantage has been taken of the free 2′-OH group of the ribose moiety of RNA (in contrast to DNA that lacks this OH group), to which various substituents were added. We have pursued two promising ones, namely 2′-O-methyl (2′-O-Me) and 2′-fluoride (2′-F). The latter modification is placed on pyrimidine nucleosides (C, U), leading to 2′-FC and 2′-FU residues. In a number of previous studies [16–19], these substitutions were introduced to various extents in the antisense strand (“guide” strand) or both strands of the siRNA and were shown to enhance stability and potency, although intranasal application was not tested. Additionally, they tend to reduce siRNA-driven innate immune response [20, 21]. In systematic studies in cell culture, the biochemical and functional activity of the siRNA was vindicated but found to be affected by the position of the modifications in the sequence [16–19]. Generally speaking, those with the modified ribonucleotides at the 5′-end of the antisense strand were less active relative to the 3′-modified ones. Internally, while 2′-F was generally well-tolerated on the antisense strand, 2′-O-Me showed significant shift in activity depending on the position. In contrast, incorporation of 2′-O-Me in the sense strand of siRNA did not show a strong positional preference. In a comprehensive study, however, internal 2′-O-Me modifications in either or both strands actually made the siRNA less active [22]. In an animal experiment [23], all the 2′-OH residues in siRNAs against hepatitis B virus were substituted with 2′-F and 2′-O-Me. When administered intravenously (i.v.) as lipid complexes, the 2′-O-Me, 2′-F siRNAs showed improved efficacy and longer half-life in plasma and liver. When these siRNAs were additionally Cy3-labeled, it revealed their accumulation in the liver and spleen, but not in the lung, explaining the success of the i.v. administration against hepatitis while suggesting that it is an ineffective route against lung infections.
Based on the absence of a uniform pattern in these studies, we reasoned that siRNAs against respiratory viruses should be individually optimized through the following steps: (1) Design the first-generation siRNA following the generally accepted sequence rules or an algorithm of your choice. (2) Select the ones with lowest IC50 (preferably below ~20 nM) in a cell culture assay for virus growth, (3) If desired, as we did for RSV (Table 1), add OMe and F substitutions in various “format” (i.e., number and placement) to generate modified, “second-generation” siRNAs. (4) Ensure that the substitutions either improved or did not reduce knockdown efficiency by screening in cell culture, (5) Confirm efficacy and lack of toxicity in animal model, and (6) test improved stability in serum and blood in vitro. Finally, if deemed necessary and resources permit, a low- or high-throughput assay for off-target effects can be performed to further ensure target specificity.
Together, the small but representative datasets cover multiple viruses, modified and unmodified siRNA. We present a consensus procedure tested in our laboratory, but essentially the same protocol can be used to test other modifications or delivery reagents as they become available. In flu, for example, we have found that the HK polymer is in fact better than a number of other reagents tested (Fig. 1).
Designing flu siRNA. siRNA was designed against sequences of M2 (left) or NP (right) genes, which are conserved in a large number of flu virus isolates. The number of flu strains showing match with the siRNA is shown on the Z-axis (vertical). The two horizontal axes represent 16 HA subtypes and 9 N subtypes. The N subunits are color-coded (N1 = brown; N2 = green, etc.), and well-known subtypes are indicated on the top (e.g., H1N1, H3N2, and H5N1). For example, the most broad-spectrum siRNAs against M2 can target 300 flu strains, including H1N1
2 Materials
The reagents described below have been used successfully by us but various equivalents are available commercially that can be optimized.
2.1 siRNAs and Their Use
-
1.
Synthetic siRNA of the chosen sequence, lyophilized, purchased without any modification, 2 nmol (Dharmacon, Lafayette, CO).
-
2.
Selected, most effective siRNA(s) from above, purchased with 2′-O-Me, 2′-FC, 2′-FU modifications in various formats (Dharmacon), in both sense and antisense strands.
-
3.
RNase-free ART aerosol-resistant pipette tips (Molecular BioProducts, San Diego, CA).
-
4.
RNase-free microfuge tubes (Ambion, Austin, TX).
-
5.
siRNA buffer: 20 mM KCl, 6 mM HEPES-KOH pH 7.5, 0.2 mM MgCl2. This is usually supplied by the siRNA manufacturer (Dharmacon) as 6× stock; dilute as and when needed.
2.2 Cell Culture and Virus Growth
We assume that the reader has access to an appropriate cell culture facility, consisting of incubators and culture hoods, and there is available expertise on virus growth and assay. In this chapter, we will only cover specific issues related to the testing of antiviral siRNA against RSV. Common respiratory viruses, such as RSV, PIV, and seasonal human influenza are biosafety level 2 (BSL2) pathogens. However, highly pathogenic influenza strains, such as avian flu H5N1 or the swine flu H1N1 virus, may require BSL3 or higher facilities. Obey all institutional regulations. The following materials are needed:
-
1.
Suitable cell line, e.g., A549 or HEp-2 cells for RSV; MDCK for flu.
-
2.
Standard cell culture media. RSV and the host cell lines grow virtually in any cell culture media. Use D-MEM with glucose, supplemented with glutamine and 10 % fetal bovine serum (heat-inactivated) with or without penicillin/streptomycin.
-
3.
Standard disposable sterile plastic ware for cell culture.
2.3 Animals and Related
-
1.
BALB/c mice, 8–10 weeks old, weighing 16–20 g (Charles River Laboratories, Wilmington, MA).
-
2.
5 mg/ml sodium pentobarbital (Nembutal).
-
3.
25-G single-use hypodermic needles (VWR, West Chester, PA).
-
4.
1 cc single-use syringes with BD Luer-Lok tip (VWR).
2.4 siRNA Transfection
-
1.
TransIT-TKO siRNA transfection reagent (Mirus Bio Corporation, Madison, WI).
-
2.
Opti-MEM I Reduced Serum Medium (Gibco, Invitrogen Corporation, Carlsbad, CA).
-
3.
RNase-free gel-loading microcapillary tips (VWR).
-
4.
RNase-free microfuge tubes (Ambion).
3 Methods
3.1 siRNA Sequence Design
-
1.
Use a free online siRNA design program for your target mRNA sequence (see Note 1 ). We generally use the one available on the Whitehead Institute (MIT) server at the following URL (http://sirna.wi.mit.edu/); registration is required but free. Copy and paste the target sequence in the box and initially choose AAN19TT in the “recommended patterns.” Usually, there is no need to change the “Filter criteria” below, i.e., leave them as in the default. Click on the Search button. With RSV L gene, we found many prospective ones (Table 1), but if no siRNA is obtained with your gene sequence, repeat the procedure, this time choosing NAN21.
With flu, a proprietary algorithm was used and thus, the siRNA sequences (Fig. 2) are also intellectual property. In general, for a wider coverage of multiple virus strains and to minimize resistance, choose viral sequences that are most conserved.
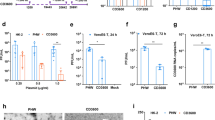
Fig. 2 High efficiency of the HK polymer in siRNA transfection. A propreitary siRNA against human cyclophilin B mRNA was tested for delivery into HEp-2 cells using the following reagents: His-Lys polymer (HKP) with selected branching [27, 28]; the cationic lipid, 1,2-dioleoyl-3-trimethylammonium-propane (DOTAP) [29]; and 5 % glucose (dextrose) in water (D5W), a common physiological solution used in the clinic. Blank is siRNA administered with no transfection reagent. Note the strong performance of HKP, especially discernible with 10 μg siRNA
-
2.
This will lead to a table of prospective siRNA sequences that are ordered according to “Thermodynamic values” by the default setting. The goal is to select sequences with high negative values, i.e., sequences closer to the top of the table. In addition, visually examine the central N19 part (i.e., ignore the first two and the last two nucleotides) and pick the ones that are AT-rich at the 3′-end (right hand side) and GC-rich at the 5′-end (left hand side). The number of sequences one can test really depends on available resources, but order at least 3 for a given target. Order the sense strand as N19dTdT and the antisense strand as (N′)19dTdT, such that the N′ part is complementary to the N19 sequence. Here, we provide an example of tested functional siRNA sequences based on the L polymerase gene of RSV (Table 1). Assume that you will receive the siRNA roughly a week after ordering. Purchase the smallest amount initially (to save money), generally 2 nmol of each.
-
3.
After receiving the lyophilized RNA (2 nmol), briefly spin the tubes to ensure that the RNA is at the bottom. Add 42 μl 1× siRNA buffer to each RNA. Pipette the solution up and down 8–10 times, avoiding the introduction of bubbles.
-
4.
Place the solution on an orbital mixer/shaker for 30 min at room temperature for complete mixing. Briefly centrifuge the tubes containing siRNA to ensure that the solution is collected at the bottom of the tube.
-
5.
Combine the volumes of the complementary strands of RNA (42 + 42 = 84 μl), vortex for 10 s and centrifuge for 30 s. Add 16 μl of 6× siRNA buffer to make 100 μl, mix.
-
6.
Heat the mixture at 60 °C for 45 min.
-
7.
Remove from heat and centrifuge briefly, 5–10 s.
-
8.
Allow solution to cool to room temperature over 30 min to allow formation of the double-stranded siRNA, which is now ready to use. The siRNA concentration of this stock solution is 20 μM (see Notes 2 and 3 ).
-
9.
Aliquot the siRNA into small volumes and store at −20 to −80 °C. For best results, do not freeze-thaw more than four times.
3.2 Reiterated Testing of Modified and Unmodified siRNA
-
1.
Trypsinize cells and seed ~5 × 104 cells in 500 μl complete growth medium per well of a 24-well plate. Incubate for 24 h to achieve a confluency of 60–70 %.
-
2.
Immediately before transfection, add 50 μl of Opti-MEM I to a sterile Eppendorf tube. To this, add 2.5 μl TransIT-TKO reagent, mix thoroughly by vortexing, and incubate at room temperature for 10–20 min.
-
3.
Dilute the siRNA 20-fold by mixing 3.5 μl of the stock with 66.5 μl siRNA buffer. Test three concentrations of each siRNA, and each concentration at least in duplicate, and if possible, triplicate. The final concentrations and the corresponding volumes are: 5 nM (1.5 μl), 10 nM (3 μl), and 50 nM (15 μl). Add these volumes to the diluted transfection reagent made in step 2 above and mix by gentle pipetting. Incubate at room temperature for 10–20 min.
-
4.
Adjust the medium volume in each well of the 24-well cell monolayer (step 1 above) to 250 μl by removing half of the original volume (250 μl).
-
5.
Add the transfection-ready siRNA mixture (from step 3) dropwise to the cells. Gently tilt and rock the plate to evenly distribute the complexes.
-
6.
Incubate for 18 h in a cell culture incubator.
-
7.
Add challenge virus to the cells. For RSV, add about 104 virions (1 μl of stock 107 pfu/ml) in each well. For each siRNA concentration, keep an uninfected well as control (to check if the siRNA itself will cause cellular death by off-target effects).
-
8.
Incubate another 24 h if RSV growth is monitored by Western blot or another 72 h if it is to be monitored by extracellular titer. For the latter, replace old media with equal volume of fresh prewarmed media (for better virus growth). For Western, remove media, wash the monolayer with 0.5 ml PBS twice, and add 20 μL of 2× SDS sample buffer. Scrap the monolayer, mix well by pipetting, boil, analyze in SDS-PAGE and Western with anti-RSV antibody. For titer, collect media from the wells. Expect ~106 pfu/ml from uninhibited cells and 1–3 log10 lower if the siRNA works well. Do serial dilutions of each sample accordingly (in fresh complete medium), plate on either A549 or HEp-2 monolayers, incubate for 48 h, count plaques under microscope (20× magnification). Take the average of triplicate plaque assays for each sample.
-
9.
Select the most effective siRNA sequences (strongest antiviral effect) as candidates for 2′-O-Me and 2′-F modifications. For reasons mentioned in the Introduction, there is no universal consensus regarding the format. Thus, the guidelines offered here should be considered suggestions only, although the sequences shown have been tested to work (Table 1). The reader is encouraged to conduct more exhaustive modifications and test other sequences if resources permit.
-
(a)
Introduce 2′-O-Me modifications in all sites in the sense strand initially, but then try less extensive modifications (at various positions), which are sometimes more effective.
-
(b)
Introduce alternate modifications in sense and antisense strands. Start with 2′-F (in available C’s and U’s) followed by 2′-O-Me in the sense strand, and reciprocate in the antisense strand (Table 1). Order the modified siRNA from commercial sources (e.g., Dharmacon) and then process them as in steps 3–9 (Subheading 3.2).
-
(a)
3.3 siRNA Testing in Mice
Prior to the intranasal administration of siRNA, the mouse (see Note 2 ) must be anesthetized by using any standard procedure available in the laboratory, e.g., by administering Nembutal via intraperitoneal (IP) injection. The recommended drug dosage for mice is 50 mg/kg (see Note 3 ).
-
1.
Gently lift the mouse by the tail and place it on a cage lid.
-
2.
Grip the loose skin of the neck to immobilize the head of the mouse. Extend the tail to draw the skin tight over the abdomen by gripping the tail with your little finger.
-
3.
Hold mouse in a head-down position and disinfect injection site (the lower right or left quadrant of the abdomen); use a hypodermic needle to administer the anesthetic. The mouse is ready for siRNA administration when no voluntary movement is observed.
-
4.
Place the anesthetized mouse on a lab towel facing up; with its head immobilized, insert microcapillary tip containing siRNA/transfection reagent complexes (see Note 4 ) into the nostril. Use a range of 2–20 nmol siRNA per mouse (e.g., 2, 6, 10 nmol). Instill solution slowly over a 2–3 min period, allowing the animal to breathe the liquid in (see Note 5 ).
-
5.
Return the mouse to the cage and monitor for at least 45 min to avoid depression of cardiac and/or respiratory functions.
-
6.
Test for the desired RNAi effect at appropriate intervals. For antiviral studies, instill virus through the nostril as well. For human RSV, which does not infect mice well, use 107–108 virus particles per animal, and measure standard lung titer assay and/or clinical symptoms (such as body weight, respiration rate, etc.) (see Note 6 ).
4 Notes
-
1.
Recently, relatively large (26–28 nt) long double-stranded RNAs that act as Dicer substrates (D-siRNA) have been shown to be more potent than the regular 19 nt siRNAs used here [24, 25]. We have also found them to be at least as potent as the first and second generation 19-mer siRNA in terms of intranasal anti-RSV activity without increased immune reactions. In preliminary experiments, they also lent well to 2′-O-Me and 2′-F modifications for intranasal antiviral activity (data not shown), but it is recommended that the exact format be optimized for each sequence. Follow the published D-siRNA design guidelines [24, 26].
-
2.
Although we have described the laboratory mouse model here, intranasal dosage and delivery can be easily scaled up or down for other laboratory animals.
-
3.
siRNA concentration: The pharmaceutical industry prefers expressing drug concentrations in wt/vol or wt/body weight (e.g., mg/kg). For the researcher, however, it is easier and more useful to express siRNA concentrations in molar units (e.g., μM or nM), since this allows direct comparison between the potency of different siRNAs even when they differ in base composition or modifications (and hence formula weight).
-
4.
Transfection reagent: siRNAs may respond differently to different reagents. Experiment with delivery reagent of your choice (Fig. 2).
-
5.
Avoid using excessive liquid because it may suffocate the animal and cause death. As a rule, keep the total instilled volume under 45 μl for BALB/c mice, but higher volumes may be tolerated by larger species.
-
6.
The protocol described here may be modified for aerosolized siRNA using an enclosure to house the anesthetized animal and a handheld nebulizer (the common type used as an inhaler by asthmatics). A larger amount of siRNA is needed because only a fraction of the aerosol is actually inhaled by the animal. If used routinely, consider optimizing a commercial motorized nebulizer. Check with the local pediatricians for the exact model, vendor, and usage. Modify the system by removing the facial mask at the delivery end and inserting the tube directly into the enclosure. A snug fit of the mask should reduce siRNA waste.
References
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Elbashir SM, Martinez J, Patkaniowska A, Lendeckel W, Tuschl T (2001) Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J 20:6877–6888
Bitko V, Barik S (2001) Phenotypic silencing of cytoplasmic genes using sequence-specific double-stranded short interfering RNA and its application in the reverse genetics of wild type negative-strand RNA viruses. BMC Microbiol 1:34
Bitko V, Musiyenko A, Shulyayeva O, Barik S (2004) Inhibition of respiratory viruses by nasally administered siRNA. Nat Med 11:50–55
Sioud M, Skorstad G, Mobergslien A, Sæbøe-Larssen S (2013) A novel peptide carrier for efficient targeting of antigens and nucleic acids to dendritic cells. FASEB J 27:3272–3283
Barik S (2004) Control of nonsegmented negative-strand RNA virus replication by siRNA. Virus Res 102:27–35
Barik S (2005) Silence of the transcripts: RNA interference in medicine. J Mol Med 83:764–773
Bitko V, Barik S (2007) Intranasal antisense therapy: preclinical models with a clinical future? Curr Opin Mol Ther 9:119–125
Tompkins SM, Lo CY, Tumpey TM, Epstein SL (2004) Protection against lethal influenza virus challenge by RNA interference in vivo. Proc Natl Acad Sci U S A 101:8682–8686
Ge Q, Filip L, Bai A, Nguyen T, Eisen HN, Chen J (2004) Inhibition of influenza virus production in virus-infected mice by RNA interference. Proc Natl Acad Sci U S A 101:8676–8681
Li BJ, Tang Q, Cheng D, Qin C, Xie FY, Wei Q, Xu J, Liu Y, Zheng BJ, Woodle MC, Zhong N, Lu PY (2005) Using siRNA in prophylactic and therapeutic regimens against SARS coronavirus in Rhesus macaque. Nat Med 11:944–951
Barik S (2012) New treatments for influenza. BMC Med 10:104
Collins PL, Melero JA (2011) Progress in understanding and controlling respiratory syncytial virus: still crazy after all these years. Virus Res 162:80–99
Maggon K, Barik S (2004) New drugs and treatment for respiratory syncytial virus. Rev Med Virol 14:149–168
Weinberg GA, Hall CB, Iwane MK, Poehling KA, Edwards KM, Griffin MR, Staat MA, Curns AT, Erdman DD, Szilagyi PG (2009) New vaccine surveillance network. Parainfluenza virus infection of young children: estimates of the population-based burden of hospitalization. J Pediatr 154:694–699
De Paula D, Bentley MV, Mahato RI (2007) Hydrophobization and bioconjugation for enhanced siRNA delivery and targeting. RNA 13:431–456
Oller E, Propp S, Murray H, Lima W, Bhat B, Prakash TP, Allerson CR, Swayze EE, Marcusson EG, Dean NM (2006) Competition for RISC binding predicts in vitro potency of siRNA. Nucleic Acids Res 34:4467–4476
Kraynack BA, Baker BF (2006) Small interfering RNAs containing full 2′-O-methylribonucleotide-modified sense strands display Argonaute2/eIF2C2-dependent activity. RNA 12:163–176
Czauderna F, Fechtner M, Dames S, Aygün H, Klippel A, Pronk GJ, Giese K, Kaufmann J (2003) Structural variations and stabilising modifications of synthetic siRNAs in mammalian cells. Nucleic Acids Res 31:2705–2716
Sioud M, Furset G, Cekaite L (2007) Suppression of immunostimulatory siRNA-driven innate immune activation by 2′-modified RNAs. Biochem Biophys Res Commun 361:122–126
Sioud M (2011) Promises and challenges in developing RNAi as a research tool and therapy. Methods Mol Biol 703:173–187
Chiu YL, Rana TM (2003) siRNA function in RNAi: a chemical modification analysis. RNA 9:1034–1048
Morrissey DV, Lockridge JA, Shaw L, Blanchard K, Jensen K, Breen W, Hartsough K, Machemer L, Radka S, Jadhav V, Vaish N, Zinnen S, Vargeese C, Bowman K, Shaffer CS, Jeffs LB, Judge A, MacLachlan I, Polisky B (2005) Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nat Biotechnol 23:1002–1007
Kim DH, Behlke MA, Rose SD, Chang MS, Choi S, Rossi JJ (2005) Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat Biotechnol 23:222–226
Siolas D, Lerner C, Burchard J, Ge W, Linsley PS, Paddison PJ, Hannon GJ, Cleary MA (2005) Synthetic shRNAs as potent RNAi triggers. Nat Biotechnol 23:227–231
Amarzguioui M, Rossi JJ (2008) Principles of Dicer substrate (D-siRNA) design and function. Methods Mol Biol 442:3–10
Leng Q, Woodle MC, Lu PY, Mixson AJ (2009) Advances in systemic siRNA delivery. Drugs Future 34:721
Chou ST, Leng Q, Scaria P, Woodle M, Mixson AJ (2011) Cancer Gene Ther 18:707–716
Simberg D, Weisman S, Talmon Y, Barenholz Y (2004) DOTAP (and other cationic lipids): chemistry, biophysics, and transfection. Crit Rev Ther Drug Carrier Syst 21:257–317
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Barik, S., Lu, P. (2015). Therapy of Respiratory Viral Infections with Intranasal siRNAs. In: Sioud, M. (eds) RNA Interference. Methods in Molecular Biology, vol 1218. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1538-5_14
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1538-5_14
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1537-8
Online ISBN: 978-1-4939-1538-5
eBook Packages: Springer Protocols